








Biotechnology Bulletin ›› 2024, Vol. 40 ›› Issue (10): 172-180.doi: 10.13560/j.cnki.biotech.bull.1985.2024-0657
Previous Articles Next Articles
ZHOU Jia-wei( ), WU Zhi-qiang(
), WU Zhi-qiang( )
)
Received:2024-07-10
Online:2024-10-26
Published:2024-11-20
Contact:
WU Zhi-qiang
E-mail:jiaweizhou@webmail.hzau.edu.cn;wuzhiqiang@caas.cn
ZHOU Jia-wei, WU Zhi-qiang. Construction Method of mitoTALENs Mitochondrial Gene Editing Vector in Plants[J]. Biotechnology Bulletin, 2024, 40(10): 172-180.
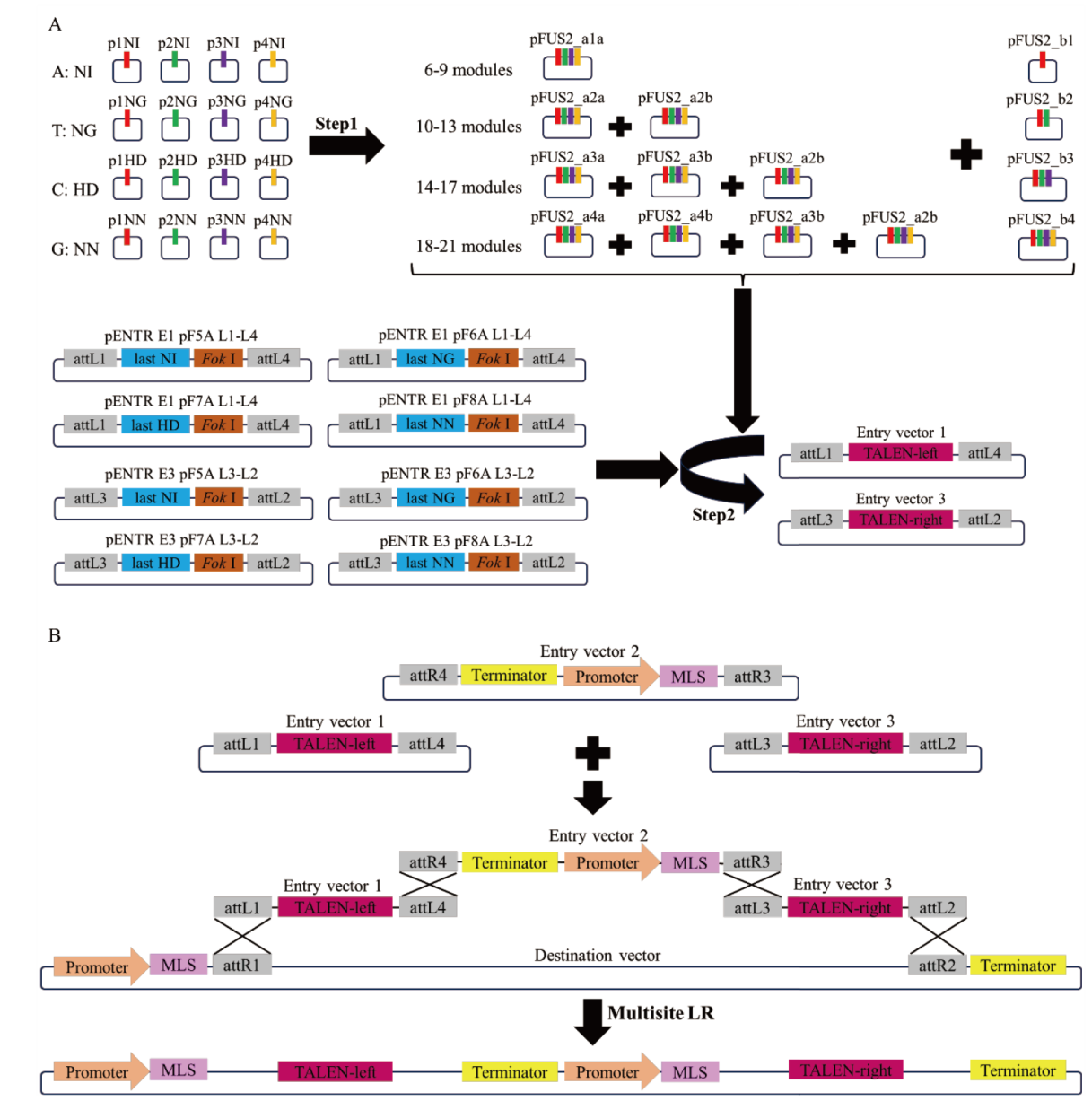
Fig. 3 Schematic diagram of mitoTALENs vector construction A: Two-step assembly schematic diagram of TALE array. B: Schematic diagram multisite LR reaction
| 载体类型 Vector type | 载体内容 Vector content | 载体原核抗性 Vector prokaryotic resistance |
|---|---|---|
| 模块质粒Module plasmids | p1NI,p2NI,p3NI,p4NI,p1NG,p2NG,p3NG,p4NG,p1HD,p2HD,p3HD,p4HD,p1NN,p2NN,p3NN,p4NN | AmpR |
| 中间阵列质粒 Intermediate array plasmids | pFUS2_a1a,pFUS2_a2a,pFUS2_a2b,pFUS2_a3a,pFUS2_a3b,pFUS2_a4a,pFUS2_a4b,pFUS2_b1,pFUS2_b2,pFUS2_b3,pFUS2_b4 | SpeR |
| 进入载体1 Entry vectors 1 | pENTR E1 pF5A L1-L4,pENTR E1 pF6A L1-L4, pENTRE1 pF7A L1-L4,pENTR E1 pF8A L1-L4 | KanR |
| 进入载体2 Entry vectors 2 | pENTER E2 HSPter-35Sp-mito | KanR |
| 进入载体3 Entry vectors 3 | pENTR E3 pR5A L3-L2,pENTR E3 pR6A L3-L2, pENTR E3 pR7A L3-L2,pENTR E3 pR8A L3-L2 | KanR |
| 目的载体 Destination vector | pDESTpK7WG2mito | SpeR |
Table 1 Plasmid vectors required for the construction of mitoTALENs vector
| 载体类型 Vector type | 载体内容 Vector content | 载体原核抗性 Vector prokaryotic resistance |
|---|---|---|
| 模块质粒Module plasmids | p1NI,p2NI,p3NI,p4NI,p1NG,p2NG,p3NG,p4NG,p1HD,p2HD,p3HD,p4HD,p1NN,p2NN,p3NN,p4NN | AmpR |
| 中间阵列质粒 Intermediate array plasmids | pFUS2_a1a,pFUS2_a2a,pFUS2_a2b,pFUS2_a3a,pFUS2_a3b,pFUS2_a4a,pFUS2_a4b,pFUS2_b1,pFUS2_b2,pFUS2_b3,pFUS2_b4 | SpeR |
| 进入载体1 Entry vectors 1 | pENTR E1 pF5A L1-L4,pENTR E1 pF6A L1-L4, pENTRE1 pF7A L1-L4,pENTR E1 pF8A L1-L4 | KanR |
| 进入载体2 Entry vectors 2 | pENTER E2 HSPter-35Sp-mito | KanR |
| 进入载体3 Entry vectors 3 | pENTR E3 pR5A L3-L2,pENTR E3 pR6A L3-L2, pENTR E3 pR7A L3-L2,pENTR E3 pR8A L3-L2 | KanR |
| 目的载体 Destination vector | pDESTpK7WG2mito | SpeR |
| 模块质粒的个数 Number of module plasmids | 中间阵列质粒 Intermediate array plasmids | 模块质粒 Module plasmids | T4 DNA ligase reaction buffer | Bsa I-HF | Quick ligase | ddH2O | 总体积 Total volume/µL |
|---|---|---|---|---|---|---|---|
| 1 | 0.3 | 0.3×1 | 0.2 | 0.1 | 0.1 | 1 | 2 |
| 2 | 0.3 | 0.3×2 | 0.2 | 0.1 | 0.1 | 0.7 | 2 |
| 3 | 0.3 | 0.3×3 | 0.2 | 0.1 | 0.1 | 0.4 | 2 |
| 4 | 0.3 | 0.3×4 | 0.2 | 0.1 | 0.1 | 0.1 | 2 |
Table 2 Reaction system in the first step assembly
| 模块质粒的个数 Number of module plasmids | 中间阵列质粒 Intermediate array plasmids | 模块质粒 Module plasmids | T4 DNA ligase reaction buffer | Bsa I-HF | Quick ligase | ddH2O | 总体积 Total volume/µL |
|---|---|---|---|---|---|---|---|
| 1 | 0.3 | 0.3×1 | 0.2 | 0.1 | 0.1 | 1 | 2 |
| 2 | 0.3 | 0.3×2 | 0.2 | 0.1 | 0.1 | 0.7 | 2 |
| 3 | 0.3 | 0.3×3 | 0.2 | 0.1 | 0.1 | 0.4 | 2 |
| 4 | 0.3 | 0.3×4 | 0.2 | 0.1 | 0.1 | 0.1 | 2 |
| 模块质粒的个数 Number of module plasmids | pFUS2_a质粒 pFUS2_a plasmids | pFUS2_b质粒 pFUS2_b plasmids | 入门载体 Entry vectors | T4 DNA ligase reaction buffer | Esp3 I | Quick ligase | ddH2O | 总体积 Total volume/µL |
|---|---|---|---|---|---|---|---|---|
| 6-9 | 0.6×1 | 0.6 | 0.3 | 0.4 | 0.2 | 0.2 | 1.7 | 4 |
| 10-13 | 0.6×2 | 0.6 | 0.3 | 0.4 | 0.2 | 0.2 | 1.1 | 4 |
| 14-17 | 0.6×3 | 0.6 | 0.3 | 0.4 | 0.2 | 0.2 | 0.5 | 4 |
| 18-21 | 0.6×4 | 0.6 | 0.3 | 0.4 | 0.2 | 0.2 | 0 | 4.1 |
Table 3 Reaction system in the second step assembly
| 模块质粒的个数 Number of module plasmids | pFUS2_a质粒 pFUS2_a plasmids | pFUS2_b质粒 pFUS2_b plasmids | 入门载体 Entry vectors | T4 DNA ligase reaction buffer | Esp3 I | Quick ligase | ddH2O | 总体积 Total volume/µL |
|---|---|---|---|---|---|---|---|---|
| 6-9 | 0.6×1 | 0.6 | 0.3 | 0.4 | 0.2 | 0.2 | 1.7 | 4 |
| 10-13 | 0.6×2 | 0.6 | 0.3 | 0.4 | 0.2 | 0.2 | 1.1 | 4 |
| 14-17 | 0.6×3 | 0.6 | 0.3 | 0.4 | 0.2 | 0.2 | 0.5 | 4 |
| 18-21 | 0.6×4 | 0.6 | 0.3 | 0.4 | 0.2 | 0.2 | 0 | 4.1 |
| 引物名称Primer name | 引物序列Primer sequence(5'-3') | 目的Purpose |
|---|---|---|
| Step1_F | TTGATGCCTGGCAGTTCCCT | 第一步组装结果检测Step 1 assembly result detection |
| Step1_R | CGAACCGAACAGGCTTATGT | |
| Step2_F | GGACCGTCGCTGTCAAGTATCA | 第二步组装结果检测 Step 2 assembly result detection |
| Step2_R | AAGAACTCCATCACCTTCATCTCCAG | |
| attB1_F | ACAAGTTTGTACAAAAAAGCAGGCT | 终载体组装结果检测 Result detection of final vectorr assembly |
| attB4_R | CAACTTTGTATAGAAAAGTTGGGTGTCT | |
| attB4_F | AGACACCCAACTTTTCTATACAAAGTTG | |
| attB3_R | CAACTTTATTATACAAAGTTGTGGAATCGAGC | |
| attB3_F | GCTCGATTCCACAACTTTGTATAATAAAGTTG | |
| attB2_R | ACCACTTTGTACAAGAAAGCTGG |
Table 4 Primers list
| 引物名称Primer name | 引物序列Primer sequence(5'-3') | 目的Purpose |
|---|---|---|
| Step1_F | TTGATGCCTGGCAGTTCCCT | 第一步组装结果检测Step 1 assembly result detection |
| Step1_R | CGAACCGAACAGGCTTATGT | |
| Step2_F | GGACCGTCGCTGTCAAGTATCA | 第二步组装结果检测 Step 2 assembly result detection |
| Step2_R | AAGAACTCCATCACCTTCATCTCCAG | |
| attB1_F | ACAAGTTTGTACAAAAAAGCAGGCT | 终载体组装结果检测 Result detection of final vectorr assembly |
| attB4_R | CAACTTTGTATAGAAAAGTTGGGTGTCT | |
| attB4_F | AGACACCCAACTTTTCTATACAAAGTTG | |
| attB3_R | CAACTTTATTATACAAAGTTGTGGAATCGAGC | |
| attB3_F | GCTCGATTCCACAACTTTGTATAATAAAGTTG | |
| attB2_R | ACCACTTTGTACAAGAAAGCTGG |
| 目的基因 Target gene | 靶点 Target | 第一步组装 Step 1 | 第二步组装 Step 2 | 多位点LR Multisite LR |
|---|---|---|---|---|
| WA352 | TAL | pFUS2_a4a(L1), pFUS2_a4b(L2), pFUS2_a3b(L3), pFUS2_a2b(L4), pFUS2_b3(L5) | pENTR E1 pF5A L1-L4(last HD) | WA352::mitoTALEN |
| pFUS2_a4a(R1), pFUS2_a4b(R2), pFUS2_a3b(R3), pFUS2_a2b(R4), pFUS2_b3(R5) | pENTR E3 pR5A L3-L2(last HD) | |||
| 总数Total | 10 | 2 | 1 |
Table 5 Vectors constructed in this study
| 目的基因 Target gene | 靶点 Target | 第一步组装 Step 1 | 第二步组装 Step 2 | 多位点LR Multisite LR |
|---|---|---|---|---|
| WA352 | TAL | pFUS2_a4a(L1), pFUS2_a4b(L2), pFUS2_a3b(L3), pFUS2_a2b(L4), pFUS2_b3(L5) | pENTR E1 pF5A L1-L4(last HD) | WA352::mitoTALEN |
| pFUS2_a4a(R1), pFUS2_a4b(R2), pFUS2_a3b(R3), pFUS2_a2b(R4), pFUS2_b3(R5) | pENTR E3 pR5A L3-L2(last HD) | |||
| 总数Total | 10 | 2 | 1 |
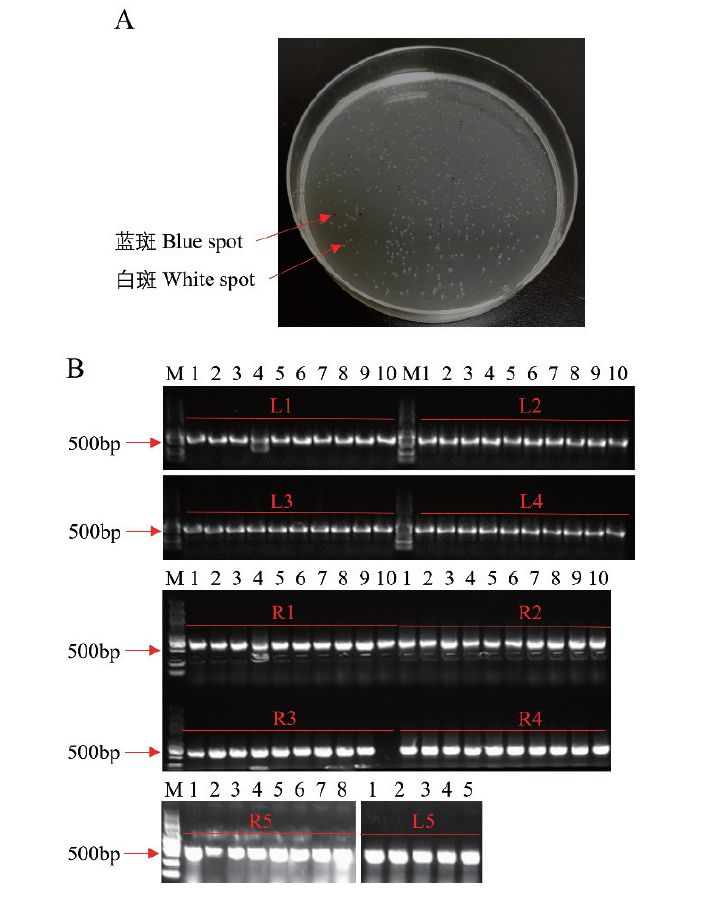
Fig. 5 Positive detection of assembly results in the first step A: Screening positive monoclonal via blue and white spot. B: The ten vectors L1-L5 and R1-R5 were detected by PCR

Fig. 7 Enzyme digestion of the assembly results in the second step A: The repeat variable diresidues(RVDs)corresponding to TAL-left(L)and TAL-right(R)of WA352 gene, where NI is the digestion site of Msc I. B: Gel image of the products of TAL-left(L)and TAL-right(R)digested by Msc I
| [1] |
Gao CX. Genome engineering for crop improvement and future agriculture[J]. Cell, 2021, 184(6): 1621-1635.
doi: 10.1016/j.cell.2021.01.005 pmid: 33581057 |
| [2] | Chang YZ, Liu BL, Jiang YY, et al. Induce male sterility by CRISPR/Cas9-mediated mitochondrial genome editing in tobacco[J]. Funct Integr Genomics, 2023, 23(3): 205. |
| [3] |
Kazama T, Okuno M, Watari Y, et al. Curing cytoplasmic male sterility via TALEN-mediated mitochondrial genome editing[J]. Nat Plants, 2019, 5(7): 722-730.
doi: 10.1038/s41477-019-0459-z pmid: 31285556 |
| [4] | Arimura SI, Ayabe H, Sugaya H, et al. Targeted gene disruption of ATP synthases 6-1 and 6-2 in the mitochondrial genome of Arabidopsis thaliana by mitoTALENs[J]. Plant J, 2020, 104(6): 1459-1471. |
| [5] |
Omukai S, Arimura SI, Toriyama K, et al. Disruption of mitochondrial open reading frame 352 partially restores pollen development in cytoplasmic male sterile rice[J]. Plant Physiol, 2021, 187(1): 236-246.
doi: 10.1093/plphys/kiab236 pmid: 34015134 |
| [6] | Takatsuka A, Kazama T, Arimura SI, et al. TALEN-mediated depletion of the mitochondrial gene orf312 proves that it is a Tadukan-type cytoplasmic male sterility-causative gene in rice[J]. Plant J, 2022, 110(4): 994-1004. |
| [7] |
Kuwabara K, Arimura SI, Shirasawa K, et al. orf137 triggers cytoplasmic male sterility in tomato[J]. Plant Physiol, 2022, 189(2): 465-468.
doi: 10.1093/plphys/kiac082 pmid: 35212743 |
| [8] |
Forner J, Kleinschmidt D, Meyer EH, et al. Targeted knockout of a conserved plant mitochondrial gene by genome editing[J]. Nat Plants, 2023, 9(11): 1818-1831.
doi: 10.1038/s41477-023-01538-2 pmid: 37814021 |
| [9] | Ayabe H, Toyoda A, Iwamoto A, et al. Mitochondrial gene defects in Arabidopsis can broadly affect mitochondrial gene expression through copy number[J]. Plant Physiol, 2023, 191(4): 2256-2275. |
| [10] | Xu FY, Su TB, Zhang XC, et al. Editing of ORF138 restores fertility of Ogura cytoplasmic male sterile broccoli via mitoTALENs[J]. Plant Biotechnol J, 2024, 22(5): 1325-1334. |
| [11] |
Nicolia A, Scotti N, D'Agostino N, et al. Mitochondrial DNA editing in potato through mitoTALEN and mitoTALECD: molecular characterization and stability of editing events[J]. Plant Methods, 2024, 20(1): 4.
doi: 10.1186/s13007-023-01124-9 pmid: 38183104 |
| [12] | Zhou JW, Nie LY, Zhang S, et al. Mitochondrial genome editing of WA352 via mitoTALENs restore fertility in cytoplasmic male sterile rice[J]. Plant Biotechnol J, 2024, 22(7): 1960-1962. |
| [13] |
Gaj T, Gersbach CA, Barbas CF III. ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering[J]. Trends Biotechnol, 2013, 31(7): 397-405.
doi: 10.1016/j.tibtech.2013.04.004 pmid: 23664777 |
| [14] |
Khalil AM. The genome editing revolution: review[J]. J Genet Eng Biotechnol, 2020, 18(1): 68.
doi: 10.1186/s43141-020-00078-y pmid: 33123803 |
| [15] | Arimura SI. MitoTALENs: a method for targeted gene disruption in plant mitochondrial genomes[J]. Methods Mol Biol, 2022, 2363: 335-340. |
| [16] |
Sakuma T, Ochiai H, Kaneko T, et al. Repeating pattern of non-RVD variations in DNA-binding modules enhances TALEN activity[J]. Sci Rep, 2013, 3: 3379.
doi: 10.1038/srep03379 pmid: 24287550 |
| [17] |
Sakuma T, Yamamoto T. Engineering customized TALENs using the platinum gate TALEN kit[J]. Methods Mol Biol, 2016, 1338: 61-70.
doi: 10.1007/978-1-4939-2932-0_6 pmid: 26443214 |
| [18] |
Guo JY, Zhang X, Chen XX, et al. Precision modeling of mitochondrial diseases in zebrafish via DdCBE-mediated mtDNA base editing[J]. Cell Discov, 2021, 7(1): 78.
doi: 10.1038/s41421-021-00307-9 pmid: 34480028 |
| No related articles found! |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||