








• 研究报告 •
杨悦1( ), 李昌宁1, 孙晓露1, 李雅迁1, 杜维俊1, 王利祥1, 王敏1(
), 李昌宁1, 孙晓露1, 李雅迁1, 杜维俊1, 王利祥1, 王敏1( ), 姬月梅2
), 姬月梅2
收稿日期:2025-04-02
出版日期:2025-12-11
通讯作者:
王敏,女,副教授,研究方向 :大豆遗传与种质创新;E-mail: wangmin3502@126.com作者简介:杨悦,女,硕士研究生,研究方向 :大豆遗传育种;E-mail: 1297720641@qq.com
基金资助:
YANG Yue1( ), LI Chang-ning1, SUN Xiao-lu1, LI Ya-qian1, DU Wei-jun1, WANG Li-xiang1, WANG Min1(
), LI Chang-ning1, SUN Xiao-lu1, LI Ya-qian1, DU Wei-jun1, WANG Li-xiang1, WANG Min1( ), JI Yue-mei2
), JI Yue-mei2
Received:2025-04-02
Published:2025-12-11
摘要:
目的 MYB(v-MYB avian myeloblastosis viral oncogene homolog)转录因子在植物生长发育及盐胁迫响应过程中具有重要的调控作用。大豆GmMYB069参与盐胁迫响应过程。克隆大豆GmMYB069并分析其耐盐功能,为大豆耐盐育种提供分子基础和基因资源。 方法 克隆GmMYB069的全长CDS序列,利用生物信息学软件对其基因序列和氨基酸序列特征进行分析。构建GmMYB069过表达载体,利用农杆菌介导法转化大豆。对过表达株系的耐盐表型和生理生化指标进行分析。 结果 成功克隆了长度为1 059 bp的GmMYB069 CDS序列,编码352个氨基酸,相对分子量为39 309.26 Da,预测等电点为6.76;其蛋白为不稳定的亲水性蛋白,多肽链中无规则卷曲占比最大,没有跨膜结构与信号肽,与密花豆的亲缘关系最近,与蔓花生的亲缘关系最远。GmMYB069在根中的表达量较高,其次为茎、叶。在盐胁迫逆境条件下,GmMYB069基因在胁迫前期表达量呈上升趋势,可达到峰值,随着胁迫时间的延长,其表达量下降,表明其在早期可以响应盐胁迫。成功构建了GmMYB069过表达载体,并进行大豆根毛转化,获得过表达嵌合株系。在盐胁迫下,与空载植株相比,过表达株系的相对表达量显著提高,过表达嵌合植株叶片中抗氧化酶(SOD、POD)升高,MDA减少,Na+、K+含量在过表达嵌合植株根中无显著差异,Na+含量在过表达嵌合植株叶片中显著降低,K+则相反。 结论 过表达大豆GmMYB069可以增强抗氧化酶活性,阻止Na+从根向叶的运输,减少植物的盐害,提高转基因植株的耐盐性。
杨悦, 李昌宁, 孙晓露, 李雅迁, 杜维俊, 王利祥, 王敏, 姬月梅. 大豆转录因子GmMYB069的克隆与耐盐功能鉴定[J]. 生物技术通报, doi: 10.13560/j.cnki.biotech.bull.1985.2025-0348.
YANG Yue, LI Chang-ning, SUN Xiao-lu, LI Ya-qian, DU Wei-jun, WANG Li-xiang, WANG Min, JI Yue-mei. Cloning and Identification of Salt Tolerance Function of Soybean Transcription Factor GmMYB069[J]. Biotechnology Bulletin, doi: 10.13560/j.cnki.biotech.bull.1985.2025-0348.
引物名称 Primer name | 引物序列 Primer sequence(5′-3′) |
|---|---|
| GmMYB069-F | ATGGGAAGGTCACCGTGTTG |
| GmMYB069-R | CCACAAAATTGGTGACGCAGAA |
| qRT-GmCYP2-F | CGGGACCAGTGTGCTTCTTCA |
| qRT-GmCYP2-R | CCCCTCCACTACAAAGGCTCG |
| qRT-GmMYB069-F | GCCTCATGTGGTGTTGAGTG |
| qRT-GmMYB069-R | TACAGCCACTGTCCATTTGGT |
| GmMYB069-pUBi-F | TTGATGTGATTACAGTCTAGAATGGGAAGGTCACCGTGTTG |
| GmMYB069-pUBi-R | GTTAATTAACCCGCTGGTACCCTACCACAAAATTGGTGACGCA |
表1 GmMYB069引物序列
Table 1 GmMYB069 primer sequence
引物名称 Primer name | 引物序列 Primer sequence(5′-3′) |
|---|---|
| GmMYB069-F | ATGGGAAGGTCACCGTGTTG |
| GmMYB069-R | CCACAAAATTGGTGACGCAGAA |
| qRT-GmCYP2-F | CGGGACCAGTGTGCTTCTTCA |
| qRT-GmCYP2-R | CCCCTCCACTACAAAGGCTCG |
| qRT-GmMYB069-F | GCCTCATGTGGTGTTGAGTG |
| qRT-GmMYB069-R | TACAGCCACTGTCCATTTGGT |
| GmMYB069-pUBi-F | TTGATGTGATTACAGTCTAGAATGGGAAGGTCACCGTGTTG |
| GmMYB069-pUBi-R | GTTAATTAACCCGCTGGTACCCTACCACAAAATTGGTGACGCA |
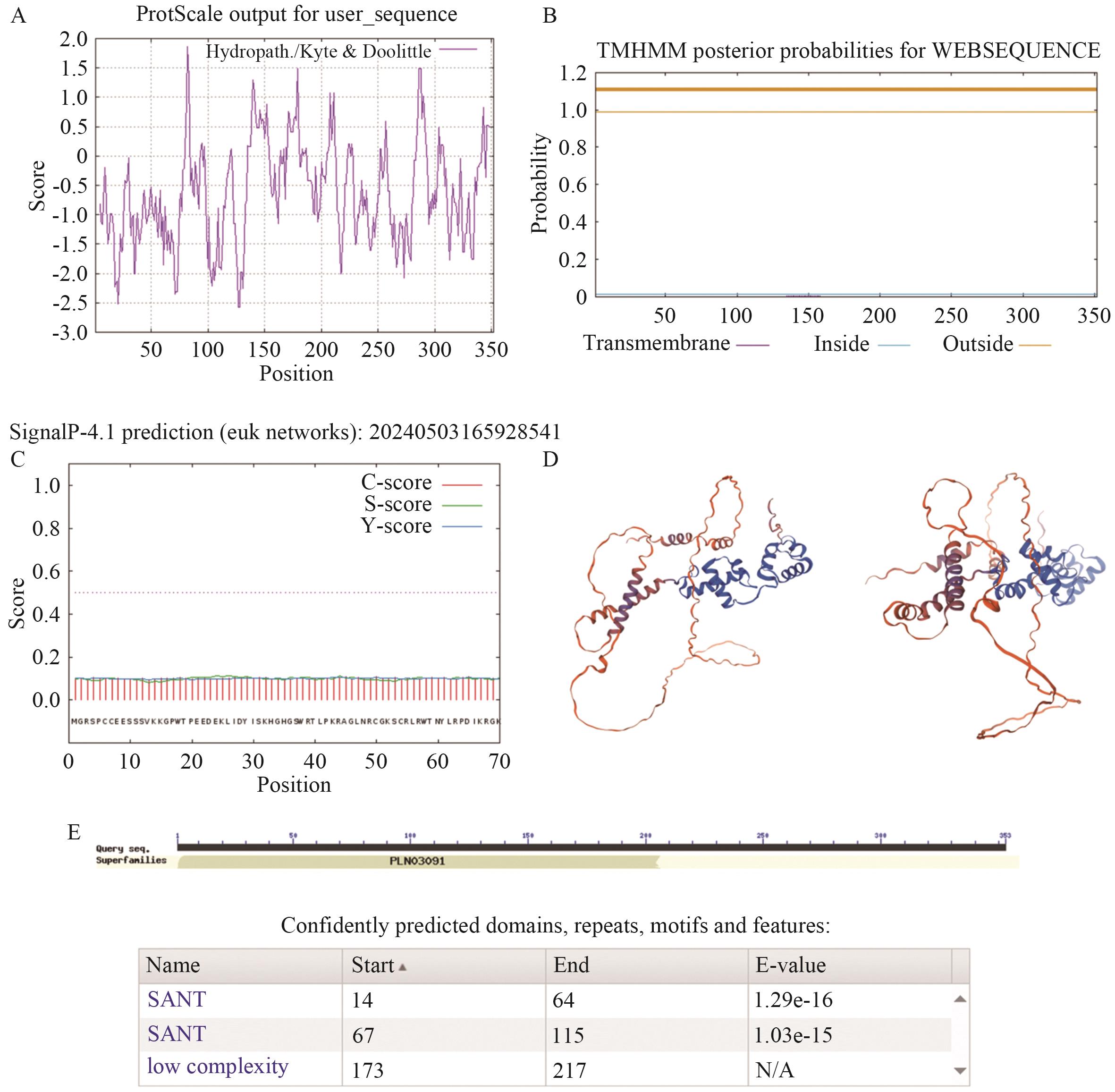
图5 GmMYB069蛋白特性分析A. GmMYB069蛋白疏水性预测图;B. GmMYB069蛋白跨膜结构域预测图;C. GmMYB069蛋白的信号肽分析(横坐标表示氨基酸的位置,纵坐标表示信号肽的概率);D. GmMYB069蛋白的保守结构域;E. GmMYB069蛋白质三级结构
Fig. 5 Characterization of GmMYB069 proteinA. GmMYB069 protein hydrophobicity prediction map. B. GmMYB069 protein transmembrane domain prediction diagram. C. Signal peptide analysis of GmMYB069 protein (The horizontal coordinate indicates the position of the amino acid, and the vertical coordinate indicates the probability of the signal peptide). D. Conservative domains of GmMYB069 protein. E. Tertiary structure of GmMYB069 protein
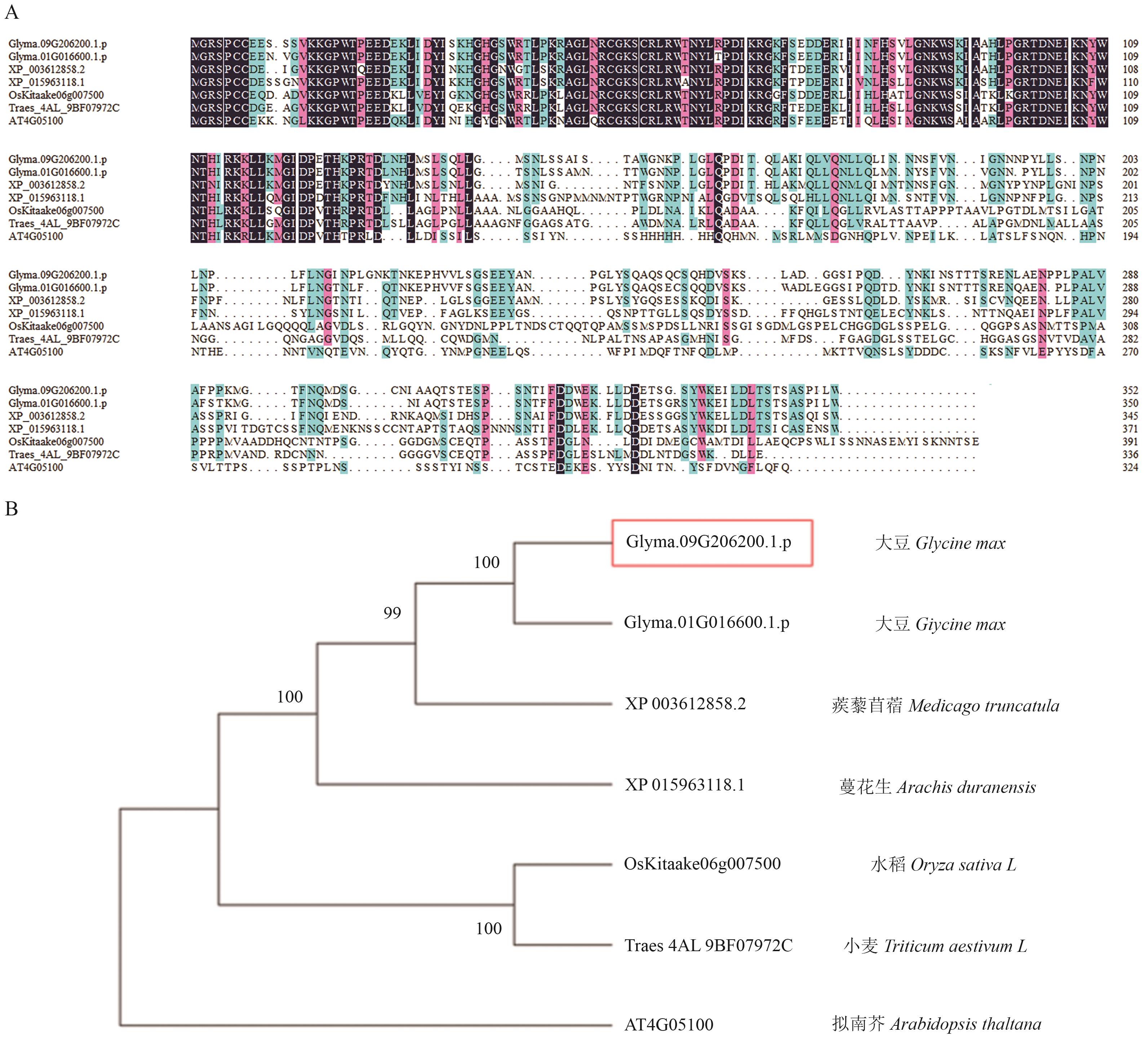
图6 GmMYB069与其他物种MYB蛋白多重序列比对(A)以及系统发育树(B)
Fig. 6 Multiple sequence alignment of GmMYB069 with MYB proteins from other species (A) and phylogenetic tree (B)
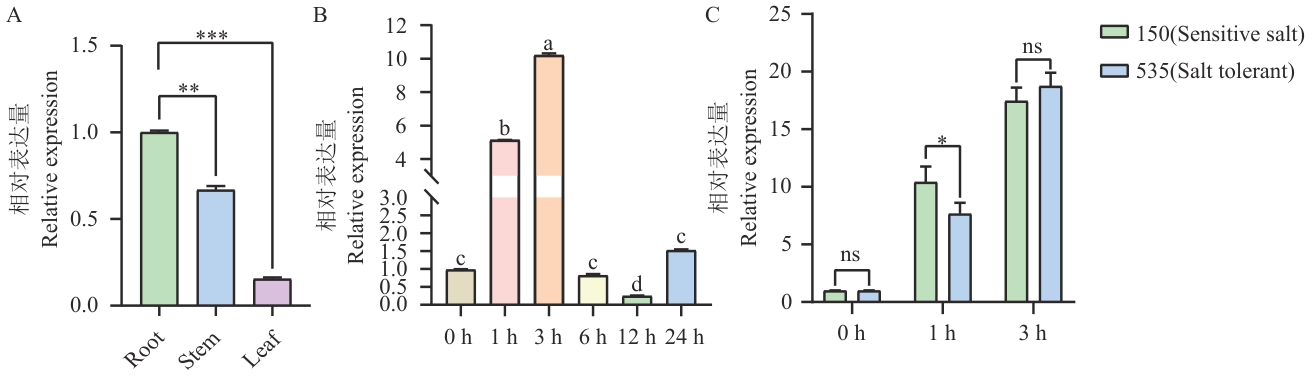
图7 大豆时空表达模式图A. 大豆不同组织GmMYB069基因表达量;B. 盐胁迫下大豆W82不同时间段GmMYB069基因表达量;C. 在150和535两种材料的不同时间段GmMYB069基因表达量。所示数据为平均数±方差,显著性差异使用t检验分析,ns表示无显著差异,*P<0.05,**P<0.01,***P<0.001,下同
Fig. 7 Temporal and spatial expression pattern diagram of soybeanA. Expressions of GmMYB069 gene in different tissues of soybean. B. The expressions of GmMYB069 gene in soybean W82 under salt stress at different time periods. C. The expressions of GmMYB069 gene at different time periods in material 150 and 535. The data shown is mean±variance, and significant differences were analyzed using t-test. ns indicates no significant difference, *P<0.05, **P<0.01, and ***P<0.001, the same below
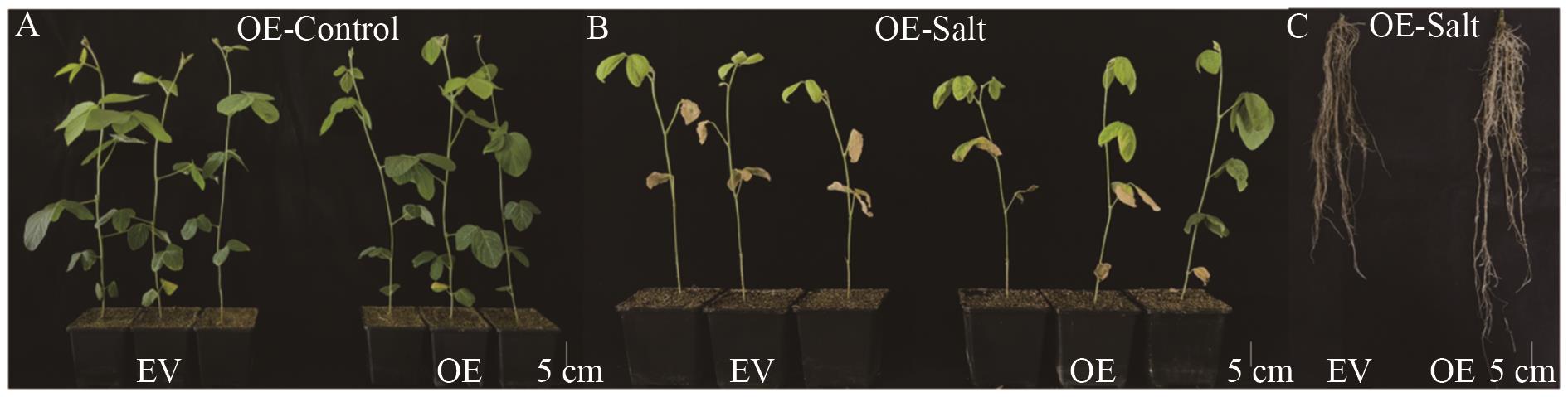

图9 基因GmMYB069过表达表型分析A:水处理下GmMYB069-OE和对照EV的表型(OE-Control:水处理下的过表达嵌合植株);B:盐处理下GmMYB069-OE和对照EV的表型(OE-Salt:盐处理下的过表达嵌合植株);C:盐胁迫下GmMYB069-OE和对照EV地下部表型(OE-Salt:盐处理下的过表达嵌合植株)
Fig. 9 Phenotypic analysis of GmMYB069 OverexpressionA. Phenotypes of GmMYB069-OE and control EV under water treatment (OE-Control: Overexpressed chimeric plants under water treatment). B. Phenotypes of GmMYB069-OE and control EV under salt treatment (OE-Salt: Overexpressed chimeric plants under salt treatment). C. Subterranean phenotype of GmMYB069-OE and control EV under salt stress (OE-Salt: Overexpressed chimeric plants under salt treatment)
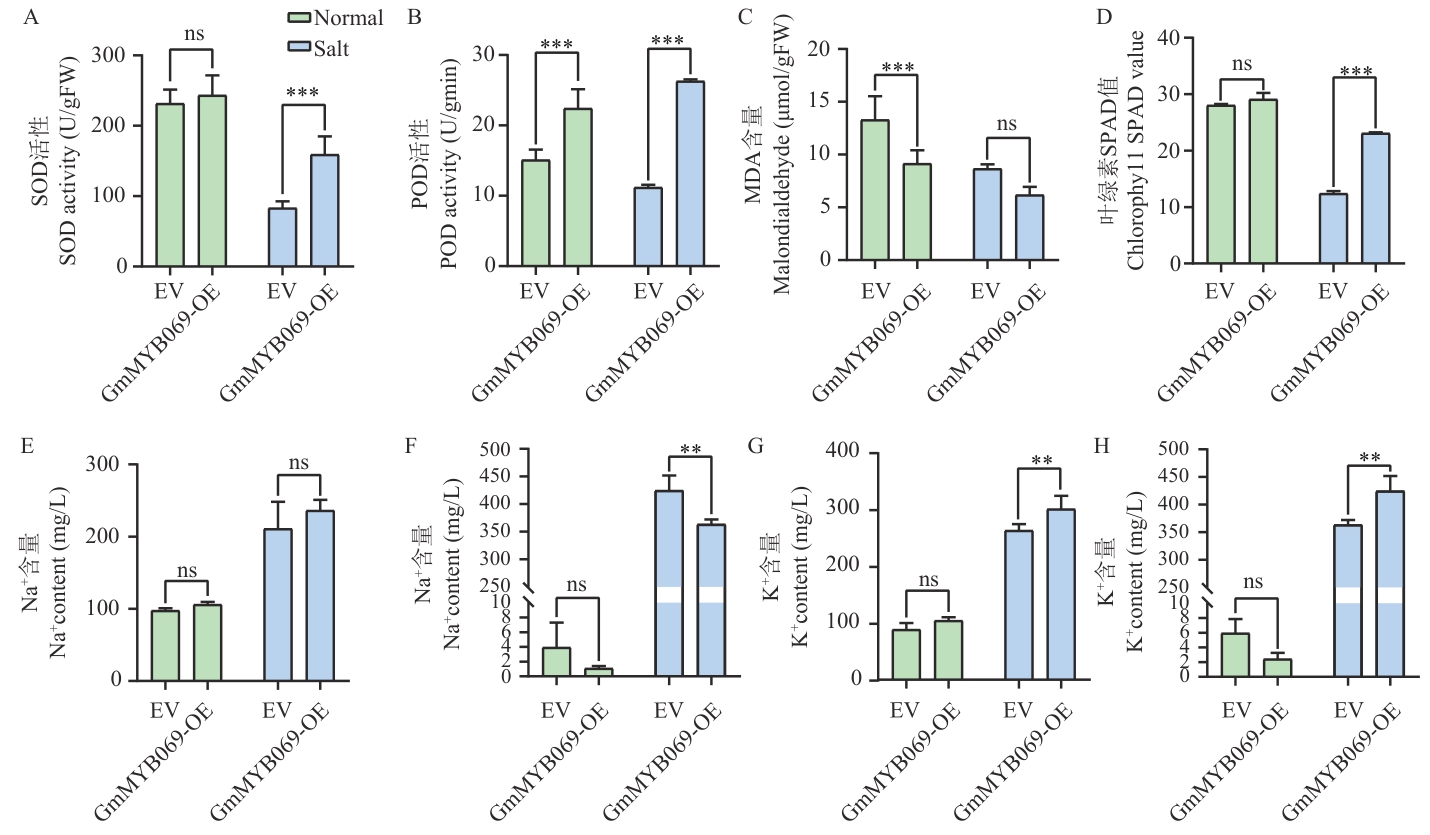
图10 盐胁迫下GmMYB069过表达嵌合植株的生理指标和Na+、K+含量A:GmMYB069-OE叶片SOD活性;B:GmMYB069-OE叶片POD活性;C:GmMYB069-OE叶片MDA含量;D:GmMYB069-OE叶片SPAD值;E:GmMYB069-OE植株在根中的Na+含量;F:GmMYB069-OE植株在叶中的Na+含量;G:GmMYB069-OE植株在根中的K+含量;H:GmMYB069-OE植株在叶中的K+含量
Fig. 10 Physiological indicators and Na+, K+ contents of GmMYB069 overexpressing chimeric plants under salt stressA: SOD activity in GmMYB069-OE leaves. B: POD activity in GmMYB069-OE leaves. C: MDA content in GmMYB069-OE leaves. D: SPAD value of GmMYB069-OE leaves. E: Na+ content in the roots of GmMYB069-OE plants. F: Na+ content in the leaves of GmMYB069-OE plants. G: K+ content in the roots of GmMYB069-OE plants. H: K+ content in the leaves of GmMYB069-OE plants
| [1] | 朱子坤, 马超, 王锦辉, 等. 大豆GmNARK基因家族鉴定及分析 [J]. 大豆科技, 2023(2): 10-20. |
| Zhu ZK, Ma C, Wang JH, et al. Identification and functional analysis of GmNARK gene family in soybean [J]. Soybean Sci & Technol, 2023(2): 10-20. | |
| [2] | Cheng Q, Gan ZR, Wang YP, et al. The soybean gene J contributes to salt stress tolerance by up-regulating salt-responsive genes [J]. Front Plant Sci, 2020, 11: 272. |
| [3] | 王燕. 樱桃李(Prunus cerasifera Ehrh .)果实主要花色苷组分及相关特性分析 [D]. 泰安: 山东农业大学, 2013. |
| Wang Y. Analysis of the main anthocyanin components and related characteristics of cherry plum(Prunus cerasifera Ehrh .)fruit [D]. Taian: Shandong Agricultural University, 2013. | |
| [4] | Aoyagi LN, Lopes-Caitar VS, de Carvalho MCCG, et al. Genomic and transcriptomic characterization of the transcription factor family R2R3-MYB in soybean and its involvement in the resistance responses to Phakopsora pachyrhizi [J]. Plant Sci, 2014, 229: 32-42. |
| [5] | Liao Y, Zou HF, Wang HW, et al. Soybean GmMYB76, GmMYB92, and GmMYB177 genes confer stress tolerance in transgenic Arabidopsis plants [J]. Cell Res, 2008, 18(10): 1047-1060. |
| [6] | Niu CF, Wei W, Zhou QY, et al. Wheat WRKY genes TaWRKY2 and TaWRKY19 regulate abiotic stress tolerance in transgenic Arabidopsis plants [J]. Plant Cell Environ, 2012, 35(6): 1156-1170. |
| [7] | Hao YJ, Wei W, Song QX, et al. Soybean NAC transcription factors promote abiotic stress tolerance and lateral root formation in transgenic plants [J]. Plant J, 2011, 68(2): 302-313. |
| [8] | Seo YJ, Park JB, Cho YJ, et al. Overexpression of the ethylene-responsive factor gene BrERF4 from Brassica rapa increases tolerance to salt and drought in Arabidopsis plants [J]. Mol Cells, 2010, 30(3): 271-278. |
| [9] | Jeong JS, Kim YS, Baek KH, et al. Root-specific expression of OsNAC10 improves drought tolerance and grain yield in rice under field drought conditions [J]. Plant Physiol, 2010, 153(1): 185-197. |
| [10] | Zou XL, Neuman D, Shen QJ. Interactions of two transcriptional repressors and two transcriptional activators in modulating gibberellin signaling in aleurone cells [J]. Plant Physiol, 2008, 148(1): 176-186. |
| [11] | Liao Y, Zou HF, Wei W, et al. Soybean GmbZIP44, GmbZIP62 and GmbZIP78 genes function as negative regulator of ABA signaling and confer salt and freezing tolerance in transgenic Arabidopsis [J]. Planta, 2008, 228(2): 225-240. |
| [12] | Nakashima K, Tran LP, Van Nguyen D, et al. Functional analysis of a NAC-type transcription factor OsNAC6 involved in abiotic and biotic stress-responsive gene expression in rice [J]. Plant J, 2007, 51(4): 617-630. |
| [13] | Hu HH, Dai MQ, Yao JL, et al. Overexpressing a NAM, ATAF, and CUC (NAC) transcription factor enhances drought resistance and salt tolerance in rice [J]. Proc Natl Acad Sci U S A, 2006, 103(35): 12987-12992. |
| [14] | Tran LP, Nakashima K, Sakuma Y, et al. Isolation and functional analysis of Arabidopsis stress-Inducible NAC transcription factors that bind to a drought-responsivecis-element in theearly responsive to dehydration stress 1 promoter W [J]. Plant Cell, 2004, 16(9): 2481-2498. |
| [15] | DuanMu HZ, Wang Y, Bai X, et al. Wild soybean roots depend on specific transcription factors and oxidation reduction related genesin response to alkaline stress [J]. Funct Integr Genom, 2015, 15(6): 651-660. |
| [16] | Gonzalez A, Zhao MZ, Leavitt JM, et al. Regulation of the anthocyanin biosynthetic pathway by the TTG1/bHLH/Myb transcriptional complex in Arabidopsis seedlings [J]. Plant J, 2008, 53(5): 814-827. |
| [17] | Rabinowicz PD, Braun EL, Wolfe AD, et al. Maize R2R3 myb genes: sequence analysis reveals amplification in the higher plants [J]. Genetics, 1999, 153(1): 427-444. |
| [18] | Du H, Yang SS, Liang Z, et al. Genome-wide analysis of the MYB transcription factor superfamily in soybean [J]. BMC Plant Biol, 2012, 12(1): 106. |
| [19] | 赵盼盼. 番茄R2R3MYB转录因子家族鉴定及SlMYB41和SlMYB64基因功能研究 [D]. 泰安: 山东农业大学, 2017. |
| Zhao PP. Identification of tomato R2R3MYB transcription factor family and functional study of SlMYB41 and SlMYB64 genes [D]. Tai’an: Shandong Agricultural University, 2017. | |
| [20] | 王欣怡, 王晓倩, 刘亚玲, 等. 紫花苜蓿MsMYB33基因克隆、亚细胞定位及转录自激活检测 [J]. 草地学报, 2023, 31(5): 1331-1337. |
| Wang XY, Wang XQ, Liu YL, et al. Cloning, subcellular localization and autonomous transcriptional activation testing of MsMYB33 gene from Medicago sativa [J]. Acta Agrestia Sin, 2023, 31(5): 1331-1337. | |
| [21] | Stracke R, Werber M, Weisshaar B. The R2R3-MYB gene family in Arabidopsis thaliana [J]. Curr Opin Plant Biol, 2001, 4(5): 447-456. |
| [22] | Dubos C, Stracke R, Grotewold E, et al. MYB transcription factors in Arabidopsis [J]. Trends Plant Sci, 2010, 15(10): 573-581. |
| [23] | Zhang LC, Zhao GY, Jia JZ, et al. Molecular characterization of 60 isolated wheat MYB genes and analysis of their expression during abiotic stress [J]. J Exp Bot, 2012, 63(1): 203-214. |
| [24] | 牛义岭, 姜秀明, 许向阳. 植物转录因子MYB基因家族的研究进展 [J]. 分子植物育种, 2016, 14(8): 2050-2059. |
| Niu YL, Jiang XM, Xu XY. Reaserch advances on transcription factor MYB gene family in plant [J]. Mol Plant Breed, 2016, 14(8): 2050-2059. | |
| [25] | Wang XP, Niu YL, Zheng Y. Multiple functions of MYB transcription factors in abiotic stress responses [J]. Int J Mol Sci, 2021, 22(11): 6125. |
| [26] | Zhang P, Wang RL, Yang XP, et al. The R2R3-MYB transcription factor AtMYB49 modulates salt tolerance in Arabidopsis by modulating the cuticle formation and antioxidant defence [J]. Plant Cell Environ, 2020, 43(8): 1925-1943. |
| [27] | Tang YH, Bao XX, Zhi YL, et al. Overexpression of a MYB family gene, OsMYB6, increases drought and salinity stress tolerance in transgenic rice [J]. Front Plant Sci, 2019, 10: 168. |
| [28] | 石广成, 杨万明, 杜维俊, 等. 大豆耐盐种质的筛选及其耐盐生理特性分析 [J]. 生物技术通报, 2022, 38(4): 174-183. |
| Shi GC, Yang WM, Du WJ, et al. Screening of salt-tolerant soybean germplasm and physiological characteristics analysis of its salt tolerance [J]. Biotechnol Bull, 2022, 38(4): 174-183. | |
| [29] | 孙群, 胡景江. 植物生理学研究技术 [M]. 杨凌: 西北农林科技大学出版社, 2006. |
| Sun Q, Hu JJ. Research technology of plant physiology [M]. Yangling: Northwest A&F University Press, 2006. | |
| [30] | Matsushita N, Matoh T. Characterization of Na+ exclusion mechanisms of salt-tolerant reed plants in comparison with salt-sensitive rice plants [J]. Physiol Plant, 1991, 83(1): 170-176. |
| [31] | 赵团结, 盖钧镒. 栽培大豆起源与演化研究进展 [J]. 中国农业科学, 2004, 37(7): 954-962. |
| Zhao TJ, Gai JY. The origin and evolution of cultivated soybean [Glycine max (L.) merr. [J]. Sci Agric Sin, 2004, 37(7): 954-962. | |
| [32] | Qiu PC, Du YC, Song MG, et al. Genetic overlap between salt and low-temperature tolerance loci at germination stage of soybean [J]. Eng Sci, 2013, 11(5): 37-40. |
| [33] | Rossi M, Borromeo I, Capo C, et al. PGPB improve photosynthetic activity and tolerance to oxidative stress in Brassica napus grown on salinized soils [J]. Applied Sciences, 2021, 11(23): 11442. |
| [34] | Lotkowska ME, Tohge T, Fernie AR, et al. The Arabidopsis transcription factor MYB112 promotes anthocyanin formation during salinity and under high light stress [J]. Plant Physiol, 2015, 169(3): 1862-1880. |
| [35] | 王春荣, 王庆杰, 李晓玲, 等. 葡萄中盐诱导的R2R3-MYB基因的筛选与表达分析 [J]. 园艺学报, 2014, 41(3): 529-535. |
| Wang CR, Wang QJ, Li XL, et al. Screening and expression analysis of salt-induced R2R3-MYB genes in grapes [J]. Acta Horticulturae Sinica, 2014, 41(3): 529-535. | |
| [36] | Wang L, Qiu TQ, Yue JR, et al. Arabidopsis ADF1 is regulated by MYB73 and is involved in response to salt stress affecting actin filament organization [J]. Plant Cell Physiol, 2021, 62(9): 1387-1395. |
| [37] | He YX, Dong YS, Yang XD, et al. Functional activation of a novel R2R3-MYB protein gene, GmMYB68, confers salt-alkali resistance in soybean (Glycine max L.) [J]. Genome, 2020, 63(1): 13-26. |
| [38] | Zhang WX, Wang N, Yang JT, et al. The salt-induced transcription factor GmMYB84 confers salinity tolerance in soybean [J]. Plant Sci, 2020, 291: 110326. |
| [39] | Du YT, Zhao MJ, Wang CT, et al. Identification and characterization of GmMYB118 responses to drought and salt stress [J]. BMC Plant Biol, 2018, 18(1): 320. |
| [40] | Pi EX, Xu J, Li HH, et al. Enhanced salt tolerance of rhizobia-inoculated soybean correlates with decreased phosphorylation of the transcription factor GmMYB183 and altered flavonoid biosynthesis [J]. Mol Cell Proteom, 2019, 18(11): 2225-2243. |
| [41] | Pi EX, Zhu CM, Fan W, et al. Quantitative phosphoproteomic and metabolomic analyses reveal GmMYB173 optimizes flavonoid metabolism in soybean under salt stress [J]. Mol Cell Proteom, 2018, 17(6): 1209-1224. |
| [42] | Wang C, Wang LJ, Lei J, et al. IbMYB308, a sweet potato R2R3-MYB gene, improves salt stress tolerance in transgenic tobacco [J]. Genes, 2022, 13(8): 1476. |
| [43] | Zhang X, Chen LC, Shi QH, et al. SlMYB102, an R2R3-type MYB gene, confers salt tolerance in transgenic tomato [J]. Plant Sci, 2020, 291: 110356. |
| [44] | Ullah A, Ul Qamar MT, Nisar M, et al. Characterization of a novel cotton MYB gene, GhMYB108-like responsive to abiotic stresses [J]. Mol Biol Rep, 2020, 47(3): 1573-1581. |
| [45] | 刘佳欣, 刘慧子, 石晶静, 等. 白桦MYB基因响应激素及盐旱处理的表达研究 [J]. 植物研究, 2020, 40(5): 743-750. |
| Liu JX, Liu HZ, Shi JJ, et al. Expression of MYB genes of birch in response to hormones, salt and drought [J]. Bull Bot Res, 2020, 40(5): 743-750. | |
| [46] | Roy SJ, Negrão S, Tester M. Salt resistant crop plants [J]. Curr Opin Biotechnol, 2014, 26: 115-124. |
| [47] | Deinlein U, Stephan AB, Horie T, et al. Plant salt-tolerance mechanisms [J]. Trends Plant Sci, 2014, 19(6): 371-379. |
| [48] | Kar RK. Plant responses to water stress: Role of reactive oxygen species [J]. Plant Signal Behav, 2011, 6(11): 1741-1745. |
| [49] | You J, Chan ZL. ROS regulation during abiotic stress responses in crop plants [J]. Front Plant Sci, 2015, 6: 1092. |
| [50] | Xu ZY, Wang CY, Xue F, et al. Wheat NAC transcription factor TaNAC29 is involved in response to salt stress [J]. Plant Physiol Biochem, 2015, 96: 356-363. |
| [51] | 李妍. 盐和PEG胁迫对丝瓜幼苗抗氧化酶活性及丙二醛含量的影响 [J]. 干旱地区农业研究, 2009, 27(2): 159-162, 178. |
| Li Y. Effect of salt and PEG to antioxidant enzymes activity and MDA concentration of Luffa cylindrical Roem [J]. Agric Res Arid Areas, 2009, 27(2): 159-162, 178. |
| [1] | 程婷婷, 刘俊, 王利丽, 练从龙, 魏文君, 郭辉, 吴尧琳, 杨晶凡, 兰金旭, 陈随清. 杜仲查尔酮异构酶基因家族全基因组鉴定及其表达模式分析[J]. 生物技术通报, 2025, 41(9): 242-255. |
| [2] | 徐小萍, 杨成龙, 和兴, 郭文杰, 吴健, 方少忠. 百合LoAPS1克隆及其在休眠解除过程的功能分析[J]. 生物技术通报, 2025, 41(9): 195-206. |
| [3] | 董向向, 缪百灵, 许贺娟, 陈娟娟, 李亮杰, 龚守富, 朱庆松. 森林草莓FveBBX20基因的生物信息学分析及开花调控功能[J]. 生物技术通报, 2025, 41(9): 115-123. |
| [4] | 李珊, 马登辉, 马红义, 姚文孔, 尹晓. 葡萄SKP1基因家族鉴定与表达分析[J]. 生物技术通报, 2025, 41(9): 147-158. |
| [5] | 黄国栋, 邓宇星, 程宏伟, 但焱南, 周会汶, 吴兰花. 大豆ZIP基因家族鉴定及响应铝胁迫的表达分析[J]. 生物技术通报, 2025, 41(9): 71-81. |
| [6] | 巩慧玲, 邢玉洁, 马俊贤, 蔡霞, 冯再平. 马铃薯LAC基因家族的鉴定及盐胁迫下表达分析[J]. 生物技术通报, 2025, 41(9): 82-93. |
| [7] | 关陟昊, 单治易, 熊赫, 赵瑞雪. 基于计算文献的大豆耦合性状知识发现研究[J]. 生物技术通报, 2025, 41(9): 345-356. |
| [8] | 张永艳, 郭思健, 李晶, 郝思怡, 李瑞得, 刘嘉鹏, 程春振. 蓝莓花青素相关VcGSTF19基因的克隆及功能研究[J]. 生物技术通报, 2025, 41(9): 139-146. |
| [9] | 李亚涛, 张志鹏, 赵梦瑶, 吕镇, 甘恬, 魏浩, 吴书凤, 马玉超. 根瘤菌Bd1的全基因组分析及TetR3对细胞生长和结瘤的负调控功能[J]. 生物技术通报, 2025, 41(9): 289-301. |
| [10] | 腊贵晓, 赵玉龙, 代丹丹, 余永亮, 郭红霞, 史贵霞, 贾慧, 杨铁钢. 红花质膜H+-ATPase基因家族成员鉴定及响应低氮低磷胁迫的表达分析[J]. 生物技术通报, 2025, 41(8): 220-233. |
| [11] | 程雪, 付颖, 柴晓娇, 王红艳, 邓欣. 谷子LHC基因家族鉴定及非生物胁迫表达分析[J]. 生物技术通报, 2025, 41(8): 102-114. |
| [12] | 朱丽娟, 张锴, 温晓蕾, 褚佳豪, 史凤玉, 王艳丽. 基于WGCNA挖掘野生大豆耐镉关键基因[J]. 生物技术通报, 2025, 41(8): 124-136. |
| [13] | 赖诗雨, 梁巧兰, 魏列新, 牛二波, 陈应娥, 周鑫, 杨思正, 王博. NbJAZ3在苜蓿花叶病毒侵染本氏烟过程中的作用[J]. 生物技术通报, 2025, 41(8): 186-196. |
| [14] | 李雅琼, 格桑拉毛, 陈启迪, 杨宇环, 何花转, 赵耀飞. 异源过表达高粱SbSnRK2.1增强拟南芥对盐胁迫的抗性[J]. 生物技术通报, 2025, 41(8): 115-123. |
| [15] | 康琴, 汪霞, 谌明洋, 徐静天, 陈诗兰, 廖平杨, 许文志, 吴卫, 徐东北. 薄荷UV-B受体基因McUVR8的克隆与表达分析[J]. 生物技术通报, 2025, 41(8): 255-266. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||