








生物技术通报 ›› 2026, Vol. 42 ›› Issue (2): 338-350.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0603
• 研究报告 • 上一篇
孙焱森1( ), 魏立翔1, 李若冰1, 张程志1, 聂宇航1, 李杰1, 才学鹏2, 乔军1, 孟庆玲1(
), 魏立翔1, 李若冰1, 张程志1, 聂宇航1, 李杰1, 才学鹏2, 乔军1, 孟庆玲1( )
)
收稿日期:2025-06-12
出版日期:2026-02-26
发布日期:2026-03-17
通讯作者:
孟庆玲,女,博士,教授,研究方向 :动物寄生虫学;E-mail: xjmqlqj@163.com作者简介:孙焱森,男,硕士研究生,研究方向 :动物寄生虫学;E-mail: 763203488@qq.com
基金资助:
SUN Yan-sen1( ), WEI Li-xiang1, LI Ruo-bing1, ZHANG Cheng-zhi1, NIE Yu-hang1, LI Jie1, CAI Xue-peng2, QIAO Jun1, MENG Qing-ling1(
), WEI Li-xiang1, LI Ruo-bing1, ZHANG Cheng-zhi1, NIE Yu-hang1, LI Jie1, CAI Xue-peng2, QIAO Jun1, MENG Qing-ling1( )
)
Received:2025-06-12
Published:2026-02-26
Online:2026-03-17
摘要:
目的 为探究少孢节丛孢菌(Arthrobotrys oligospora)Zn(Ⅱ)2Cys6型转录因子AOL-113的生物学功能。 方法 利用同源重组的方法构建Δ AOL-113缺失株和CΔ AOL-113回补株,从菌丝生长特性、营养物质利用、环境胁迫响应、分生孢子产生及萌发和线虫捕食能力等表型进行分析,同时结合转录组学技术,在乙酸钠诱导条件下对WT菌株与Δ AOL-113缺失株进行差异表达基因分析,探究转录因子AOL-113的分子调控机制。 结果 转录因子AOL-113编码GAL4型锌指蛋白,与转录因子FacB有较高的同源性。与WT菌株相比,Δ AOL-113缺失株生长速率、碳源利用特别是脂肪酸的利用能力显著下降,对H2O2和甲萘醌这类氧化胁迫剂更敏感。此外,∆ AOL-113缺失株产生捕器数量及捕器形成速度显著下降,对线虫的降解消化利用能力也显著下降。回补株表型与WT株无显著差异。转录组分析证实,AOL-113缺失导致多个代谢通路(包括氧化磷酸化、脂质代谢、乙醛酸循环、过氧化物酶体、半乳糖代谢和糖酵解)中关键基因表达显著下调,从而影响菌丝生长、碳源利用、抗胁迫能力、捕食能力等多个生物学过程。 结论 转录因子AOL-113不仅通过调控细胞代谢(尤其是乙酸代谢和脂肪酸β氧化)为菌丝生长和捕食器形成提供能量支持,同时也参与调控捕食线虫和消化线虫的效率,为揭示少孢节丛孢菌从腐生向寄生生活方式转变的分子机制和家畜线虫病生防制剂的研发提供了新思路。
孙焱森, 魏立翔, 李若冰, 张程志, 聂宇航, 李杰, 才学鹏, 乔军, 孟庆玲. AOL-113转录因子对少孢节丛孢菌菌丝生长、胁迫响应及捕食能力的调控作用[J]. 生物技术通报, 2026, 42(2): 338-350.
SUN Yan-sen, WEI Li-xiang, LI Ruo-bing, ZHANG Cheng-zhi, NIE Yu-hang, LI Jie, CAI Xue-peng, QIAO Jun, MENG Qing-ling. Regulatory Roles of Transcription Factor AOL-113 in Mycelial Growth, Stress Response, and Predatory Ability of Arthrobotrys oligospora[J]. Biotechnology Bulletin, 2026, 42(2): 338-350.
引物名称 Primer name | 引物序列 Primer sequence (5′-3′) | 长度 Length (bp) | 基因目的 Gene description |
|---|---|---|---|
| AOL-113-F/AOL-113-R | ATGTCGTTGAAACGGTCG/GTCATTCCAGCTTGATAGAACCG | 3 213 | AOL-113基因全长 |
| AOL-113-5F-pUC19 | 1 861 | AOL-113基因5′端基因序列 | |
| AOL-113-5R-HygR | |||
| HygR-F-113-5R | 1 441 | 潮霉素抗性基因 | |
| HygR-R-113-3F | |||
| AOL-113-3F-HygR | 1 865 | AOL-113基因3′端基因序列 | |
| AOL-113-3R-pUC19 | |||
| CΔ AOL-113-5F-pUC19 | 3 482 | CΔ AOL-113基因回补全长及5′端侧翼序列 | |
| CΔ AOL-113-5R-NeoR | |||
| NeoR-F-113-5R | 919 | G-418抗性基因 | |
| NeoR-R-113-3F | |||
| CΔ AOL-113-3F-pUC19 | 1 382 | CΔ AOL-113 3′端基因序列 | |
| CΔ AOL-113-3R-pUC19 | |||
| Δ AOL-113YZ-F/Δ AOL-113YZ-R | GTATACTTACCAACCTGCTCC/ GCTACTTCCTGATGATATCCTTC | 3 827/2 209 | 野生型和缺失转化子菌株的验证 |
| CΔ AOL-113YZ-F/CΔ AOL-113YZ-R | CTTCCTCTTCAATTCCTCTTACAAGG/ GGTCATGGTGAAGGATATACTTGT | 3 445/2 301 | 回补株转化子菌株的验证 |
表1 研究中使用的特异性引物
Table 1 Specific primers used in the study
引物名称 Primer name | 引物序列 Primer sequence (5′-3′) | 长度 Length (bp) | 基因目的 Gene description |
|---|---|---|---|
| AOL-113-F/AOL-113-R | ATGTCGTTGAAACGGTCG/GTCATTCCAGCTTGATAGAACCG | 3 213 | AOL-113基因全长 |
| AOL-113-5F-pUC19 | 1 861 | AOL-113基因5′端基因序列 | |
| AOL-113-5R-HygR | |||
| HygR-F-113-5R | 1 441 | 潮霉素抗性基因 | |
| HygR-R-113-3F | |||
| AOL-113-3F-HygR | 1 865 | AOL-113基因3′端基因序列 | |
| AOL-113-3R-pUC19 | |||
| CΔ AOL-113-5F-pUC19 | 3 482 | CΔ AOL-113基因回补全长及5′端侧翼序列 | |
| CΔ AOL-113-5R-NeoR | |||
| NeoR-F-113-5R | 919 | G-418抗性基因 | |
| NeoR-R-113-3F | |||
| CΔ AOL-113-3F-pUC19 | 1 382 | CΔ AOL-113 3′端基因序列 | |
| CΔ AOL-113-3R-pUC19 | |||
| Δ AOL-113YZ-F/Δ AOL-113YZ-R | GTATACTTACCAACCTGCTCC/ GCTACTTCCTGATGATATCCTTC | 3 827/2 209 | 野生型和缺失转化子菌株的验证 |
| CΔ AOL-113YZ-F/CΔ AOL-113YZ-R | CTTCCTCTTCAATTCCTCTTACAAGG/ GGTCATGGTGAAGGATATACTTGT | 3 445/2 301 | 回补株转化子菌株的验证 |
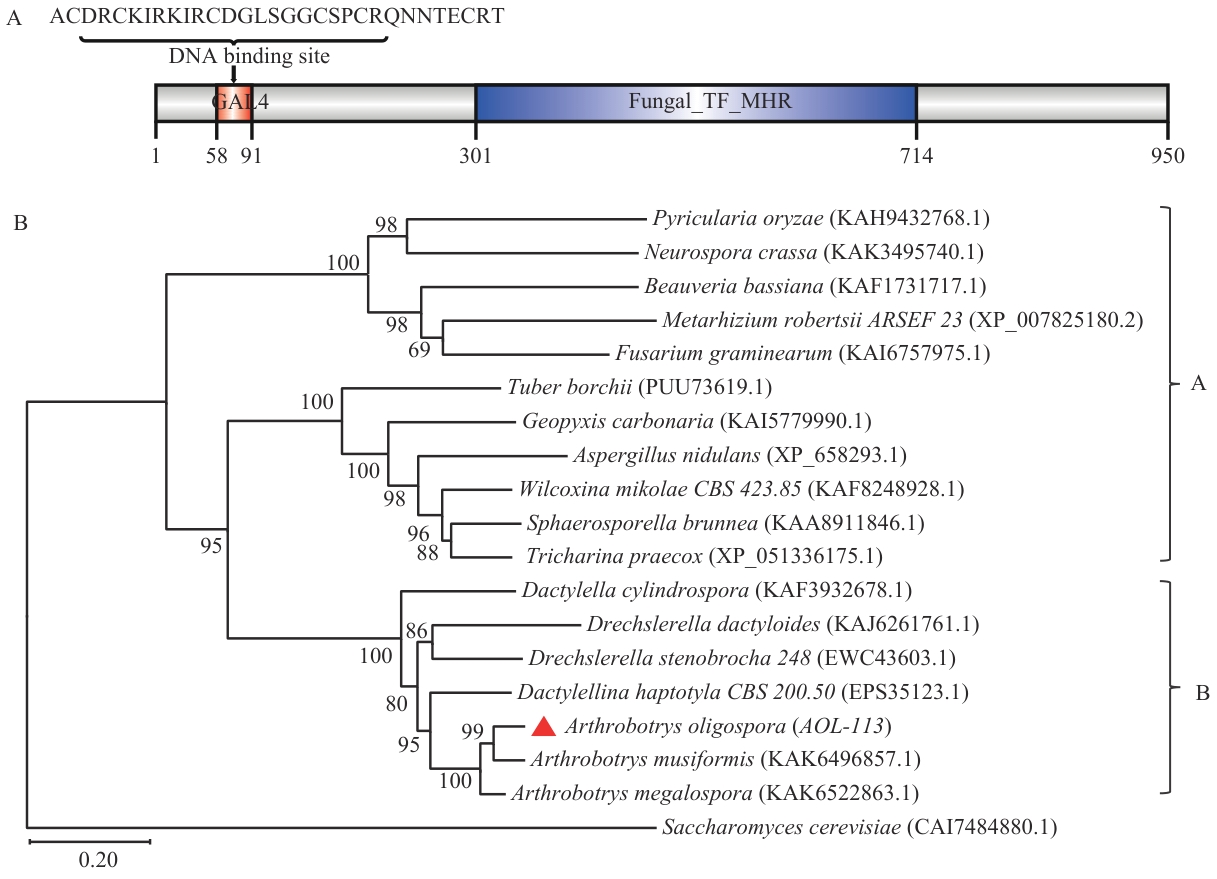
图1 少孢节丛孢菌Zn(II)₂Cys₆型转录因子AOL-113的分子特征和系统发育分析A:转录因子AOL-113结构域特征。B:转录因子AOL-113同源基因的真菌进化树分析
Fig. 1 Molecular characteristics and phylogenetic analysis of the Zn2(II)Cys6-type transcription factor AOL-113 of A. oligosporaA: Domain characteristic of transcription factor AOL-113. B: Phylogenetic analysis of different fungi based on transcription factor AOL-113orthologs
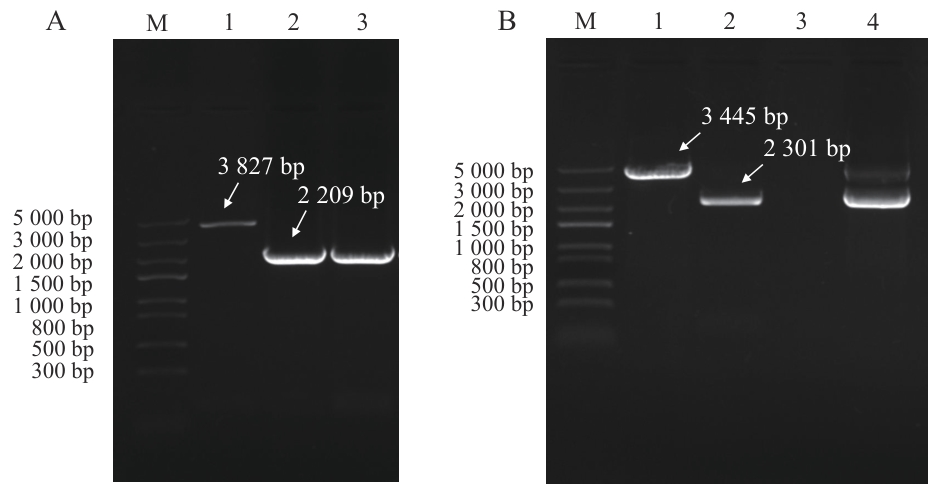
图2 缺失株与回补株的构建与筛选A:缺失菌株电泳检测结果。M:DL-5000 DNA marker;1:阴性对照;2-3:缺失菌株阳性克隆。B:回补株电泳检测结果。M:DL-5000 DNA marker;1:回补菌株阳性克隆;2-4:阴性对照
Fig. 2 Construction and screening of deletion strains and complementary strainsA: Electrophoresis detection results of deletion strains. M: DL-5000 DNA marker; 1: negative control; 2-3: positive clones of deletion strains. B: Electrophoresis detection results of complementary strains. M: DL-5000 DNA marker; 1: positive clones of complementary strains; 2-4: negative control
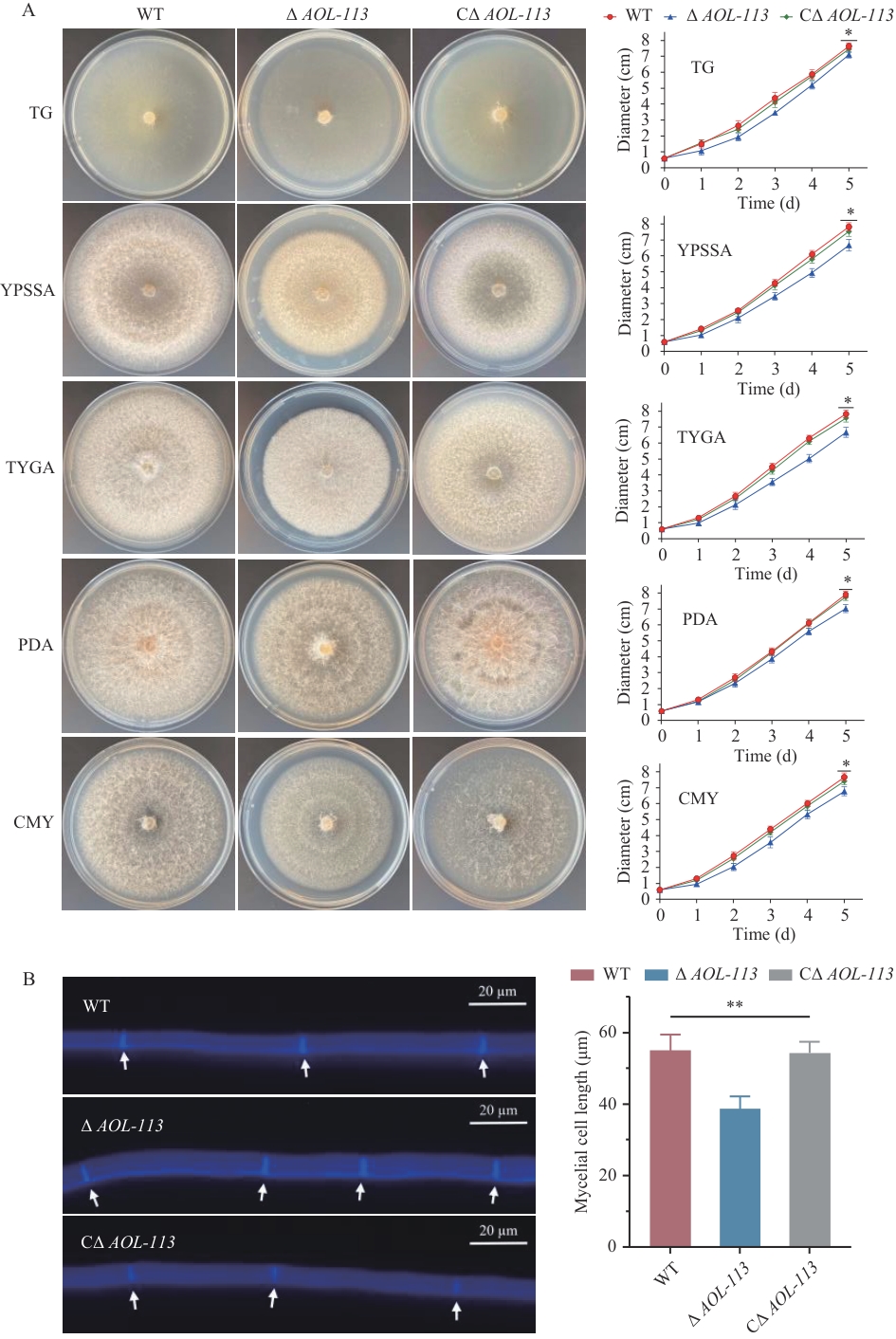
图3 生长速率测定及菌丝细胞长度的比较A:TG、YPSSA、TYGA、PDA和CMY培养基上28 ℃培养5 d菌落形态观察及生长速率测定。B:菌丝细胞的CFW染色,比例尺=20 μm,菌丝隔膜用白色箭头表示
Fig. 3 Comparison of growth rate and mycelial cell lengthA: TG, YPSSA, TYGA, PDA and CMY media were incubated at 28 ℃ for 5 d for colony morphology observation and growth rate measurement. B: CFW staining of mycelia. Scale = 20 μm, the hyphal septa are denoted by white arrows. *P<0.05, **P<0.01. The same below
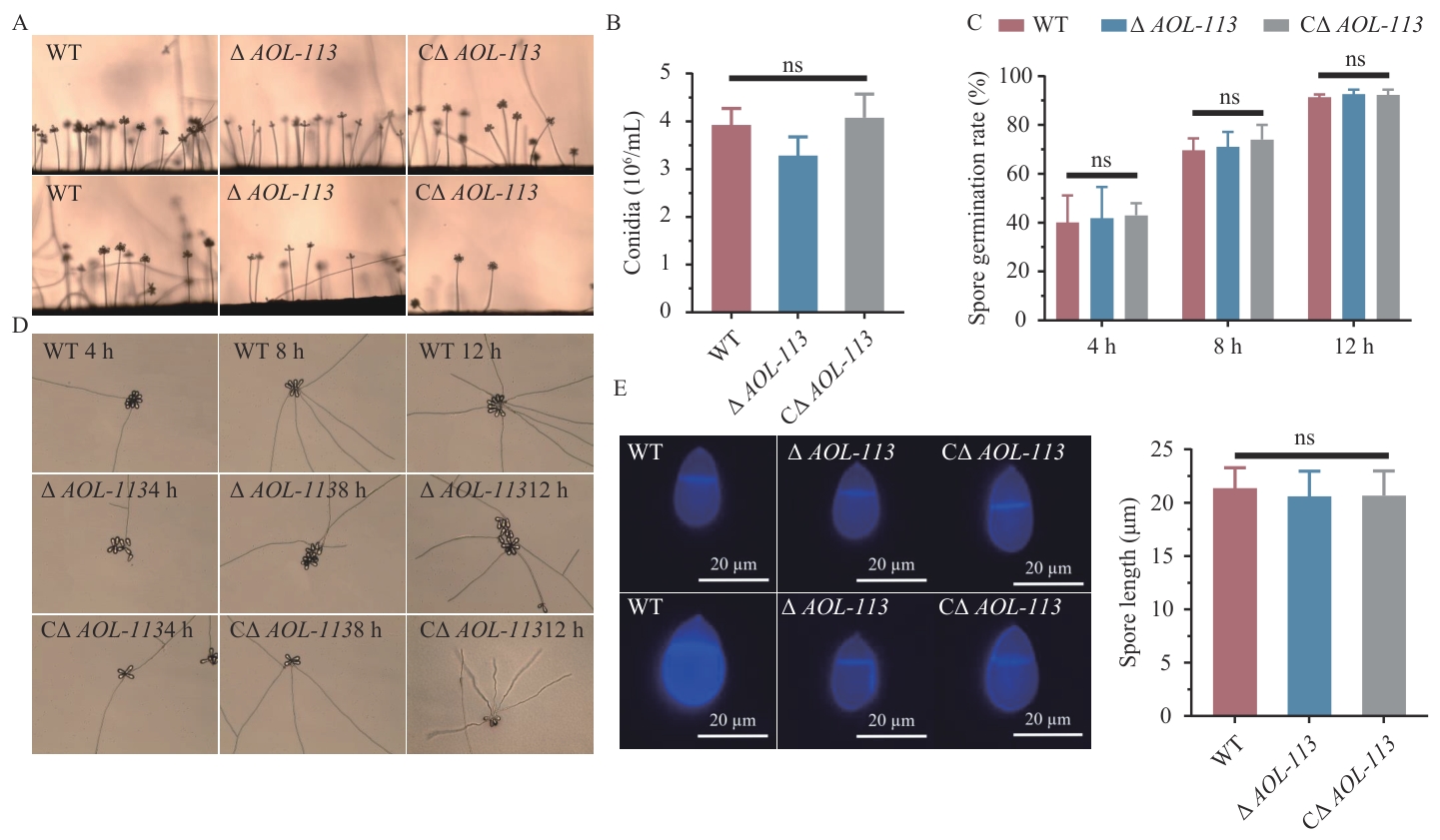
图6 产孢率、孢子萌发率的比较及分生孢子的荧光观察A:分生孢子梗侧拍观察。B:产孢率测定。C:孢子萌发测定。D:孢子萌发观察。E:分生孢子荧光观察及孢子长度测定
Fig. 6 Comparison of sporulation rate, spore germination rate and fluorescence observation of conidiaA: Side view of the phloem peduncle. B: Determination of sporulation rate. C: Spore germination rate. D: Observation of conidial germination. E: Fluorescence observation of conidia and spore length determination
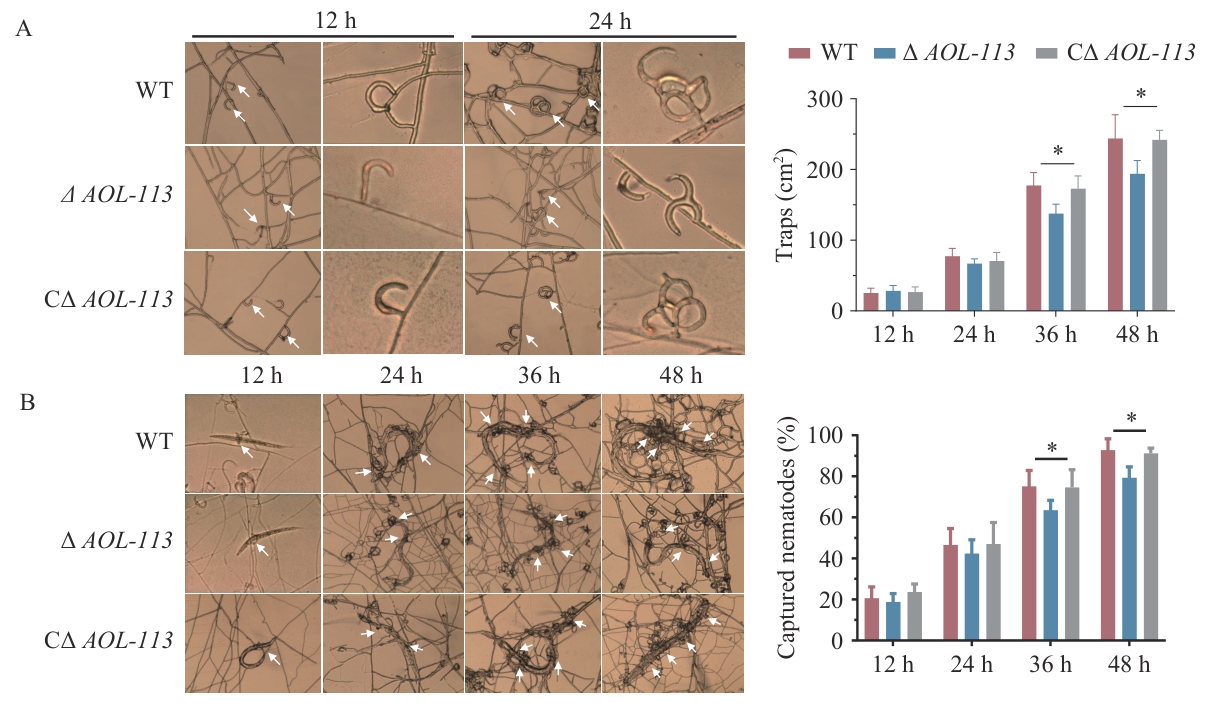
图7 捕器形成和捕食能力比较A:不同时间点捕器形成情况及捕器计数。B:线虫捕食情况的观察及捕食率测定
Fig. 7 Comparison of the trap formation and predatory abilityA: Trap formation and traps counts at different time points. B: Observations of nematode predation and determination of predation rate
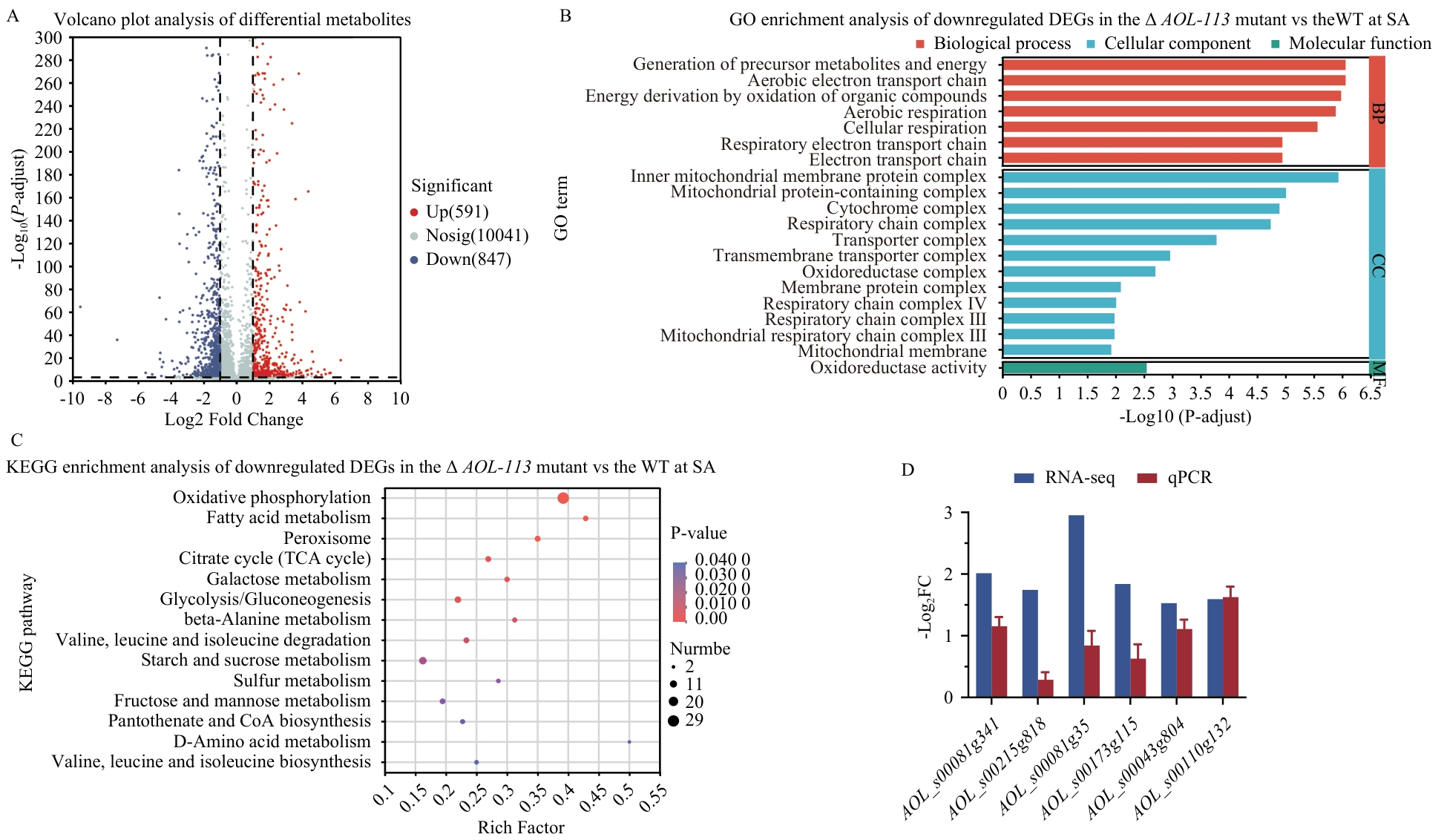
图8 乙酸钠诱导条件下少孢节丛孢菌WT株和Δ AOL-113缺失株DEGs分析A:DEGs火山图。上调DEGs为红色;下调的DEGs为蓝色。B:下调DEGs的GO富集分析。C:下调DEGs的KEGG富集分析。D:差异表达基因RT-qPCR验证
Fig. 8 Analysis of DEGs between WT and Δ AOL-113 of Arthrobotrys oligospora exposed to sodium acetate (SA)A: Volcano plot of DEGs. Up-regulated DEGs in red; down-regulated DEGs in blue. B: GO enrichment analysis of downregulated DEGs. C: KEGG enrichment analysis of downregulated DEGs. D: Validation of transcriptomic sequencing using RT-qPCR
| [1] | Gong SS, Meng QL, Qiao J, et al. Biological characteristics of recombinant Arthrobotrys oligospora chitinase AO-801 [J]. Korean J Parasitol, 2022, 60(5): 345-352. |
| [2] | Li NX, Sun YS, Liu YC, et al. Expression profiles and characterization of microRNAs responding to chitin in Arthrobotrys oligospora [J]. Arch Microbiol, 2024, 206(5): 220. |
| [3] | Larsen M. Prospects for controlling animal parasitic nematodes by predacious micro fungi [J]. Parasitology, 2000, 120(7): 121-131. |
| [4] | Cai KZ, Wang FH, Wang KY, et al. In vitro predatory activity of Arthrobotrys oligospora and after passing through gastrointestinal tract of small ruminants on infective larvae of trichostrongylides [J]. Exp Parasitol, 2017, 177: 104-111. |
| [5] | Wang WS, Meng QL, Qiao J, et al. Isolation of Arthrobotrys oligospora from soil of the Chinese Northern Tianshan Mountain slope pasture show predatory ability against Haemonchus contortus larvae [J]. Biocontrol Sci Technol, 2014, 24(2): 170-179. |
| [6] | 张嘉华, 张慧梅, 马喜喜, 等. 少孢节丛孢菌几丁质酶AO-492对线虫的降解作用研究 [J]. 生物技术通报, 2024, 40(5): 261-268. |
| Zhang JH, Zhang HM, Ma XX, et al. Degradation of nematodes by chitinase AO-492 from arthrospora oligospora [J]. Biotechnol Bull, 2024, 40(5): 261-268. | |
| [7] | Bai N, Xie MH, Liu QQ, et al. AoMedA has a complex regulatory relationship with AoBrlA, AoAbaA, and AoWetA in conidiation, trap formation, and secondary metabolism in the nematode-trapping fungusArthrobotrys oligospora [J]. Appl Environ Microbiol, 2023, 89(9): e00983-23. |
| [8] | Liu YL, Li ZY, Liu JJ, et al. Electron-transferring flavoprotein and its dehydrogenase required for fungal pathogenicity in Arthrobotrys oligospora [J]. Int J Mol Sci, 2024, 25(20): 10934. |
| [9] | Hynes MJ, Murray SL, Duncan AN, et al. Regulatory genes controlling fatty acid catabolism and peroxisomal functions in the filamentous FungusAspergillus nidulans [J]. Eukaryot Cell, 2006, 5(5): 794-805. |
| [10] | Hagiwara D, Miura D, Shimizu K, et al. A Novel Zn2-Cys6 Transcription Factor AtrR Plays a Key Role in an Azole Resistance Mechanism of Aspergillus fumigatus by co-regulating cyp51A and cdr1B Expressions [J]. PLoS Pathog, 2017, 13(1): e1006096. |
| [11] | Zhu CH, Sun JX, Tian FF, et al. The Bbotf1 Zn(Ⅱ)2Cys6 transcription factor contributes to antioxidant response, fatty acid assimilation, peroxisome proliferation and infection cycles in insect pathogenic fungus Beauveria bassiana [J]. J Invertebr Pathol, 2024, 204: 108083. |
| [12] | Sun XY, Wang F, Lan N, et al. The Zn(II)2Cys6-type transcription factor ADA-6 regulates conidiation, sexual development, and oxidative stress response in Neurospora crassa [J]. Front Microbiol, 2019, 10: 750. |
| [13] | Bai N, Xie MH, Liu QQ, et al. AoSte12 is required for mycelial development, conidiation, trap morphogenesis, and secondary metabolism by regulating hyphal fusion in nematode-trapping Fungus Arthrobotrys oligospora [J]. Microbiol Spectr, 2023, 11(2): e03957-22. |
| [14] | Yang L, Li XM, Ma YX, et al. The arf-GAP proteins AoGcs1 and AoGts1 regulate mycelial development, endocytosis, and pathogenicity in Arthrobotrys oligospora [J]. J Fungi, 2022, 8(5): 463. |
| [15] | Zhang Y, Wang X, Ran Y, et al. AfLaeA, a global regulator of mycelial growth, chlamydospore production, pathogenicity, secondary metabolism, and energy metabolism in the nematode-trapping FungusArthrobotrys flagrans [J]. Microbiol Spectr, 2023, 11(4): e00186-23. |
| [16] | 韩改革, 贾晓东, 韩雪琳, 等. 长片段融合PCR在构建烟曲霉rho1基因回补株中的应用 [J]. 中国真菌学杂志, 2013, 8(3): 129-133. |
| Han GG, Jia XD, Han XL, et al. The application of long-fragment fusion PCR in the construction of rho1-compelement strain of Aspergillus fumigatus [J]. Chin J Mycol, 2013, 8(3): 129-133. | |
| [17] | Zhu MC, Zhao N, Liu YK, et al. The cAMP-PKA signalling pathway regulates hyphal growth, conidiation, trap morphogenesis, stress tolerance, and autophagy in Arthrobotrys oligospora [J]. Environ Microbiol, 2022, 24(12): 6524-6538. |
| [18] | Liu QQ, Jiang KX, Duan SP, et al. Identification of a transcription factor AoMsn2 of the Hog1 signaling pathway contributes to fungal growth, development and pathogenicity in Arthrobotrys oligospora [J]. J Adv Res, 2025, 68: 1-15. |
| [19] | Peng H, Dong XY, Lu HQ, et al. A putative F-box-domain-encoding gene AOL_s00076g207 regulates the development and pathogenicity of Arthrobotrys oligospora [J]. J Basic Microbiol, 2022, 62(1): 74-81. |
| [20] | Li J, Wu RN, Wang M, et al. The pH sensing receptor AopalH plays important roles in the nematophagous fungus Arthrobotrys oligospora [J]. Fungal Biol, 2019, 123(7): 547-554. |
| [21] | Zhang GS, Zheng YQ, Ma YX, et al. The velvet proteins VosA and VelB play different roles in conidiation, trap formation, and pathogenicity in the nematode-trapping fungus Arthrobotrys oligospora [J]. Front Microbiol, 2019, 10: 1917. |
| [22] | Liu QQ, Bai N, Duan SP, et al. Characterizing the role of AosfgA and AofluG in mycelial and conidial development in Arthrobotrys oligospora and their role in secondary metabolism [J]. Microorganisms, 2024, 12(3): 615. |
| [23] | 李良德, 李金玉, 李慧玲, 等. 一株灰茶尺蠖高毒力金龟子绿僵菌的筛选、鉴定与培养条件 [J]. 茶叶科学, 2025, 45(3): 427-438. |
| Li LD, Li JY, Li HL, et al. Screening, identification and culture conditions of a highly virulent Metarhizium anisopliae strain against Ectropis grisescens warre [J]. J Tea Sci, 2025, 45(3): 427-438. | |
| [24] | Zhao XY, Wang YC, Zhao Y, et al. Malate synthase gene AoMls in the nematode-trapping fungus Arthrobotrys oligospora contributes to conidiation, trap formation, and pathogenicity [J]. Appl Microbiol Biotechnol, 2014, 98(6): 2555-2563. |
| [25] | Yang L, Li XM, Xie MH, et al. Pleiotropic roles of Ras GTPases in the nematode-trapping fungus Arthrobotrys oligospora identified through multi-omics analyses [J]. iScience, 2021, 24(8): 102820. |
| [26] | Liu QQ, Li DN, Bai N, et al. Peroxin Pex14/17 is required for trap formation, and plays pleiotropic roles in mycelial development, stress response, and secondary metabolism in Arthrobotrys oligospora [J]. mSphere, 2023, 8(2): e00012-23. |
| [27] | Lu JP, Cao HJ, Zhang LL, et al. Systematic analysis of Zn2Cys6 transcription factors required for development and pathogenicity by high-throughput gene knockout in the rice blast fungus [J]. PLoS Pathog, 2014, 10(10): e1004432. |
| [28] | Carrillo AJ, Schacht P, Cabrera IE, et al. Functional profiling of transcription factor genes in Neurospora crassa [J]. G3, 2017, 7(9): 2945-2956. |
| [29] | Flaherty JE, Woloshuk CP. Regulation of fumonisin biosynthesis in Fusarium verticillioides by a zinc binuclear cluster-type gene, ZFR1 [J]. Appl Environ Microbiol, 2004, 70(5): 2653-2659. |
| [30] | Todd RB, Murphy RL, Martin HM, et al. The acetate regulatory gene facB of Aspergillus nidulans encodes a Zn(II)2Cys6 transcriptional activator [J]. Mol Gen Genet, 1997, 254(5): 495-504. |
| [31] | Todd RB, Andrianopoulos A, Davis MA, et al. FacB, the Aspergillus nidulans activator of acetate utilization genes, binds dissimilar DNA sequences [J]. EMBO J, 1998, 17(7): 2042-2054. |
| [32] | bin Yusof MT, Kershaw MJ, Soanes DM, et al. FAR1 and FAR2 regulate the expression of genes associated with lipid metabolism in the rice blast fungus Magnaporthe oryzae [J]. PLoS One, 2014, 9(6): e99760. |
| [33] | Bibbins M, Crepin VF, Cummings NJ, et al. A regulator gene for acetate utilisation from Neurospora crassa [J]. Mol Genet Genomics, 2002, 267(4): 498-505. |
| [34] | Ries LNA, Alves de Castro P, Pereira Silva L, et al. Aspergillus fumigatus acetate utilization impacts virulence traits and pathogenicity [J]. mBio, 2021, 12(4): e0168221. |
| [35] | Liu QQ, Li DN, Jiang KX, et al. AoPEX1 and AoPEX6 are required for mycelial growth, conidiation, stress response, fatty acid utilization, and trap formation in Arthrobotrys oligospora [J]. Microbiol Spectr, 2022, 10(2): e0027522. |
| [36] | Zhou DX, Zhu YM, Bai N, et al. Aolatg1 and Aolatg13 regulate autophagy and play different roles in conidiation, trap formation, and pathogenicity in the nematode-trapping fungus Arthrobotrys oligospora [J]. Front Cell Infect Microbiol, 2021, 11: 824407. |
| [37] | Zhao N, Zhu MC, Liu QQ, et al. AoPrdx2 regulates oxidative stress, reactive oxygen species, trap formation, and secondary metabolism in Arthrobotrys oligospora [J]. JoF, 2024, 10(2): 110. |
| [1] | 张嘉华, 张慧梅, 马喜喜, 孙焱森, 李若冰, 李柠杏, 才学鹏, 乔军, 孟庆玲. 少孢节丛孢菌几丁质酶AO-492对线虫的降解作用研究[J]. 生物技术通报, 2024, 40(5): 261-268. |
| [2] | 姚姿婷, 曹雪颖, 肖雪, 李瑞芳, 韦小妹, 邹承武, 朱桂宁. 火龙果溃疡病菌实时荧光定量PCR内参基因的筛选[J]. 生物技术通报, 2023, 39(5): 92-102. |
| [3] | 赵佳, 孙毅, 梁宏, 黄静, 杜建中. 现代生物技术在根际微生物群落研究中的应用[J]. 生物技术通报, 2012, 0(12): 65-70. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||