








Biotechnology Bulletin ›› 2023, Vol. 39 ›› Issue (10): 1-16.doi: 10.13560/j.cnki.biotech.bull.1985.2023-0243
AN Chang1,2( ), LU Lin1, SHEN Meng-qian3, CHEN Sheng-zhen3, YE Kang-zhuo1, QIN Yuan1,2,3(
), LU Lin1, SHEN Meng-qian3, CHEN Sheng-zhen3, YE Kang-zhuo1, QIN Yuan1,2,3( ), ZHENG Ping1(
), ZHENG Ping1( )
)
Received:2023-03-17
Online:2023-10-26
Published:2023-11-28
Contact:
QIN Yuan, ZHENG Ping
E-mail:ancher0928@163.com;zhengping13@mails.ucas.ac.cn;yuanqin@fafu.edu.cn
AN Chang, LU Lin, SHEN Meng-qian, CHEN Sheng-zhen, YE Kang-zhuo, QIN Yuan, ZHENG Ping. Research Progress of bHLH Gene Family in Plants and Its Application Prospects in Medical Plants[J]. Biotechnology Bulletin, 2023, 39(10): 1-16.
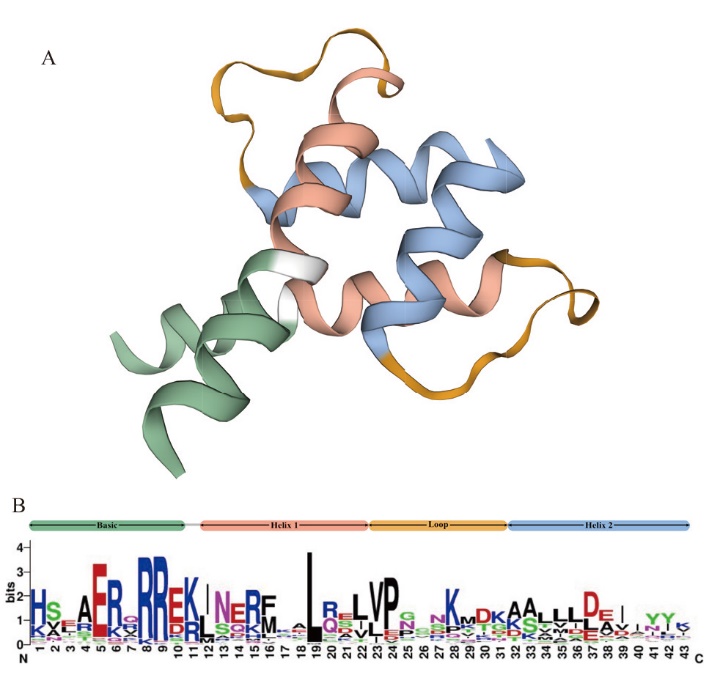
Fig. 1 Domain characteristics and distribution of bHLH family members from A. annua A: Three-dimensional protein structure of bHLH. B: Sequence logo of the bHLH domain generated in WebLogo
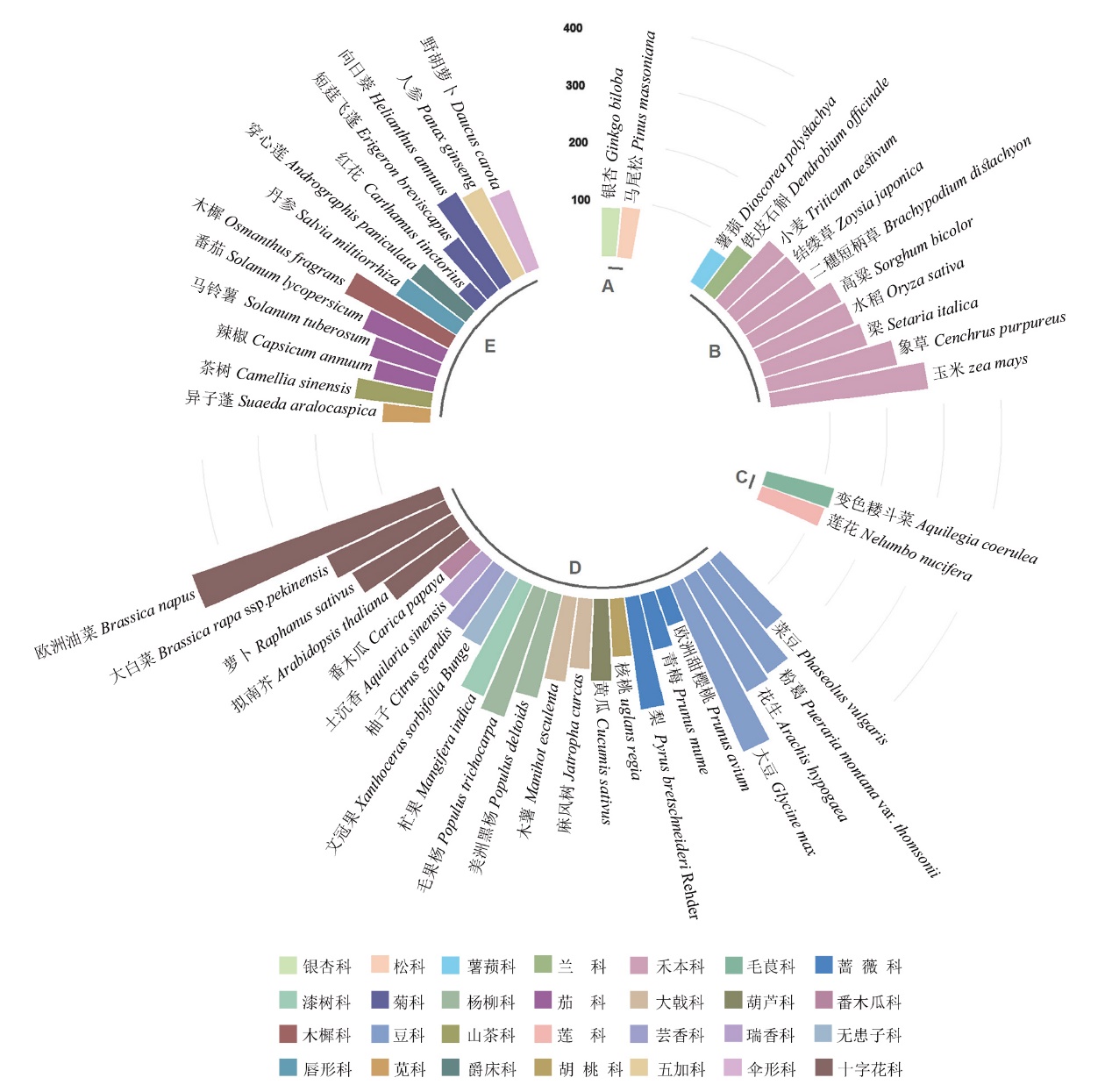
Fig. 2 Statistics of the number of bHLH family members in vascular plants A: Gymnosperms. B: Monocots. C: Basal groups of eudicots. D: Core eudicots(superrosids). E: Core eudicots(superasterids)
| [1] |
Rowe N, Speck T. Plant growth forms: an ecological and evolutionary perspective[J]. New Phytol, 2005, 166(1): 61-72.
pmid: 15760351 |
| [2] |
Riechmann JL, Ratcliffe OJ. A genomic perspective on plant transcription factors[J]. Curr Opin Plant Biol, 2000, 3(5): 423-434.
pmid: 11019812 |
| [3] | 郭兰萍, 康传志, 周涛, 等. 中药生态农业最新进展及展望[J]. 中国中药杂志, 2021, 46(8): 1851-1857. |
| Guo LP, Kang CZ, Zhou T, et al. Ecological agriculture of Chinese materia medica: update and future perspectives[J]. China J Chin Mater Med, 2021, 46(8): 1851-1857. | |
| [4] | 郭兰萍, 周良云, 康传志, 等. 药用植物适应环境胁迫的策略及道地药材“拟境栽培”[J]. 中国中药杂志, 2020, 45(9): 1969-1974. |
| Guo LP, Zhou LY, Kang CZ, et al. Strategies for medicinal plants adapting environmental stress and “simulative habitat cultivation” of Dao-di herbs[J]. China J Chin Mater Med, 2020, 45(9): 1969-1974. | |
| [5] | 蒋待泉, 王红阳, 康传志, 等. 复合胁迫对药用植物次生代谢的影响及机制[J]. 中国中药杂志, 2020, 45(9): 2009-2016. |
| Jiang DQ, Wang HY, Kang CZ, et al. Influence and mechanism of stress combination on medicinal plants secondary metabolism[J]. China J Chin Mater Med, 2020, 45(9): 2009-2016. | |
| [6] | Sun W, Xu ZC, Song C, et al. Herbgenomics: decipher molecular genetics of medicinal plants[J]. Innovation(Camb), 2022, 3(6): 100322. |
| [7] |
Feller A, Machemer K, Braun EL, et al. Evolutionary and comparative analysis of MYB and bHLH plant transcription factors[J]. Plant J, 2011, 66(1): 94-116.
doi: 10.1111/tpj.2011.66.issue-1 URL |
| [8] | 鄢丹, 王伽伯, 李俊贤, 等. 论道地药材品质辨识及其与生态环境的相关性研究策略[J]. 中国中药杂志, 2012, 37(17): 2672-2675. |
| Yan D, Wang JB, Li JX, et al. Strategy for research on quality identification and ecological environment-related of Dao-di herb[J]. China J Chin Mater Med, 2012, 37(17): 2672-2675. | |
| [9] |
Ledent V, Vervoort M. The basic helix-loop-helix protein family: comparative genomics and phylogenetic analysis[J]. Genome Res, 2001, 11(5): 754-770.
doi: 10.1101/gr.177001 pmid: 11337472 |
| [10] |
Jones S. An overview of the basic helix-loop-helix proteins[J]. Genome Biol, 2004, 5(6): 226.
doi: 10.1186/gb-2004-5-6-226 pmid: 15186484 |
| [11] |
Ludwig SR, Habera LF, Dellaporta SL, et al. Lc, a member of the maize R gene family responsible for tissue-specific anthocyanin production, encodes a protein similar to transcriptional activators and contains the myc-homology region[J]. Proc Natl Acad Sci USA, 1989, 86(18): 7092-7096.
doi: 10.1073/pnas.86.18.7092 pmid: 2674946 |
| [12] |
Zhang TT, Lyu W, Zhang HS, et al. Genome-wide analysis of the basic helix-loop-helix(bHLH)transcription factor family in maize[J]. BMC Plant Biol, 2018, 18(1): 235.
doi: 10.1186/s12870-018-1441-z |
| [13] |
Skinner MK, Rawls A, Wilson-Rawls J, et al. Basic helix-loop-helix transcription factor gene family phylogenetics and nomenclature[J]. Differentiation, 2010, 80(1): 1-8.
doi: 10.1016/j.diff.2010.02.003 pmid: 20219281 |
| [14] |
Toledo-Ortiz G, Huq E, Quail PH. The Arabidopsis basic/helix-loop-helix transcription factor family[J]. Plant Cell, 2003, 15(8): 1749-1770.
doi: 10.1105/tpc.013839 pmid: 12897250 |
| [15] |
Roig-Villanova I, Bou-Torrent J, Galstyan A, et al. Interaction of shade avoidance and auxin responses: a role for two novel atypical bHLH proteins[J]. EMBO J, 2007, 26(22): 4756-4767.
doi: 10.1038/sj.emboj.7601890 pmid: 17948056 |
| [16] |
Medina-Puche L, Martínez-Rivas FJ, Molina-Hidalgo FJ, et al. An atypical HLH transcriptional regulator plays a novel and important role in strawberry ripened receptacle[J]. BMC Plant Biol, 2019, 19(1): 586.
doi: 10.1186/s12870-019-2092-4 pmid: 31881835 |
| [17] |
Carretero-Paulet L, Galstyan A, Roig-Villanova I, et al. Genome-wide classification and evolutionary analysis of the bHLH family of transcription factors in Arabidopsis, poplar, rice, moss, and algae[J]. Plant Physiol, 2010, 153(3): 1398-1412.
doi: 10.1104/pp.110.153593 pmid: 20472752 |
| [18] | 位志坤, 许自成, 贾国涛. bHLH转录因子家族调控植物非生物逆境胁迫响应的研究进展[J/OL]. 分子植物育种, 2022. https://kns.cnki.net/kcms/detail/46.1068.S.20220615.0859.002.html. |
| Wei ZK, Xu ZC, Jia GT, et al. BHLH transcription factor family regulates abiotic stress response in plants research progress[J/OL]. Mol Breed, 2022. https://kns.cnki.net/kcms/detail/46.1068.S.20220615.0859.002.html. | |
| [19] |
Hong YQ, Ahmad N, Tian YY, et al. Genome-wide identification, expression analysis, and subcellular localization of Carthamus tinctorius bHLH transcription factors[J]. Int J Mol Sci, 2019, 20(12): 3044.
doi: 10.3390/ijms20123044 URL |
| [20] |
Shen W, Cui X, Li H, et al. Genome-wide identification and analyses of bHLH family genes in Brassica napus[J]. Can J Plant Sci, 2019, 99(5): 589-598.
doi: 10.1139/cjps-2018-0230 |
| [21] |
Pires N, Dolan L. Origin and diversification of basic-helix-loop-helix proteins in plants[J]. Mol Biol Evol, 2010, 27(4): 862-874.
doi: 10.1093/molbev/msp288 pmid: 19942615 |
| [22] |
Gao F, Robe K, Gaymard F, et al. The transcriptional control of iron homeostasis in plants: a tale of bHLH transcription factors?[J]. Front Plant Sci, 2019, 10: 6.
doi: 10.3389/fpls.2019.00006 pmid: 30713541 |
| [23] |
Gao C, Sun JL, Wang CQ, et al. Genome-wide analysis of basic/helix-loop-helix gene family in peanut and assessment of its roles in pod development[J]. PLoS One, 2017, 12(7): e0181843.
doi: 10.1371/journal.pone.0181843 URL |
| [24] |
Mao TY, Liu YY, Zhu HH, et al. Genome-wide analyses of the bHLH gene family reveals structural and functional characteristics in the aquatic plant Nelumbo nucifera[J]. PeerJ, 2019, 7: e7153.
doi: 10.7717/peerj.7153 URL |
| [25] |
Chu Y, Xiao SM, Su H, et al. Genome-wide characterization and analysis of bHLH transcription factors in Panax ginseng[J]. Acta Pharm Sin B, 2018, 8(4): 666-677.
doi: 10.1016/j.apsb.2018.04.004 URL |
| [26] |
Mauxion JP, Chevalier C, Gonzalez N. Complex cellular and molecular events determining fruit size[J]. Trends Plant Sci, 2021, 26(10): 1023-1038.
doi: 10.1016/j.tplants.2021.05.008 URL |
| [27] |
Zhu ZG, Chen GP, Guo XH, et al. Overexpression of SlPRE2, an atypical bHLH transcription factor, affects plant morphology and fruit pigment accumulation in tomato[J]. Sci Rep, 2017, 7(1): 5786.
doi: 10.1038/s41598-017-04092-y |
| [28] |
Zhu ZG, Liang HL, Chen GP, et al. The bHLH transcription factor SlPRE2 regulates tomato fruit development and modulates plant response to gibberellin[J]. Plant Cell Rep, 2019, 38(9): 1053-1064.
doi: 10.1007/s00299-019-02425-x pmid: 31123809 |
| [29] |
Tan C, Qiao HL, Ma M, et al. Genome-wide identification and characterization of melon bHLH transcription factors in regulation of fruit development[J]. Plants, 2021, 10(12): 2721.
doi: 10.3390/plants10122721 URL |
| [30] |
Yu JQ, Gu KD, Sun CH, et al. The apple bHLH transcription factor MdbHLH3 functions in determining the fruit carbohydrates and malate[J]. Plant Biotechnol J, 2021, 19(2): 285-299.
doi: 10.1111/pbi.v19.2 URL |
| [31] |
Zhang Y, Mitsuda N, Yoshizumi T, et al. Two types of bHLH transcription factor determine the competence of the pericycle for lateral root initiation[J]. Nat Plants, 2021, 7(5): 633-643.
doi: 10.1038/s41477-021-00919-9 pmid: 34007039 |
| [32] |
Groszmann M, Paicu T, Smyth DR. Functional domains of SPATULA, a bHLH transcription factor involved in carpel and fruit development in Arabidopsis[J]. Plant J, 2008, 55(1): 40-52.
doi: 10.1111/tpj.2008.55.issue-1 URL |
| [33] |
Ichihashi Y, Horiguchi G, Gleissberg S, et al. The bHLH transcription factor SPATULA controls final leaf size in Arabidopsis thaliana[J]. Plant Cell Physiol, 2010, 51(2): 252-261.
doi: 10.1093/pcp/pcp184 pmid: 20040585 |
| [34] |
Groszmann M, Paicu T, Alvarez JP, et al. SPATULA and ALCATRAZ, are partially redundant, functionally diverging bHLH genes required for Arabidopsis gynoecium and fruit development[J]. Plant J, 2011, 68(5): 816-829.
doi: 10.1111/tpj.2011.68.issue-5 URL |
| [35] |
Makkena S, Lamb RS. The bHLH transcription factor SPATULA is a key regulator of organ size in Arabidopsis thaliana[J]. Plant Signal Behav, 2013, 8(5): e24140.
doi: 10.4161/psb.24140 URL |
| [36] |
Makkena S, Lamb RS. The bHLH transcription factor SPATULA regulates root growth by controlling the size of the root meristem[J]. BMC Plant Biol, 2013, 13: 1.
doi: 10.1186/1471-2229-13-1 URL |
| [37] |
Li HM, Sun JQ, Xu YX, et al. The bHLH-type transcription factor AtAIB positively regulates ABA response in Arabidopsis[J]. Plant Mol Biol, 2007, 65(5): 655-665.
doi: 10.1007/s11103-007-9230-3 URL |
| [38] |
Liu WW, Tai HH, Li SS, et al. bHLH122 is important for drought and osmotic stress resistance in Arabidopsis and in the repression of ABA catabolism[J]. New Phytol, 2014, 201(4): 1192-1204.
doi: 10.1111/nph.2014.201.issue-4 URL |
| [39] |
Yang TR, Yao SF, Hao L, et al. Wheat bHLH-type transcription factor gene TabHLH1 is crucial in mediating osmotic stresses tolerance through modulating largely the ABA-associated pathway[J]. Plant Cell Rep, 2016, 35(11): 2309-2323.
doi: 10.1007/s00299-016-2036-5 URL |
| [40] |
Seo JS, Joo J, Kim MJ, et al. OsbHLH148, a basic helix-loop-helix protein, interacts with OsJAZ proteins in a jasmonate signaling pathway leading to drought tolerance in rice[J]. Plant J, 2011, 65(6): 907-921.
doi: 10.1111/tpj.2011.65.issue-6 URL |
| [41] |
Castilhos G, Lazzarotto F, Spagnolo-Fonini L, et al. Possible roles of basic helix-loop-helix transcription factors in adaptation to drought[J]. Plant Sci, 2014, 223: 1-7.
doi: 10.1016/j.plantsci.2014.02.010 pmid: 24767109 |
| [42] |
Li CJ, Yan CX, Sun QX, et al. The bHLH transcription factor AhbHLH112 improves the drought tolerance of peanut[J]. BMC Plant Biol, 2021, 21(1): 540.
doi: 10.1186/s12870-021-03318-6 pmid: 34784902 |
| [43] |
Zhao Q, Fan ZH, Qiu LN, et al. MdbHLH130, an apple bHLH transcription factor, confers water stress resistance by regulating stomatal closure and ROS homeostasis in transgenic tobacco[J]. Front Plant Sci, 2020, 11: 543696.
doi: 10.3389/fpls.2020.543696 URL |
| [44] |
Dong Y, Wang CP, Han X, et al. A novel bHLH transcription factor PebHLH35 from Populus euphratica confers drought tolerance through regulating stomatal development, photosynthesis and growth in Arabidopsis[J]. Biochem Biophys Res Commun, 2014, 450(1): 453-458.
doi: 10.1016/j.bbrc.2014.05.139 URL |
| [45] |
Qiu JR, Huang Z, Xiang XY, et al. MfbHLH38, a Myrothamnus flabellifolia bHLH transcription factor, confers tolerance to drought and salinity stresses in Arabidopsis[J]. BMC Plant Biol, 2020, 20(1): 542.
doi: 10.1186/s12870-020-02732-6 |
| [46] | Ji XY, Nie XG, Liu YJ, et al. A bHLH gene from Tamarix hispida improves abiotic stress tolerance by enhancing osmotic potential and decreasing reactive oxygen species accumulation[J]. Tree Physiol, 2016, 36(2): 193-207. |
| [72] |
Wang WJ, Shinwari KI, Zhang H, et al. The bHLH transcription factor OsbHLH057 regulates iron homeostasis in rice[J]. Int J Mol Sci, 2022, 23(23): 14869.
doi: 10.3390/ijms232314869 URL |
| [73] |
Wang WJ, Ye J, Xu H, et al. OsbHLH061 links topless/topless-related repressor proteins with positive regulator of iron homeostasis 1 to maintain iron homeostasis in rice[J]. New Phytol, 2022, 234(5): 1753-1769.
doi: 10.1111/nph.18096 pmid: 35288933 |
| [74] |
Cui YC, Xu GY, Wang ML, et al. Expression of OsMSR3 in Arabidopsis enhances tolerance to cadmium stress[J]. Plant Cell Tiss Organ Cult, 2013, 113(2): 331-340.
doi: 10.1007/s11240-012-0275-x URL |
| [75] |
Xu ZL, Liu XQ, He XL, et al. The soybean basic hlix-loop-hlix transcription factor ORG3-like enhances cadmium tolerance via increased iron and reduced cadmium uptake and transport from roots to shoots[J]. Front Plant Sci, 2017, 8: 1098.
doi: 10.3389/fpls.2017.01098 URL |
| [76] | 宋倩. 铝胁迫下大豆GmbHLH30转录因子调控相关基因的分离与鉴定[D]. 昆明: 昆明理工大学, 2014. |
| Song Q. Isolation and characterization of relate genes regulated by soybean GmbHLH30 transcription factor under Al stress[D]. Kunming: Kunming University of Science and Technology, 2014. | |
| [77] |
Sun KL, Wang HY, Xia ZL. The maize bHLH transcription factor bHLH105 confers manganese tolerance in transgenic tobacco[J]. Plant Sci, 2019, 280: 97-109.
doi: S0168-9452(18)31005-7 pmid: 30824033 |
| [78] |
Hichri I, Barrieu F, Bogs J, et al. Recent advances in the transcriptional regulation of the flavonoid biosynthetic pathway[J]. J Exp Bot, 2011, 62(8): 2465-2483.
doi: 10.1093/jxb/erq442 pmid: 21278228 |
| [79] |
McClean PE, Lee RA, Howe K, et al. The common bean V gene encodes flavonoid 3'5' hydroxylase: a major mutational target for flavonoid diversity in angiosperms[J]. Front Plant Sci, 2022, 13: 869582.
doi: 10.3389/fpls.2022.869582 URL |
| [80] |
Baudry A, Caboche M, Lepiniec L. TT8 controls its own expression in a feedback regulation involving TTG1 and homologous MYB and bHLH factors, allowing a strong and cell-specific accumulation of flavonoids in Arabidopsis thaliana[J]. Plant J, 2006, 46(5): 768-779.
doi: 10.1111/tpj.2006.46.issue-5 URL |
| [47] |
Liang BB, Wan SG, Ma QL, et al. A novel bHLH transcription factor PtrbHLH66 from trifoliate orange positively regulates plant drought tolerance by mediating root growth and ROS scavenging[J]. Int J Mol Sci, 2022, 23(23): 15053.
doi: 10.3390/ijms232315053 URL |
| [48] |
Liu H, Yang Y, Liu DD, et al. Transcription factor TabHLH49 positively regulates dehydrin WZY2 gene expression and enhances drought stress tolerance in wheat[J]. BMC Plant Biol, 2020, 20(1): 259.
doi: 10.1186/s12870-020-02474-5 |
| [49] |
Gu XY, Gao SX, Li J, et al. The bHLH transcription factor regulated gene OsWIH2 is a positive regulator of drought tolerance in rice[J]. Plant Physiol Biochem, 2021, 169: 269-279.
doi: 10.1016/j.plaphy.2021.11.031 URL |
| [50] |
Kim J, Kim HY. Functional analysis of a calcium-binding transcription factor involved in plant salt stress signaling[J]. FEBS Lett, 2006, 580(22): 5251-5256.
pmid: 16962584 |
| [51] |
Krishnamurthy P, Vishal B, Khoo K, et al. Expression of AoN-HX1 increases salt tolerance of rice and Arabidopsis, and bHLH transcription factors regulate AtNHX1 and AtNHX6 in Arabidop-sis[J]. Plant Cell Rep, 2019, 38(10): 1299-1315.
doi: 10.1007/s00299-019-02450-w pmid: 31350571 |
| [52] |
Verma D, Jalmi SK, Bhagat PK, et al. A bHLH transcription factor, MYC2, imparts salt intolerance by regulating proline biosynthesis in Arabidopsis[J]. FEBS J, 2020, 287(12): 2560-2576.
doi: 10.1111/febs.v287.12 URL |
| [53] |
Zhang HF, Guo JB, Chen XQ, et al. Pepper bHLH transcription factor CabHLH035 contributes to salt tolerance by modulating ion homeostasis and proline biosynthesis[J]. Hortic Res, 2022, 9: uhac203.
doi: 10.1093/hr/uhac203 URL |
| [54] |
Wang FB, Zhu H, Chen DH, et al. A grape bHLH transcription factor gene, VvbHLH1, increases the accumulation of flavonoids and enhances salt and drought tolerance in transgenic Arabidopsis thaliana[J]. Plant Cell Tiss Organ Cult, 2016, 125(2): 387-398.
doi: 10.1007/s11240-016-0953-1 URL |
| [55] |
Yu CM, Yan M, Dong HZ, et al. Maize bHLH55 functions positively in salt tolerance through modulation of AsA biosynthesis by directly regulating GDP-mannose pathway genes[J]. Plant Sci, 2021, 302: 110676.
doi: 10.1016/j.plantsci.2020.110676 URL |
| [56] |
Song YS, Li JL, Sui Y, et al. The sweet sorghum SbWRKY50 is negatively involved in salt response by regulating ion homeostasis[J]. Plant Mol Biol, 2020, 102(6): 603-614.
doi: 10.1007/s11103-020-00966-4 pmid: 32052233 |
| [57] |
Zhang KY, Zhang ZP, Lu F, et al. Bulked segregant analysis-sequencing identification of candidate genes for salt tolerance at the seedling stage of sorghum(Sorghum bicolor)[J]. Plant Breed, 2022, 141(3): 366-378.
doi: 10.1111/pbr.v141.3 URL |
| [58] |
Song YS, Li SM, Sui Y, et al. SbbHLH85, a bHLH member, modulates resilience to salt stress by regulating root hair growth in sorghum[J]. Theor Appl Genet, 2022, 135(1): 201-216.
doi: 10.1007/s00122-021-03960-6 |
| [59] |
Chinnusamy V, Ohta M, Kanrar S, et al. ICE1: a regulator of cold-induced transcriptome and freezing tolerance in Arabidopsis[J]. Genes Dev, 2003, 17(8): 1043-1054.
doi: 10.1101/gad.1077503 URL |
| [60] |
Xu WR, Jiao YT, Li RM, et al. Chinese wild-growing Vitis amurensis ICE1 and ICE2 encode MYC-type bHLH transcription activators that regulate cold tolerance in Arabidopsis[J]. PLoS One, 2014, 9(7): e102303.
doi: 10.1371/journal.pone.0102303 URL |
| [61] |
Jin R, Kim HS, Yu T, et al. Identification and function analysis of bHLH genes in response to cold stress in sweetpotato[J]. Plant Physiol Biochem, 2021, 169: 224-235.
doi: 10.1016/j.plaphy.2021.11.027 URL |
| [62] |
Verma RK, Kumar VVS, Yadav SK, et al. Overexpression of Arabidopsis ICE1 enhances yield and multiple abiotic stress tolerance in indica rice[J]. Plant Signal Behav, 2020, 15(11): 1814547.
doi: 10.1080/15592324.2020.1814547 URL |
| [63] |
Yao PF, Sun ZX, Li CL, et al. Overexpression of Fagopyrum tataricum FtbHLH2 enhances tolerance to cold stress in transgenic Arabidopsis[J]. Plant Physiol Biochem, 2018, 125: 85-94.
doi: 10.1016/j.plaphy.2018.01.028 URL |
| [64] |
Yang XY, Wang R, Hu QL, et al. DlICE1, a stress-responsive gene from Dimocarpus longan, enhances cold tolerance in transgenic Arabidopsis[J]. Plant Physiol Biochem, 2019, 142: 490-499.
doi: 10.1016/j.plaphy.2019.08.007 URL |
| [65] |
Thao NP, Khan MI, Thu NBA, et al. Role of ethylene and its cross talk with other signaling molecules in plant responses to heavy metal stress[J]. Plant Physiol, 2015, 169(1): 73-84.
doi: 10.1104/pp.15.00663 pmid: 26246451 |
| [66] |
Kim SA, LaCroix IS, Gerber SA, et al. The iron deficiency response in Arabidopsis thaliana requires the phosphorylated transcription factor URI[J]. Proc Natl Acad Sci USA, 2019, 116(50): 24933-24942.
doi: 10.1073/pnas.1916892116 URL |
| [81] |
Deng J, Li JJ, Su MY, et al. A bHLH gene NnTT8 of Nelumbo nucifera regulates anthocyanin biosynthesis[J]. Plant Physiol Biochem, 2021, 158: 518-523.
doi: 10.1016/j.plaphy.2020.11.038 URL |
| [82] |
Jia N, Wang JJ, Liu JM, et al. DcTT8, a bHLH transcription factor, regulates anthocyanin biosynthesis in Dendrobium candidum[J]. Plant Physiol Biochem, 2021, 162: 603-612.
doi: 10.1016/j.plaphy.2021.03.006 URL |
| [83] |
Qi Y, Zhou L, Han LL, et al. PsbHLH1, a novel transcription factor involved in regulating anthocyanin biosynthesis in tree peony(Paeonia suffruticosa)[J]. Plant Physiol Biochem, 2020, 154: 396-408.
doi: 10.1016/j.plaphy.2020.06.015 URL |
| [84] |
Li H, Yang Z, Zeng QW, et al. Abnormal expression of bHLH3 disrupts a flavonoid homeostasis network, causing differences in pigment composition among mulberry fruits[J]. Hortic Res, 2020, 7(1): 83.
doi: 10.1038/s41438-020-0302-8 |
| [85] |
Zhao R, Song XX, Yang N, et al. Expression of the subgroup IIIf bHLH transcription factor CpbHLH1 from Chimonanthus praecox(L.) in transgenic model plants inhibits anthocyanin accumulation[J]. Plant Cell Rep, 2020, 39(7): 891-907.
doi: 10.1007/s00299-020-02537-9 |
| [86] | Zhang JH, Lyu HZ, Liu WJ, et al. bHLH transcription factor SmbHLH92 negatively regulates biosynthesis of phenolic acids and tanshinones in Salvia miltiorrhiza[J]. Chin Herb Med, 2020, 12(3): 237-246. |
| [87] |
Xing BC, Yang DF, Yu HZ, et al. Overexpression of SmbHLH10 enhances tanshinones biosynthesis in Salvia miltiorrhiza hairy roots[J]. Plant Sci, 2018, 276: 229-238.
doi: 10.1016/j.plantsci.2018.07.016 URL |
| [88] |
Mertens J, Pollier J, Vanden Bossche R, et al. The bHLH transcription factors TSAR1 and TSAR2 regulate triterpene saponin biosynthesis in Medicago truncatula[J]. Plant Physiol, 2016, 170(1): 194-210.
doi: 10.1104/pp.15.01645 pmid: 26589673 |
| [89] |
De Boer K, Tilleman S, Pauwels L, et al. apetala2/ethylene response factor and basic helix-loop-helix tobacco transcription factors cooperatively mediate jasmonate-elicited nicotine biosynthesis[J]. Plant J, 2011, 66(6): 1053-1065.
doi: 10.1111/tpj.2011.66.issue-6 URL |
| [90] |
Xiao L, Huang D, Wu ZD, et al. Genome-wide identification of the bHLH transcription factor family and analysis of bHLH genes related to puerarin biosynthesis in Pueraria lobata var. thomsonii(Benth.)[J]. Plant Gene, 2023, 33: 100390.
doi: 10.1016/j.plgene.2022.100390 URL |
| [67] |
Gao F, Robe K, Bettembourg M, et al. The transcription factor bHLH121 interacts with bHLH105(ILR3)and its closest homologs to regulate iron homeostasis in Arabidopsis[J]. Plant Cell, 2020, 32(2): 508-524.
doi: 10.1105/tpc.19.00541 URL |
| [68] | Akmakjian GZ, Riaz N, Guerinot ML. Photoprotection during iron deficiency is mediated by the bHLH transcription factors PYE and ILR3[J]. Proc Natl Acad Sci USA, 2021, 118(40): e2024918118. |
| [69] |
Li YY, Sui XY, Yang JS, et al. A novel bHLH transcription factor, NtbHLH1, modulates iron homeostasis in tobacco(Nicotiana tabacum L.)[J]. Biochem Biophys Res Commun, 2020, 522(1): 233-239.
doi: 10.1016/j.bbrc.2019.11.063 URL |
| [70] |
Li L, Gao WW, Peng Q, et al. Two soybean bHLH factors regulate response to iron deficiency[J]. J Integr Plant Biol, 2018, 60(7): 608-622.
doi: 10.1111/jipb.12651 |
| [71] |
Wang SD, Li L, Ying YH, et al. A transcription factor OsbHLH156 regulates Strategy II iron acquisition through localising IRO2 to the nucleus in rice[J]. New Phytol, 2020, 225(3): 1247-1260.
doi: 10.1111/nph.16232 pmid: 31574173 |
| [91] |
Sánchez-Pérez R, Pavan S, Mazzeo R, et al. Mutation of a bHLH transcription factor allowed almond domestication[J]. Science, 2019, 364(6445): 1095-1098.
doi: 10.1126/science.aav8197 pmid: 31197015 |
| [92] |
Zheng H, Fu XQ, Shao J, et al. Transcriptional regulatory network of high-value active ingredients in medicinal plants[J]. Trends Plant Sci, 2023, 28(4): 429-446.
doi: 10.1016/j.tplants.2022.12.007 pmid: 36621413 |
| [93] |
Zhang X, Luo HM, Xu ZC, et al. Genome-wide characterisation and analysis of bHLH transcription factors related to tanshinone biosynthesis in Salvia miltiorrhiza[J]. Sci Rep, 2015, 5: 11244.
doi: 10.1038/srep11244 pmid: 26174967 |
| [94] | 李林. 基于茉莉素信号介导的丹参转录因子SmbHLH59、SmMYB97和SmWRKY14的功能研究[D]. 西安: 陕西师范大学, 2021. |
| Li L. Functional study on transcription factors SmbHLH59, SmMYB97 and SmWRKY14 of Salvia Miltiorrhiza based on Jasmine signal[D]. Xi'an: Shaanxi Normal University, 2021. | |
| [95] |
Wang H, Li SY, Li YA, et al. MED25 connects enhancer-promoter looping and MYC2-dependent activation of jasmonate signalling[J]. Nat Plants, 2019, 5(6): 616-625.
doi: 10.1038/s41477-019-0441-9 pmid: 31182849 |
| [96] |
Liu SC, Wang Y, Shi M, et al. SmbHLH60 and SmMYC2 antagonistically regulate phenolic acids and anthocyanins biosynthesis in Salvia miltiorrhiza[J]. J Adv Res, 2022, 42: 205-219.
doi: 10.1016/j.jare.2022.02.005 URL |
| [97] | 方庆. 基于SmbHLH124转录因子的丹参次生代谢工程研究[D]. 成都: 电子科技大学, 2021. |
| Fang Q. Metabolic engineering with transcription factor SmbHLH124 in Salvia miltiorrhiza[D]. Chengdu: University of Electronic Science and Technology of China, 2021. | |
| [98] | 甘雨, 吴端, 张栋, 等. 黄花蒿bHLH转录因子基因家族鉴定及光调控分析[J]. 中国现代中药, 2021, 23(3): 441-452. |
| Gan Y, Wu D, Zhang D, et al. Identification and light regulation analysis of bHLH transcription factor gene family in Artemisia annua[J]. Mod Chin Med, 2021, 23(3): 441-452. | |
| [99] |
Kayani SI, Shen Q, Ma YN, et al. The YABBY family transcription factor AaYABBY5 directly targets cytochrome P450 monooxygenase(CYP71AV1)and double-bond reductase 2(DBR2)involved in artemisinin biosynthesis in Artemisia annua[J]. Front Plant Sci, 2019, 10: 1084.
doi: 10.3389/fpls.2019.01084 URL |
| [100] |
Liao BS, Shen XF, Xiang L, et al. Allele-aware chromosome-level genome assembly of Artemisia annua reveals the correlation between ADS expansion and artemisinin yield[J]. Mol Plant, 2022, 15(8): 1310-1328.
doi: 10.1016/j.molp.2022.05.013 URL |
| [101] |
Gordân R, Shen N, Dror I, et al. Genomic regions flanking E-box binding sites influence DNA binding specificity of bHLH transcription factors through DNA shape[J]. Cell Rep, 2013, 3(4): 1093-1104.
doi: 10.1016/j.celrep.2013.03.014 pmid: 23562153 |
| [102] |
Le Dréau G, Escalona R, Fueyo R, et al. E proteins sharpen neurogenesis by modulating proneural bHLH transcription factors'activity in an E-box-dependent manner[J]. eLife, 2018, 7: e37267.
doi: 10.7554/eLife.37267 URL |
| [103] |
Shi P, Fu XQ, Shen Q, et al. The roles of AaMIXTA1 in regulating the initiation of glandular trichomes and cuticle biosynthesis in Artemisia annua[J]. New Phytol, 2018, 217(1): 261-276.
doi: 10.1111/nph.14789 pmid: 28940606 |
| [104] |
Ji YP, Xiao JW, Shen YL, et al. Cloning and characterization of AabHLH1, a bHLH transcription factor that positively regulates artemisinin biosynthesis in Artemisia annua[J]. Plant Cell Physiol, 2014, 55(9): 1592-1604.
doi: 10.1093/pcp/pcu090 URL |
| [105] |
Zhang QZ, Wu NY, Jian DQ, et al. Overexpression of AaPIF3 promotes artemisinin production in Artemisia annua[J]. Ind Crops Prod, 2019, 138: 111476.
doi: 10.1016/j.indcrop.2019.111476 URL |
| [106] |
Xiang LE, Jian DQ, Zhang FY, et al. The cold-induced transcription factor bHLH112 promotes artemisinin biosynthesis indirectly via ERF1 in Artemisia annua[J]. J Exp Bot, 2019, 70(18): 4835-4848.
doi: 10.1093/jxb/erz220 pmid: 31087059 |
| [107] |
Bredow M, Vanderbeld B, Walker VK. Ice-binding proteins confer freezing tolerance in transgenic Arabidopsis thaliana[J]. Plant Biotechnol J, 2017, 15(1): 68-81.
doi: 10.1111/pbi.2017.15.issue-1 URL |
| [108] |
Shen Q, Huang HY, Xie LH, et al. Basic helix-loop-helix transcription factors AabHLH2 and AabHLH3 function antagonistically with AaMYC2 and are negative regulators in artemisinin biosynthesis[J]. Front Plant Sci, 2022, 13: 885622.
doi: 10.3389/fpls.2022.885622 URL |
| [109] | 周琪. AabHLH106调控青蒿非青蒿素倍半萜生物合成的功能研究[D]. 重庆: 西南大学, 2022. |
| Zhou Q. Functional characterization of AabHLH106 in the regulation on non-artemisinin sesquiterpenes biosynthesis in Artemisia annua[D]. Chongqing: Southwest University, 2022. | |
| [110] |
Heim MA, Jakoby M, Werber M, et al. The basic helix-loop-helix transcription factor family in plants: a genome-wide study of protein structure and functional diversity[J]. Mol Biol Evol, 2003, 20(5): 735-747.
doi: 10.1093/molbev/msg088 pmid: 12679534 |
| [111] |
Lamb P, McKnight SL. Diversity and specificity in transcriptional regulation: the benefits of heterotypic dimerization[J]. Trends Biochem Sci, 1991, 16(11): 417-422.
pmid: 1776171 |
| [112] |
Amoutzias GD, Robertson DL, Van de Peer Y, et al. Choose your partners: dimerization in eukaryotic transcription factors[J]. Trends Biochem Sci, 2008, 33(5): 220-229.
doi: 10.1016/j.tibs.2008.02.002 pmid: 18406148 |
| [113] |
Ben Rejeb K, Abdelly C, Savouré A. How reactive oxygen species and proline face stress together[J]. Plant Physiol Biochem, 2014, 80: 278-284.
doi: 10.1016/j.plaphy.2014.04.007 URL |
| [114] |
Mittler R. Abiotic stress, the field environment and stress combination[J]. Trends Plant Sci, 2006, 11(1): 15-19.
doi: 10.1016/j.tplants.2005.11.002 pmid: 16359910 |
| [115] |
Zhu L, Zhao MZ, Chen MY, et al. The bHLH gene family and its response to saline stress in Jilin ginseng, Panax ginseng C.A. Meyer[J]. Mol Genet Genomics, 2020, 295(4): 877-890.
doi: 10.1007/s00438-020-01658-w |
| [1] | LIN Hong-yan, GUO Xiao-rui, LIU Di, LI Hui, LU Hai. Molecular Mechanism of Transcriptional Factor AtbHLH68 in Regulating Cell Wall Development by Transcriptome Analysis [J]. Biotechnology Bulletin, 2023, 39(9): 105-116. |
| [2] | LYU Qiu-yu, SUN Pei-yuan, RAN Bin, WANG Jia-rui, CHEN Qing-fu, LI Hong-you. Cloning, Subcellular Localization and Expression Analysis of the Transcription Factor Gene FtbHLH3 in Fagopyrum tataricum [J]. Biotechnology Bulletin, 2023, 39(8): 194-203. |
| [3] | LI Bo, LIU He-xia, CHEN Yu-ling, ZHOU Xing-wen, ZHU Yu-lin. Cloning, Subcellular Localization and Expression Analysis of CnbHLH79 Transcription Factor from Camellia nitidissima [J]. Biotechnology Bulletin, 2023, 39(8): 241-250. |
| [4] | HU Hai-lin, XU Li, LI Xiao-xu, WANG Chen-can, MEI Man, DING Wen-jing, ZHAO Yuan-yuan. Advances in the Regulation of Plant Growth, Development and Stress Physiology by Small Peptide Hormones [J]. Biotechnology Bulletin, 2023, 39(7): 13-25. |
| [5] | LI Yu, LI Su-zhen, CHEN Ru-mei, LU Hai-qiang. Advances in the Regulation of Iron Homeostasis by bHLH Transcription Factors in Plant [J]. Biotechnology Bulletin, 2023, 39(7): 26-36. |
| [6] | ZHAO Xue-ting, GAO Li-yan, WANG Jun-gang, SHEN Qing-qing, ZHANG Shu-zhen, LI Fu-sheng. Cloning and Expression of AP2/ERF Transcription Factor Gene ShERF3 in Sugarcane and Subcellular Localization of Its Encoded Protein [J]. Biotechnology Bulletin, 2023, 39(6): 208-216. |
| [7] | LI Yuan-hong, GUO Yu-hao, CAO Yan, ZHU Zhen-zhou, WANG Fei-fei. Research Progress in the Microalgal Growth and Accumulation of Target Products Regulated by Exogenous Phytohormone [J]. Biotechnology Bulletin, 2023, 39(6): 61-72. |
| [8] | FENG Shan-shan, WANG Lu, ZHOU Yi, WANG You-ping, FANG Yu-jie. Research Progresses on WOX Family Genes in Regulating Plant Development and Abiotic Stress Response [J]. Biotechnology Bulletin, 2023, 39(5): 1-13. |
| [9] | ZHAI Ying, LI Ming-yang, ZHANG Jun, ZHAO Xu, YU Hai-wei, LI Shan-shan, ZHAO Yan, ZHANG Mei-juan, SUN Tian-guo. Heterologous Expression of Soybean Transcription Factor GmNF-YA19 Improves Drought Resistance of Transgenic Tobacco [J]. Biotechnology Bulletin, 2023, 39(5): 224-232. |
| [10] | XUE Jiao ZHU Qing-feng FENG Yan-zhao CHEN Pei LIU Wen-hua ZHANG Ai-xia LIU Qin-jian ZHANG Qi YU Yang. Advances in Upstream Open Reading Frame in Plant Genes [J]. Biotechnology Bulletin, 2023, 39(4): 157-165. |
| [11] | WEI Ming WANG Xin-yu WU Guo-qiang ZHAO Meng. The Role of NAD-dependent Deacetylase SRT in Plant Epigenetic Inheritance Regulation [J]. Biotechnology Bulletin, 2023, 39(4): 59-70. |
| [12] | SANG Tian, WANG Peng-cheng. Research Progress in Plant SUMOylation [J]. Biotechnology Bulletin, 2023, 39(3): 1-12. |
| [13] | YANG Chun-hong, DONG Lu, CHEN Lin, SONG Li. Characterization of Soybean VAS1 Gene Family and Its Involvement in Lateral Root Development [J]. Biotechnology Bulletin, 2023, 39(3): 133-142. |
| [14] | LIU Cheng-xia, SUN Zong-yan, LUO Yun-bo, ZHU Hong-liang, QU Gui-qin. Multifaceted Roles of bHLH Phosphorylation in Regulation of Plant Physiological Functions [J]. Biotechnology Bulletin, 2023, 39(3): 26-34. |
| [15] | ZHENG Min-min, LIU Jie, ZHAO Qing. Research Progress and Prospects of Biological Studies on the Medicinal Plant Scutellaria baicalensis [J]. Biotechnology Bulletin, 2023, 39(2): 10-23. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||