








生物技术通报 ›› 2024, Vol. 40 ›› Issue (1): 160-167.doi: 10.13560/j.cnki.biotech.bull.1985.2023-0363
梁晋军1( ), 朱溯远1, 张宇琴1, 张鹏飞1, 温鹏飞1(
), 朱溯远1, 张宇琴1, 张鹏飞1, 温鹏飞1( ), 杨运良2(
), 杨运良2( )
)
收稿日期:2023-04-19
出版日期:2024-01-26
发布日期:2024-02-06
通讯作者:
温鹏飞,男,博士,教授,研究方向:葡萄逆境生理与分子生物学和果实品质形成与调控;E-mail: wenpengfei@126.com;作者简介:梁晋军,男,博士,讲师,研究方向:果树分子生物学;E-mail: liangjinjun1989@163.com;朱溯远为本文共同第一作者
基金资助:
LIANG Jin-jun1( ), ZHU Su-yuan1, ZHANG Yu-qin1, ZHANG Peng-fei1, WEN Peng-fei1(
), ZHU Su-yuan1, ZHANG Yu-qin1, ZHANG Peng-fei1, WEN Peng-fei1( ), YANG Yun-liang2(
), YANG Yun-liang2( )
)
Received:2023-04-19
Published:2024-01-26
Online:2024-02-06
摘要:
【目的】为了鉴别栽培柿品种间的遗传差异,开发一种基于核糖体DNA内转录间隔区(nrDNA internally transcribed spacer)序列简单、高效的SNP分子标记新方法,为种质的收集、利用及推广应用提供参考。【方法】以18种六倍体栽培柿品种叶片为试材,对其ITS区进行扩增测序,并分析序列差异,最后用Sau96 I限制性内切酶对151处杂合位点进行酶切验证。【结果】18份柿品种ITS长度均为730 bp,共存在6个杂合位点,分别在151、168、205、278、279和622 bp处。六倍体柿杂合位点处碱基峰图面积比例存在一定的规律,即C:T=2:1、C:T=1:1、C:T=1:2和A:G=1:1,利用出现的杂合位点及其峰图面积比例规律差异将18份栽培柿品种分成11类。【结论】151处位点的酶切结果验证了酶切产物浓度与峰图面积比例一致。这种基于nrDNA ITS序列SNP分子标记的新方法能将18份栽培柿品种分成11类。
梁晋军, 朱溯远, 张宇琴, 张鹏飞, 温鹏飞, 杨运良. 一种鉴别栽培柿品种SNP标记的新方法[J]. 生物技术通报, 2024, 40(1): 160-167.
LIANG Jin-jun, ZHU Su-yuan, ZHANG Yu-qin, ZHANG Peng-fei, WEN Peng-fei, YANG Yun-liang. A Novel SNP Marker for the Identification of Persimmon(Diospyros kaki)Cultivars[J]. Biotechnology Bulletin, 2024, 40(1): 160-167.
| 编号Code | 学名Scientific | 倍性水平Ploidy level | 种/品种名Species/Cultivar name | 脱涩类型Astringent type | 来源Source |
|---|---|---|---|---|---|
| 1 | D. kaki Thunb | 2n=6X=90 | 骏河 Suruga | PCNA | 日本Japan |
| 2 | D. kaki Thunb | 2n=6X=90 | 晚御所 Oku-gosho | PCNA | 日本Japan |
| 3 | D. kaki Thunb | 2n=6X =90 | 太秋 Taishuu | PCNA | 日本Japan |
| 4 | D. kaki Thunb | 2n=6X =90 | 花御所 Hanagosho | PCNA | 日本Japan |
| 5 | D. kaki Thunb | 2n=6X =90 | 兴津20 Okitsu 20 | PCNA | 日本Japan |
| 6 | D. kaki Thunb | 2n=6X =90 | 前川次郎Jirou | PCNA | 日本Japan |
| 7 | D. kaki Thunb | 2n=6X=90 | 伊豆 Yidou | PCNA | 日本Japan |
| 8 | D. kaki Thunb | 2n=6X=90 | 富有 Fuyuu | PCNA | 日本Japan |
| 9 | D. kaki Thunb | 2n=6X=90 | 早秋 Soshu | PCNA | 日本Japan |
| 10 | D. kaki Thunb | 2n=6X=90 | 阳丰 Youhou | PCNA | 日本Japan |
| 11 | D. kaki Thunb | 2n=6X=90 | 海库曼 Haikuman | PVNA | 美国America |
| 12 | D. kaki Thunb | 2n=6X=90 | 斤柿 Jinshi | PCA | 日本Japan |
| 13 | D. kaki Thunb | 2n=6X=90 | 法莲坊 Falianfang | PCA | 日本Japan |
| 14 | D. kaki Thunb | 2n=6X=90 | 火葫芦 Huohulu | PCA | 中国山西Shanxi, China |
| 15 | D. kaki Thunb | 2n=6X=90 | 小涩柿 Xiaoseshi | PCA | 中国山西Shanxi, China |
| 16 | D. kaki Thunb | 2n=6X=90 | 橘蜜柿 Jumishi | PCA | 中国山西Shanxi, China |
| 17 | D. kaki Thunb | 2n=6X=90 | 黑柿 Heishi | PCA | 中国山西Shanxi, China |
| 18 | D. kaki Thunb | 2n=6X=90 | 水化柿 Shuihuashi | PCA | 中国山西Shanxi, China |
表1 试验材料
Table 1 Materials in the experiment
| 编号Code | 学名Scientific | 倍性水平Ploidy level | 种/品种名Species/Cultivar name | 脱涩类型Astringent type | 来源Source |
|---|---|---|---|---|---|
| 1 | D. kaki Thunb | 2n=6X=90 | 骏河 Suruga | PCNA | 日本Japan |
| 2 | D. kaki Thunb | 2n=6X=90 | 晚御所 Oku-gosho | PCNA | 日本Japan |
| 3 | D. kaki Thunb | 2n=6X =90 | 太秋 Taishuu | PCNA | 日本Japan |
| 4 | D. kaki Thunb | 2n=6X =90 | 花御所 Hanagosho | PCNA | 日本Japan |
| 5 | D. kaki Thunb | 2n=6X =90 | 兴津20 Okitsu 20 | PCNA | 日本Japan |
| 6 | D. kaki Thunb | 2n=6X =90 | 前川次郎Jirou | PCNA | 日本Japan |
| 7 | D. kaki Thunb | 2n=6X=90 | 伊豆 Yidou | PCNA | 日本Japan |
| 8 | D. kaki Thunb | 2n=6X=90 | 富有 Fuyuu | PCNA | 日本Japan |
| 9 | D. kaki Thunb | 2n=6X=90 | 早秋 Soshu | PCNA | 日本Japan |
| 10 | D. kaki Thunb | 2n=6X=90 | 阳丰 Youhou | PCNA | 日本Japan |
| 11 | D. kaki Thunb | 2n=6X=90 | 海库曼 Haikuman | PVNA | 美国America |
| 12 | D. kaki Thunb | 2n=6X=90 | 斤柿 Jinshi | PCA | 日本Japan |
| 13 | D. kaki Thunb | 2n=6X=90 | 法莲坊 Falianfang | PCA | 日本Japan |
| 14 | D. kaki Thunb | 2n=6X=90 | 火葫芦 Huohulu | PCA | 中国山西Shanxi, China |
| 15 | D. kaki Thunb | 2n=6X=90 | 小涩柿 Xiaoseshi | PCA | 中国山西Shanxi, China |
| 16 | D. kaki Thunb | 2n=6X=90 | 橘蜜柿 Jumishi | PCA | 中国山西Shanxi, China |
| 17 | D. kaki Thunb | 2n=6X=90 | 黑柿 Heishi | PCA | 中国山西Shanxi, China |
| 18 | D. kaki Thunb | 2n=6X=90 | 水化柿 Shuihuashi | PCA | 中国山西Shanxi, China |
| 类型 Type | 品种编号 Cultivar No. | ITS变异位点Nucleotide of variable sites of ITS | |||||
|---|---|---|---|---|---|---|---|
| 151 | 168 | 205 | 278 | 279 | 622 | ||
| D1 | 14 | T | T | C | T | T | G |
| D2 | 1, 2, 3, 4, 15, 16 | C | T | C | T | T | G |
| D3 | 17 | T | T | C | C | T | G |
| D4 | 18 | T | T | C | T | Y3 | G |
| D5 | 12 | T | T | C | Y2 | Y3 | G |
| D6 | 5 | Y1 | Y1 | C | T | T | G |
| D7 | 13 | Y1 | T | Y1 | T | Y2 | R |
| D8 | 6,7 | Y1 | T | C | Y3 | T | G |
| D9 | 8,9 | Y2 | T | C | Y2 | T | G |
| D10 | 10 | Y3 | T | C | Y1 | T | G |
| D11 | 11 | Y3 | T | C | Y2 | T | G |
表2 基于柿种内ITS区的杂合位点现象聚类
Table 2 Clustering of heterozygous sites based on ITS region in persimmon species
| 类型 Type | 品种编号 Cultivar No. | ITS变异位点Nucleotide of variable sites of ITS | |||||
|---|---|---|---|---|---|---|---|
| 151 | 168 | 205 | 278 | 279 | 622 | ||
| D1 | 14 | T | T | C | T | T | G |
| D2 | 1, 2, 3, 4, 15, 16 | C | T | C | T | T | G |
| D3 | 17 | T | T | C | C | T | G |
| D4 | 18 | T | T | C | T | Y3 | G |
| D5 | 12 | T | T | C | Y2 | Y3 | G |
| D6 | 5 | Y1 | Y1 | C | T | T | G |
| D7 | 13 | Y1 | T | Y1 | T | Y2 | R |
| D8 | 6,7 | Y1 | T | C | Y3 | T | G |
| D9 | 8,9 | Y2 | T | C | Y2 | T | G |
| D10 | 10 | Y3 | T | C | Y1 | T | G |
| D11 | 11 | Y3 | T | C | Y2 | T | G |
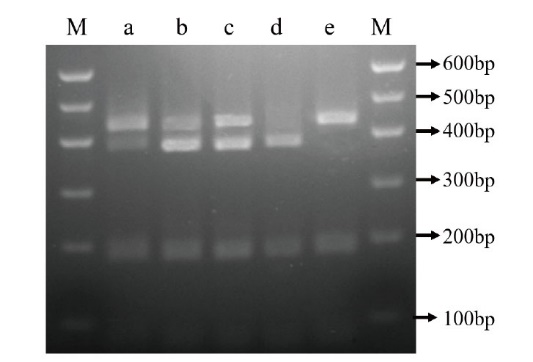
图2 151位点处特异酶切图 M:DNA maker;a:‘阳丰’;b:‘兴津20’;c:‘早秋’;d:‘晚御所’;e:‘黑柿’
Fig. 2 151 specific enzyme cut at sites M: DNA maker; a: ‘Youhou’; b: ‘Okitsu 20’; c: ‘Soshu’; d: ‘Oku-gosho’; e: ‘Heishi’
| [1] |
Zhuang DH, Kitajima A, Ishida M, et al. Chromosome numbers of Diospyros kaki cultivars[J]. Engei Gakkai Zasshi, 1990, 59(2): 289-297.
doi: 10.2503/jjshs.59.289 URL |
| [2] | 罗正荣, 张青林, 徐莉清, 等. 柿的遗传多样性及育种研究进展[J]. 落叶果树, 2021, 53(3): 1-5. |
| Luo ZR, Zhang QL, Xu LQ, et al. Advances in genetic diversity and breeding of persimmon[J]. Deciduous Fruits, 2021, 53(3): 1-5. | |
| [3] | 艾呈祥, 秦志华, 陶吉寒, 等. 32个柿主栽品种SSR图谱构建及遗传变异分析[J]. 西北植物学报, 2011, 31(11): 2185-2191. |
| Ai CX, Qin ZH, Tao JH, et al. SSR fingerprints and genetic variations of the 32 persimmon major cultivars[J]. Acta Botanica Boreali-Occidentalia Sinica, 2011, 31(11): 2185-2191. | |
| [4] | 陈星, 高子厚. DNA分子标记技术的研究与应用[J]. 分子植物育种, 2019, 17(6): 1970-1977. |
| Chen X, Gao ZH. The study and application of DNA molecular marker technique[J]. Mol Plant Breed, 2019, 17(6): 1970-1977. | |
| [5] |
Liang YQ, Han WJ, Sun P, et al. Genetic diversity among germplasms of Diospyros kaki based on SSR markers[J]. Sci Hortic, 2015, 186: 180-189.
doi: 10.1016/j.scienta.2015.02.015 URL |
| [6] |
Guan CF, Zhang PX, Hu CQ, et al. Genetic diversity, germplasm identification and population structure of Diospyros kaki Thunb. from different geographic regions in China using SSR markers[J]. Sci Hortic, 2019, 251: 233-240.
doi: 10.1016/j.scienta.2019.02.062 URL |
| [7] | 戚建锋, 李晓鹏, 王文然, 等. 利用基于SSR标记的MCID法鉴定72个柿地方品种[J]. 植物遗传资源学报, 2018, 19(5): 895-903. |
| Qi JF, Li XP, Wang WR, et al. Identification of 72 persimmon landraces by using SSR markers-based MCID metheod[J]. J Plant Genet Resour, 2018, 19(5): 895-903. | |
| [8] | 熊春艳. 浙江省柿农家品种SSR亲缘关系分析与鉴别[D]. 北京: 中国林业科学研究院, 2013. |
| Xiong CY. Analysis and identification of SSR genetic relationship of persimmon farmers in Zhejiang Province[D]. Beijing: Chinese Academy of Forestry, 2013. | |
| [9] | 傅建敏, 索玉静, 张嘉嘉, 等. 柿属植物cpDNA分子标记开发及遗传变异研究[J]. 中南林业科技大学学报, 2017, 37(6): 1-6. |
| Fu JM, Suo YJ, Zhang JJ, et al. Exploitation of cpDNA molecular markers and genetic variation of Diospyros spp[J]. J Central South Univ For Technol, 2015, 37(6): 1-6. | |
| [10] | 梁晋军, 梁玉琴, 傅建敏. 基于叶绿体DNA PCR-RFLP分析柿及其近缘种亲缘关系[J]. 园艺学报, 2014, 41(12): 2474-2480. |
| Liang JJ, Liang YQ, Fu JM. Genetic relationships of Diospyros kaki and related Diospyros species using chloroplast DNA PCR-RFLP markers[J]. Acta Hortic Sin, 2014, 41(12): 2474-2480. | |
| [11] |
Samarina LS, Malyarovskaya VI, Reim S, et al. Genetic diversity in Diospyros germplasm in the western caucasus based on SSR and ISSR polymorphism[J]. Biology, 2021, 10(4): 341.
doi: 10.3390/biology10040341 URL |
| [12] |
Guo DL, Luo ZR. Genetic relationships of some PCNA persimmons(Diospyros kaki Thunb.) from China and Japan revealed by SRAP analysis[J]. Genet Resour Crop Evol, 2006, 53(8): 1597-1603.
doi: 10.1007/s10722-005-8717-5 URL |
| [13] |
Jing ZB, Ruan XF, Wang RZ, et al. Genetic diversity and relationships between and within persimmon(Diospyros L.) wild species and cultivated varieties by SRAP markers[J]. Plant Syst Evol, 2013, 299(8): 1485-1492.
doi: 10.1007/s00606-013-0810-1 URL |
| [14] | Akbulut M, Ercişli S, Yildirim N, et al. The comparison of persimmon genotypes(Diospyros kaki Thunb.) by using RAPD and FAME data[J]. Roman Biotechnol Lett, 2008, 13(4): 3851-3858. |
| [15] |
Suo YJ, Sun P, Cheng HH, et al. A high-quality chromosomal genome assembly of Diospyros oleifera Cheng[J]. GigaScience, 2020, 9(1): giz164.
doi: 10.1093/gigascience/giz164 URL |
| [16] |
Akagi T, Shirasawa K, Nagasaki H, et al. The persimmon genome reveals clues to the evolution of a lineage-specific sex determination system in plants[J]. PLoS Genet, 2020, 16(2): e1008566.
doi: 10.1371/journal.pgen.1008566 URL |
| [17] | Mao WT, Yao GX, Wang SD, et al. Chromosome-level genomes of seeded and seedless date plum based on third-generation DNA sequencing and Hi-C analysis[J]. F, 2021, 1(1): 1-9. |
| [18] | 普天磊, 金杰, 何璐, 等. 基于SNP和InDel标记的余甘子群体遗传分析[J]. 果树学报, 2023, 40(5): 875-883. |
| Pu TL, Jing J, He L, et al. Population and genetic analysis of Phyllanthus emblica by SNP and InDel markers[J]. J Fruit Sci, 2023, 40(5): 875-883. | |
| [19] |
贺囡囡, 冯云敢, 蒙云飞, 等. 利用SNP标记及配合力划分超甜玉米自交系的杂种优势群[J]. 植物遗传资源学报, 2021, 22(1): 165-173.
doi: 10.13430/j.cnki.jpgr.20200512004 |
| He NN, Feng YG, Meng YF, et al. Classifying heterotic groups of super-sweet corn inbred lines by combining ability and SNP markers[J]. J Plant Genet Res, 2021, 22(1): 165-173. | |
| [20] |
卢茂昂, 彭小爱, 张玲, 等. 基于55K SNP芯片揭示小麦育种亲本遗传多样性[J]. 作物学报, 2023, 49(6): 1708-1704.
doi: 10.3724/SP.J.1006.2023.21047 |
| Lu MA, Peng XA, Zhang L, et al. Genetic diversity of wheat breeding parents revealed by 55K SNP-based microarray[J]. Acat Agronomica Sinica, 2023, 49(6): 1708-1704. | |
| [21] |
郑向华, 叶俊华, 程朝平, 等. 利用SNP标记进行水稻品种籼粳鉴定[J]. 作物学报, 2022, 48(2): 342-352.
doi: 10.3724/SP.J.1006.2022.02085 |
| Zheng XH, Ye JH, Cheng CP, et al. Xian-geng identification by SNP markers in Oryza sativa L.[J]. Acat Agron Sin, 2022, 48(2): 342-352. | |
| [22] |
Ma KB, Yang SJ, Jo YS, et al. Development of Kompetitive Allele Specific PCR markers for identification of persimmon varieties using genotyping-by-sequencing[J]. Electron J Biotechnol, 2021, 49: 72-81.
doi: 10.1016/j.ejbt.2020.11.003 URL |
| [23] | 杜改改, 孙鹏, 索玉静, 等. 基于柿雌雄花芽转录组测序的SSR和SNP多态性分析[J]. 中国农业大学学报, 2017, 22(10): 45-55. |
| Du GG, Sun P, Suo YJ, et al. SSR and SNP polymorphism analyisis based on persimmon(Diospyros kaki)transcriptome[J]. J China Agric Univ, 2017, 22(10): 45-55. | |
| [24] | 王艺儒, 索玉静, 傅建敏. 小果甜柿果实转录组的SSR、SNP和InDel特征分析[J]. 西北农林科技大学学报: 自然科学版, 2022, 50(7): 147-154. |
| Wang YR, Suo YJ, Fu JM. SSR, SNP and lnDeI analysis based on transcriptome data of Diospyros kaki ‘Xiaoguo-tianshi’ fruit[J]. J Northwest A&F Univ Nat Sci Ed, 2022, 50(7): 147-154. | |
| [25] | 刘一心, 杜运鹏, 崔凯峰, 等. 基于ITS序列的百合属野生资源与栽培品种亲缘关系评价[J]. 分子植物育种, 2019, 17(11): 3761-3768. |
| Liu YX, Du YP, Cui KF, et al. Evaluation of genetic relationship between wild resources and cultivars of Lilium based on ITS sequence[J]. Mol Plant Breed, 2019, 17(11): 3761-3768. | |
| [26] | 王建波, 张文驹, 陈家宽. 核rDNA的ITS序列在被子植物系统与进化研究中的应用[J]. 植物分类学报, 1999, 37(4): 407-416. |
| Wang JB, Zhang WJ, Chen JK. Application of ITS sequence of nuclear rDNA in angiosperm system and evolution research[J]. Acta Phytotaxon Sin, 1999, 37(4): 407-416. | |
| [27] |
Chen ZD, Li JH. Phylogenetics and biogeography of Alnus(Betulaceae)inferred from sequences of nuclear ribosomal DNA ITS region[J]. Int J Plant Sci, 2004, 165(2): 325-335.
doi: 10.1086/382795 URL |
| [28] | 杨梅花, 苗江林, 厚富霞, 等. 基于ITS和形态学系统分析新疆桦木属种间系统发育关系[J]. 西南林业大学学报: 自然科学, 2023, 43(1): 179-185. |
| Yang MH, Miao JL, Hou FX, et al. Phylogenetic relationship analysis based on ITS sequences and morphological characteristics for Betula species in Xinjiang[J]. J Southwest for Univ Nat Sci, 2023, 43(1): 179-185. | |
| [29] |
Qi W, Tuerxun J, Li J, et al. Identification of Bupleurum(Apiaceae)seeds by allele-specific PCR based on ITS sequences[J]. 3 Biotech, 2020, 10(6): 240.
doi: 10.1007/s13205-020-02233-1 |
| [30] |
Park SY, Jeon MJ, Joung YH, et al. Genetic variations of Narcissus tazetta and selected Narcissus cultivars based on the sequence analysis of nrITS and trnL-IS-trnF regions[J]. Genet Resour Crop Evol, 2023, 70(6): 1585-1603.
doi: 10.1007/s10722-022-01521-4 |
| [31] | 刘达, 王相瑾, 柴军红, 等. 玄参属种间ITS序列的遗传多样性分析[J/OL]. 分子植物育种, 2022. http://kns.cnki.net/kcms/detail/46.1068.S.20221215.1739.017.html. |
| Liu D, Wang XJ, Chai JH, et al. Genetic diversity analysis of ITS sequences of scrophularia[J/OL]. Mol Plant Breed, 2022. http://kns.cnki.net/kcms/detail/46.1068.S.20221215.1739.017.html. | |
| [32] | 朱玲, 唐式敏, 韩硕, 等. 基于ITS序列的新疆野苹果系统发育及遗传多样性分析[J]. 分子植物育种, 2022, 20(11): 3715-3721. |
| Zhu L, Tang SM, Han S, et al. Analysis of phylogeny and genetic diversity of Malus sieversii(ledeb.) M. Roem. based on ITS sequence[J]. Mol Plant Breed, 2022, 20(11): 3715-3721. | |
| [33] | 易庆平. 柿属植物DNA提取纯化检测技术体系的建立[D]. 武汉: 华中农业大学, 2006. |
| Yi QP. Establishment of DNA extraction, purification and detection technology system for persimmon plants[D]. Wuhan: Huazhong Agricultural University, 2006. | |
| [34] | Zhang Q, Dai W, Jin X, et al. Bioinformatics analysis on pathogenic genes of Botrytis cinerea Pers.[J]. Agr Biotechnol, 2018, 7(1): 57-61, 86. |
| [35] | 陈炜烨, 刘冬冬, 徐建华, 等. ImageJ软件在重组质粒pET32a-CDK2中蛋白表达的应用[J]. 中国热带医学, 2014, 14(1): 23-25. |
| Chen WY, Liu DD, Xu JH, et al. Application of image J software in analyzing protein expression of recombinant plasmid p ET32a-CDK2[J]. China Trop Med, 2014, 14(1): 23-25. | |
| [36] | 其乐木格. 肿瘤细胞新分子标记的探索[D]. 呼和浩特: 内蒙古大学, 2016. |
| Qi LMG. Screening for novel molecular markers of tumor cells[D]. Hohhot: Inner Mongolia University, 2016. | |
| [37] |
Ma KB, Yang SJ, Jo YS, et al. Development of kompetitive allele specific PCR markers for identification of persimmon varieties using genotyping-by-sequencing[J]. Electron J Biotechnol, 2021, 49: 72-81.
doi: 10.1016/j.ejbt.2020.11.003 URL |
| [38] | 郭大龙. 几种分子标记技术的建立及其在部分柿属植物亲缘关系研究中的应用[D]. 武汉: 华中农业大学, 2006. |
| Guo DL. Establishment of several molecular marker techniques and their application in the study of genetic relationship of some persimmon plants[D]. Wuhan: Huazhong Agricultural University, 2006. | |
| [39] | 梁晋军. 柿及其近缘种亲缘关系及柿品种遗传多样性的研究[D]. 北京: 中国林业科学研究院, 2015. |
| Liang JJ. Study on the genetic relationship of persimmon and its related species and the genetic diversity of persimmon varieties[D]. Beijing: Chinese Academy of Forestry, 2015. |
| [1] | 崔学强, 黄昌艳, 邓杰玲, 李先民, 李秀玲, 张自斌. 基于SLAF-seq技术的石斛兰SNP标记开发及亲缘关系分析[J]. 生物技术通报, 2023, 39(6): 141-148. |
| [2] | 肖小军, 陈明, 韩德鹏, 余跑兰, 郑伟, 肖国滨, 周庆红, 周会汶. 甘蓝型油菜每角果粒数全基因组关联分析[J]. 生物技术通报, 2023, 39(3): 143-151. |
| [3] | 周晓楠, 徐金青, 雷雨晴, 王海庆. 基于GBS-seq的青藏扁蓿豆SNP标记开发[J]. 生物技术通报, 2022, 38(4): 303-310. |
| [4] | 洪军, 卫夏怡, 吉冰洁, 叶延欣, 程天赐. 铜绿假单胞菌对鲎素耐药前后的差异表达基因及SNP变化研究[J]. 生物技术通报, 2021, 37(9): 191-202. |
| [5] | 陈一丹, 张昱, 杨洁, 张勤, 姜力. 基于转录组测序的奶牛产奶性状重要功能基因挖掘[J]. 生物技术通报, 2020, 36(9): 244-252. |
| [6] | 余钧剑, 迟美丽, 贾永义, 刘士力, 竺俊全, 顾志敏. 四引物扩增受阻突变体系PCR技术及其在动植物遗传育种研究中的应用[J]. 生物技术通报, 2020, 36(5): 32-38. |
| [7] | 邹坤, 崔红艳, 薛缘, 张少伟, 路丽丽, 赵志辉, 苏瑛. 雷州黑鸭FSHR和ESR1与繁殖性状关联分析[J]. 生物技术通报, 2019, 35(8): 118-126. |
| [8] | 郭萌萌, 周延清, 段红英, 杨珂, 邵露营. 基于地黄转录组数据的SNP标记开发与地黄指纹图谱构建[J]. 生物技术通报, 2019, 35(11): 224-230. |
| [9] | 张欢, 张全伟, 王琪, 马友记, 张勇, 赵兴绪. 牦牛TMEM-18基因多态性及其与生产性能关联分析[J]. 生物技术通报, 2017, 33(2): 89-96. |
| [10] | 张先文,, 贺治洲, 江南, 邓华凤, 李继明. 高通量基因型分型技术及其在水稻中的应用[J]. 生物技术通报, 2017, 33(12): 67-73. |
| [11] | 李宏博, 庞斌双, 刘丽华, 刘阳娜, 赵昌平, 陈景堂. 河北区试小麦品种(系)DNA指纹图谱构建及遗传差异分析[J]. 生物技术通报, 2015, 31(6): 93-99. |
| [12] | 唐红杰, 赵生国, 雷赵民, 王欣荣, 王建福, 蔡原, 吴建平. 早胜牛类群A-FABP基因SNPs及其与胴体品质和肉质性状的相关性分析[J]. 生物技术通报, 2014, 0(5): 96-101. |
| [13] | 张晓萌, 马普, 王洪迪, 王秀利. SNPs在水产动物中的研究进展[J]. 生物技术通报, 2013, 0(8): 7-11. |
| [14] | 龚俞, 杨永强, 焦仁刚, 惠嫣婷, 刘若余. 牛JAK2基因启动子区多态及生物信息学研究[J]. 生物技术通报, 2013, 0(6): 104-109. |
| [15] | 冯娜娜;徐真;马洪雨;马春艳;乔振国;马凌波;. 凡纳滨对虾7个不同家系遗传差异的微卫星标记分析[J]. , 2012, 0(11): 133-138. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||