








生物技术通报 ›› 2020, Vol. 36 ›› Issue (10): 237-246.doi: 10.13560/j.cnki.biotech.bull.1985.2020-0335
王丹蕊1,3( ), 沈文丽2, 魏子艳2, 王尚1, 邓晔1,2,3(
), 沈文丽2, 魏子艳2, 王尚1, 邓晔1,2,3( )
)
收稿日期:2020-03-27
出版日期:2020-10-26
发布日期:2020-11-02
作者简介:王丹蕊,女,博士研究生,研究方向:微生物生态学;E-mail: 基金资助:
WANG Dan-rui1,3( ), SHEN Wen-li2, WEI Zi-yan2, WANG Shang1, DENG Ye1,2,3(
), SHEN Wen-li2, WEI Zi-yan2, WANG Shang1, DENG Ye1,2,3( )
)
Received:2020-03-27
Published:2020-10-26
Online:2020-11-02
摘要:
单细胞测序技术是指在单细胞水平上对核酸分子进行测序的技术。近年来,单细胞测序技术异军突起,成为分子生物学研究的热点,在医学、生物化学、生命科学等领域取得了卓越的成果,也成为了单细胞生态学的重要组成部分。单细胞测序与扩增子或宏基因组技术结合可以更准确地识别微生物物种,探究种群异质性,进一步研究特定物种的功能,获得稀有物种的完整基因组。简述了单细胞测序的产生及发展历程,重点介绍了细胞分离和基因组扩增的前沿技术,并详细论述了单细胞测序技术在微生物生态领域中的应用。
王丹蕊, 沈文丽, 魏子艳, 王尚, 邓晔. 单细胞测序技术在微生物生态领域中的应用[J]. 生物技术通报, 2020, 36(10): 237-246.
WANG Dan-rui, SHEN Wen-li, WEI Zi-yan, WANG Shang, DENG Ye. Applications of Single-cell Sequencing Technology in Microbial Ecology[J]. Biotechnology Bulletin, 2020, 36(10): 237-246.
| 细胞分离技术 | 技术简介 | 优点 | 局限性 | 应用案例 |
|---|---|---|---|---|
| 有限稀释法 | 培养液稀释至约每0.1mL含有1个菌或细胞,根据泊松分布计算 | 简单廉价 | 准确性差,不具有针对性 | 对HIV-1病毒的全长序列进行测序[ |
| 显微操作法 | 机械显微操作,用毛细管从各类样品中捕获单个细胞 | 对细胞进行可视性评估 | 通量极低,易对细胞造成机械损伤 | 分离并检测食物中的致病菌[ |
| 激光捕获显微分离技术 | 将细胞进行固定和显色处理后直接分离 | 显色标记与分离结合,可分离复杂基质中的单细胞 | 通量低,易引入杂质或导致遗传信息丢失 | 研究动植物宿主-微生物的相互作用,鉴定未培养细菌,对单核原核细胞进行分析[ |
| 拉曼镊子 | 通过拉曼显微镜区分生化特性不同的细胞,再用激光将其捕获 | 不需标记处理 | 只能分离生化特性显著不同的细胞 | 结合拉曼光镊和单细胞芯片分离癌细胞、红细胞、淋巴细胞和大肠杆菌[ |
| 涡旋与相分隔 | 通过高速涡旋形成油包水体系 | 通量高,效率高,成本低 | 难以保证过程的精确性 | 用于探究硫酸盐还原功能类群[ |
| 荧光细胞分选技术 | 基于所需细胞属性,结合多参数对特定细胞进行检测分离,同时评估生理学和分类学特性 | 灵敏度高,自动化程度高,可依据大小、粒度、荧光等属性分选 | 通量中等,成本高 | 分离丝状真菌并进行分类[ |
| 微流控技术 | 通过微流控芯片原件等设备包裹和分离单细胞样品 | 通量高,精确稳定 | 需要特定仪器装备,开发周期较长 | 单细胞全基因组测序及抗性基因分析[ |
表1 常见单细胞分离技术一览
| 细胞分离技术 | 技术简介 | 优点 | 局限性 | 应用案例 |
|---|---|---|---|---|
| 有限稀释法 | 培养液稀释至约每0.1mL含有1个菌或细胞,根据泊松分布计算 | 简单廉价 | 准确性差,不具有针对性 | 对HIV-1病毒的全长序列进行测序[ |
| 显微操作法 | 机械显微操作,用毛细管从各类样品中捕获单个细胞 | 对细胞进行可视性评估 | 通量极低,易对细胞造成机械损伤 | 分离并检测食物中的致病菌[ |
| 激光捕获显微分离技术 | 将细胞进行固定和显色处理后直接分离 | 显色标记与分离结合,可分离复杂基质中的单细胞 | 通量低,易引入杂质或导致遗传信息丢失 | 研究动植物宿主-微生物的相互作用,鉴定未培养细菌,对单核原核细胞进行分析[ |
| 拉曼镊子 | 通过拉曼显微镜区分生化特性不同的细胞,再用激光将其捕获 | 不需标记处理 | 只能分离生化特性显著不同的细胞 | 结合拉曼光镊和单细胞芯片分离癌细胞、红细胞、淋巴细胞和大肠杆菌[ |
| 涡旋与相分隔 | 通过高速涡旋形成油包水体系 | 通量高,效率高,成本低 | 难以保证过程的精确性 | 用于探究硫酸盐还原功能类群[ |
| 荧光细胞分选技术 | 基于所需细胞属性,结合多参数对特定细胞进行检测分离,同时评估生理学和分类学特性 | 灵敏度高,自动化程度高,可依据大小、粒度、荧光等属性分选 | 通量中等,成本高 | 分离丝状真菌并进行分类[ |
| 微流控技术 | 通过微流控芯片原件等设备包裹和分离单细胞样品 | 通量高,精确稳定 | 需要特定仪器装备,开发周期较长 | 单细胞全基因组测序及抗性基因分析[ |
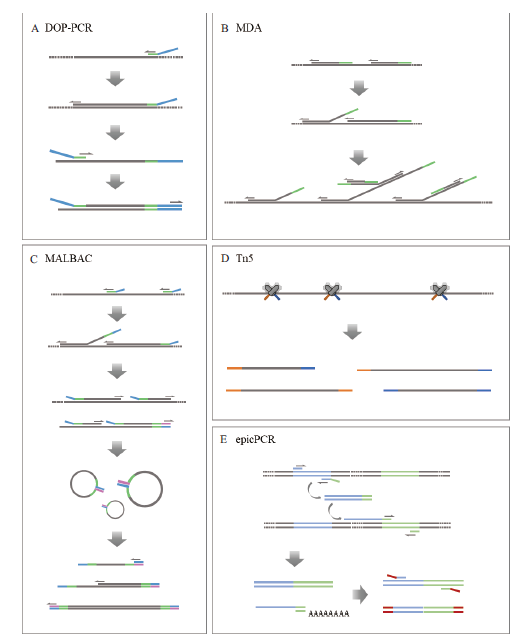
图1 常见单细胞基因组扩增技术原理示意图 A:简并寡核苷酸引物PCR技术(DOP-PCR)。随机引物的3' 端含6bp的随机序列,可以随机和基因组DNA结合,实现对全基因组的扩增;B:多重置换扩增技术(MDA)。随机六聚体引物与模板DNA结合,并在φ29 DNA聚合酶的作用下延伸;随后引物与延伸链结合,以多分支结构的形式延伸扩增;C:多次退火环状循环扩增技术(MALBAC)。首先引物与模板DNA结合,在具有置换活性的DNA聚合酶作用下延伸产生半扩增产物;随后随机引物与半扩增产物结合并延伸形成完整产物;最后对尾部互补成环的完整产物进行扩增;D:Tn5转座酶技术。Tn5转座酶随机将样品DNA片段化,并在小片段DNA两端加上特定的接头,便于后续的扩增和测序;E:细胞内融合基因技术(epicPCR)。两段目标基因被封装在同一微球中,在3条特殊引物的作用下产生融合片段;随后通过巢式PCR消除半扩增产物的影响,特异性扩增融合片段,并缩短其长度供二代测序
| [1] | 朱忠旭, 陈新. 单细胞测序技术及应用进展[J]. 基因组学与应用生物学, 2015(5):9-15. |
| Zhu Z, Chen X. Single cell sequencing technology and its applications progress[J]. Genomics and Applied Biology, 2015(5):9-15. | |
| [2] |
Gawad C, Koh W, Quake SR, et al. Single-cell genome sequencing:current state of the science[J]. Nature Reviews Genetics, 2016,17(3):175-188.
URL pmid: 26806412 |
| [3] |
Iscove N. Haematopoiesis. Searching for stem cells[J]. Nature, 1990,347(6289):126-127.
URL pmid: 2395468 |
| [4] |
Telenius H, Carter NP, Bebb CE, et al. Degenerate oligonucleotide-primed PCR:General amplification of target DNA by a single degenerate primer[J]. Genomics, 1992,13(3):718-725.
URL pmid: 1639399 |
| [5] |
Dean FB, Nelson JR, Giesler TL, et al. Rapid amplification of plasmid and phage DNA using Phi 29 DNA polymerase and multiply-primed rolling circle amplification[J]. Genome Res, 2001,11(6):1095-1099.
URL pmid: 11381035 |
| [6] |
Raghunathan A, Ferguson HR, Bornarth CJ, et al. Genomic DNA amplification from a single bacterium[J]. Applied and Environmental Microbiology, 2005,71(6):3342-3347.
URL pmid: 15933038 |
| [7] |
Lasken RS. Single-cell genomic sequencing using Multiple Displacement Amplification.[J]. Current Opinion in Microbiology, 2007,10(5):510-516.
URL pmid: 17923430 |
| [8] |
Stepanauskas R, Fergusson EA, Brown J, et al. Improved genome recovery and integrated cell-size analyses of individual uncultured microbial cells and viral particles[J]. Nature Communications, 2017,8(1):84.
URL pmid: 28729688 |
| [9] |
Zong CH, Lu SJ, Chapman AR, et al. Genome-wide detection of single-nucleotide and copy-number variations of a single human cell[J]. Science, 2012,338(6114):1622-1626.
URL pmid: 23258894 |
| [10] | 张慧, 徐楚帆, 江来. 单细胞测序相关技术及其在生物医学研究中的应用[J]. 实用医学杂志, 2020,36(3):410-413. |
| Zhang H, Xu C, Jiang L. Development of single-cell sequencing and its biomedical applications[J]. The Journal of Practical Medicine, 2020,36(3):410-413. | |
| [11] | Sanschagrin S, Yergeau E. Next-generation sequencing of 16S ribosomal RNA gene amplicons[J]. Journal of Visualized Experiments, 2014(90):e51709. |
| [12] |
Ranjan R, Rani A, Metwally A, et al. Analysis of the microbiome:Advantages of whole genome shotgun versus 16S amplicon sequencing[J]. Biochemical and Biophysical Research Communications, 2016,469(4):967-977.
URL pmid: 26718401 |
| [13] |
Sims D, Sudbery I, Ilott NE, et al. Sequencing depth and coverage:key considerations in genomic analyses[J]. Nature Reviews Genetics, 2014,15(2):121-132.
URL pmid: 24434847 |
| [14] |
Luo C, Rodriguez-RLM, Konstantinidis KT. A user’s guide to quantitative and comparative analysis of metagenomic datasets[J]. Methods in Enzymology, 2013,531:525-547.
URL pmid: 24060135 |
| [15] | 魏子艳, 金德才, 邓晔. 环境微生物宏基因组学研究中的生物信息学方法[J]. 微生物学通报, 2015,42(5):890-901. |
| Wei Z, Jin D, Deng Y. Bioinformatics tools and applications in the study of environmental microbial metagenomics[J]. Microbiology China, 2015,42(5):890-901. | |
| [16] |
Thomas T, Gilbert J, Meyer F. Metagenomics-a guide from sampling to data analysis[J]. Microbial Informatics and Experimentation, 2012,2(1):3-14.
doi: 10.1186/2042-5783-2-3 URL pmid: 22587947 |
| [17] |
Stepanauskas R. Single cell genomics:an individual look at microbes[J]. Current Opinion in Microbiology, 2012,15(5):613-620.
URL pmid: 23026140 |
| [18] |
Bowers RM, Lee J, Woyke T. Sequencing of genomes from environmental single cells[J]. Methods Mol Biol, 2018,1712:97-111.
doi: 10.1007/978-1-4939-7514-3_8 URL pmid: 29224071 |
| [19] |
Ishii S, Tago K, Senoo K. Single-cell analysis and isolation for microbiology and biotechnology:methods and applications[J]. Appl Microbiol Biotechnol, 2010,86(5):1281-1292.
doi: 10.1007/s00253-010-2524-4 URL pmid: 20309540 |
| [20] |
Spencer SJ, Tamminen MV, Preheim SP, et al. Massively parallel sequencing of single cells by epicPCR links functional genes with phylogenetic markers[J]. ISME Journal, 2016,10(2):427-436.
doi: 10.1038/ismej.2015.124 URL pmid: 26394010 |
| [21] |
Lan F, Demaree B, Ahmed N, et al. Single-cell genome sequencing at ultra-high-throughput with microfluidic droplet barcoding[J]. Nat Biotechnol, 2017,35(7):640-646.
doi: 10.1038/nbt.3880 URL pmid: 28553940 |
| [22] | Sanchez Barea J, Lee J, Kang DK. Recent advances in droplet-based microfluidic technologies for biochemistry and molecular biology[J]. Micromachines(Basel), 2019,10(6):412. |
| [23] |
Hebberecht L, Vancoillie L, Schauvliege M, et al. Single genome sequencing of near full-length HIV-1 RNA using a limiting dilution approach[J]. J Virol Methods, 2019,274:113737.
URL pmid: 31562885 |
| [24] |
Marisa H, Myriam M, Renaud C, et al. Development of a micromanipulation method for single cell isolation of prokaryotes and its application in food safety[J]. PLoS One, 2018, 13(5):e0198208-.
doi: 10.1371/journal.pone.0241865 URL pmid: 33152039 |
| [25] |
Podgorny OV, Lazarev VN. Laser microdissection:A promising tool for exploring microorganisms and their interactions with hosts[J]. Journal of Microbiological Methods, 2017,138:82-92.
URL pmid: 26775287 |
| [26] |
Fang T, Shang WH, Liu C, et al. Nondestructive identification and accurate isolation of single cells through a chip with Raman optical tweezers[J]. Analytical Chemistry, 2019,91(15):9932-9939.
URL pmid: 31251569 |
| [27] | Bleichrodt RJ, Read ND. Flow cytometry and FACS applied to filamentous fungi[J]. Fungal Biol Rev, 2019,33(1):1-15. |
| [28] |
Huang L, Ma F, Chapman A, et al. Single-cell whole-genome amplification and sequencing:Methodology and applications[J]. Annu Rev Genomics Hum Genet, 2015,16:79-102.
doi: 10.1146/annurev-genom-090413-025352 URL pmid: 26077818 |
| [29] | Blagodatskikh KA, Kramarov VM, Barsova EV, et al. Improved DOP-PCR(iDOP-PCR):A robust and simple WGA method for efficient amplification of low copy number genomic DNA[J]. PLoS One, 2017,12(9):12. |
| [30] |
Pugh TJ, Delaney AD, Farnoud N, et al. Impact of whole genome amplification on analysis of copy number variants[J]. Nucleic Acids Research, 2008,36(13):e80.
doi: 10.1093/nar/gkn378 URL pmid: 18559357 |
| [31] |
Xu Y, Zhao F. Single-cell metagenomics:challenges and applications[J]. Protein & Cell, 2018,9(5):501-510.
URL pmid: 29696589 |
| [32] |
Woyke T, Doud DFR, Schulz F. The trajectory of microbial single-cell sequencing[J]. Nature Methods, 2017,14(11):1045-1054.
URL pmid: 29088131 |
| [33] |
Clingenpeel S, Clum A, Schwientek P, et al. Reconstructing each cell’s genome within complex microbial communities-dream or reality?[J]. Frontiers in Microbiology, 2015,5:771.
doi: 10.3389/fmicb.2014.00771 URL pmid: 25620966 |
| [34] | Povilaitis T, Alzbutas G, Sukackaite R, et al. In vitro evolution of phi29 DNA polymerase using isothermal compartmentalized self replication technique[J]. Protein Engineering Design & Selection, 2016,29(12):617-628. |
| [35] | Chen M, Song P, Zou D, et al. Comparison of multiple displacement amplification(MDA)and multiple annealing and looping-based amplification cycles(MALBAC)in single-cell sequencing[J]. PLoS One, 2014,9(12):e0124990. |
| [36] |
de Bourcy CFA, De Vlaminck I, Kanbar JN, et al. A quantitative comparison of single-cell whole genome amplification methods[J]. PLoS One, 2014,9(8):e105585.
URL pmid: 25136831 |
| [37] |
Chen C, Xing D, Tan L, et al. Single-cell whole-genome analyses by Linear Amplification via Transposon Insertion(LIANTI)[J]. Science, 2017,356(6334):189-194.
URL pmid: 28408603 |
| [38] | Florenza J, Tamminen M, Bertilsson S. Uncovering microbial inter-domain interactions in complex communities[J]. Philosophical Transactions of the Royal Society B-Biological Sciences, 2019,374(1786):20190087. |
| [39] | Lovett M. The applications of single-cell genomics[J]. Hum Mol Genet, 2013,22:22-26. |
| [40] | Dubey RK, Tripathi V, Prabha R, et al. Single-cell genomics and metagenomics for microbial diversity analysis[M]. Unravelling the Soil Microbiome. Switzerland:Springer, 2020: 33-49. |
| [41] |
Munson-McGee JH, Field EK, Bateson M, et al. Nanoarchaeota, their Sulfolobales host, and Nanoarchaeota virus distribution across yellowstone national park hot springs[J]. Applied and Environmental Microbiology, 2015,81(22):7860-7868.
URL pmid: 26341207 |
| [42] |
Woyke T, Jarett J. Function-driven single-cell genomics[J]. Microbial Biotechnology, 2015,8(1):38-39.
doi: 10.1111/1751-7915.12247 URL pmid: 25627845 |
| [43] |
Trapnell C. Defining cell types and states with single-cell genomics[J]. Genome Research, 2015,25(10):1491-1498.
URL pmid: 26430159 |
| [44] |
Pamp SJ, Harrington ED, Quake SR, et al. Single-cell sequencing provides clues about the host interactions of segmented filamentous bacteria(SFB)[J]. Genome Research, 2012,22(6):1107-1119.
URL pmid: 22434425 |
| [45] |
Lasken RS. Genomic sequencing of uncultured microorganisms from single cells[J]. Nature Reviews Microbiology, 2012,10(9):631-640.
doi: 10.1038/nrmicro2857 URL pmid: 22890147 |
| [46] |
Locey KJ, Lennon JT. Scaling laws predict global microbial diversity[J]. Proceedings of the National Academy of Sciences of the United States of America, 2016,113(21):5970-5975.
doi: 10.1073/pnas.1521291113 URL pmid: 27140646 |
| [47] |
Rinke C, Schwientek P, Sczyrba A, et al. Insights into the phylogeny and coding potential of microbial dark matter[J]. Nature, 2013,499(7459):431-437.
URL pmid: 23851394 |
| [48] |
Tyson GW, Chapman J, Hugenholtz P, et al. Community structure and metabolism through reconstruction of microbial genomes from the environment[J]. Nature, 2004,428(6978):37-43.
doi: 10.1038/nature02340 URL pmid: 14961025 |
| [49] |
Sogin ML, Morrison HG, Huber JA, et al. Microbial diversity in the deep sea and the underexplored “rare biosphere”[J]. Proceedings of the National Academy of Sciences of the United States of America, 2006,103(32):12115-12120.
doi: 10.1073/pnas.0605127103 URL pmid: 16880384 |
| [50] |
Schloss PD, Girard RA, Martin T, et al. Status of the archaeal and bacterial census:an update[J]. mBio, 2016,7(3):e00201-16.
doi: 10.1128/mBio.00201-16 URL pmid: 27190214 |
| [51] |
Wu D, Hugenholtz P, Mavromatis K, et al. A phylogeny-driven genomic encyclopaedia of Bacteria and Archaea[J]. Nature, 2009,462(7276):1056-1060.
doi: 10.1038/nature08656 URL pmid: 20033048 |
| [52] |
Lasken RS, McLean JS. Recent advances in genomic DNA sequencing of microbial species from single cells[J]. Nature Reviews Genetics, 2014,15(9):577-584.
doi: 10.1038/nrg3785 URL pmid: 25091868 |
| [53] |
Yu FB, Blainey PC, Schulz F, et al. Microfluidic-based mini-metagenomics enables discovery of novel microbial lineages from complex environmental samples[J]. eLife, 2017,6:e26580.
URL pmid: 28678007 |
| [54] |
Sieracki ME, Poulton NJ, Jaillon O, et al. Single cell genomics yields a wide diversity of small planktonic protists across major ocean ecosystems[J]. Scientific Reports, 2019,9:6025.
doi: 10.1038/s41598-019-42487-1 URL pmid: 30988337 |
| [55] |
Ahrendt SR, Quandt CA, Ciobanu D, et al. Leveraging single-cell genomics to expand the fungal tree of life[J]. Nature Microbiology, 2018,3(12):1417-1428.
URL pmid: 30297742 |
| [56] | 李俊锋. 基于16S rRNA和宏基因组高通量测序的微生物多样性研究[D]. 北京:清华大学, 2015. |
| Li J. Microbial diversity research based on high-throughput sequencing data of 16S rRNA and metagenome[D]. Beijing:Tsinghua University, 2015. | |
| [57] |
Pop M, Salzberg SL. Bioinformatics challenges of new sequencing technology[J]. Trends in Genetics, 2008,24(3):142-149.
doi: 10.1016/j.tig.2007.12.006 URL pmid: 18262676 |
| [58] | 许亚昆, 马越, 胡小茜, 等. 基于三代测序技术的微生物组学研究进展[J]. 生物多样性, 2019,27(5):534-542. |
|
Xu Y, Ma Y, Hu X, et al. Analysis of prospective microbiology research using third-generation sequencing technology[J]. Biodiversity Science, 2019,27(5):534-542.
doi: 10.17520/biods.2018201 URL |
|
| [59] | 唐勇, 刘旭. SMRT测序技术及其在微生物研究中的应用[J]. 生物技术通报, 2018,34(6):48-53. |
| Tang Y, Liu X. SMRT Sequencing and Its application in microorganism studies[J]. Biotechnology Bulletin, 2018,34(6):48-53. | |
| [60] |
Kashtan N, Roggensack SE, Rodrigue S, et al. Single-cell genomics reveals hundreds of coexisting subpopulations in wild Prochlorococcus[J]. Science, 2014,344(6182):416-420.
doi: 10.1126/science.1248575 URL pmid: 24763590 |
| [61] |
Yoon HS, Price DC, Stepanauskas R, et al. Single-cell genomics reveals organismal interactions in uncultivated marine protists[J]. Science, 2011,332(6030):714-717.
doi: 10.1126/science.1203163 URL pmid: 21551060 |
| [62] |
Engel P, Stepanauskas R, Moran NA. Hidden diversity in honey bee gut symbionts detected by single-cell genomics[J]. PLoS Genetics, 2014,10(9):e1004596.
URL pmid: 25210772 |
| [63] |
Jochum LM, Schreiber L, Marshall IPG, et al. Single-cell genomics reveals a diverse metabolic potential of uncultivated Desulfatiglans-related deltaproteobacteria widely distributed in marine sediment[J]. Frontiers in Microbiology, 2018,9:2038.
doi: 10.3389/fmicb.2018.02038 URL pmid: 30233524 |
| [64] | Magdanova LA, Golyasnaya NV. Heterogeneity as an adaptive trait of microbial populations[J]. Microbiology, 2013,82(1):1-10. |
| [65] |
Stepanauskas R. Wiretapping into microbial interactions by single cell genomics[J]. Frontiers in Microbiology, 2015,6:258.
doi: 10.3389/fmicb.2015.00258 URL pmid: 25904902 |
| [66] |
Paez-Espino D, Eloe-Fadrosh EA, Pavlopoulos GA, et al. Uncovering earth’s virome[J]. Nature, 2016,536(7617):425-430.
doi: 10.1038/nature19094 URL pmid: 27533034 |
| [67] |
Abergel C, Legendre M, Claverie JM. The rapidly expanding universe of giant viruses:Mimivirus, Pandoravirus, Pithovirus and Mollivirus[J]. FEMS Microbiology Reviews, 2015,39(6):779-796.
doi: 10.1093/femsre/fuv037 URL pmid: 26391910 |
| [68] |
Roux S, Hawley AK, Beltran MT, et al. Ecology and evolution of viruses infecting uncultivated SUP05 bacteria as revealed by single-cell- and meta- genomics[J]. eLife, 2014,3:e03125.
doi: 10.7554/eLife.03125 URL pmid: 25171894 |
| [69] |
Martinez-Hernandez F, Fornas O, Lluesma Gomez M, et al. Single-cell genomics uncover Pelagibacter as the putative host of the extremely abundant uncultured 37-F6 viral population in the ocean[J]. ISME Journal, 2019,13(1):232-236.
URL pmid: 30228380 |
| [70] |
Labonte JM, Pachiadaki M, Fergusson E, et al. Single cell genomics-based analysis of gene content and expression of prophages in a diffuse-flow deep-sea hydrothermal system[J]. Frontiers in Microbiology, 2019,10:1262.
URL pmid: 31244796 |
| [71] |
Huber H, Hohn MJ, Rachel R, et al. A new phylum of Archaea represented by a nanosized hyperthermophilic symbiont[J]. Nature, 2002,417(6884):63-67.
doi: 10.1038/417063a URL pmid: 11986665 |
| [72] |
He X, McLean JS, Edlund A, et al. Cultivation of a human-associated TM7 phylotype reveals a reduced genome and epibiotic parasitic lifestyle[J]. Proceedings of the National Academy of Sciences of the United States of America, 2015,112(1):244-249.
URL pmid: 25535390 |
| [73] |
Comolli LR, Banfield JF. Inter-species interconnections in acid mine drainage microbial communities[J]. Frontiers in Microbiology, 2014,5:367.
URL pmid: 25120533 |
| [74] |
Nakayama T, Nomura M, Takano Y, et al. Single-cell genomics unveiled a cryptic cyanobacterial lineage with a worldwide distribution hidden by a dinoflagellate host[J]. Proceedings of the National Academy of Sciences of the United States of America, 2019,116(32):15973-15978.
doi: 10.1073/pnas.1902538116 URL pmid: 31235587 |
| [75] |
Farnelid HM, Turk-Kubo KA, Zehr JP. Identification of associations between bacterioplankton and photosynthetic picoeukaryotes in coastal waters[J]. Frontiers in Microbiology, 2016,7:339.
URL pmid: 27148165 |
| [76] | Seymour JR, Amin SA, Raina JB, et al. Zooming in on the phycosphere:the ecological interface for phytoplankton-bacteria relationships[J]. Nature Microbiology, 2017,2(7):17065. |
| [77] |
Abd H, Weintraub A, Sandstrom G. Intracellular survival and replication of Vibrio cholerae O139 in aquatic free-living amoebae[J]. Environ Microbiol, 2005,7(7):1003-1008.
doi: 10.1111/j.1462-2920.2005.00771.x URL pmid: 15946296 |
| [78] |
Qin HY, Wang S, Feng K, et al. Unraveling the diversity of sedimentary sulfate-reducing prokaryotes(SRP)across Tibetan saline lakes using epicPCR[J]. Microbiome, 2019,7(1):71.
URL pmid: 31054577 |
| [79] |
Hultman J, Tamminen M, Parnanen K, et al. Host range of antibiotic resistance genes in wastewater treatment plant influent and effluent[J]. FEMS Microbiology Ecology, 2018, 94(4):fiy038.
doi: 10.1093/femsle/fnaa156 URL pmid: 33068111 |
| [80] |
Chijiiwa R, Hosokawa M, Kogawa M, et al. Single-cell genomics of uncultured bacteria reveals dietary fiber responders in the mouse gut microbiota[J]. Microbiome, 2020,8(1):5.
URL pmid: 31969191 |
| [81] |
Doud DFR, Bowers RM, Schulz F, et al. Function-driven single-cell genomics uncovers cellulose-degrading bacteria from the rare biosphere[J]. The ISME Journal, 2019,14(3):659-675.
doi: 10.1038/s41396-019-0557-y URL pmid: 31754206 |
| [82] | Richards TA, Massana R, Pagliara S, et al. Single cell ecology[J]. Philosophical Transactions of the Royal Society B-Biological Sciences, 2019,374(1786):20190076. |
| [83] |
Nurk S, Bankevich A, Antipov D, et al. Assembling single-cell genomes and mini-metagenomes from chimeric MDA products[J]. Journal of Computational Biology, 2013,20(10):714-737.
URL pmid: 24093227 |
| [1] | 吴昊, 刘紫微, 郑颖, 戴雅文, 时权. 单细胞水平解析人牙龈间充质干细胞异质性[J]. 生物技术通报, 2023, 39(7): 325-332. |
| [2] | 鲁兆祥, 王夕冉, 连新磊, 廖晓萍, 刘雅红, 孙坚. 基于功能宏基因组学挖掘抗生素耐药基因研究进展[J]. 生物技术通报, 2022, 38(9): 17-27. |
| [3] | 钟辉, 刘亚军, 王滨花, 和梦洁, 吴兰. 分析方法对细菌群落16S rRNA基因扩增测序分析结果的影响[J]. 生物技术通报, 2022, 38(6): 81-92. |
| [4] | 张淼, 孙祥瑞, 徐春明. 单细胞RNA测序数据分析方法研究进展[J]. 生物技术通报, 2021, 37(1): 52-59. |
| [5] | 过冬冬, 孙芬, 贺轩昂, 羊东晔, 黄来强. 单细胞测序技术在肝脏疾病的应用与展望[J]. 生物技术通报, 2021, 37(1): 137-144. |
| [6] | 朱庆元, 李天晴. 单细胞转录组测序技术在心脏发育、疾病以及医学中的应用[J]. 生物技术通报, 2021, 37(1): 145-154. |
| [7] | 杨秀清, 陈彦梅, 吴瑞薇, 王保玉, 韩作颖. 煤地质环境微生物总基因组DNA提取方法的优化[J]. 生物技术通报, 2018, 34(9): 177-183. |
| [8] | 刘捷孟, 戚继. 元基因组学方法在环境微生物中的研究进展[J]. 生物技术通报, 2015, 31(11): 51-59. |
| [9] | 潘孝明 ,梁兴国. 全基因组扩增技术原理及研究进展[J]. 生物技术通报, 2014, 0(12): 47-54. |
| [10] | 金浩;李柏林;潘迎捷;欧杰;陈兰明;. 宏基因组学技术在微生物功能酶开发中的应用[J]. , 2012, 0(09): 46-50. |
| [11] | 徐晓宇;刘和;. 454测序法在环境微生物生态研究中的应用[J]. , 2010, 0(01): 73-77. |
| [12] | 徐晓宇;刘和;. 环境微生物基因组技术研究进展[J]. , 2006, 0(04): 54-58. |
| [13] | 顾继东;hkucc.hku;hk. 国外环境生物技术的发展和展望[J]. , 1999, 0(06): 8-12. |
| [14] | 孙雷心. 英国科学家称蝎病毒安全有效[J]. , 1996, 0(05): 19-20. |
| [15] | 孙雷心. 生物技术简讯(二)[J]. , 1995, 0(01): 30-30. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||