








Biotechnology Bulletin ›› 2022, Vol. 38 ›› Issue (5): 74-83.doi: 10.13560/j.cnki.biotech.bull.1985.2021-0981
Previous Articles Next Articles
LEI Jun1,2( ), CHEN Qin1,2, DENG Bing1,2, ZHANG Jin-yu3, LIU Di-qiu1,2, CUI Xiu-ming1,2, GE Feng1,2(
), CHEN Qin1,2, DENG Bing1,2, ZHANG Jin-yu3, LIU Di-qiu1,2, CUI Xiu-ming1,2, GE Feng1,2( )
)
Received:2021-08-02
Online:2022-05-26
Published:2022-06-10
Contact:
GE Feng
E-mail:353597247@qq.com;panaxcordyceps@163.com
LEI Jun, CHEN Qin, DENG Bing, ZHANG Jin-yu, LIU Di-qiu, CUI Xiu-ming, GE Feng. Biosynthesis of Panax notoginseng Saponins Regulated by R2R3-MYB Transcription Factor PnMYB1[J]. Biotechnology Bulletin, 2022, 38(5): 74-83.
| 引物名称 Primer name | 序列 Sequence(5'-3') |
|---|---|
| PnGAPDH-F | CTTTGGTTTAAGGAACCCAGAGG |
| PnGAPDH-R | AAGGGGAGCAAGGCAGTTAGTAG |
| PnSS-F | CGAGCACTTGACACTGTTGAGGAT |
| PnSS-R | CTATTGCCTCCTGGTAACCGTTTC |
| PnDS-F | CAAGCACACGATGGTCACTGGC |
| PnDS-R | CATTTTGATGGTTGTAAACGAAGCG |
| PnSE-F | AGGTGAACTTCTACAACCAGGAGGC |
| PnSE-R | CTCAACCAGAGATGTAACAGTCCCC |
| PnCAS-F | GAAATTATACCCTGACCACCGT |
| PnCAS-R | CCAAACCTTTTACACCGAACC |
| PnMYB1-F | GCAGGTTTAAAGAGATGTGGGAAG |
| PnMYB1-R | CTGAAGCAGAGGAGGAGTGATTG |
Table 1 Primers used for qRT-PCR
| 引物名称 Primer name | 序列 Sequence(5'-3') |
|---|---|
| PnGAPDH-F | CTTTGGTTTAAGGAACCCAGAGG |
| PnGAPDH-R | AAGGGGAGCAAGGCAGTTAGTAG |
| PnSS-F | CGAGCACTTGACACTGTTGAGGAT |
| PnSS-R | CTATTGCCTCCTGGTAACCGTTTC |
| PnDS-F | CAAGCACACGATGGTCACTGGC |
| PnDS-R | CATTTTGATGGTTGTAAACGAAGCG |
| PnSE-F | AGGTGAACTTCTACAACCAGGAGGC |
| PnSE-R | CTCAACCAGAGATGTAACAGTCCCC |
| PnCAS-F | GAAATTATACCCTGACCACCGT |
| PnCAS-R | CCAAACCTTTTACACCGAACC |
| PnMYB1-F | GCAGGTTTAAAGAGATGTGGGAAG |
| PnMYB1-R | CTGAAGCAGAGGAGGAGTGATTG |
| 引物名称 Primer name | 序列 Sequence(5'-3') |
|---|---|
| PnCAS-GSP1 | AGCTTTCTCGATGTCCGCAAGCTCTTC |
| PnCAS-GSP2 | GGATTTCCTCCCTCTGCAATTTTAAGC |
| PnDS-GSP1 | TCCCAAAATGCTTAAAGCCTCGATCCA |
| PnDS-GSP2 | CTTGTAGGGCTTATTGTTATGCAGATTGTG |
| PnSS-GSP1 | CATTTCATGTCCCGTTTTCCTGTAAGAAC |
| PnSS-GSP2 | ACAGTGTTTCTTTCCAGCGAGGACTCC |
| PnDSP-F | ATCGATTCAATACCGTGTGCTACTATGCAAC |
| PnDSP-R | GGATCCTGGTTATGTGGTGTACATAGATGGC |
| PnCASP-F | AAGCTTTGAGGGGCCAAATTCGTTG |
| PnCASP-R | TCTAGACACTCTGCACACAAATTTAGCTCC |
| PnSEP-F | AAGCTTTTGTGGGTCAGATCAGATGGA |
| PnSEP-R | GGATCCGGTGTTGGTTGGACGTTCAC |
Table 2 Primers used for promoter clone
| 引物名称 Primer name | 序列 Sequence(5'-3') |
|---|---|
| PnCAS-GSP1 | AGCTTTCTCGATGTCCGCAAGCTCTTC |
| PnCAS-GSP2 | GGATTTCCTCCCTCTGCAATTTTAAGC |
| PnDS-GSP1 | TCCCAAAATGCTTAAAGCCTCGATCCA |
| PnDS-GSP2 | CTTGTAGGGCTTATTGTTATGCAGATTGTG |
| PnSS-GSP1 | CATTTCATGTCCCGTTTTCCTGTAAGAAC |
| PnSS-GSP2 | ACAGTGTTTCTTTCCAGCGAGGACTCC |
| PnDSP-F | ATCGATTCAATACCGTGTGCTACTATGCAAC |
| PnDSP-R | GGATCCTGGTTATGTGGTGTACATAGATGGC |
| PnCASP-F | AAGCTTTGAGGGGCCAAATTCGTTG |
| PnCASP-R | TCTAGACACTCTGCACACAAATTTAGCTCC |
| PnSEP-F | AAGCTTTTGTGGGTCAGATCAGATGGA |
| PnSEP-R | GGATCCGGTGTTGGTTGGACGTTCAC |
| 人参皂苷标准品 Ginsenoside standards | 回归方程 Regression | R2 |
|---|---|---|
| Rg1 | y = 3944.9x +69.409 | 0.9997 |
| Re | y = 2646.8x+22.808 | 0.9999 |
| Rb1 | y = 3227.3x+49.436 | 0.9998 |
| Rd | y = 2953.5x+41.408 | 0.9998 |
| R1 | y = 2705.6x+50.697 | 0.9996 |
Table 3 Linear regression equations for five major mono-mer saponins
| 人参皂苷标准品 Ginsenoside standards | 回归方程 Regression | R2 |
|---|---|---|
| Rg1 | y = 3944.9x +69.409 | 0.9997 |
| Re | y = 2646.8x+22.808 | 0.9999 |
| Rb1 | y = 3227.3x+49.436 | 0.9998 |
| Rd | y = 2953.5x+41.408 | 0.9998 |
| R1 | y = 2705.6x+50.697 | 0.9996 |
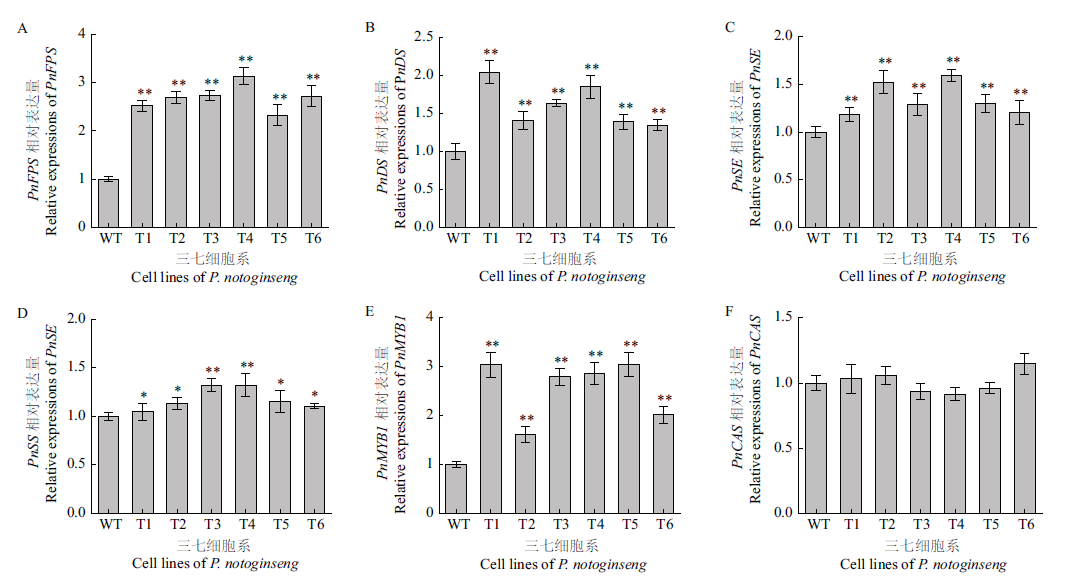
Fig. 3 Relative expressions of gene PnFPS(A),PnDS(B),PnSE(C),PnSS(D),PnMYB1(E)and PnCAS(F)in the T1-T6 cell lines of P. notoginseng WT:Wild type cell of P. notoginseng;**P<0.01 vs control group;*P<0.05 vs control group. The same below
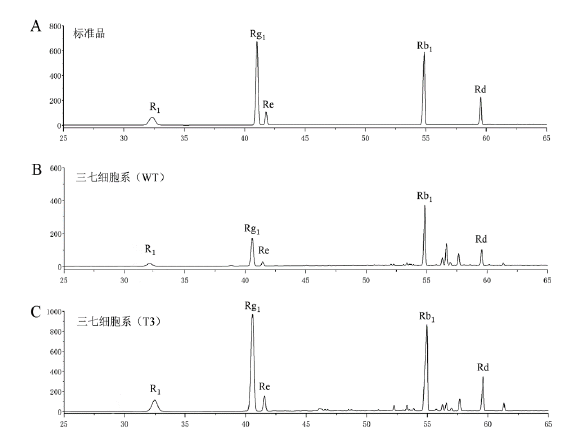
Fig. 5 HPLC analysis of transformed PnMYB1 P. notogin-seng cell line A:Standard. B:Wild-type P. notoginseng cell. C:Transformed PnMYB1 T3 P. notoginseng cell line
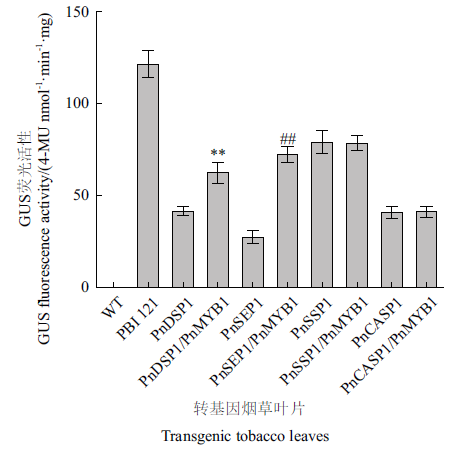
Fig. 7 GUS fluorescence activity analysis PnDSP1,PnSEP1,PnSSP1 and PnCASP1 refer to the tobacco leaves transgenic with PnDSP1,PnSEP1,PnSSP1 and PnCASP1 promoter fragments,respectively. PnSSP1/PnMYB1,PnSEP1/PnMYB1,PnDSP1/PnMYB1 and PnCASP1/PnMYB1 refers to the tobacco leaves that PnSSP1,PnSEP1,PnDSP1 and PnCASP1 promoter fragments co-transfected with PnMYB1,respectively. PBI 121:PBI 121-transfected wild-type tobacco leaves. WT:Wild tobacco leaves.(** P < 0.01 vs PnDSP1;and ## P <0.01 vs PnSEP1)
| [1] |
Ng TB. Pharmacological activity of Sanchi ginseng(Panax notoginseng)[J]. J Pharm Pharmacol, 2010, 58(8):1007-1019.
doi: 10.1211/jpp.58.8.0001 URL |
| [2] |
Kuzuyama T. Mevalonate and nonmevalonate pathways for the biosynthesis of isoprene units[J]. Biosci Biotechnol Biochem, 2002, 66(8):1619-1627.
doi: 10.1271/bbb.66.1619 URL |
| [3] |
Kim DH. Chemical Diversity of Panax ginseng, Panax quinquifolium, and Panax notoginseng[J]. J Ginseng Res, 2012, 36(1):1-15.
doi: 10.5142/jgr.2012.36.1.1 URL |
| [4] |
Zhang JZ. Overexpression analysis of plant transcription factors[J]. Curr Opin Plant Biol, 2003, 6(5):430-440.
doi: 10.1016/S1369-5266(03)00081-5 URL |
| [5] |
Heim MA, Jakoby M, Werber M, et al. The basic helix-loop-helix transcription factor family in plants:a genome-wide study of protein structure and functional diversity[J]. Mol Biol Evol, 2003, 20(5):735-747.
doi: 10.1093/molbev/msg088 URL |
| [6] |
Ma D, Pu G, Lei C, et al. Isolation and characterization of AaWRKY1, an Artemisia annua transcription factor that regulates the Amorpha-4, 11-diene synthase gene, a key gene of artemisinin biosynthesis[J]. Plant Cell Physiol, 2009, 50(12):2146-2161.
doi: 10.1093/pcp/pcp149 URL |
| [7] |
Dubos C, Stracke R, Grotewold E, et al. MYB transcription factors in Arabidopsis[J]. Trends Plant Sci, 2010, 15(10):573-581.
doi: 10.1016/j.tplants.2010.06.005 URL |
| [8] |
Bedon F, Bomal C, Caron S, et al. Subgroup 4 R2R3-MYBs in conifer trees:gene family expansion and contribution to the isoprenoid- and flavonoid-oriented responses[J]. J Exp Bot, 2010, 61(14):3847-3864.
doi: 10.1093/jxb/erq196 URL |
| [9] | Li CY, Leopold AL, Sander GW, et al. CrBPF1 overexpression alters transcript levels of terpenoid indole alkaloid biosynthetic and regulatory genes[J]. Front Plant Sci, 2015, 6:818. |
| [10] | 徐俊雄, 覃建兵, 王岩岩, 等. 虎杖MYB转录因子PcMYB1的表达特性和功能研究[J]. 中草药, 2020, 51(3):726-732. |
| Xu JX, Qin JB, Wang YY, et al. Expression and function analysis of transcription factor PcMYB1 from Polygonum cuspidatum[J]. Chin Tradit Herb Drugs, 2020, 51(3):726-732. | |
| [11] | 应宇翔, 何凤明, 张云峰, 等. 灯盏花MYB基因克隆及其荧光表达载体的构建[J]. 中草药, 2017, 48(20):4306-4315. |
| Ying YX, He FM, Zhang YF, et al. Cloning of MYB gene and construction of greenfluorescent protein expression vector in Erigeron breviscapus[J]. Chin Tradit Herb Drugs, 2017, 48(20):4306-4315. | |
| [12] |
Puissant C, Houdebine LM. An improvement of the single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction[J]. BioTechniques, 1990, 8(2):148-149.
pmid: 1690559 |
| [13] |
Chomczynski P, Sacchi N. Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction[J]. Anal Biochem, 1987, 162(1):156-159.
doi: 10.1006/abio.1987.9999 pmid: 2440339 |
| [14] |
Dai Y, Qin Q, Dai D, et al. Isolation and characterization of a novel cDNA encoding methyl jasmonate-responsive transcription factor TcAP2 from Taxus cuspidata[J]. Biotechnol Lett, 2009, 31(11):1801-1809.
doi: 10.1007/s10529-009-0068-4 URL |
| [15] |
Höfgen R, Willmitzer L. Storage of competent cells for Agrobacterium transformation[J]. Nucleic Acids Res, 1988, 16(20):9877.
pmid: 3186459 |
| [16] |
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method[J]. Methods, 2001, 25(4):402-408.
doi: 10.1006/meth.2001.1262 pmid: 11846609 |
| [17] |
Wako T, Kimura S, Chikagawa Y, et al. Characterization of MYB proteins acting as transcriptional regulatory factors for carrot phenylalanine ammonia-lyase gene(DcPAL3)[J]. Plant Biotechnol, 2010, 27(2):131-139.
doi: 10.5511/plantbiotechnology.27.131 URL |
| [18] |
Wang N, Xu H, Jiang S, et al. MYB12 and MYB22 play essential roles in proanthocyanidin and flavonol synthesis in red-fleshed apple(Malus sieversii f. niedzwetzkyana)[J]. Plant J, 2017, 90(2):276-292.
doi: 10.1111/tpj.13487 URL |
| [19] |
Gonzalez A, Zhao M, Leavitt JM, et al. Regulation of the anthocyanin biosynthetic pathway by the TTG1/bHLH/Myb transcriptional complex in Arabidopsis seedlings[J]. Plant J, 2008, 53(5):814-827.
doi: 10.1111/j.1365-313X.2007.03373.x URL |
| [20] |
Dai X, Xu Y, Ma Q, et al. Overexpression of an R1R2R3 MYB gene, OsMYB3R-2, increases tolerance to freezing, drought, and salt stress in transgenic Arabidopsis[J]. Plant Physiol, 2007, 143(4):1739-1751.
doi: 10.1104/pp.106.094532 URL |
| [21] |
Elomaa P, Uimari A, Mehto M, et al. Activation of anthocyanin biosynthesis in Gerbera hybrida(Asteraceae)suggests conserved protein-protein and protein-promoter interactions between the anciently diverged monocots and eudicots[J]. Plant Physiol, 2003, 133(4):1831-1842.
pmid: 14605235 |
| [22] |
Deluc L, Bogs J, Walker AR, et al. The transcription factor VvMYB5b contributes to the regulation of anthocyanin and proanthocyanidin biosynthesis in developing grape berries[J]. Plant Physiol, 2008, 147(4):2041-2053.
doi: 10.1104/pp.108.118919 URL |
| [23] |
Li CH, Qiu J, Yang GS, et al. Isolation and characterization of a R2R3-MYB transcription factor gene related to anthocyanin biosynthesis in the spathes of Anthurium andraeanum(Hort. )[J]. Plant Cell Rep, 2016, 35(10):2151-2165.
doi: 10.1007/s00299-016-2025-8 URL |
| [24] |
Lin WK, Bolitho K, Grafton K, et al. An R2R3 MYB transcription factor associated with regulation of the anthocyanin biosynthetic pathway in Rosaceae[J]. BMC Plant Biol, 2010, 10:50.
doi: 10.1186/1471-2229-10-50 pmid: 20302676 |
| [25] |
Lee MH, Jeong JH, Seo JW, et al. Enhanced triterpene and phytosterol biosynthesis in Panax ginseng overexpressing squalene synthase gene[J]. Plant Cell Physiol, 2004, 45(8):976-984.
doi: 10.1093/pcp/pch126 URL |
| [26] |
Deng B, Zhang P, Ge F, et al. Enhancement of triterpenoid saponins biosynthesis in Panax notoginseng cells by co-overexpressions of 3-hydroxy-3-methylglutaryl CoA reductase and squalene synthase genes[J]. Biochem Eng J, 2017, 122:38-46.
doi: 10.1016/j.bej.2017.03.001 URL |
| [27] |
Laitinen RA, Ainasoja M, Broholm SK, et al. Identification of target genes for a MYB-type anthocyanin regulator in Gerbera hybrida[J]. J Exp Bot, 2008, 59(13):3691-3703.
doi: 10.1093/jxb/ern216 pmid: 18725377 |
| [28] |
Leung KW, Wong AS. Pharmacology of ginsenosides:a literature review[J]. Chin Med, 2010, 5:20.
doi: 10.1186/1749-8546-5-20 URL |
| [29] |
Lee JJ, Kwon HK, Jung IH, et al. Anti-cancer activities of ginseng extract fermented with Phellinus linteus[J]. Mycobiology, 2009, 37(1):21-27.
doi: 10.4489/MYCO.2009.37.1.021 URL |
| [30] |
Ni N, Liu Q, Ren H, et al. Ginsenoside Rb1 protects rat neural progenitor cells against oxidative injury[J]. Molecules, 2014, 19(3):3012-3024.
doi: 10.3390/molecules19033012 URL |
| [31] | 陈勤, 雷君, 葛锋, 等. 人参属三萜皂苷骨架修饰的研究进展[J]. 中药材, 2020, 11:2831-2837. |
| Chen Q, Lei J, Ge F, et al. The research progress of ginseng triterpenoid saponin skeleton modifying enzyme[J]. Journal of Chinese Medicinal Materials, 2020, 11:2831-2837. |
| [1] | HUANG Xiao-long, SUN Gui-lian, MA Dan-dan, YAN Hui-qing. Construction of Yeast One-hybrid Library and Screening of Factors Regulating LAZY1 Expression in Rice [J]. Biotechnology Bulletin, 2023, 39(9): 126-135. |
| [2] | HAN Hao-zhang, ZHANG Li-hua, LI Su-hua, ZHAO Rong, WANG Fang, WANG Xiao-li. Construction of cDNA Library of Cinnamomun bodinieri Induced by Saline-alkali Stress and Screening of CbP5CS Upstream Regulators [J]. Biotechnology Bulletin, 2023, 39(9): 236-245. |
| [3] | LYU Qiu-yu, SUN Pei-yuan, RAN Bin, WANG Jia-rui, CHEN Qing-fu, LI Hong-you. Cloning, Subcellular Localization and Expression Analysis of the Transcription Factor Gene FtbHLH3 in Fagopyrum tataricum [J]. Biotechnology Bulletin, 2023, 39(8): 194-203. |
| [4] | XU Jing, ZHU Hong-lin, LIN Yan-hui, TANG Li-qiong, TANG Qing-jie, WANG Xiao-ning. Cloning of IbHQT1 Promoter and Identification of Upstream Regulatory Factors in Sweet Potato [J]. Biotechnology Bulletin, 2023, 39(8): 213-219. |
| [5] | LI Bo, LIU He-xia, CHEN Yu-ling, ZHOU Xing-wen, ZHU Yu-lin. Cloning, Subcellular Localization and Expression Analysis of CnbHLH79 Transcription Factor from Camellia nitidissima [J]. Biotechnology Bulletin, 2023, 39(8): 241-250. |
| [6] | CHEN Xiao, YU Ming-lan, WU Long-kun, ZHENG Xiao-ming, PANG Hong-bo. Research Progress in lncRNA and Their Responses to Low Temperature Stress in Plant [J]. Biotechnology Bulletin, 2023, 39(7): 1-12. |
| [7] | ZHAO Lin-yan, XU Wu-mei, WANG Hao-ji, WANG Kun-yan, WEI Fu-gang, YANG Shao-zhou, GUAN Hui-lin. Effects of Applying Biochar on the Rhizosphere Fungal Community and Survival Rate of Panax notoginseng Under Continuous Cropping [J]. Biotechnology Bulletin, 2023, 39(7): 219-227. |
| [8] | GUO Yi-ting, ZHAO Wen-ju, REN Yan-jing, ZHAO Meng-liang. Identification and Analysis of NAC Transcription Factor Family Genes in Helianthus tuberosus L. [J]. Biotechnology Bulletin, 2023, 39(6): 217-232. |
| [9] | FENG Shan-shan, WANG Lu, ZHOU Yi, WANG You-ping, FANG Yu-jie. Research Progresses on WOX Family Genes in Regulating Plant Development and Abiotic Stress Response [J]. Biotechnology Bulletin, 2023, 39(5): 1-13. |
| [10] | WANG Bing, ZHAO Hui-na, YU Jing, YU Shi-zhou, LEI Bo. Research Progress in the Regulation of Plant Branch Development [J]. Biotechnology Bulletin, 2023, 39(5): 14-22. |
| [11] | ZHANG Xin-bo, CUI Hao-liang, SHI Pei-hua, GAO Jin-chun, ZHAO Shun-ran, TAO Chen-yu. Research Progress in Low-input Chromatin Immunoprecipitation Assay [J]. Biotechnology Bulletin, 2023, 39(4): 227-235. |
| [12] | GE Yan-rui, ZHAO Ran, XU Jing, LI Ruo-fan, HU Yun-tao, LI Rui-li. Advances in the Development and Regulation of Vascular Cambium [J]. Biotechnology Bulletin, 2023, 39(3): 13-25. |
| [13] | LIU Cheng-xia, SUN Zong-yan, LUO Yun-bo, ZHU Hong-liang, QU Gui-qin. Multifaceted Roles of bHLH Phosphorylation in Regulation of Plant Physiological Functions [J]. Biotechnology Bulletin, 2023, 39(3): 26-34. |
| [14] | ZHAO Meng-liang, GUO Yi-ting, REN Yan-jing. Identification and Analysis of WRKY Transcription Factor Family Genes in Helianthus tuberosus [J]. Biotechnology Bulletin, 2023, 39(2): 116-125. |
| [15] | HAN Fang-ying, HU Xin, WANG Nan-nan, XIE Yu-hong, WANG Xiao-yan, ZHU Qiang. Research Progress in Response of DREBs to Abiotic Stress in Plant [J]. Biotechnology Bulletin, 2023, 39(11): 86-98. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||