








生物技术通报 ›› 2024, Vol. 40 ›› Issue (1): 186-193.doi: 10.13560/j.cnki.biotech.bull.1985.2023-0336
刘星雨1( ), 李洁2, 朱龙佼3, 李相阳2, 许文涛3(
), 李洁2, 朱龙佼3, 李相阳2, 许文涛3( )
)
收稿日期:2023-04-11
出版日期:2024-01-26
发布日期:2024-02-06
通讯作者:
许文涛,博士,教授,研究方向:生物安全、功能核酸及功能食品;E-mail: xuwentao@cau.edu.cn作者简介:刘星雨,硕士研究生,研究方向:基于功能核酸的生物传感;E-mail: liuxingyu0321@163.com
基金资助:
LIU Xing-yu1( ), LI Jie2, ZHU Long-jiao3, LI Xiang-yang2, XU Wen-tao3(
), LI Jie2, ZHU Long-jiao3, LI Xiang-yang2, XU Wen-tao3( )
)
Received:2023-04-11
Published:2024-01-26
Online:2024-02-06
摘要:
铜绿假单胞菌是一种专性需氧的革兰氏阴性菌,是引起人类机会性感染的最主要的微生物,对人类造成多种健康威胁。因此,快速、灵敏、特异的检测对预防及治疗铜绿假单胞菌感染十分关键。目前常规的检测方法仍存在耗时久、成本高、操作难度大等问题,因此需要更加高效经济的检测方法。适配体是一种从随机文库中筛选出来的寡核苷酸,能够形成多种复杂的二级结构,近年来,由于其易于合成和修饰,并且具有高亲和力和强特异性而受到了广泛的关注,也涌现出了大量基于适配体的铜绿假单胞菌检测和防治的研究。本文总结了目前铜绿假单胞菌适配体的筛选和裁剪方面的研究进展,并从铜绿假单胞菌的检测和防治两个方面对其适配体的应用进行了介绍,以期为铜绿假单胞菌适配体的开发和应用提供参考。
刘星雨, 李洁, 朱龙佼, 李相阳, 许文涛. 铜绿假单胞菌适配体的获得及应用[J]. 生物技术通报, 2024, 40(1): 186-193.
LIU Xing-yu, LI Jie, ZHU Long-jiao, LI Xiang-yang, XU Wen-tao. Aptamer of Pseudomonas aeruginosa: Acquiring and Application[J]. Biotechnology Bulletin, 2024, 40(1): 186-193.
| 靶标 Target | 筛选方式SELEX type | 序列 Sequence(5'-3') | Kd/(nmol·L-1) | 亲和力检测方式 Affinity detection method | 长度Length/nt | 参考文献Reference |
|---|---|---|---|---|---|---|
| 全菌 | Cell-SELEX | CCCCCGTTGCTTTCGCTTTTCCTTTCGCTTGT-GTTCGTTTCGTCCCTGCTTCCTTCCTTCCTTC-CTTTCTTG | 17.27 ± 5.00 | 流式细胞术 | 72 | [ |
| 全菌 | Cell-SELEX | ATGCACTCTTCTATCGGTAGTTGAGGGTGCGG | 12.5±2.4 | 流式细胞术 | 32 | [ |
| 外毒素A | Decoy-SELEX | TGTACCGTCTGAGCGATTCGTACCATAGGGTGCTTTTCAAGGCCACACGTTAGTGTAAGCCAGTCAGTGTTAAGGAGTGC | 4 200-4 500 | 表面等离子体共振 | 80 | [ |
| 脂多糖 | Capture-SELEX | CCTTCTAACAGAATGTTGTTAGATAGC | 46.2±9.5 | 等热滴定量热 | 27 | [ |
| 全菌 | Cell-SELEX | GGGAGGACFGAUFGCFGGGAACFUFGAAGAGUFUFUFGAUFCFAUFGGCFUFCFAGAUFUFGAACFGCFUFGGCFGGCFAGACFGACFUCFGCFGCFGA | — | — | 73 | [ |
| 全菌 | WB-SELEX | TGGACCTTGCGATTGACAGCAGACATGAGTCAGGACGTGACCGCTGGACCTTGCGATTGACAGCAGACATGAGTCAGGAC | 15.16±3.62 | 流式细胞术 | 80 | [ |
| 全菌 | Cell-SELEX | CCCCCGTTGCTTTCGCTTTTCCTTTCGCTTTTGTTCGTTTCGTCCCTGCTTCCTTTCTTG | 14.55 | 地高辛标记适配体 | 60 | [ |
| 丁酰基-高丝氨酸 | Structure-switching SELEX | CCATCCACACTCCGCAAGTGGGGAGGGGAGAGACGACGATCCTGTGGGTTTTCTGCAGTGAGTCGTGTTTTCGACTTATTGCGTCGGCTGCCTCTACAT | 28.47 | 流式细胞术 | 99 | [ |
表1 铜绿假单胞菌适配体的筛选
Table 1 SELEX of Pseudomonas aeruginosa aptamers
| 靶标 Target | 筛选方式SELEX type | 序列 Sequence(5'-3') | Kd/(nmol·L-1) | 亲和力检测方式 Affinity detection method | 长度Length/nt | 参考文献Reference |
|---|---|---|---|---|---|---|
| 全菌 | Cell-SELEX | CCCCCGTTGCTTTCGCTTTTCCTTTCGCTTGT-GTTCGTTTCGTCCCTGCTTCCTTCCTTCCTTC-CTTTCTTG | 17.27 ± 5.00 | 流式细胞术 | 72 | [ |
| 全菌 | Cell-SELEX | ATGCACTCTTCTATCGGTAGTTGAGGGTGCGG | 12.5±2.4 | 流式细胞术 | 32 | [ |
| 外毒素A | Decoy-SELEX | TGTACCGTCTGAGCGATTCGTACCATAGGGTGCTTTTCAAGGCCACACGTTAGTGTAAGCCAGTCAGTGTTAAGGAGTGC | 4 200-4 500 | 表面等离子体共振 | 80 | [ |
| 脂多糖 | Capture-SELEX | CCTTCTAACAGAATGTTGTTAGATAGC | 46.2±9.5 | 等热滴定量热 | 27 | [ |
| 全菌 | Cell-SELEX | GGGAGGACFGAUFGCFGGGAACFUFGAAGAGUFUFUFGAUFCFAUFGGCFUFCFAGAUFUFGAACFGCFUFGGCFGGCFAGACFGACFUCFGCFGCFGA | — | — | 73 | [ |
| 全菌 | WB-SELEX | TGGACCTTGCGATTGACAGCAGACATGAGTCAGGACGTGACCGCTGGACCTTGCGATTGACAGCAGACATGAGTCAGGAC | 15.16±3.62 | 流式细胞术 | 80 | [ |
| 全菌 | Cell-SELEX | CCCCCGTTGCTTTCGCTTTTCCTTTCGCTTTTGTTCGTTTCGTCCCTGCTTCCTTTCTTG | 14.55 | 地高辛标记适配体 | 60 | [ |
| 丁酰基-高丝氨酸 | Structure-switching SELEX | CCATCCACACTCCGCAAGTGGGGAGGGGAGAGACGACGATCCTGTGGGTTTTCTGCAGTGAGTCGTGTTTTCGACTTATTGCGTCGGCTGCCTCTACAT | 28.47 | 流式细胞术 | 99 | [ |

图2 用于铜绿假单胞菌检测的适配体传感器 A:用于铜绿假单胞菌检测的比色传感器;B:基于金纳米颗粒的电化学传感器用于铜绿假单胞菌检测;C:用于铜绿假单胞菌检测的夹心型电化学传感器; D:用于铜绿假单胞菌检测的荧光生物传感器
Fig. 2 Aptamer sensor for P. aeruginosa detection A: Colorimetric biosensor for P. aeruginosa detection. B: Gold nanoparticle-based electrochemical biosensor for P. aeruginosa detection. C: Sandwich electrochemical biosensor for P. aeruginosa detection. D: Fluorescent biosensor for P. aeruginosa detection
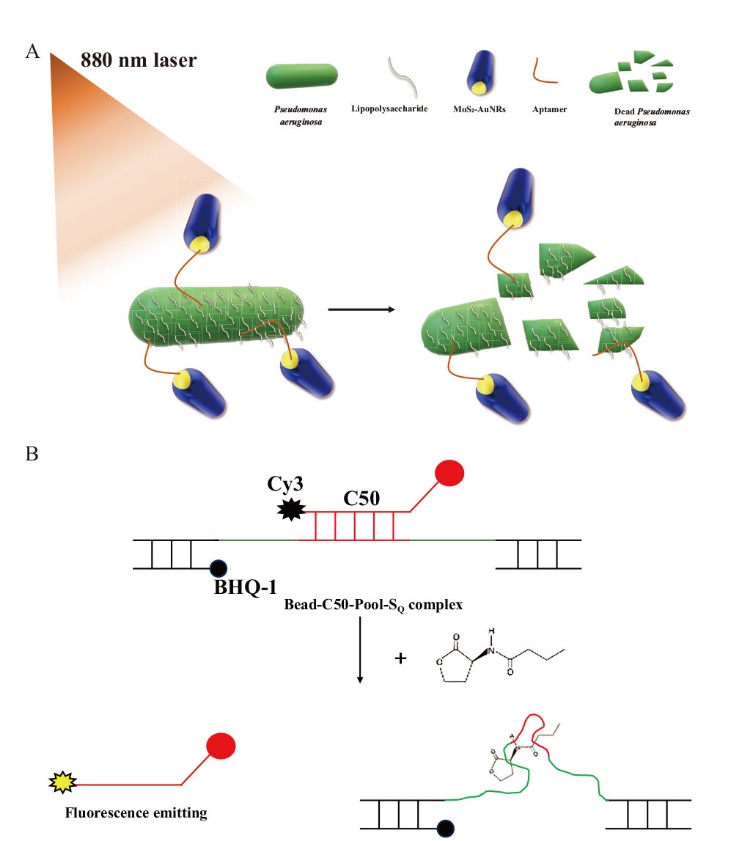
图3 基于适配体的铜绿假单胞菌防治 A:适配体偶联的金纳米棒靶向抑制铜绿假单胞菌;B:抑制铜绿假单胞菌的生物膜形成的适配体
Fig. 3 Aptamer-based control of P. aeruginosa A: Aptamer-conjugated gold nanorods that inhibit P. aeruginosa. B: Aptamers that inhibit biofilm formation of P. aeruginosa
| [1] |
Panayidou S, Georgiades K, Christofi T, et al. Pseudomonas aeruginosa core metabolism exerts a widespread growth-independent control on virulence[J]. Sci Rep, 2020, 10(1): 9505.
doi: 10.1038/s41598-020-66194-4 pmid: 32528034 |
| [2] |
Hu PW, Chen JY, Chen YH, et al. Molecular epidemiology, resistance, and virulence properties of Pseudomonas aeruginosa cross-colonization clonal isolates in the non-outbreak setting[J]. Infect Genet Evol, 2017, 55: 288-296.
doi: 10.1016/j.meegid.2017.09.010 URL |
| [3] |
Ugwuanyi FC, Ajayi A, Ojo DA, et al. Evaluation of efflux pump activity and biofilm formation in multidrug resistant clinical isolates of Pseudomonas aeruginosa isolated from a Federal Medical Center in Nigeria[J]. Ann Clin Microbiol Antimicrob, 2021, 20(1): 11.
doi: 10.1186/s12941-021-00417-y |
| [4] |
Patel BM, Paratz JD, Mallet A, et al. Characteristics of bloodstream infections in burn patients: an 11-year retrospective study[J]. Burns, 2012, 38(5): 685-690.
doi: 10.1016/j.burns.2011.12.018 pmid: 22348799 |
| [5] |
Thi MTT, Wibowo D, Rehm BHA. Pseudomonas aeruginosa biofilms[J]. Int J Mol Sci, 2020, 21(22): 8671.
doi: 10.3390/ijms21228671 URL |
| [6] |
Jurado-Martín I, Sainz-Mejías M, McClean S. Pseudomonas aeruginosa: an audacious pathogen with an adaptable arsenal of virulence factors[J]. Int J Mol Sci, 2021, 22(6): 3128.
doi: 10.3390/ijms22063128 URL |
| [7] |
Sainz-Mejías M, Jurado-Martín I, McClean S. Understanding Pseudomonas aeruginosa-host interactions: the ongoing quest for an efficacious vaccine[J]. Cells, 2020, 9(12): 2617.
doi: 10.3390/cells9122617 URL |
| [8] |
Ellington AD, Szostak JW. In vitro selection of RNA molecules that bind specific ligands[J]. Nature, 1990, 346(6287): 818-822.
doi: 10.1038/346818a0 |
| [9] |
Qian SW, Chang DR, He SS, et al. Aptamers from random sequence space: Accomplishments, gaps and future considerations[J]. Anal Chim Acta, 2022, 1196: 339511.
doi: 10.1016/j.aca.2022.339511 URL |
| [10] |
Xu WT, He WC, Du ZH, et al. Functional nucleic acid nanomaterials: development, properties, and applications[J]. Angew Chem Int Ed Engl, 2021, 60(13): 6890-6918.
doi: 10.1002/anie.v60.13 URL |
| [11] |
李天顺, 李宸葳, 王佳, 等. 功能核酸筛选过程中次级文库的有效制备[J]. 生物技术通报, 2023, 39(3): 116-122.
doi: 10.13560/j.cnki.biotech.bull.1985.2022-0743 |
| Li TS, Li CW, Wang J, et al. Efficient generation of secondary libraries during functional nucleic acids screening[J]. Biotechnol Bull, 2023, 39(3): 116-122. | |
| [12] |
Qi S, Duan N, Khan IM, et al. Strategies to manipulate the performance of aptamers in SELEX, post-SELEX and microenvironment[J]. Biotechnol Adv, 2022, 55: 107902.
doi: 10.1016/j.biotechadv.2021.107902 URL |
| [13] |
Darmostuk M, Rimpelova S, Gbelcova H, et al. Current approaches in SELEX: an update to aptamer selection technology[J]. Biotechnol Adv, 2015, 33(6 Pt 2): 1141-1161.
doi: 10.1016/j.biotechadv.2015.02.008 pmid: 25708387 |
| [14] |
Wang KY, Zeng YL, Yang XY, et al. Utility of aptamer-fluorescence in situ hybridization for rapid detection of Pseudomonas aeruginosa[J]. Eur J Clin Microbiol Infect Dis, 2011, 30(2): 273-278.
doi: 10.1007/s10096-010-1074-0 URL |
| [15] |
Soundy J, Day D. Selection of DNA aptamers specific for live Pseudomonas aeruginosa[J]. PLoS One, 2017, 12(9): e0185385.
doi: 10.1371/journal.pone.0185385 URL |
| [16] | Hong KL, Yancey K, Battistella L, et al. Selection of single-stranded DNA molecular recognition elements against exotoxin A using a novel decoy-SELEX method and sensitive detection of exotoxin A in human serum[J]. Biomed Res Int, 2015, 2015: 417641. |
| [17] |
Ye H, Duan N, Gu HJ, et al. Fluorometric determination of lipopolysaccharides via changes of the graphene oxide-enhanced fluorescence polarization caused by truncated aptamers[J]. Mikrochim Acta, 2019, 186(3): 173.
doi: 10.1007/s00604-019-3261-8 pmid: 30771102 |
| [18] |
Davydova AS, Vorobyeva MA, Kabilov MR, et al. In vitro selection of cell-internalizing 2'-modified RNA aptamers against Pseudomonas aeruginosa[J]. Russ J Bioorg Chem, 2017, 43(1): 58-63.
doi: 10.1134/S1068162016060030 URL |
| [19] |
Wang HY, Chi Z, Cong Y, et al. Development of a fluorescence assay for highly sensitive detection of Pseudomonas aeruginosa based on an aptamer-carbon dots/graphene oxide system[J]. RSC Adv, 2018, 8(57): 32454-32460.
doi: 10.1039/C8RA04819C URL |
| [20] | 曾燕丽, 兰小鹏, 江丽, 等. 灭活铜绿假单胞菌适体的筛选[J]. 中国生物化学与分子生物学报, 2009, 25(1): 90-97. |
| Zeng YL, Lan XP, Jiang L, et al. Selection of aptamers specifically binding to inactivate Pseudomonas aeruginosa[J]. Chin J Biochem Mol Biol, 2009, 25(1): 90-97. | |
| [21] |
Zhao M, Li WB, Liu KC, et al. C4-HSL aptamers for blocking qurom sensing and inhibiting biofilm formation in Pseudomonas aeruginosa and its structure prediction and analysis[J]. PLoS One, 2019, 14(2): e0212041.
doi: 10.1371/journal.pone.0212041 URL |
| [22] |
Tang YJ, Ali Z, Zou J, et al. Detection methods for Pseudomonas aeruginosa: history and future perspective[J]. RSC Adv, 2017, 7(82): 51789-51800.
doi: 10.1039/C7RA09064A URL |
| [23] |
Deschaght P, Van Daele S, De Baets F, et al. PCR and the detection of Pseudomonas aeruginosa in respiratory samples of CF patients. A literature review[J]. J Cyst Fibros, 2011, 10(5): 293-297.
doi: 10.1016/j.jcf.2011.05.004 pmid: 21684819 |
| [24] |
Schmitz FRW, Cesca K, Valério A, et al. Colorimetric detection of Pseudomonas aeruginosa by aptamer-functionalized gold nanoparticles[J]. Appl Microbiol Biotechnol, 2023, 107(1): 71-80.
doi: 10.1007/s00253-022-12283-5 |
| [25] |
Das R, Dhiman A, Kapil A, et al. Aptamer-mediated colorimetric and electrochemical detection of Pseudomonas aeruginosa utilizing peroxidase-mimic activity of gold NanoZyme[J]. Anal Bioanal Chem, 2019, 411(6): 1229-1238.
doi: 10.1007/s00216-018-1555-z |
| [26] |
Abedi R, Bakhsh Raoof J, Mohseni M, et al. Sandwich-type electrochemical aptasensor for highly sensitive and selective detection of Pseudomonas aeruginosa bacteria using a dual signal amplification strategy[J]. Bioelectrochemistry, 2023, 150:108332.
doi: 10.1016/j.bioelechem.2022.108332 URL |
| [27] |
Roushani M, Sarabaegi M, Pourahmad F. Impedimetric aptasensor for Pseudomonas aeruginosa by using a glassy carbon electrode modified with silver nanoparticles[J]. Mikrochim Acta, 2019, 186(11): 725.
doi: 10.1007/s00604-019-3858-y pmid: 31655899 |
| [28] |
Xie YX, Xie GM, Yuan JS, et al. A novel fluorescence biosensor based on double-stranded DNA branch migration-induced HCR and DNAzyme feedback circuit for sensitive detection of Pseudomonas aeruginosa(clean version)[J]. Anal Chim Acta, 2022, 1232: 340449.
doi: 10.1016/j.aca.2022.340449 URL |
| [29] |
Hu JY, Fu KY, Bohn PW. Whole-cell Pseudomonas aeruginosa localized surface plasmon resonance aptasensor[J]. Anal Chem, 2018, 90(3): 2326-2332.
doi: 10.1021/acs.analchem.7b04800 URL |
| [30] |
Zhong ZT, Gao XM, Gao R, et al. Selective capture and sensitive fluorometric determination of Pseudomonas aeruginosa by using aptamer modified magnetic nanoparticles[J]. Mikrochim Acta, 2018, 185(8): 377.
doi: 10.1007/s00604-018-2914-3 |
| [31] |
Mayr FB, Yende S, Angus DC. Epidemiology of severe sepsis[J]. Virulence, 2014, 5(1): 4-11.
doi: 10.4161/viru.27372 pmid: 24335434 |
| [32] |
Li J, Zhang YZ, Zhu LJ, et al. Smart nucleic acid hydrogels with high stimuli-responsiveness in biomedical fields[J]. Int J Mol Sci, 2022, 23(3): 1068.
doi: 10.3390/ijms23031068 URL |
| [33] |
Yang M, Chen X, Zhu LJ, et al. Aptamer-functionalized DNA-silver nanocluster nanofilm for visual detection and elimination of bacteria[J]. ACS Appl Mater Interfaces, 2021, 13(32): 38647-38655.
doi: 10.1021/acsami.1c05751 URL |
| [34] |
Rolband L, Yourston L, Chandler M, et al. DNA-templated fluorescent silver nanoclusters inhibit bacterial growth while being non-toxic to mammalian cells[J]. Molecules, 2021, 26(13): 4045.
doi: 10.3390/molecules26134045 URL |
| [35] | Lv J, Li B, Luo TT, et al. Selective photothermal therapy based on lipopolysaccharide aptamer functionalized MoS2 nanosheet-coated gold nanorods for multidrug-resistant Pseudomonas aeruginosa infection[J]. Adv Healthc Mater, 2023, 12(15): e2202794. |
| [36] |
Kraemer M, Bellion M, Kissmann AK, et al. Aptamers as novel binding molecules on an antimicrobial peptide-armored composite hydrogel wound dressing for specific removal and efficient eradication of Pseudomonas aeruginosa[J]. Int J Mol Sci, 2023, 24(5):4800.
doi: 10.3390/ijms24054800 URL |
| [37] |
Soundy J, Day D. Delivery of antibacterial silver nanoclusters to Pseudomonas aeruginosa using species-specific DNA aptamers[J]. J Med Microbiol, 2020, 69(4): 640-652.
doi: 10.1099/jmm.0.001174 URL |
| [1] | 谢宏, 周丽莹, 李舒文, 王梦迪, 艾晔, 晁跃辉. 蒺藜苜蓿MtCIM基因结构和功能分析[J]. 生物技术通报, 2024, 40(1): 262-269. |
| [2] | 温晓蕾, 李建嫄, 李娜, 张娜, 杨文香. 小麦叶锈菌与小麦互作的酵母双杂交cDNA文库构建与应用[J]. 生物技术通报, 2023, 39(9): 136-146. |
| [3] | 吴巧茵, 施友志, 李林林, 彭政, 谭再钰, 刘利平, 张娟, 潘勇. 类胡萝卜素降解菌株的原位筛选及其在雪茄提质增香中的应用[J]. 生物技术通报, 2023, 39(9): 192-201. |
| [4] | 江海溶, 崔若琪, 王玥, 白淼, 张明露, 任连海. NH3和H2S降解功能菌的分离鉴定及降解特性研究[J]. 生物技术通报, 2023, 39(9): 246-254. |
| [5] | 苗永美, 苗翠苹, 于庆才. 枯草芽孢杆菌BBs-27发酵液性质及脂肽对黄色镰刀菌的抑菌作用[J]. 生物技术通报, 2023, 39(9): 255-267. |
| [6] | 薛宁, 王瑾, 李世新, 刘叶, 程海娇, 张玥, 毛雨丰, 王猛. 多基因同步调控结合高通量筛选构建高产L-苯丙氨酸的谷氨酸棒杆菌工程菌株[J]. 生物技术通报, 2023, 39(9): 268-280. |
| [7] | 张岳一, 兰社益, 裴海闰, 封棣. 多菌种联用发酵燕麦麸皮工艺优化及发用功效评价[J]. 生物技术通报, 2023, 39(9): 58-70. |
| [8] | 马俊秀, 吴皓琼, 姜威, 闫更轩, 胡基华, 张淑梅. 蔬菜软腐病菌广谱拮抗细菌菌株筛选鉴定及防效研究[J]. 生物技术通报, 2023, 39(7): 228-240. |
| [9] | 谢东, 汪流伟, 李宁健, 李泽霖, 徐子航, 张庆华. 一株多功能菌株的发掘、鉴定及解磷条件优化[J]. 生物技术通报, 2023, 39(7): 241-253. |
| [10] | 陈彩萍, 任昊, 龙腾飞, 何冰, 鲁兆祥, 孙坚. 大肠杆菌Nissle 1917对炎症性肠病治疗作用的研究进展[J]. 生物技术通报, 2023, 39(6): 109-118. |
| [11] | 李典典, 粟元, 李洁, 许文涛, 朱龙佼. 抗菌适配体的筛选与应用进展[J]. 生物技术通报, 2023, 39(6): 126-132. |
| [12] | 董聪, 高庆华, 王玥, 罗同阳, 王庆庆. 基于联合策略提高FAD依赖的葡萄糖脱氢酶的酵母表达[J]. 生物技术通报, 2023, 39(6): 316-324. |
| [13] | 张雪萍, 鲁雨晴, 张月倩, 李晓娟. 植物细胞外囊泡及其分析技术的进展[J]. 生物技术通报, 2023, 39(5): 32-43. |
| [14] | 李天顺, 李宸葳, 王佳, 朱龙佼, 许文涛. 功能核酸筛选过程中次级文库的有效制备[J]. 生物技术通报, 2023, 39(3): 116-122. |
| [15] | 钟朝滨, 朱龙佼, 张博洋, 张洋子, 陈可仁, 许文涛. 细胞特异性核酸适配体在疾病治疗中的应用[J]. 生物技术通报, 2023, 39(12): 118-127. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||