








生物技术通报 ›› 2025, Vol. 41 ›› Issue (12): 82-94.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0500
田云1( ), 孔辰1, 杨冲2, 刘统高2, 高红瑞1, 李佳慧1, 马云1, 蔡蓓1(
), 孔辰1, 杨冲2, 刘统高2, 高红瑞1, 李佳慧1, 马云1, 蔡蓓1( )
)
收稿日期:2025-05-14
出版日期:2025-12-26
发布日期:2026-01-06
通讯作者:
蔡蓓,女,讲师,研究方向 :羊遗传育种及毛囊发育生物学;E-mail: caibei1115@163.com作者简介:田云,女,硕士研究生,研究方向 :动物遗传育种;E-mail: 2294928895@qq.com
基金资助:
TIAN Yun1( ), KONG Chen1, YANG Chong2, LIU Tong-gao2, GAO Hong-rui1, LI Jia-hui1, MA Yun1, CAI Bei1(
), KONG Chen1, YANG Chong2, LIU Tong-gao2, GAO Hong-rui1, LI Jia-hui1, MA Yun1, CAI Bei1( )
)
Received:2025-05-14
Published:2025-12-26
Online:2026-01-06
摘要:
畜牧业是农业的重要组成部分,其经济价值与畜禽表型性状(如产肉量、产蛋率等)密切相关。这些性状由复杂的遗传网络调控,并且受到环境因素的影响,导致挖掘决定这些性状的功能基因较为困难。遗传筛选技术的出现使基因型-表型关系的发现变得简单,但早期的遗传筛选技术通量低并且难以分析产生的大量数据致使效率低下。随着生物技术的迭代更新,遗传筛选技术进入高通量筛选时代,为解析基因型-表型关系提供了高效手段。传统的高通量筛选方法主要包括RNA干扰(RNAi)和cDNA过表达等,但RNAi易脱靶且无法完全抑制基因表达,cDNA文库构建成本高且可能引发细胞毒性。CRISPR/Cas系统的开发促使为遗传筛选提供了新的策略。该系统基于细菌的适应性免疫机制,通过向导RNA靶向DNA序列并利用Cas蛋白精准编辑基因,具有操作简便、特异性高和适用范围广的优势。CRISPR高通量筛选技术主要包括功能缺失型(CRISPRko)、功能获得型(CRISPRa/CRISPRi)、碱基编辑等多种模式,能够直接在DNA水平进行扰动,克服了传统方法的不足。本文介绍了遗传筛选技术的类型、特点以及在畜牧学中的应用,并展望了CRISPR高通量筛选技术在畜牧业育种中的巨大潜力。
田云, 孔辰, 杨冲, 刘统高, 高红瑞, 李佳慧, 马云, 蔡蓓. CRISPR高通量筛选技术及其在畜牧研究中的应用[J]. 生物技术通报, 2025, 41(12): 82-94.
TIAN Yun, KONG Chen, YANG Chong, LIU Tong-gao, GAO Hong-rui, LI Jia-hui, MA Yun, CAI Bei. CRISPR-based High-throughput Screening Technology and Its Applications in Livestock Research[J]. Biotechnology Bulletin, 2025, 41(12): 82-94.
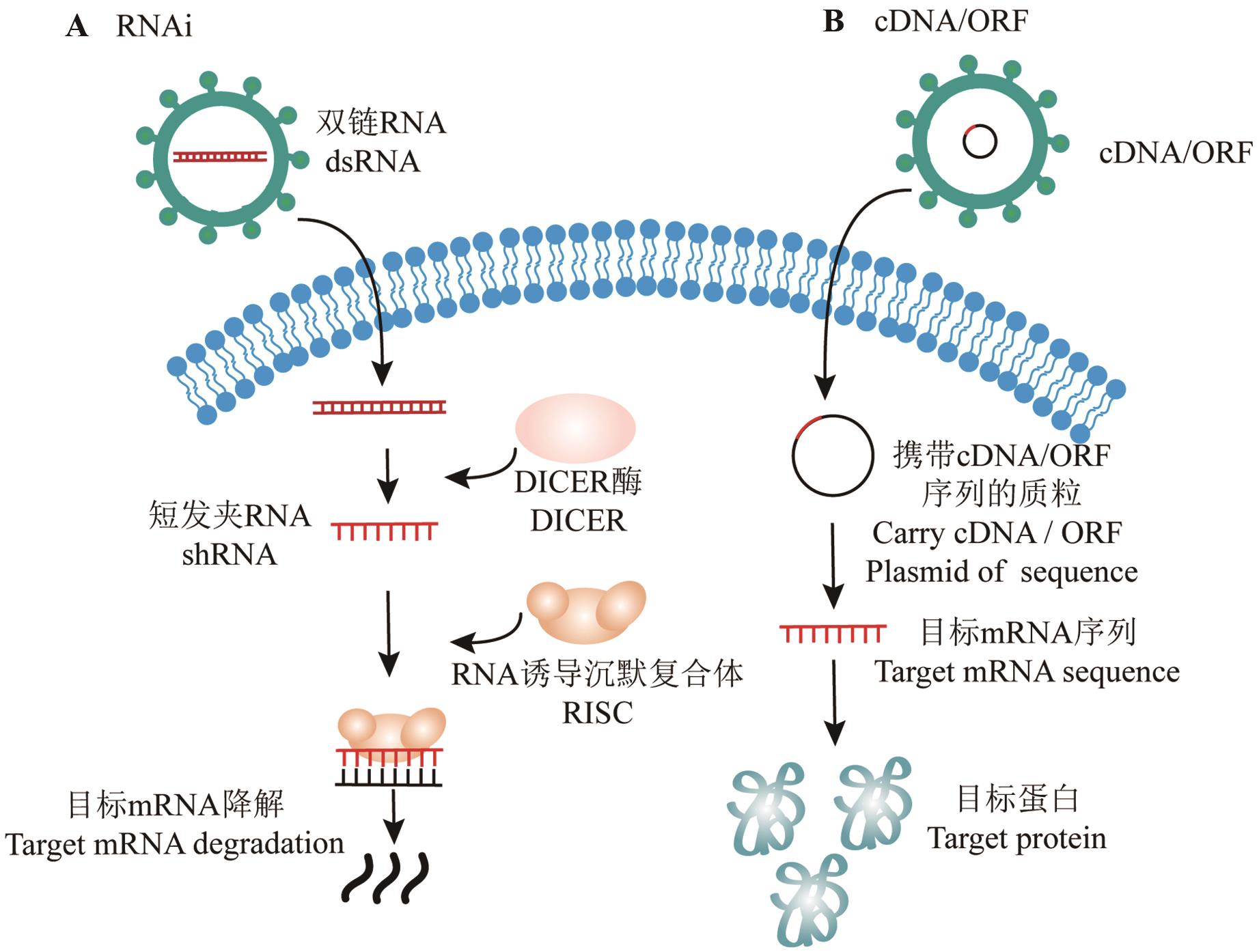
图1 RNAi及cDNA/ORF过表达示意图A:RNAi干扰过程;B:cDNA/ORF过表达过程
Fig. 1 Schematic diagram of RNAi and cDNA/ORF overexpressionA: RNAi interference process; B: cDNA/ORF overexpression process
特征 Feature | 全基因组sgRNA文库 Genome-wide sgRNA library | 单一基因的sgRNA Single gene sgRNA |
|---|---|---|
| 目标范围 | 全基因组范围(包含数万的基因) | 单一或少数基因 |
| sgRNA数量级 | 数万或数十万条sgRNA | 4-10条sgRNA |
| 设计深度 | 每个基因4-6条sgRNA | 每个基因可设计数10条sgRNA |
| 覆盖范围 | 广度优先(尽可能的覆盖多的基因) | 深度优先(聚焦特定基因) |
| 应用场景 | 未知基因的挖掘 | 基因功能的精细验证 |
表1 全基因组sgRNA文库与单一基因sgRNA设计的比较
Table 1 Comparison of genome-wide sgRNA library and single gene sgRNA design
特征 Feature | 全基因组sgRNA文库 Genome-wide sgRNA library | 单一基因的sgRNA Single gene sgRNA |
|---|---|---|
| 目标范围 | 全基因组范围(包含数万的基因) | 单一或少数基因 |
| sgRNA数量级 | 数万或数十万条sgRNA | 4-10条sgRNA |
| 设计深度 | 每个基因4-6条sgRNA | 每个基因可设计数10条sgRNA |
| 覆盖范围 | 广度优先(尽可能的覆盖多的基因) | 深度优先(聚焦特定基因) |
| 应用场景 | 未知基因的挖掘 | 基因功能的精细验证 |
sgRNA文库类型 Type of sgRNA library | sgRNA位点要求 Requirements of sgRNA site | 位点要求的原因及目的 Reason and purpose of site requirements |
|---|---|---|
| 全基因组文库 | 一般位于编码序列的前25% | 确保蛋白功能的完全丧失 |
| 激活文库 | TSS上游200 bp左右 | 确保激活结构域能够靶向TSS位点,以实现基因表达的增强 |
| 干扰文库 | TSS下游+25-75 nts左右 | 确保转录抑制结构域能够靶向TSS位点,以实现沉默基因 |
| 敲除文库 | 选择编码区前50%(靠近ATG),避开内含子剪切位点 | 确保蛋白功能的丧失,避免截留的蛋白残基仍存有功能 |
| STOP文库 | TSS下游100 bp左右 | 确保终止密码子的提前引入,以丧失沉默基因。 |
表2 不同CRISPR筛选技术中sgRNA文库的设计要求
Table 2 sgRNA site design requirements
sgRNA文库类型 Type of sgRNA library | sgRNA位点要求 Requirements of sgRNA site | 位点要求的原因及目的 Reason and purpose of site requirements |
|---|---|---|
| 全基因组文库 | 一般位于编码序列的前25% | 确保蛋白功能的完全丧失 |
| 激活文库 | TSS上游200 bp左右 | 确保激活结构域能够靶向TSS位点,以实现基因表达的增强 |
| 干扰文库 | TSS下游+25-75 nts左右 | 确保转录抑制结构域能够靶向TSS位点,以实现沉默基因 |
| 敲除文库 | 选择编码区前50%(靠近ATG),避开内含子剪切位点 | 确保蛋白功能的丧失,避免截留的蛋白残基仍存有功能 |
| STOP文库 | TSS下游100 bp左右 | 确保终止密码子的提前引入,以丧失沉默基因。 |
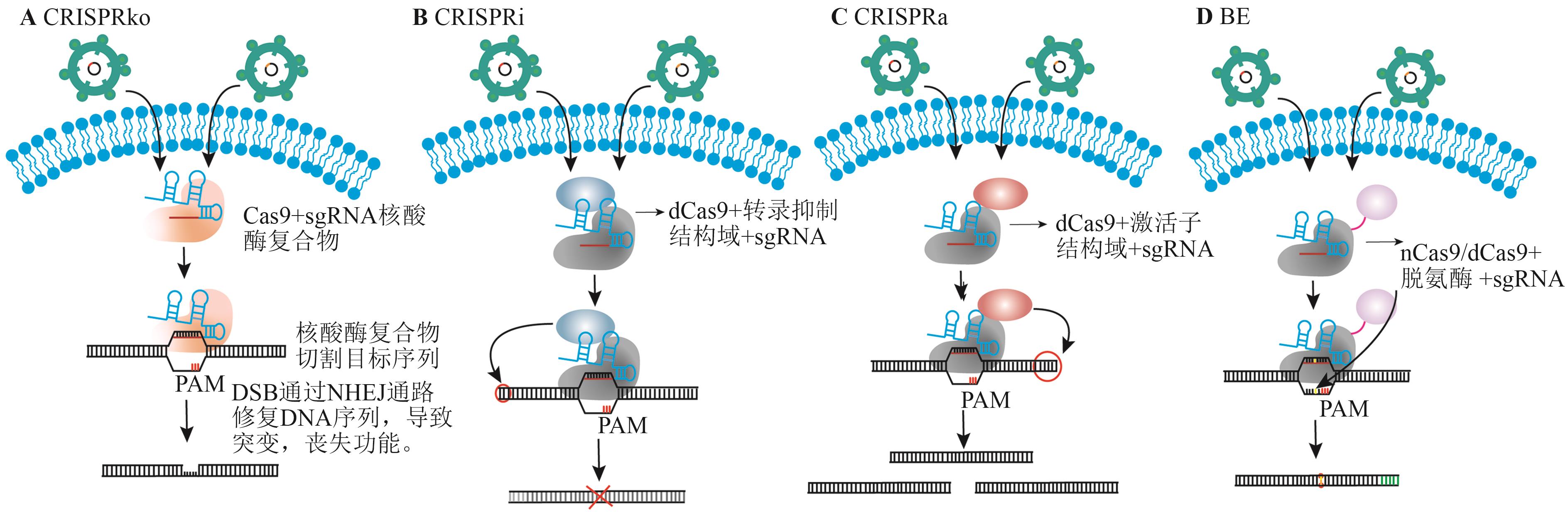
图3 不同类型CRISPR筛选工具示意图A:CRISPRko筛选示意图;B:CRISPRi筛选示意图;C:CRISPRa筛选示意图;D:BE筛选示意图
Fig. 3 Schematic diagrams of different types of CRISPR screeening toolsA: CRISPRko screening diagram. B: CRISPRi screening diagram. C: CRISPRa screening diagram. D: BE screening diagram
筛选工具 Screening tool | 扰动方式 Disturbed mode | 优点 Merit | 局限性 Limitation |
|---|---|---|---|
| RNAi | 通过降解mRNA抑制基因表达 | 简化了筛选方法,降低筛选文库的构建成本 | 无法针对基因组基因进行筛选;基因沉默不完全,只能在细胞质中行使功能 |
| cDNA/ORF | 提高目标蛋白的表达 | 第一个功能获得型筛选方法,是发现冗余基因功能的方法 | 文库构建困难,并且成本高;某些基因过表达后会导致细胞死亡 |
| CRISPRko | 利用gRNA引导Cas9酶切割目标DNA序列,通过细胞的DNA修复机制导致目标基因敲除 | 可以最大程度地避免基因的假沉默;可实现非编码基因的筛选 | 存在脱靶效应;断裂DNA双链引起的DNA毒性;无法研究基因恢复后的情况 |
| CRISPRi | 通过阻断转录起始实现基因沉默 | 不引起DNA双链断裂,造成的损伤是可逆的,脱靶效应小,特异性强 | 对基因结构有依赖性 |
| CRISPRa | 激活转录因子,增强基因转录活性 | 不引起DNA双链断裂;造成的损伤是可逆的 | 不能针对特定转录本激活;激活效率受限 |
| BE | 将脱氨酶与dCas9结合,实现碱基的转换,从而实现起始密码子和终止密码子的沉默和提前终止,以实现基因的沉默 | 不引起DNA双链断裂的情况下,实现特定碱基的转换 | 编辑范围有限;在编辑窗口中发生旁观者编辑 |
表3 筛选工具
Table 3 Screening tool
筛选工具 Screening tool | 扰动方式 Disturbed mode | 优点 Merit | 局限性 Limitation |
|---|---|---|---|
| RNAi | 通过降解mRNA抑制基因表达 | 简化了筛选方法,降低筛选文库的构建成本 | 无法针对基因组基因进行筛选;基因沉默不完全,只能在细胞质中行使功能 |
| cDNA/ORF | 提高目标蛋白的表达 | 第一个功能获得型筛选方法,是发现冗余基因功能的方法 | 文库构建困难,并且成本高;某些基因过表达后会导致细胞死亡 |
| CRISPRko | 利用gRNA引导Cas9酶切割目标DNA序列,通过细胞的DNA修复机制导致目标基因敲除 | 可以最大程度地避免基因的假沉默;可实现非编码基因的筛选 | 存在脱靶效应;断裂DNA双链引起的DNA毒性;无法研究基因恢复后的情况 |
| CRISPRi | 通过阻断转录起始实现基因沉默 | 不引起DNA双链断裂,造成的损伤是可逆的,脱靶效应小,特异性强 | 对基因结构有依赖性 |
| CRISPRa | 激活转录因子,增强基因转录活性 | 不引起DNA双链断裂;造成的损伤是可逆的 | 不能针对特定转录本激活;激活效率受限 |
| BE | 将脱氨酶与dCas9结合,实现碱基的转换,从而实现起始密码子和终止密码子的沉默和提前终止,以实现基因的沉默 | 不引起DNA双链断裂的情况下,实现特定碱基的转换 | 编辑范围有限;在编辑窗口中发生旁观者编辑 |
| [1] | 王文月, 米晓钰, 孙康泰, 等. 畜禽重要性状遗传调控机制与分子设计育种 [J]. 中国农业科技导报, 2022, 24(12): 39-47. |
| Wang WY, Mi XY, Sun KT, et al. Genetic regulation mechanisms of important traits and molecular design breeding in livestock and poultry [J]. J Agric Sci Technol, 2022, 24(12): 39-47. | |
| [2] | Martiny JBH, Jones SE, Lennon JT, et al. Microbiomes in light of traits: a phylogenetic perspective [J]. Science, 2015, 350(6261): aac9323. |
| [3] | Yang CT, Lei Y, Ren TL, et al. The current situation and development prospect of whole-genome screening [J]. Int J Mol Sci, 2024, 25(1): 658. |
| [4] | Yang WN, Feng H, Zhang XH, et al. Crop phenomics and high-throughput phenotyping: past decades, current challenges, and future perspectives [J]. Mol Plant, 2020, 13(2): 187-214. |
| [5] | Hu SJ, Gan ML, Wei ZA, et al. Identification of host factors for livestock and poultry viruses: genome-wide screening technology based on the CRISPR system [J]. Front Microbiol, 2024, 15: 1498641. |
| [6] | Unniyampurath U, Pilankatta R, Krishnan MN. RNA interference in the age of CRISPR: will CRISPR interfere with RNAi? [J]. Int J Mol Sci, 2016, 17(3): 291. |
| [7] | Vercauteren S, Fiesack S, Maroc L, et al. The rise and future of CRISPR-based approaches for high-throughput genomics [J]. FEMS Microbiol Rev, 2024, 48(5): fuae020. |
| [8] | St Johnston D. The art and design of genetic screens: Drosophila melanogaster [J]. Nat Rev Genet, 2002, 3(3): 176-188. |
| [9] | Jorgensen EM, Mango SE. The art and design of genetic screens: Caenorhabditis elegans [J]. Nat Rev Genet, 2002, 3(5): 356-369. |
| [10] | Forsburg SL. The art and design of genetic screens: yeast [J]. Nat Rev Genet, 2001, 2(9): 659-668. |
| [11] | Shalem O, Sanjana NE, Zhang F. High-throughput functional genomics using CRISPR-Cas9 [J]. Nat Rev Genet, 2015, 16(5): 299-311. |
| [12] | Josi C, Bürki S, Vidal S, et al. Large-scale analysis of the Mycoplasma bovis genome identified non-essential, adhesion- and virulence-related genes [J]. Front Microbiol, 2019, 10: 2085. |
| [13] | Hai T, Cao C, Shang H, et al. Pilot study of large-scale production of mutant pigs by ENU mutagenesis [J]. eLife, 2017, 6: e26248. |
| [14] | Tanner AR, Kennedy VC, Lynch CS, et al. In vivo investigation of ruminant placenta function and physiology—a review [J]. J Anim Sci, 2022, 100(6): skac045. |
| [15] | Qian Q, Xu RY, Wang YP, et al. The NS4A protein of classical swine fever virus suppresses RNA silencing in mammalian cells [J]. J Virol, 2022, 96(15): e01874-21. |
| [16] | Wang DP, Xiu JB, Zhao JY, et al. miR-AB, a miRNA-based shRNA viral toolkit for multicolor-barcoded multiplex RNAi at a single-cell level [J]. EMBO Rep, 2022, 23(4): e53691. |
| [17] | Li Y, Mensah EO, Fordjour E, et al. Recent advances in high-throughput metabolic engineering: Generation of oligonucleotide-mediated genetic libraries [J]. Biotechnol Adv, 2022, 59: 107970. |
| [18] | Boutros M, Ahringer J. The art and design of genetic screens: RNA interference [J]. Nat Rev Genet, 2008, 9(7): 554-566. |
| [19] | Li D, Yang JH, Malik V, et al. An RNAi screen of RNA helicases identifies eIF4A3 as a regulator of embryonic stem cell identity [J]. Nucleic Acids Res, 2022, 50(21): 12462-12479. |
| [20] | Žemaitis K, Subramaniam A, Galeev R, et al. RNAi screen identifies MTA1 as an epigenetic modifier of differentiation commitment in human HSPCs [J]. Exp Hematol, 2022, 115: 20-29. |
| [21] | Wu HY, Lv PY, et al. Genetic screen identified PRMT5 as a neuroprotection target against cerebral ischemia [J]. eLife, 2024, 12: RP89754. |
| [22] | Qi SH, Sun C, Wang J, et al. Identification of NECTIN1 as a novel restriction factor for flavivirus infection [J]. mBio, 2024, 15(12) |
| [23] | Li Q, Wang YH, Hu X, et al. Transcriptional states and chromatin accessibility during bovine myoblasts proliferation and myogenic differentiation [J]. Cell Prolif, 2022, 55(5): e13219. |
| [24] | Wang Q, Zhang QW, Wang XY, et al. Yak FOXO1 and FOXO3 SNPs and association with production traits, and their promotes cells apoptosis via RNAi [J]. Gene, 2020, 743: 144592. |
| [25] | He JN, Liu QY, Yu SY, et al. Expression and functional analysis of the Follistatin-like 3 (FSTL3) gene in the sheep ovary during the oestrous cycle [J]. Reprod Domest Anim, 2021, 56(3): 427-436. |
| [26] | Booker M, Samsonova AA, Kwon Y, et al. False negative rates in Drosophila cell -based-based RNAi screens: a case study [J]. BMC Genom, 2011, 12(1): 50. |
| [27] | Zeng Y, Cullen BR. RNA interference in human cells is restricted to the cytoplasm [J]. RNA, 2002, 8(7): 855-860. |
| [28] | Grimm D. The dose can make the poison: lessons learned from adverse in vivo toxicities caused by RNAi overexpression [J]. Silence, 2011, 2(1): 8. |
| [29] | Frecot DI, Froehlich T, Rothbauer U. 30 years of nanobodies-an ongoing success story of small binders in biological research [J]. J Cell Sci, 2023, 136(21): jcs261395. |
| [30] | Sone M, Mitsuhashi N, Sugiura Y, et al. Identification of genes supporting cold resistance of mammalian cells: lessons from a hibernator [J]. Cell Death Dis, 2024, 15: 685. |
| [31] | Yamada Y, Miyamoto T, Higuchi S, et al. cDNA expression library screening revealed novel functional genes involved in clear cell carcinogenesis of the ovary in vitro [J]. J Obstet Gynaecol, 2021, 41(1): 100-105. |
| [32] | Zhang P, Kratz AS, Salama M, et al. Expression screening using a Medaka cDNA library identifies evolutionarily conserved regulators of the p53/Mdm2 pathway [J]. BMC Biotechnol, 2015, 15(1): 92. |
| [33] | Mita M. Relaxin-like gonad-stimulating peptides in Asteroidea [J]. Biomolecules, 2023, 13(5): 781. |
| [34] | Kannan S, Miyamoto M, Zhu RJ, et al. Trajectory reconstruction identifies dysregulation of perinatal maturation programs in pluripotent stem cell-derived cardiomyocytes [J]. Cell Rep, 2023, 42(4): 112330. |
| [35] | Ng AHM, Khoshakhlagh P, Rojo Arias JE, et al. A comprehensive library of human transcription factors for cell fate engineering [J]. Nat Biotechnol, 2021, 39(4): 510-519. |
| [36] | Legut M, Gajic Z, Guarino M, et al. A genome-scale screen for synthetic drivers of T cell proliferation [J]. Nature, 2022, 603(7902): 728-735. |
| [37] | 姜光飞, 何维琪, 余蕊, 等. 猪静脉血管内皮细胞cDNA文库构建及与猪非典型瘟病毒云南株关键变异基因互作蛋白的筛选 [J]. 动物医学进展, 2023, 44(7): 1-9. |
| Jiang GF, He WQ, Yu R, et al. Construction of SVEC cDNA library and screening of interacting proteins with key variant gene of APPV-YN [J]. Prog Vet Med, 2023, 44(7): 1-9. | |
| [38] | Wei P, Xue W, Zhao Y, et al. CRISPR-based modular assembly of a UAS-cDNA/ORF plasmid library for more than 5500 Drosophila genes conserved in humans [J]. Genome Res, 2020, 30(1): 95-106. |
| [39] | Tyumentseva M, Tyumentsev A, Akimkin V. CRISPR/Cas9 landscape: current state and future perspectives [J]. Int J Mol Sci, 2023, 24(22): 16077. |
| [40] | Hwang S, Maxwell KL. Diverse mechanisms of CRISPR-Cas9 inhibition by type II anti-CRISPR proteins [J]. J Mol Biol, 2023, 435(7): 168041. |
| [41] | Khoshandam M, Soltaninejad H, Ali Hamidieh A, et al. CRISPR, CAR-T, and NK: Current applications and future perspectives [J]. Genes Dis, 2024, 11(4): 101121. |
| [42] | Makarova KS, Wolf YI, Iranzo J, et al. Evolutionary classification of CRISPR-Cas systems: a burst of class 2 and derived variants [J]. Nat Rev Microbiol, 2020, 18(2): 67-83. |
| [43] | Ishibashi R, Maki R, Toyoshima F. Gene targeting in adult organs using in vivo cleavable donor plasmids for CRISPR-Cas9 and CRISPR-Cas12a [J]. Sci Rep, 2024, 14: 7615. |
| [44] | Zhao LX, Qiu MY, Li XJ, et al. CRISPR-Cas13a system: a novel tool for molecular diagnostics [J]. Front Microbiol, 2022, 13: 1060947. |
| [45] | Rees HA, Liu DR. Base editing: precision chemistry on the genome and transcriptome of living cells [J]. Nat Rev Genet, 2018, 19(12): 770-788. |
| [46] | Doman JL, Raguram A, Newby GA, et al. Evaluation and minimization of Cas9-independent off-target DNA editing by cytosine base editors [J]. Nat Biotechnol, 2020, 38(5): 620-628. |
| [47] | Huang TP, Newby GA, Liu DR. Precision genome editing using cytosine and adenine base editors in mammalian cells [J]. Nat Protoc, 2021, 16(2): 1089-1128. |
| [48] | Agrotis A, Ketteler R. A new age in functional genomics using CRISPR/Cas9 in arrayed library screening [J]. Front Genet, 2015, 6: 300. |
| [49] | Shi YJ, Katdare KA, Kim H, et al. An arrayed CRISPR knockout screen identifies genetic regulators of GLUT1 expression [J]. Sci Rep, 2023, 13: 21038. |
| [50] | Barry T, Mason K, Roeder K, et al. Robust differential expression testing for single-cell CRISPR screens at low multiplicity of infection [J]. Genome Biol, 2024, 25(1): 124. |
| [51] | Shang WJ, Wang F, Fan GF, et al. Key elements for designing and performing a CRISPR/Cas9-based genetic screen [J]. J Genet Genom, 2017, 44(9): 439-449. |
| [52] | Otten ABC, Sun BK. Research techniques made simple: CRISPR genetic screens [J]. J Investig Dermatol, 2020, 140(4): 723-728.e1. |
| [53] | Liang YH, Xie JK, Zhang QJ, et al. AGBE: a dual deaminase-mediated base editor by fusing CGBE with ABE for creating a saturated mutant population with multiple editing patterns [J]. Nucleic Acids Res, 2022, 50(9): 5384-5399. |
| [54] | Zhou YX, Zhu SY, Cai CZ, et al. High-throughput screening of a CRISPR/Cas9 library for functional genomics in human cells [J]. Nature, 2014, 509(7501): 487-491. |
| [55] | Sanjana NE, Shalem O, Zhang F. Improved vectors and genome-wide libraries for CRISPR screening [J]. Nat Meth, 2014, 11(8): 783-784. |
| [56] | Wang XJ, Liu ZW, Li GL, et al. Efficient gene silencing by adenine base editor-mediated start codon mutation [J]. Mol Ther, 2020, 28(2): 431-440. |
| [57] | Liang YJ, Yao XX, Han JX, et al. Establishment of a CRISPR-based lentiviral activation library for transcription factor screening in porcine cells [J]. Animals, 2025, 15(1): 19. |
| [58] | Doench JG, Fusi N, Sullender M, et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR-Cas9 [J]. Nat Biotechnol, 2016, 34(2): 184-191. |
| [59] | Joung J, Konermann S, Gootenberg JS, et al. Author Correction: Genome-scale CRISPR-Cas9 knockout and transcriptional activation screening [J]. Nat Protoc, 2019, 14(7): 2259. |
| [60] | Shalem O, Sanjana NE, Hartenian E, et al. Genome-scale CRISPR-Cas9 knockout screening in human cells [J]. Science, 2014, 343(6166): 84-87. |
| [61] | Evers B, Jastrzebski K, Heijmans JPM, et al. CRISPR knockout screening outperforms shRNA and CRISPRi in identifying essential genes [J]. Nat Biotechnol, 2016, 34(6): 631-633. |
| [62] | Morgens DW, Deans RM, Li A, et al. Systematic comparison of CRISPR/Cas9 and RNAi screens for essential genes [J]. Nat Biotechnol, 2016, 34(6): 634-636. |
| [63] | Merk DJ, Paul L, Tsiami F, et al. CRISPR-Cas9 screens reveal common essential miRNAs in human cancer cell lines [J]. Genome Med, 2024, 16(1): 82. |
| [64] | Su GN, Liu J, Duan CR, et al. Enteric coronavirus PDCoV evokes a non-Warburg effect by hijacking pyruvic acid as a metabolic hub [J]. Redox Biol, 2024, 71: 103112. |
| [65] | Ma NN, Zhang MJ, Zhou JR, et al. Genome-wide CRISPR/Cas9 library screen identifies C16orf62 as a host dependency factor for porcine deltacoronavirus infection [J]. Emerg Microbes Infect, 2024, 13(1): 2400559. |
| [66] | Shen DD, Zhang GG, Weng XG, et al. A genome-wide CRISPR/Cas9 knockout screen identifies TMEM239 as an important host factor in facilitating African swine fever virus entry into early endosomes [J]. PLoS Pathog, 2024, 20(7): e1012256. |
| [67] | 李艳. 基于CRISPR/Cas9文库筛选绒山羊毛囊干细胞增殖必需基因 [D]. 杨凌: 西北农林科技大学, 2024. |
| Li Y. Screening of essential genes for the proliferation of Cashmere goat hair follicle stem cells based on CRISPR/Cas9 library [D]. Yangling: Northwest A & F University, 2024. | |
| [68] | Liu ZX, Dai LL, Sun TH, et al. Massively parallel CRISPR-Cas9 knockout screening in sheep granulosa cells for FSH response genes [J]. Animals, 2024, 14(6): 898. |
| [69] | Gilbert LA, Larson MH, Morsut L, et al. CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes [J]. Cell, 2013, 154(2): 442-451. |
| [70] | Kampmann M. CRISPRi and CRISPRa screens in mammalian cells for precision biology and medicine [J]. ACS Chem Biol, 2018, 13(2): 406-416. |
| [71] | Xue W, Jian WG, Meng YY, et al. Knockdown of SETD2 promotes erastin-induced ferroptosis in ccRCC [J]. Cell Death Dis, 2023, 14: 539. |
| [72] | Wagner RT, Hlady RA, Pan XY, et al. SETD2 loss-of-function uniquely sensitizes cells to epigenetic targeting of NSD1-directed H3K36 methylation [J]. Genome Biol, 2025, 26(1): 22. |
| [73] | Li E, Benitez C, Boggess SC, et al. CRISPRi-based screens in iAssembloids to elucidate neuron-Glia interactions [J]. Neuron, 2025, 113(5): 701-718.e8. |
| [74] | Kaplan SJ, Wong W, Yan JL, et al. CRISPR screening uncovers a long-range enhancer for ONECUT1 in pancreatic differentiation and links a diabetes risk variant [J]. Cell Rep, 2024, 43(8): 114640. |
| [75] | Zhu XY, Luo H, Yu XR, et al. Genome-wide CRISPRi screening of key genes for recombinant protein expression in Bacillus subtilis [J]. Adv Sci, 2024, 11(33): e2404313. |
| [76] | Gilbert LA, Horlbeck MA, Adamson B, et al. Genome-scale CRISPR-mediated control of gene repression and activation [J]. Cell, 2014, 159(3): 647-661. |
| [77] | Schmidt R, Steinhart Z, Layeghi M, et al. CRISPR activation and interference screens decode stimulation responses in primary human T cells [J]. Science, 2022, 375(6580): eabj4008. |
| [78] | Kuscu C, Parlak M, Tufan T, et al. CRISPR-STOP gene silencing through base-editing-induced nonsense mutations [J]. Nat Meth, 2017, 14(7): 710-712. |
| [79] | Schmidt R, Ward CC, Dajani R, et al. Base-editing mutagenesis maps alleles to tune human T cell functions [J]. Nature, 2024, 625(7996): 805-812. |
| [80] | Xiao MS, Damodaran AP, Kumari B, et al. Genome-scale exon perturbation screens uncover exons critical for cell fitness [J]. Mol Cell, 2024, 84(13): 2553-2572.e19. |
| [81] | Cheng WS, Liu F, Ren ZJ, et al. Parallel functional assessment of m6A sites in human endodermal differentiation with base editor screens [J]. Nat Commun, 2022, 13: 478. |
| [82] | Li YZ, Xu T, Ma HZ, et al. Functional profiling of serine, threonine and tyrosine sites [J]. Nat Chem Biol, 2025, 21(4): 532-543. |
| [83] | Hsu PD, Scott DA, Weinstein JA, et al. DNA targeting specificity of RNA-guided Cas9 nucleases [J]. Nat Biotechnol, 2013, 31(9): 827-832. |
| [84] | Gaudelli NM, Komor AC, Rees HA, et al. Programmable base editing of A·T to G·C in genomic DNA without DNA cleavage [J]. Nature, 2017, 551(7681): 464-471. |
| [85] | Chen SY, Xie WH, Liu ZQ, et al. CRISPR start-loss: a novel and practical alternative for gene silencing through base-editing-induced start codon mutations [J]. Mol Ther Nucleic Acids, 2020, 21: 1062-1073. |
| [86] | Fang L, Wang W, Li GP, et al. CIGAR-seq, a CRISPR/Cas-based method for unbiased screening of novel mRNA modification regulators [J]. Mol Syst Biol, 2020, 16(11): e10025. |
| [87] | Niu Y, Ferreira Azevedo CA, LI X, et al. Multiparametric and accurate functional analysis of genetic sequence variants using CRISPR-Select[J]. Nat Genet, 2022, 54(12): 1983-1993. |
| [88] | Dhainaut M, Rose SA, Akturk G, et al. Spatial CRISPR genomics identifies regulators of the tumor microenvironment [J]. Cell, 2022, 185(7): 1223-1239.e20. |
| [1] | 李思经. 激素禁令无事实依据[J]. , 1996, 0(06): 32-32. |
| [2] | 李馨;杨隽. PCR技术及其在畜牧业中的应用[J]. , 1996, 0(05): 1-4. |
| [3] | 孙国凤;. 日本开辟利用畜牧业副产品的产业[J]. , 1988, 0(04): 23-24. |
| [4] | 王宝珍;. 生物技术在畜牧业中的应用前景[J]. , 1987, 0(09): 8-9. |
| [5] | 马亚敏;. Embrex公司制订了将生物技术产品用于鸡的计划[J]. , 1987, 0(07): 22-23. |
| [6] | 本刊编辑部;. “K88-K99双价基因工程菌苗的构建及免疫原性试验”通过阶段成果鉴定[J]. , 1986, 0(10): 14-14. |
| [7] | 王宝珍;. 苏联和保加利亚在生物技术方面的长期合作[J]. , 1986, 0(03): 11-12. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||