








生物技术通报 ›› 2023, Vol. 39 ›› Issue (1): 104-114.doi: 10.13560/j.cnki.biotech.bull.1985.2022-0417
收稿日期:2022-04-07
出版日期:2023-01-26
发布日期:2023-02-02
作者简介:陈泉冰,女,硕士,研究方向:酶工程与生物催化;E-mail: 基金资助:
CHEN Quan-bing1( ), CAO Wei-jie1, LI Chun1,2, LV Bo1(
), CAO Wei-jie1, LI Chun1,2, LV Bo1( )
)
Received:2022-04-07
Published:2023-01-26
Online:2023-02-02
摘要:
GH79家族的糖苷水解酶在碳水化合物改性、细胞免疫识别和信号传导等方面具有广泛的生理活性和重要的应用前景。然而,目前GH79家族的多样性催化机理仍不清楚,识别底物的结构基础和分子机制尚不清晰。本文总结了近几年GH79家族的研究进展,系统分析了GH79家族酶的来源与分布,通过对酶的序列特征、分子进化关系、蛋白结构解析等方面进行深入阐述,旨在为后续的GH79家族的蛋白质工程和功能催化机制的解析奠定基础。
陈泉冰, 曹伟洁, 李春, 吕波. GH79家族糖苷水解酶分子进化关系和蛋白结构研究[J]. 生物技术通报, 2023, 39(1): 104-114.
CHEN Quan-bing, CAO Wei-jie, LI Chun, LV Bo. Molecular Evolutionary Relationship and Protein Structure of Glycoside Hydrolases from GH79 Family[J]. Biotechnology Bulletin, 2023, 39(1): 104-114.
| 水解方式 Hydrolysis method | 名称 Name | 切割位点 Cleavage site |
|---|---|---|
| 外切型 Exo-type | 唾液酸酶 | α-2,6糖苷键连接的唾液酸残基 |
| β-半乳糖苷酶 | β-1,4糖苷键连接的半乳糖残基 | |
| α-甘露糖苷酶 | α-1,2糖苷键,α-1,3糖苷键,α-1,6糖苷键连接的甘露糖型 | |
| β-甘露糖苷酶 | β-1,4糖苷键连接的甘露糖型 | |
| β-淀粉酶 | 自非还原末端依次切割麦芽糖单位,不切也不逾越α-1,6糖苷键 | |
| 内切型 Endo-type | α-淀粉酶 | 在分子内部随机切割α-1,4糖苷键,不切α-1,6糖苷键 |
| 异淀粉酶 | 水解支链淀粉或糖原中的α-1,6糖苷键 | |
| Endo F | β-1,4糖苷键连接的高甘露糖型杂合糖型 | |
| Endo H | β-1,4糖苷键连接的高甘露糖型杂合糖型 | |
| O-糖苷酶 | 水解蛋白核心1(Galβ1-3 GalNAC-Thr/Ser)和核心3(GlcNACβ1-3 GalNAC-Thr/Ser)之间的O-连接二糖单位 |
表1 糖苷水解酶的分类
Table 1 Classification of glycoside hydrolases
| 水解方式 Hydrolysis method | 名称 Name | 切割位点 Cleavage site |
|---|---|---|
| 外切型 Exo-type | 唾液酸酶 | α-2,6糖苷键连接的唾液酸残基 |
| β-半乳糖苷酶 | β-1,4糖苷键连接的半乳糖残基 | |
| α-甘露糖苷酶 | α-1,2糖苷键,α-1,3糖苷键,α-1,6糖苷键连接的甘露糖型 | |
| β-甘露糖苷酶 | β-1,4糖苷键连接的甘露糖型 | |
| β-淀粉酶 | 自非还原末端依次切割麦芽糖单位,不切也不逾越α-1,6糖苷键 | |
| 内切型 Endo-type | α-淀粉酶 | 在分子内部随机切割α-1,4糖苷键,不切α-1,6糖苷键 |
| 异淀粉酶 | 水解支链淀粉或糖原中的α-1,6糖苷键 | |
| Endo F | β-1,4糖苷键连接的高甘露糖型杂合糖型 | |
| Endo H | β-1,4糖苷键连接的高甘露糖型杂合糖型 | |
| O-糖苷酶 | 水解蛋白核心1(Galβ1-3 GalNAC-Thr/Ser)和核心3(GlcNACβ1-3 GalNAC-Thr/Ser)之间的O-连接二糖单位 |
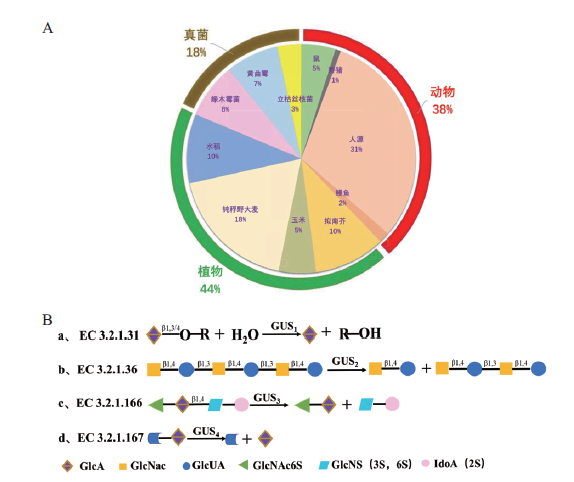
图1 GH79家族糖苷水解酶的分布和催化类型分类 A:不同来源的GH79糖苷酶分布;B:GH79糖苷酶催化的分子模型
Fig. 1 Distribution and classification of catalytic types of glycoside hydrolases in the family GH79 A: Distribution of GH79 glycosidase from different sources. B: Molecular model of GH79 glycosidase catalysis
| 蛋白名称 Protein name | 来源 Organism | 底物 Substrate | EC编号 EC# | 参考文献 Reference |
|---|---|---|---|---|
| GlcA79A | Acidobacterium capsulatum | β-D-glucuronicacid | 3.2.1.31 | [ |
| AtGUS2 | Arabidopsis thaliana | pNPGlcA | 3.2.1.31 | [ |
| Nc6GAL | Neurospora crassa | Arabinogalactan-proteins | 3.2.1.31 | [ |
| FobglcA | Fusarium oxysporum | Gum arabic | 3.2.1.31 | [ |
| BpHep | Burkholderia pseudomallei | Heparan sulfate | 3.2.1.166 | [ |
| HpsE | Rattus norvegicus | Heparan sulfate | 3.2.1.166 | [ |
| Hpse1 | Danio rerio | Heparan sulfate glycosaminoglycans | 3.2.1.166 | [ |
| Hpa1 | Homo sapiens | Heparan sulfate | 3.2.1.166 | [ |
| Heparanase | Gallus gallus | Heparan sulfate | 3.2.1.166 | [ |
| Heparanase | Homo sapiens | Heparan sulfate | 3.2.1.166 | [ |
| Heparanase | Sus scrofa | Heparan sulfate | 3.2.1.166 | [ |
| Hpa | Mus musculus | Heparan sulfate | 3.2.1.166 | [ |
| Hyaluronidase | Hirudo nipponia | Hyaluronan | 3.2.1.36 | [ |
| LHyal | Hirudo nipponia | Hyaluronan | 3.2.1.36 | [ |
| SguS | Scutellaria baicalensis | Baicalein-7-O-β-D-glucuronide-conjugated | 3.2.1.167 | [ |
| AnGlcAase | Aspergillus niger | Arabinogalactan-proteins | 3.2.1.- | [ |
| TpGUS79A | Talaromyces pinophilus Li-93 | Glycyrrhizinate | 3.2.1.- | [ |
| Glucuronidase 1 | Prunus dulcis | Almond | 3.2.1.- | [ |
| Heparanase precursor | Bos taurus | Heparan sulfate | 3.2.1.- | [ |
| Hypothetical protein TrVGV298_008121 | Trichoderma virens | - | 3.2.1.- | [ |
表2 GH79家族不同功能糖苷水解酶
Table 2 Different functional glycoside hydrolases of the GH79 family
| 蛋白名称 Protein name | 来源 Organism | 底物 Substrate | EC编号 EC# | 参考文献 Reference |
|---|---|---|---|---|
| GlcA79A | Acidobacterium capsulatum | β-D-glucuronicacid | 3.2.1.31 | [ |
| AtGUS2 | Arabidopsis thaliana | pNPGlcA | 3.2.1.31 | [ |
| Nc6GAL | Neurospora crassa | Arabinogalactan-proteins | 3.2.1.31 | [ |
| FobglcA | Fusarium oxysporum | Gum arabic | 3.2.1.31 | [ |
| BpHep | Burkholderia pseudomallei | Heparan sulfate | 3.2.1.166 | [ |
| HpsE | Rattus norvegicus | Heparan sulfate | 3.2.1.166 | [ |
| Hpse1 | Danio rerio | Heparan sulfate glycosaminoglycans | 3.2.1.166 | [ |
| Hpa1 | Homo sapiens | Heparan sulfate | 3.2.1.166 | [ |
| Heparanase | Gallus gallus | Heparan sulfate | 3.2.1.166 | [ |
| Heparanase | Homo sapiens | Heparan sulfate | 3.2.1.166 | [ |
| Heparanase | Sus scrofa | Heparan sulfate | 3.2.1.166 | [ |
| Hpa | Mus musculus | Heparan sulfate | 3.2.1.166 | [ |
| Hyaluronidase | Hirudo nipponia | Hyaluronan | 3.2.1.36 | [ |
| LHyal | Hirudo nipponia | Hyaluronan | 3.2.1.36 | [ |
| SguS | Scutellaria baicalensis | Baicalein-7-O-β-D-glucuronide-conjugated | 3.2.1.167 | [ |
| AnGlcAase | Aspergillus niger | Arabinogalactan-proteins | 3.2.1.- | [ |
| TpGUS79A | Talaromyces pinophilus Li-93 | Glycyrrhizinate | 3.2.1.- | [ |
| Glucuronidase 1 | Prunus dulcis | Almond | 3.2.1.- | [ |
| Heparanase precursor | Bos taurus | Heparan sulfate | 3.2.1.- | [ |
| Hypothetical protein TrVGV298_008121 | Trichoderma virens | - | 3.2.1.- | [ |
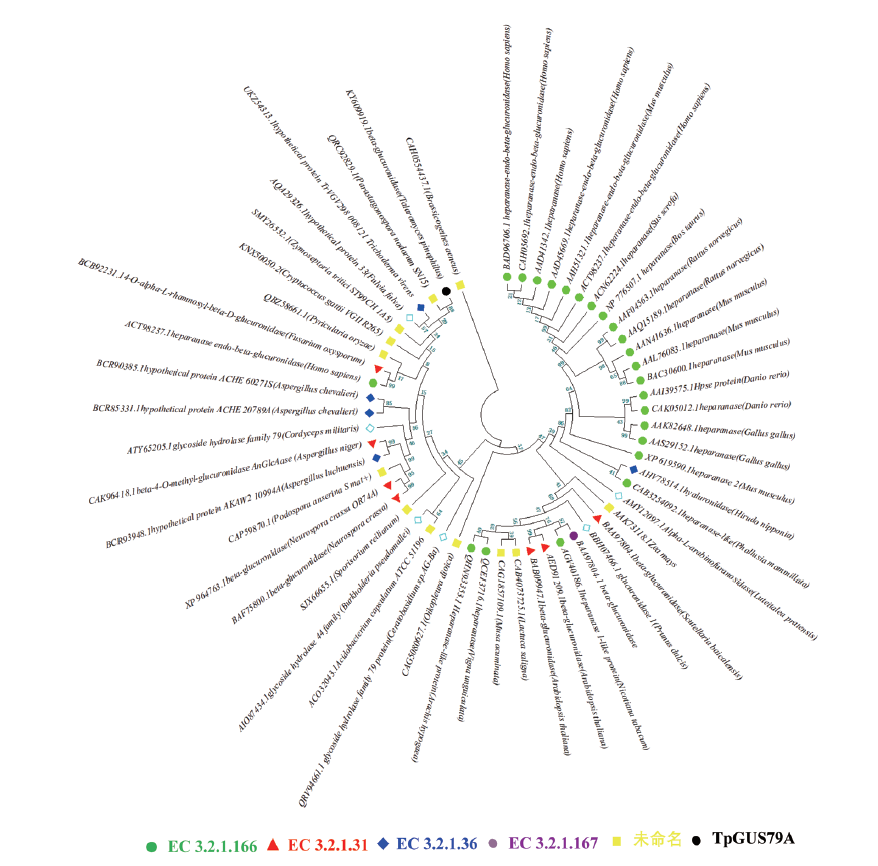
图2 GH79家族不同功能糖苷水解酶进化树 保守的催化基序突出显示,即GNE/ETNS
Fig. 2 Phylogenetic tree of glycoside hydrolases with differ-ent functions in the GH79 family conserved catalytic motif highlighted,i.e, GNE/ETNS

图6 GH79 家族蛋白结构已知的具有不同作用模式的 “exo-pocket”loop比较
Fig. 6 Comparison of “exo-pocket” loops with different modes of action for known structures of GH79 family proteins
| [1] |
OKUYAMA M. Function and structure studies of GH family 31 and 97 α-glycosidases[J]. Biosci Biotechnol Biochem, 2011, 75(12): 2269-2277.
doi: 10.1271/bbb.110610 URL |
| [2] |
Larrucea S, Butta N, Arias-Salgado EG, et al. Expression of podocalyxin enhances the adherence, migration, and intercellular communication of cells[J]. Exp Cell Res, 2008, 314(10): 2004-2015.
doi: 10.1016/j.yexcr.2008.03.009 pmid: 18456258 |
| [3] |
Singh RS, Singh RP, Kennedy JF. Recent insights in enzymatic synthesis of fructooligosaccharides from inulin[J]. Int J Biol Macromol, 2016, 85:565-572.
doi: 10.1016/j.ijbiomac.2016.01.026 pmid: 26791586 |
| [4] |
Guo LC, Katiyo W, Lu LS, et al. Glycyrrhetic acid 3-O-mono-β-d-glucuronide(GAMG):an innovative high-potency sweetener with improved biological activities[J]. Compr Rev Food Sci Food Saf, 2018, 17(4): 905-919.
doi: 10.1111/1541-4337.12353 URL |
| [5] | Lu DQ, Zhang SM, Wang J, et al. Adsorption separation of 3 beta-D-monoglucuronyl-18 beta-glycyrrhetinic acid from directional biotransformation products of glycyrrhizin[J]. African J Biotechnol, 2008, 7(22): 3995-4003. |
| [6] |
Tang WJ, Yang YA, Xu H, et al. Synthesis and discovery of 18α-GAMG as anticancer agent in vitro and in vivo via down expression of protein p65[J]. Sci Rep, 2014, 4:7106.
doi: 10.1038/srep07106 URL |
| [7] |
Mitsudome T, Xu J, Nagata Y, et al. Expression, purification, and characterization of endo-β-N-acetylglucosaminidase H using baculovirus-mediated silkworm protein expression system[J]. Appl Biochem Biotechnol, 2014, 172(8): 3978-3988.
doi: 10.1007/s12010-014-0814-5 pmid: 24599668 |
| [8] |
Komba S, Ito Y. Β-galactosidase-catalyzed intramolecular transglycosylation[J]. Tetrahedron Lett, 2001, 42(48): 8501-8505.
doi: 10.1016/S0040-4039(01)01826-3 URL |
| [9] |
Davies GJ, Williams SJ. Carbohydrate-active enzymes:sequences, shapes, contortions and cells[J]. Biochem Soc Trans, 2016, 44(1): 79-87.
doi: 10.1042/BST20150186 URL |
| [10] |
Niers TMH, Klerk CPW, DiNisio M, et al. Mechanisms of heparin induced anti-cancer activity in experimental cancer models[J]. Crit Rev Oncol Hematol, 2007, 61(3): 195-207.
doi: 10.1016/j.critrevonc.2006.07.007 URL |
| [11] |
Hamre AG, Strømnes AGS, Gustavsen D, et al. Treatment of recalcitrant crystalline polysaccharides with lytic polysaccharide monooxygenase relieves the need for glycoside hydrolase processivity[J]. Carbohydr Res, 2019, 473:66-71.
doi: 10.1016/j.carres.2019.01.001 URL |
| [12] |
Saito A, Wakao M, Deguchi H, et al. Towards the assembly of heparin and heparan sulfate oligosaccharide libraries:efficient synthesis of uronic acid and disaccharide building blocks[J]. Tetrahedron, 2010, 66(22): 3951-3962.
doi: 10.1016/j.tet.2010.03.077 URL |
| [13] |
Michikawa M, Ichinose H, Momma M, et al. Structural and biochemical characterization of glycoside hydrolase family 79 β-glucuronidase from Acidobacterium capsulatum[J]. J Biol Chem, 2012, 287(17): 14069-14077.
doi: 10.1074/jbc.M112.346288 pmid: 22367201 |
| [14] |
Bohlmann L, Tredwell GD, Yu X, et al. Functional and structural characterization of a heparanase[J]. Nat Chem Biol, 2015, 11(12): 955-957.
doi: 10.1038/nchembio.1956 pmid: 26565989 |
| [15] |
Konishi T, Kotake T, Soraya D, et al. Properties of family 79 beta-glucuronidases that hydrolyze beta-glucuronosyl and 4-O-methyl-beta-glucuronosyl residues of arabinogalactan-protein[J]. Carbohydr Res, 2008, 343(7): 1191-1201.
doi: 10.1016/j.carres.2008.03.004 URL |
| [16] |
Eudes A, Mouille G, Thévenin J, et al. Purification, cloning and functional characterization of an endogenous beta-glucuronidase in Arabidopsis thaliana[J]. Plant Cell Physiol, 2008, 49(9): 1331-1341.
doi: 10.1093/pcp/pcn108 URL |
| [17] |
Hulett MD, Freeman C, Hamdorf BJ, et al. Cloning of mammalian heparanase, an important enzyme in tumor invasion and metastasis[J]. Nat Med, 1999, 5(7): 803-809.
doi: 10.1038/10525 pmid: 10395326 |
| [18] |
Podyma-Inoue KA, Yokote H, Sakaguchi K, et al. Characterization of heparanase from a rat parathyroid cell line[J]. J Biol Chem, 2002, 277(36): 32459-32465.
doi: 10.1074/jbc.M203282200 pmid: 12077130 |
| [19] |
Toyoshima M, Nakajima M. Human heparanase. purification, characterization, cloning, and expression[J]. J Biol Chem, 1999, 274(34): 24153-24160.
doi: 10.1074/jbc.274.34.24153 pmid: 10446189 |
| [20] |
Masola V, Bellin G, Gambaro G, et al. Heparanase:a multitasking protein involved in extracellular matrix(ECM)remodeling and intracellular events[J]. Cells, 2018, 7(12): 236.
doi: 10.3390/cells7120236 URL |
| [21] |
Kondo T, Kichijo M, Nakaya M, et al. Biochemical and structural characterization of a novel 4-O-α-l-rhamnosyl-β-d-glucuronidase from Fusarium oxysporum[J]. FEBS J, 2021, 288(16): 4918-4938.
doi: 10.1111/febs.15795 URL |
| [22] |
Wei KH, Liu IH. Heparan sulfate glycosaminoglycans modulate migration and survival in zebrafish primordial germ cells[J]. Theriogenology, 2014, 81(9): 1275-1285. e1-2.
doi: 10.1016/j.theriogenology.2014.02.009 URL |
| [23] |
Goldshmidt O, Zcharia E, Aingorn H, et al. Expression pattern and secretion of human and chicken heparanase are determined by their signal peptide sequence[J]. J Biol Chem, 2001, 276(31): 29178-29187.
doi: 10.1074/jbc.M102462200 pmid: 11387326 |
| [24] |
Strausberg RL, Feingold EA, Grouse LH, et al. Generation and initial analysis of more than 15, 000 full-length human and mouse cDNA sequences[J]. Proc Natl Acad Sci USA, 2002, 99(26): 16899-16903.
doi: 10.1073/pnas.242603899 pmid: 12477932 |
| [25] |
Miles JR, Vallet JL, Freking BA, et al. Molecular cloning and characterisation of heparanase mRNA in the porcine placenta throughout gestation[J]. Reprod Fertil Dev, 2009, 21(6): 757-772.
doi: 10.1071/RD09041 URL |
| [26] |
Jin P, Kang Z, Zhang N, et al. High-yield novel leech hyaluronidase to expedite the preparation of specific hyaluronan oligomers[J]. Sci Rep, 2014, 4:4471.
doi: 10.1038/srep04471 pmid: 24667183 |
| [27] | Huang H, Hou XD, Xu RR, et al. Structure and cleavage pattern of a hyaluronate 3-glycanohydrolase in the glycoside hydrolase 79 family[J]. Carbohydr Polym, 2022, 277:118838. |
| [28] |
Sasaki K, Taura F, Shoyama Y, et al. Molecular characterization of a novel beta-glucuronidase from Scutellaria baicalensis Georgi[J]. J Biol Chem, 2000, 275(35): 27466-27472.
doi: 10.1074/jbc.M004674200 pmid: 10858442 |
| [29] | Xu YH, Feng XD, Jia JT, et al. A novel β-glucuronidase from Talaromyces pinophilus Li-93 precisely hydrolyzes glycyrrhizin into glycyrrhetinic acid 3- O-mono-β- d-glucuronide[J]. Appl Environ Microbiol, 2018, 84(19): e00755-18. |
| [30] |
Sánchez-Pérez R, Pavan S, Mazzeo R, et al. Mutation of a bHLH transcription factor allowed almond domestication[J]. Science, 2019, 364(6445): 1095-1098.
doi: 10.1126/science.aav8197 pmid: 31197015 |
| [31] |
Hambruch N, Kumstel S, Haeger JD, et al. Bovine placentomal heparanase and syndecan expression is related to placental maturation[J]. Placenta, 2017, 57:42-51.
doi: S0143-4004(17)30290-4 pmid: 28864018 |
| [32] | Li WC, Lin TC, Chen CL, et al. Complete genome sequences and genome-wide characterization of Trichoderma biocontrol agents provide new insights into their evolution and variation in genome organization, sexual development, and fungal-plant interactions[J]. Microbiol Spectr, 2021, 9(3): e0066321. |
| [33] |
Wu L, Jiang JB, Jin Y, et al. Activity-based probes for functional interrogation of retaining β-glucuronidases[J]. Nat Chem Biol, 2017, 13(8): 867-873.
doi: 10.1038/nchembio.2395 pmid: 28581485 |
| [34] |
Li QF, Jiang T, Liu R, et al. Tuning the pH profile of β-glucuronidase by rational site-directed mutagenesis for efficient transformation of glycyrrhizin[J]. Appl Microbiol Biotechnol, 2019, 103(12): 4813-4823.
doi: 10.1007/s00253-019-09790-3 pmid: 31055652 |
| [35] |
Vlodavsky I, Ilan N, Naggi A, et al. Heparanase:structure, biological functions, and inhibition by heparin-derived mimetics of heparan sulfate[J]. Curr Pharm Des, 2007, 13(20): 2057-2073.
doi: 10.2174/138161207781039742 URL |
| [36] |
Wang F, Wan A, Rodrigues B. The function of heparanase in diabetes and its complications[J]. Can J Diabetes, 2013, 37(5): 332-338.
doi: 10.1016/j.jcjd.2013.05.008 pmid: 24500561 |
| [37] |
Kuroyama H, Tsutsui N, Hashimoto Y, et al. Purification and characterization of a beta-glucuronidase from Aspergillus niger[J]. Carbohydr Res, 2001, 333(1): 27-39.
doi: 10.1016/S0008-6215(01)00114-8 URL |
| [38] | Harty DWS, Mayo JA, Cook SL, et al. Environmental regulation of glycosidase and peptidase production by Streptococcus gordonii FSS2[J]. Microbiology(Reading), 2000, 146(Pt 8): 1923-1931. |
| [39] |
Imaki H, Tomoyasu T, Yamamoto N, et al. Identification and characterization of a novel secreted glycosidase with multiple glycosidase activities in Streptococcus intermedius[J]. J Bacteriol, 2014, 196(15): 2817-2826.
doi: 10.1128/JB.01727-14 URL |
| [40] | Tomoyasu T, Yamasaki T, Chiba S, et al. Positive- and negative-control pathways by blood components for intermedilysin production in Streptococcus intermedius[J]. Infect Immun, 2017, 85(9): e00379-e00317. |
| [41] |
Fratianni F, Ombra MN, D'Acierno A, et al. Polyphenols content and in vitro α-glycosidase activity of different Italian monofloral honeys, and their effect on selected pathogenic and probiotic bacteria[J]. Microorganisms, 2021, 9(8): 1694.
doi: 10.3390/microorganisms9081694 URL |
| [42] | Weber BA, Klein JR, Henrich B. Expression of the phospho-beta-glycosidase ArbZ from Lactobacillus delbrueckii subsp. lactis in Lactobacillus helveticus:substrate induction and catabolite repression[J]. Microbiology(Reading), 2000, 146(Pt 8): 1941-1948. |
| [43] |
Chang CF, Ho CW, Wu CY, et al. Discovery of picomolar slow tight-binding inhibitors of alpha-fucosidase[J]. Chem Biol, 2004, 11(9): 1301-1306.
doi: 10.1016/j.chembiol.2004.07.009 URL |
| [44] |
Liu L, Tharmalingam T, Maischberger E, et al. A HPLC-based glycoanalytical protocol allows the use of natural O-glycans derived from glycoproteins as substrates for glycosidase discovery from microbial culture[J]. Glycoconj J, 2013, 30(8): 791-800.
doi: 10.1007/s10719-013-9483-9 URL |
| [45] |
Zhang LW, Qiu HL, Yuan S, et al. Revelation of mechanism for aqueous saponins content decrease during storage of Dioscorea zingiberensis C. H. Wright tubers:an essential prerequisite to ensure clean production of diosgenin[J]. Ind Crops Prod, 2018, 125:178-185.
doi: 10.1016/j.indcrop.2018.09.003 URL |
| [46] |
Brooks JF II, Gyllborg MC, Cronin DC, et al. Global discovery of colonization determinants in the squid symbiont Vibrio fischeri[J]. PNAS, 2014, 111(48): 17284-17289.
doi: 10.1073/pnas.1415957111 pmid: 25404340 |
| [47] | Zhang BQ, Zhang N, Zhang QQ, et al. Transcriptome profiles of Sporisorium reilianum during the early infection of resistant and susceptible maize isogenic lines[J]. J Fungi(Basel), 2021, 7(2): 150. |
| [48] |
Novakazi F, Göransson M, Stefánsson TS, et al. Virulence of Icelandic Pyrenophora teres f. teres populations and resistance of Icelandic spring barley lines[J]. J Plant Pathol, 2022, 104(1): 205-213.
doi: 10.1007/s42161-021-00972-5 URL |
| [49] |
Hall BG. Building phylogenetic trees from molecular data with MEGA[J]. Mol Biol Evol, 2013, 30(5): 1229-1235.
doi: 10.1093/molbev/mst012 pmid: 23486614 |
| [50] |
Newman L, Duffus ALJ, Lee C. Using the free program MEGA to build phylogenetic trees from molecular data[J]. Am Biol Teach, 2016, 78(7): 608-612.
doi: 10.1525/abt.2016.78.7.608 URL |
| [51] |
Vinader V, Haji-Abdullahi MH, Patterson LH, et al. Synthesis of a pseudo-disaccharide library and its application to the characterisation of the heparanase catalytic site[J]. PLoS One, 2013, 8(11): e82111.
doi: 10.1371/journal.pone.0082111 URL |
| [52] |
Okuyama M, Yoshida T, Hondoh H, et al. Catalytic role of the calcium ion in GH97 inverting glycoside hydrolase[J]. FEBS Lett, 2014, 588(17): 3213-3217.
doi: 10.1016/j.febslet.2014.07.002 pmid: 25017438 |
| [53] |
Qin Z, Yang SQ, Zhao LM, et al. Catalytic mechanism of a novel glycoside hydrolase family 16 “elongating” β-transglycosylase[J]. J Biol Chem, 2017, 292(5): 1666-1678.
doi: 10.1074/jbc.M116.762419 URL |
| [54] |
Vuong TV, Wilson DB. Glycoside hydrolases:catalytic base/nucleophile diversity[J]. Biotechnol Bioeng, 2010, 107(2): 195-205.
doi: 10.1002/bit.22838 pmid: 20552664 |
| [55] |
Davies G, Henrissat B. Structures and mechanisms of glycosyl hydrolases[J]. Structure, 1995, 3(9): 853-859.
doi: 10.1016/S0969-2126(01)00220-9 pmid: 8535779 |
| [56] |
Wu L, Viola CM, Brzozowski AM, et al. Structural characterization of human heparanase reveals insights into substrate recognition[J]. Nat Struct Mol Biol, 2015, 22(12): 1016-1022.
doi: 10.1038/nsmb.3136 pmid: 26575439 |
| [1] | 陈明雨, 倪烜, 司友斌, 孙凯. 固定化真菌漆酶在环境有机污染修复中的应用研究进展[J]. 生物技术通报, 2021, 37(6): 244-258. |
| [2] | 张梦恬, 裴娟 ,李国 ,赵辉 ,陈建权 ,祝建波, 王爱英. 新疆石河子地区棉花黄萎病菌分离鉴定及其致病力分析[J]. 生物技术通报, 2018, 34(6): 73-78. |
| [3] | 姜平,额尔敦木图,高凤山. 荷包猪SLA-DRB基因cDNA的克隆及分子进化特征分析[J]. 生物技术通报, 2015, 31(12): 150-157. |
| [4] | 吕学泽;梁玉荣;贺云霞;张培君;龚玉梅;王宏俊;. 副鸡禽杆菌aroA基因克隆及序列分析[J]. , 2012, 0(08): 83-87. |
| [5] | 何广正;徐书景;马宁;刘东;鞠建松;. 丙氨酸脱氢酶研究概况[J]. , 2011, 0(12): 27-32. |
| [6] | 彭娟;李思光;罗丹丹;罗玉萍;. miR-124基因家族的分子进化与靶基因预测[J]. , 2011, 0(10): 167-178. |
| [7] | 王波;孙君社;翟玉盼;孙立伟;张秀清;. 植物Ⅲ型聚酮合酶基因家族的分子进化分析[J]. , 2011, 0(01): 83-89. |
| [8] | . 科学出版社新书[J]. , 2010, 0(06): 137-178. |
| [9] | 吴振芳;陈惠;曾民;吴琦;. 蛋白质新功能定向进化研究策略[J]. , 2009, 0(03): 11-15. |
| [10] | 王璋瑜;. 分子技术推动热带生物学研究[J]. , 1992, 0(06): 27-27. |
| [11] | 孙国凤;. 单一触点标记系统树计算机软件[J]. , 1991, 0(07): 21-22. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||