








生物技术通报 ›› 2026, Vol. 42 ›› Issue (2): 317-324.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0792
王婷婷1( ), 何孟雅1, 盛家顺1, 高晨2, 蔡含芳1, 付彤1, 孙宇1, 高腾云1(
), 何孟雅1, 盛家顺1, 高晨2, 蔡含芳1, 付彤1, 孙宇1, 高腾云1( ), 张天留1(
), 张天留1( )
)
收稿日期:2025-07-23
出版日期:2026-02-26
发布日期:2026-03-17
通讯作者:
高腾云,男,博士生导师,研究方向 :动物生产与智慧牧业;E-mail: Dairyfarm@163.com作者简介:王婷婷,女,硕士研究生,研究方向 :分子生物学;E-mail: 16696736731@163.com
基金资助:
WANG Ting-ting1( ), HE Meng-ya1, SHENG Jia-shun1, GAO Chen2, CAI Han-fang1, FU Tong1, SUN Yu1, GAO Teng-yun1(
), HE Meng-ya1, SHENG Jia-shun1, GAO Chen2, CAI Han-fang1, FU Tong1, SUN Yu1, GAO Teng-yun1( ), ZHANG Tian-liu1(
), ZHANG Tian-liu1( )
)
Received:2025-07-23
Published:2026-02-26
Online:2026-03-17
摘要:
目的 利用CRISPR/Cas9技术构建NCAPG基因敲除的牛胎儿成纤维细胞模型,并探究NCAPG基因缺失对细胞活性的影响。 方法 在牛NCAPG基因转录本第6外显子区域设计特异性sgRNA,将其与SpCas9蛋白孵育形成核糖核蛋白(RNP)复合物,通过电转染技术将RNP复合物导入牛胎儿成纤维细胞。随后采用ClonePlus™技术进行单克隆分离培养,经PCR扩增和测序验证后成功获得NCAPG基因敲除的阳性单克隆细胞株。 结果 ICE软件分析表明,sgRNA的敲除效率达到49%;TA克隆测序结果证实,所获得的单克隆细胞均为杂合子,其靶位点存在5 bp的缺失突变;Western blot检测显示,与野生型细胞相比,敲除细胞株中NCAPG蛋白表达水平极显著降低(P<0.01);CCK-8细胞活性检测结果表明,NCAPG基因敲除显著抑制了细胞增殖能力(P<0.01),表现为细胞生长速率明显减缓。 结论 利用CRISPR/Cas9技术成功构建了牛胎儿成纤维细胞NCAPG基因高效敲除方法,并证实NCAPG基因缺失会显著抑制细胞活性,为深入研究牛NCAPG基因的功能及其分子机制提供了重要的实验材料。
王婷婷, 何孟雅, 盛家顺, 高晨, 蔡含芳, 付彤, 孙宇, 高腾云, 张天留. 基于CRISPR/Cas9技术制备NCAPG基因敲除牛成纤维细胞系[J]. 生物技术通报, 2026, 42(2): 317-324.
WANG Ting-ting, HE Meng-ya, SHENG Jia-shun, GAO Chen, CAI Han-fang, FU Tong, SUN Yu, GAO Teng-yun, ZHANG Tian-liu. Preparation of NCAPG Knockout Bovine Fibroblast Cell Lines Using CRISPR/Cas9 Technology[J]. Biotechnology Bulletin, 2026, 42(2): 317-324.
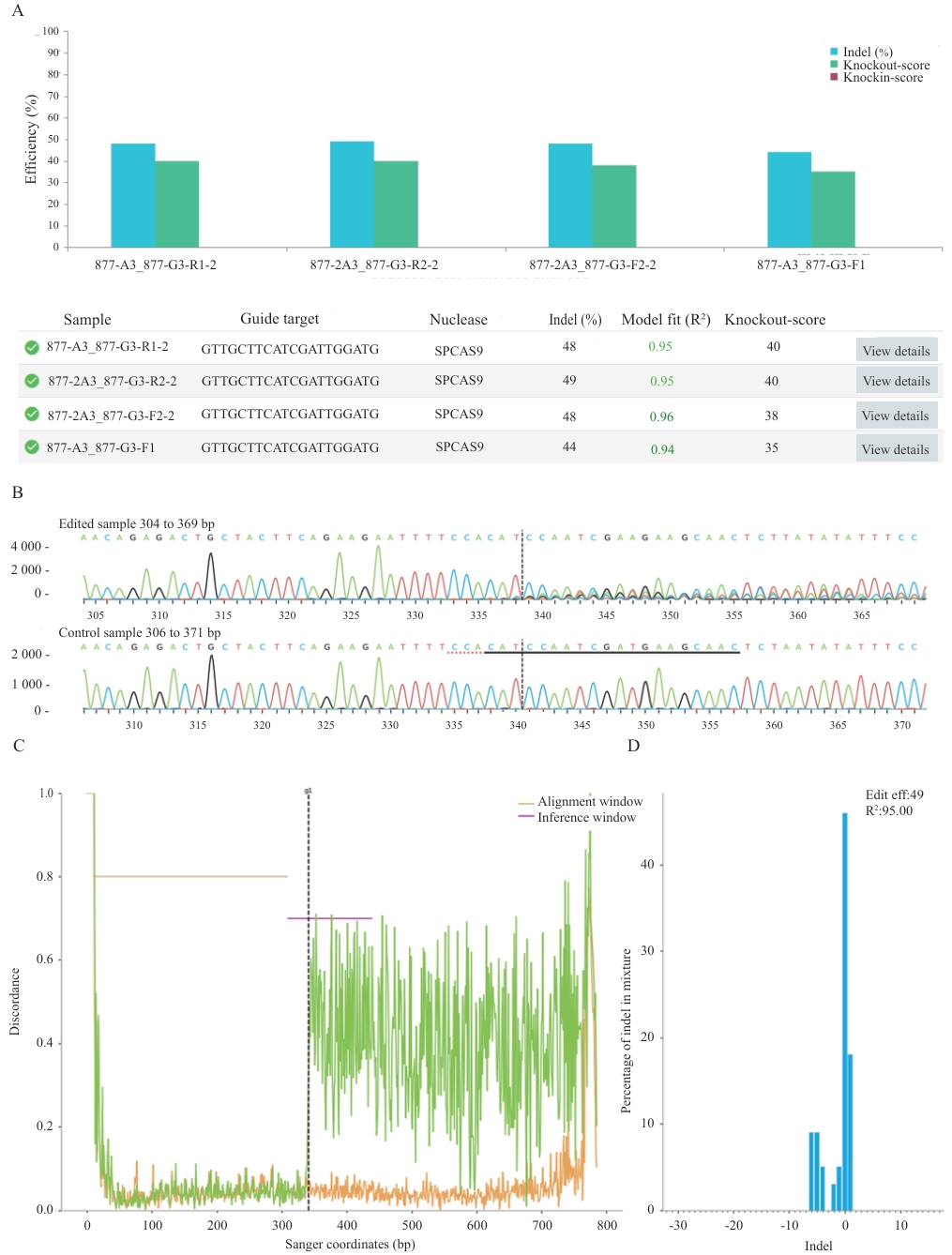
图3 ICE软件分析sgRNA的编辑效率A:sgRNA在不同单克隆中的切割效率;B:Sanger序列视图,分别代表已编辑和野生型序列;sgRNA靶序列用黑色下划线标注;PAM序列以红色虚线标出;垂直虚线指示预测的Cas9切割位点;C:序列差异度分析,显示了编辑样本(绿色)和对照样本(橙色)的测序轨迹的不一致性;D:Indel类型统计:展示了插入和缺失(Insert/Indel)大小及其相对频率
Fig. 3 Analysis of sgRNA editing efficiency using ICE softwareA: The cutting efficiency of sgRNA in different monoclonal cell lines. B: Sanger sequence view, respectively representing the edited and wild-type sequences. The sgRNA target sequence is annotated with a black underline; the PAM sequence is marked with a red dashed line; and a vertical dashed line indicates the predicted Cas9 cleavage site. C: Trace discrepancy analysis, showing the misalignment between the sequencing traces of the edited sample (green) and the control sample (orange). D: Indel type statistics: Displays the size of insertions and deletions (Insert/Deletion) and their relative frequency
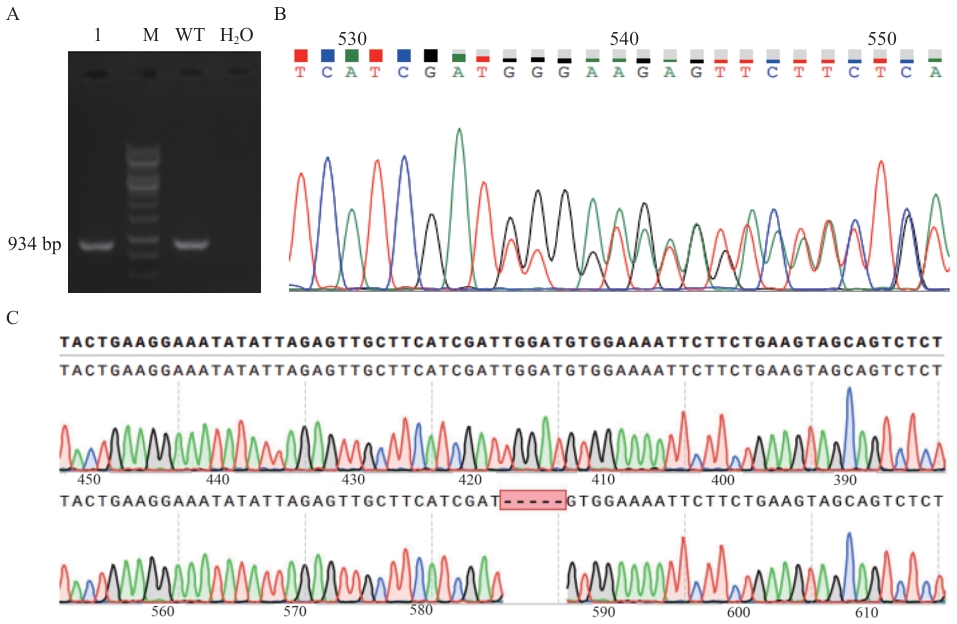
图4 敲除NCAPG基因型的鉴定A:NCAPG基因敲除靶位点PCR扩增产物琼脂凝胶电泳检测;M:DL10000 DNA marker;1:NCAPG基因敲除靶位点PCR扩增结果;WT:野生型对照组;H2O:阴性对照;B:阳性单克隆细胞峰图;C:TA克隆测序比对结果
Fig. 4 Identification of NCAPG knockout genotypesA: Agarose gel electrophoresis of PCR amplified products of NCAPG gene knockout target sites. M: DL10000 DNA marker. 1: PCR amplified results of NCAPG gene knockout target sites. WT: Wild type control. H2O: Negative control. B: Sequencing peak map of positive cell clones. C: Sequencing alignment results after TA cloning
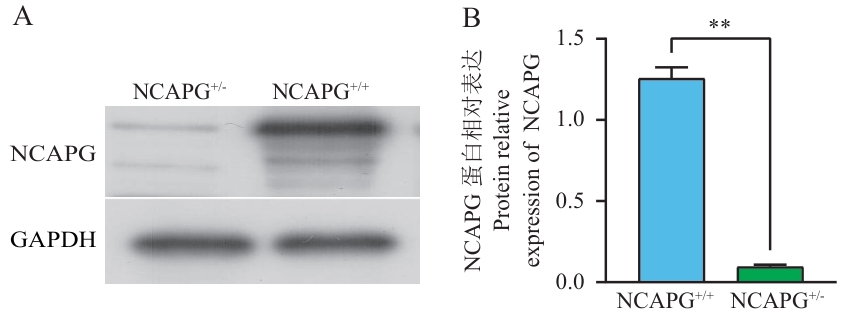
图5 Western blotting检测NCAPG蛋白表达A:野生型和基因敲除的NCAPG蛋白条带图;B:NCAPG蛋白水平的定量分析;NCAPG+/+:野生型成纤维细胞;NCAPG+/-:基因敲除的成纤维细胞;GAPDH为内参。**P<0.01,下同
Fig. 5 Expression of NCAPG protein detected using Western blottingA: Western blot analysis of NCAPG expression in wild-type and knockout cells; B: Quantitative densitometric analysis of NCAPG protein levels. NCAPG+/+: Wild type fibroblasts. NCAPG+/-: Fibroblasts with gene knockout; GAPDH was used as a loading control. *P<0.01, the same below
| [1] | 郭佳俊, 崔励柏, 卢美轩, 等. 黄牛经济性状主效基因研究进展 [J]. 中国牛业科学, 2024, 50(1): 58-64. |
| Guojia J, Cui LB, Lu MX, et al. Research progress on major genes of economic traits in cattle [J]. China Cattle Sci, 2024, 50(1): 58-64. | |
| [2] | Li ZQ, Liu HY, Wang J, et al. Retrospect and prospect: reproductive technologies in beef cattle [J]. Mol Biol Rep, 2025, 52(1): 251. |
| [3] | Hoshiba H, Setoguchi K, Watanabe T, et al. Comparison of the effects explained by variations in the bovine PLAG1 and NCAPG genes on daily body weight gain, linear skeletal measurements and carcass traits in Japanese Black steers from a progeny testing program [J]. Anim Sci J, 2013, 84(7): 529-534. |
| [4] | Liu Y, Duan XY, Chen S, et al. NCAPG is differentially expressed during longissimus muscle development and is associated with growth traits in Chinese Qinchuan beef cattle [J]. Genet Mol Biol, 2015, 38(4): 450-456. |
| [5] | Altmann S, Murani E, Schwerin M, et al. Maternal dietary protein restriction and excess affects offspring gene expression and methylation of non-SMC subunits of condensin I in liver and skeletal muscle [J]. Epigenetics, 2012, 7(3): 239-252. |
| [6] | Ma XC, Ying F, Li ZD, et al. New insights into the genetic loci related to egg weight and age at first egg traits in broiler breeder [J]. Poult Sci, 2024, 103(5): 103613. |
| [7] | Yuan ZH, Ge L, Su PW, et al. NCAPG regulates myogenesis in sheep, and SNPs located in its putative promoter region are associated with growth and development traits [J]. Animals, 2023, 13(20): 3173. |
| [8] | Jabbar A, Zulfiqar F, Mahnoor M, et al. Advances and perspectives in the application of CRISPR-Cas9 in livestock [J]. Mol Biotechnol, 2021, 63(9): 757-767. |
| [9] | Wang ST, Qu ZX, Huang QY, et al. Application of gene editing technology in resistance breeding of livestock [J]. Life, 2022, 12(7): 1070. |
| [10] | Gim GM, Kwon DH, Eom KH, et al. Production of MSTN-mutated cattle without exogenous gene integration using CRISPR-Cas9 [J]. Biotechnol J, 2022, 17(7): 2100198. |
| [11] | Tian HB, Niu HM, Luo J, et al. Knockout of stearoyl-CoA desaturase 1 decreased milk fat and unsaturated fatty acid contents of the goat model generated by CRISPR/Cas9 [J]. J Agric Food Chem, 2022, 70(13): 4030-4043. |
| [12] | Burkard C, Lillico SG, Reid E, et al. Precision engineering for PRRSV resistance in pigs: Macrophages from genome edited pigs lacking CD163 SRCR5 domain are fully resistant to both PRRSV genotypes while maintaining biological function [J]. PLoS Pathog, 2017, 13(2): e1006206. |
| [13] | Yang H, Pu LL, Li RB, et al. NCAPG is transcriptionally regulated by CBX3 and activates the Wnt/β-catenin signaling pathway to promote proliferation and the cell cycle and inhibit apoptosis in colorectal cancer [J]. J Gastrointest Oncol, 2023, 14(2): 900-912. |
| [14] | Zhang Q, Su RX, Shan C, et al. Non-SMC condensin I complex, subunit G (NCAPG) is a novel mitotic gene required for hepatocellular cancer cell proliferation and migration [J]. Oncol Res, 2018, 26(2): 269-276. |
| [15] | Setoguchi K, Watanabe T, Weikard R, et al. The SNP c.1326T>G in the non-SMC condensin Ⅰ complex, subunit G (NCAPG) gene encoding a p.Ile442Met variant is associated with an increase in body frame size at puberty in cattle [J]. Anim Genet, 2011, 42(6): 650-655. |
| [16] | 闵奇, 刘益丽, 蒋梦娟, 等. 调控牛体型大小的信号通路及候选基因研究进展 [J]. 黑龙江畜牧兽医, 2024(3): 22-30. |
| Min Q, Liu YL, Jiang MJ, et al. Research progress on signaling pathways and candidate genes for regulating body size in cattle (Bos Taurus) [J]. Heilongjiang Anim Sci Vet Med, 2024(3): 22-30. | |
| [17] | Zhang WG, Li JY, Guo Y, et al. Multi-strategy genome-wide association studies identify the DCAF16-NCAPG region as a susceptibility locus for average daily gain in cattle [J]. Sci Rep, 2016, 6: 38073. |
| [18] | Niu QH, Zhang TL, Xu L, et al. Identification of candidate variants associated with bone weight using whole genome sequence in beef cattle [J]. Front Genet, 2021, 12: 750746. |
| [19] | Widmann P, Reverter A, Weikard R, et al. Systems biology analysis merging phenotype, metabolomic and genomic data identifies non-SMC condensin Ⅰ complex, subunit G (NCAPG) and cellular maintenance processes as major contributors to genetic variability in bovine feed efficiency [J]. PLoS One, 2015, 10(4): e0124574. |
| [20] | Hu X, Xing YS, Fu X, et al. NCAPG dynamically coordinates the myogenesis of fetal bovine tissue by adjusting chromatin accessibility [J]. Int J Mol Sci, 2020, 21(4): 1248. |
| [21] | Seipold S, Priller FC, Goldsmith P, et al. Non-SMC condensin I complex proteins control chromosome segregation and survival of proliferating cells in the zebrafish neural retina [J]. BMC Dev Biol, 2009, 9(1): 40. |
| [22] | 刘思远, 易国强, 唐中林, 等. 基于CRISPR/Cas9系统在全基因组范围内筛选功能基因及调控元件研究进展 [J]. 遗传, 2020, 42(5): 435-443. |
| Liu SY, Yi GQ, Tang ZL, et al. Progress on genome-wide CRISPR/Cas9 screening for functional genes and regulatory elements [J]. Hereditas, 2020, 42(5): 435-443. | |
| [23] | Hsu PD, Scott DA, Weinstein JA, et al. DNA targeting specificity of RNA-guided Cas9 nucleases [J]. Nat Biotechnol, 2013, 31(9): 827-832. |
| [24] | 朱娜娜. 利用CRISPR/Cas9构建OCT4-EGFP-PuroR双报告功能的巴马猪胎儿成纤维细胞 [D]. 合肥: 安徽农业大学, 2023. |
| Zhu NN. Construction of OCT4-EGFP-PuroR dual-reporter functional Bama pig fetal fibroblasts using CRISPR/Cas9 [D]. Hefei: Anhui Agricultural University, 2023. | |
| [25] | Conant D, Hsiau T, Rossi N, et al. Inference of CRISPR edits from Sanger trace data [J]. CRISPR J, 2022, 5(1): 123-130. |
| [26] | Jiang FG, Doudna JA. CRISPR-Cas9 structures and mechanisms [J]. Annu Rev Biophys, 2017, 46: 505-529. |
| [27] | 苏楠, 吴正红, 祁小乐. CRISPR/Cas9系统RNP体内递送的挑战与解决策略 [J]. 沈阳药科大学学报, 2023, 40(7): 964-976. |
| Su N, Wu ZH, Qi XL. Challenges and strategies for in vivo delivery of CRISPR/Cas9-RNP [J]. J Shenyang Pharm Univ, 2023, 40(7): 964-976. | |
| [28] | 张雪萍, 刘嘉仪, 王彦芳, 等. 利用CRISPR/Cas9编辑系统构建ACTA1基因敲除的PEFs细胞系 [J]. 中国畜牧兽医, 2024, 51(6): 2273-2284. |
| Zhang XP, Liu JY, Wang YF, et al. Construction of ACTA1 gene knockout PEFs cell lines by CRISPR/Cas9 editing system [J]. China Anim Husb Vet Med, 2024, 51(6): 2273-2284. | |
| [29] | Sun HH, Zhang H, Yan Y, et al. NCAPG promotes the oncogenesis and progression of non-small cell lung cancer cells through upregulating LGALS1 expression [J]. Mol Cancer, 2022, 21(1): 55. |
| [30] | Zinshteyn B, Sinha NK, Enam SU, et al. Translational repression of NMD targets by GIGYF2 and EIF4E2 [J]. PLoS Genet, 2021, 17(10): e1009813. |
| [31] | Gim GM, Eom KH, Kwon DH, et al. Generation of double knockout cattle via CRISPR-Cas9 ribonucleoprotein (RNP) electroporation [J]. J Anim Sci Biotechnol, 2023, 14(1): 103. |
| [1] | 陈小玲, 廖东庆, 黄尚飞, 陈英, 芦志龙, 陈东. 利用CRISPR/Cas9系统改造酿酒酵母的研究进展[J]. 生物技术通报, 2023, 39(8): 148-158. |
| [2] | 施炜涛, 姚春鹏, 魏文康, 王蕾, 房元杰, 仝钰洁, 马晓姣, 蒋文, 张晓爱, 邵伟. 利用CRISPR/Cas9技术构建MDH2敲除细胞株及抗呕吐毒素效应研究[J]. 生物技术通报, 2023, 39(7): 307-315. |
| [3] | 岳鹏鹏, 郭俊璠, 于鸿浩, 付灿, 王小燕, 高进涛. 基于CRISPR/cas9系统高效编辑小鼠Galt基因[J]. 生物技术通报, 2020, 36(8): 235-342. |
| [4] | 何士俊, 万毅虹, 章嘉雯, 蔡秀潮, 刘静文, 刘叔文, 姚新刚. CRISPR/Cas9慢病毒系统敲除胰岛β细胞PKA C-α的研究[J]. 生物技术通报, 2020, 36(3): 102-109. |
| [5] | 田文佳, 窦桂铭, 王莎, 孙靖雅, 马玉超. 利用CRISPR/Cas9系统建立内生链霉菌SAT1的基因簇敲除体系[J]. 生物技术通报, 2019, 35(6): 1-8. |
| [6] | 贾启鹏,申培磊,张欢,张勇. CRISPR/Cas9系统介导敲除CSN2基因奶山羊胎儿成纤维突变细胞的制备[J]. 生物技术通报, 2017, 33(9): 131-138. |
| [7] | 杨刚刚,马诚凯,史世会,张全义,王泽,吕中原,王绪洋,许晓亚,崔晴晴,张继红,丁一,徐存拴. 豹蛙抗瘤酶与人血清白蛋白融合蛋白的毕赤酵母高效表达与活性测定[J]. 生物技术通报, 2015, 31(10): 222-229. |
| [8] | 虞飞, 李斌, 张晶晶, 丁海麦, 张学明. 重组人gdnf乳腺特异表达载体转染牛胎儿成纤维细胞的研究[J]. 生物技术通报, 2014, 30(8): 102-107. |
| [9] | 王旭静. 荷兰一公司在欧洲首次推出商业化的自体皮肤移植产品[J]. , 2002, 0(05): 48-49. |
| [10] | 王颖. Ciba-Geigy与ISIS共同开发反义技术[J]. , 1991, 0(06): 16-16. |
| [11] | . 细胞工程[J]. , 1991, 0(06): 41-46. |
| [12] | 邓永鸿;. 生血因子可否用于治疗艾滋病[J]. , 1988, 0(01): 15-16. |
| [13] | 柴勇;. 消除聚乙二醇的毒性[J]. , 1987, 0(03): 18-18. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||