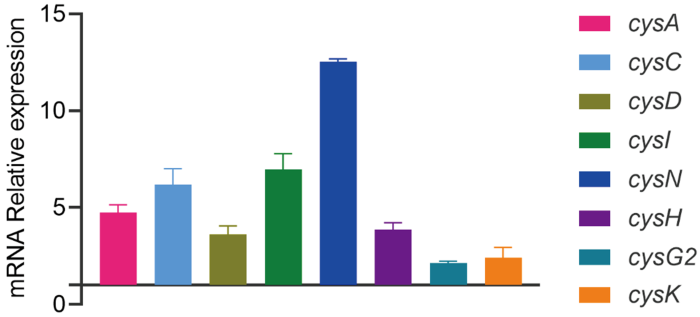嗜水气单胞菌(Aeromonas hydrophila)是一种广泛分布于自然界水体的人-畜-鱼共患病的革兰氏阴性菌。该菌的流行病爆发可导致养殖鱼类大量死亡,给渔业生产造成重大损失[1],同时该菌也能通过污染食物和伤口感染人类,引起败血症、肠胃炎等疾病危及人们的生命健康[2]。近年来,随着全球人口的增加,水产养殖成为人们饮食中鱼蛋白的主要来源,由此世界渔业增势逐年升高。但是,水产养殖业的快速发展以及大量人工养殖场的出现,也导致水产品细菌性病害越发严重,以至于人们开始大量使用抗菌药物[3]。然而,由于抗生素的大范围使用且滥用,导致耐药菌株层出不穷,细菌耐药性渐渐地变成全球公共卫生的一大难题[4]。因此,了解细菌对抗生素的耐药机制显得格外迫切。
在防治嗜水气单胞菌感染的药物中,喹诺酮类药物是最常见的抗生素之一。而恩诺沙星(enrofloxacin,ENR)属于第三代喹诺酮类抗菌药物,是一种畜禽和水产养殖常用的抗菌药物,具有广谱杀菌活性和很强的亲水性,对革兰氏阴性菌和革兰氏阳性菌都有很好的灭菌作用[5]。前期研究表明,ENR可通过与细菌DNA回旋酶亚基A结合抑制酶的连接与切割功能,阻碍细菌的复制从而体现出杀菌效果[6]。然而,近年来在全球范围内关于恩诺沙星耐药菌株的报道层出不穷,使得细菌耐药形势越来越严峻[7-8]。因为耐药菌株的出现不仅会给水产养殖业带来巨大的经济损失,还能通过污染水产品使食用的人类出现不良反应或感染抗生素耐药细菌,危害人们的生命健康[9]。为了进一步探究嗜水气单胞菌应对恩诺沙星胁迫的分子机制,本研究利用蛋白组学技术比较嗜水气单胞菌在有无恩诺沙星胁迫下蛋白表达的差异,以期为嗜水气单胞菌应对ENR胁迫的响应行为提供理论依据,并进一步了解嗜水气单胞菌对抗生素的内在适应机制。
1 材料与方法
1.1 材料
本研究使用的嗜水气单胞菌ATCC7966由广东海洋大学庞欢瑛教授惠赠。氯霉素(chloramphenicol,Chl)、氨苄青霉素(ampicillin,Amp)和恩诺沙星(enrofloxacin,ENR)购买于索莱宝生物科技有限公司;琼脂糖,琼脂粉,LB(Luria-Bertani)培养基均购买于北京拜尔迪生物技术有限公司;PCR产物纯化试剂盒、细菌DNA提取试剂盒、Taq酶和质粒提取试剂盒均购于南京诺维赞生物科技有限公司;限制性内切酶、核苷酸染料、DNA Marker(DL2000与DL5000)均购买于TaKaRa。
1.2 方法
1.2.1 最低抑菌浓度(minimal inhibitory concentration,MIC)值和生长曲线测定
采用二倍稀释法,用LB培养基对ENR进行梯度稀释,使其终浓度为0.031 2、0.015 6、0.007 8、0.003 9和0.001 95 µg/mL。挑取嗜水气单胞菌ATCC7966单克隆至5 mL LB试管中过夜培养16 h至稳定状态,次日转接培养至OD600 nm=1.0时,以1∶100的比例加入含有ENR的LB培养基中静置培养16 h(30℃培养箱中)后记录其MIC值,并利用酶标仪测定各浓度的吸光值。而后,以不加抗生素的LB为对照,测定在1/2 MIC、1/4 MIC、1/8 MIC浓度下的生长曲线。每个样品重复3次,每小时记录一次数据。
1.2.2 蛋白样品制备
将野生型嗜水气单胞菌ATCC7966(WT)菌株接种到5 mL LB液体培养基中,在30℃摇床中培养16 h至稳定期。继而把WT菌液按1∶100比例加入含有终浓度为0.007 8 µg/mL ENR的100 mL LB液体培养基中培养至OD600 nm=1.0时(以不加ENR为对照),收集菌体,8 000×g离心10 min后去除上清液再用预冷的1×PBS缓冲液洗涤2次。接着加入2 mL 裂解液(6 mol/L尿素, 2 mol/L硫脲, 100 mmol/L Tris-HCl pH 7.6,PI蛋白酶抑制剂)对菌体进行重悬,然后用超声波进行破碎15 min至样品透明。4℃离心取上清并用BCA试剂盒测定蛋白浓度。每个样品上30 µL跑SDS-PAGE而后进行考染。
1.2.3 质谱鉴定
取每组蛋白样品约50 µg至超滤管中,用 50 mmol/L二硫苏糖醇、56℃进行还原 40 min,再用25 mmol/L碘乙酰胺、避光静置进行烷基化30 min,离心弃滤液按 1∶20 加入胰蛋白酶,37℃孵育过夜降解成多肽[10]。将处理好的多肽样品用C18 柱(waters, Inc., milford, MA)除盐。使用EASY-nano-LC 色谱系统(thermo scientific Inc., waltham, MA, USA)以60 nL/min的流速分离脱盐的多肽样品[11]。分离梯度为: 0-18 min, B液(含0.1%甲酸的乙腈)从6%到12%线性上升; 18-77 min B液从12%到20%线性上升;77-109 min B液从20%到32%线性上升,然后在1 min内B液升至90%并维持到120 min。使用Orbitrap Fusion Lumos Tribrid 质谱仪(thermo scientific)分析多肽。具体参数设置为:离子源喷雾电压2.1 kV,循环时间3 s,离子转移管温度为300℃,扫描范围300-1 400 m/z,分辨率120 K, AGC靶点5e5,最大IT为50 ms。MS/MS采用离子阱法,AGC靶为5e3,最大IT为35 MS,碰撞能量为30%。质谱采集的数据依赖性采集(data dependent acquisition, DDA)原始数据导入到Spectronout Pulsar X(biognosys公司)建立DDA谱图库,建库参数使用默认最优参数“BGS factory setting”。然后将DIA原始数据导入到Spectronout Pulsar X进行蛋白质定性定量分析。
1.2.4 生物信息学分析
根据质谱数据,将P-value值小于0.05且差异倍数大于1.5或小于0.667的蛋白设为差异蛋白。借助David(
1.2.5 硫限制生长曲线分析
1.2.6 qPCR验证
表1 本研究所使用的引物对列表
Table 1
| Gene | Oligonucleotide sequence(5'-3') | Purpose |
|---|---|---|
| cysA-F | ATCGGGTGGTGCTGATGAACG | RT-qPCR |
| cysA-R | TGGGTAGCAGGCGGGTGATG | RT-qPCR |
| cysN-F | GGAGTCCGACAGCCAGAAG | RT-qPCR |
| cysN-R | GGCGGTGGAGAAATAGCG | RT-qPCR |
| cysD-F | GCTGGACATCTGGCAATACA | RT-qPCR |
| cysD-R | GCTCCTGCTTCACTTCATCTT | RT-qPCR |
| cysI-F | TCCACGCCAACGATCTCAA | RT-qPCR |
| cysI-R | GCCACGGCTGGTAAACTCAT | RT-qPCR |
| cysH-F | CCCCATCAATCGGGCTCT | RT-qPCR |
| cysH-R | TCCATCGGCTCCACCTTG | RT-qPCR |
| cysG2-F | CGCCTTTGCCTCCAACACC | RT-qPCR |
| cysG2-R | CCTTCTTGCCGACCGACACC | RT-qPCR |
| cysC-F | TATCGTGCTCACCGCCTTTA | RT-qPCR |
| cysC-R | CCTGCCCTAGCCTTCTTGTAG | RT-qPCR |
| cysK-F | ATCGGCGCCAACCTGATCT | RT-qPCR |
| cysK-R | CATTGCCAAGGCCAACGAG | RT-qPCR |
2 结果
2.1 嗜水气单胞菌对ENR的MIC值及生长曲线
通过测定嗜水气单胞菌在不同浓度的ENR下的生长情况进行测定,发现其对ENR的MIC值为0.031 2 µg/mL。因此根据测定所得的MIC值,本研究还对WT分别在1/2、1/4、1/8 MIC的ENR处理下的生长情况进行测定。结果如图1-A所示,与对照相比,嗜水气单胞菌在不同浓度的ENR胁迫下生长情况均受到抑制,但是在1/4 MIC处其生长情况受到明显抑制,并且还具有较高的活性。故本研究选择0.007 8 µg/mL ENR(1/4 MIC)作为处理浓度进行后续研究。
图1
图1
定量蛋白质组学数据分析
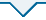
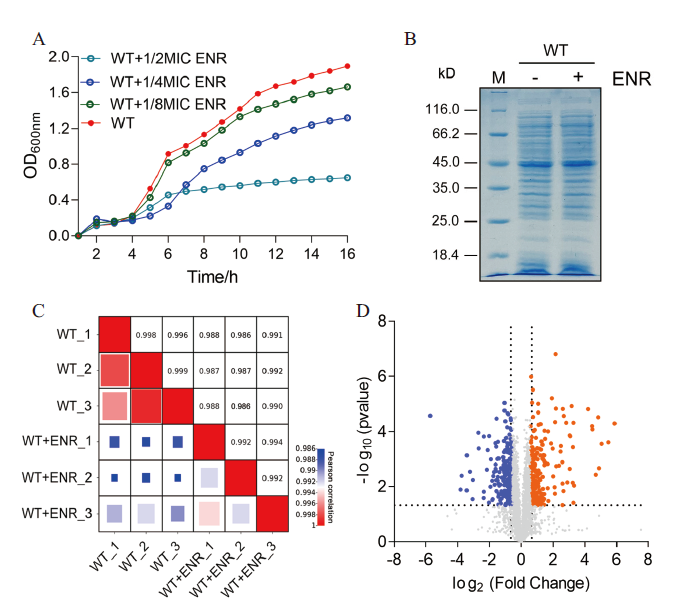
A: 不同浓度恩诺沙星胁迫下嗜水气单胞菌的生长曲线;B: 嗜水气单胞菌在有无恩诺沙星处理下的SDS-PAGE图谱;C: 通过相关系数对各组样品的3次生物学重复的蛋白表达量进行相关性分析;D: 火山图显示蛋白表达差异的丰度。其中蓝色点表示差异表达下调的蛋白、橙色点表示差异表达上调的蛋白和灰色表达非差异表达的蛋白
Fig. 1
Analysis of quantitative proteomics data
A: Growth curves of A. hydrophila under different concentrations of enrofloxacin stress. B: SDS-PAGE profiles of A. hydrophila with or without enrofloxacin treatment. C: Correlation analysis of protein expression of three biological replicates in each group was carried out by correlation coefficient. D: Volcanic plots showing the abundance of differential protein expression. The blue dots represent the differentially downregulated proteins, the orange dots represent the differentially up-regulated proteins, and the gray dots represent the non-differentially expressed proteins
2.2 嗜水气单胞菌在ENR胁迫下蛋白表达差异
将嗜水气单胞菌在有无ENR(1/4 MIC)处理下培养至OD600 nm=1.0时收集菌体,将全菌蛋白利用SDS-PAGE胶进行电泳分离。结果如图1-B所示,与对照相比,两组蛋白样品之间存在略微不同的条带。为了更好地了解嗜水气单胞菌对ENR的耐药机制,将各组中具有3个独立重复的蛋白样品用胰蛋白酶消化后进行质谱鉴定,比较有无ENR胁迫下的蛋白表达差异。通过对每个生物学重复的数据进行相关性分析发现,所有相关系数R均大于0.98,这表明蛋白质组学数据相当准确、合理,可用于进一步分析(图1-C)。同时火山图还显示(图1-D),本研究共鉴定到2 838个蛋白并且有446个差异表达蛋白,其中包括233个蛋白表达上调和213个蛋白表达下调。
2.3 GO富集分析
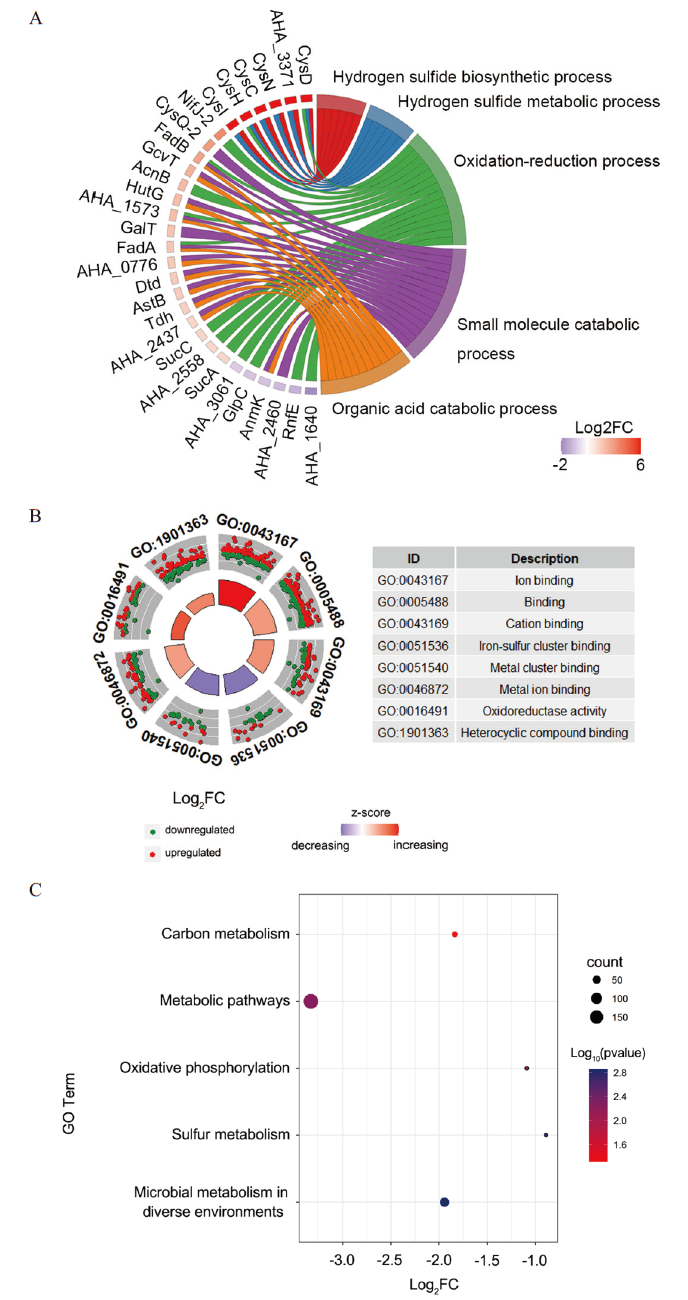
为了更深入地了解ENR对嗜水气单胞菌生物学特性的影响,本研究利用DAVID在线网站对所有差异蛋白进行富集分析,通过R语言进行可视化。结果显示在生物过程(BP)中,有5种通路受到影响,如硫化氢生物合成过程(6个,蛋白全部上调)、硫化氢代谢过程(6个,蛋白全部上调)、氧化还原过程(14个,上下调的差异蛋白比例为5∶2)、小分子分解代谢过程(14个,上下调的差异蛋白比例为6∶1)和有机酸分解代谢过程(11个,上下调的差异蛋白比例为11∶1)。其中硫化氢生物合成过程是富集程度最高的生物过程。该过程涉及到的6个差异蛋白均显著上调(图2-A)。同时,与分子功能(MF)相关的分子功能变化共有8种(图2-B),包括离子结合(111个,上下调的差异蛋白比例为65∶46)、结合(145个,上下调的差异蛋白比例为78∶67)、阳离子结合(66个,上下调的差异蛋白比例为37∶29)、铁硫簇结合(23个,上下调的差异蛋白比例为9∶14)、金属簇结合(23个,上下调的差异蛋白比例为9∶14)、金属离子结合(65个,上下调的差异蛋白比例为36∶29)、氧化还原酶活性(46个,上下调的差异蛋白比例为28∶18)和杂环化合物结合(101个,上下调的差异蛋白比例为56∶45)(图2-B)。
图2
图2
ENR胁迫下差异表达蛋白的GO和KEGG富集分析
A:使用DAVID在线网站和R语言分析差异表达蛋白的生物过程;B:分子功能;C:KEGG代谢通路富集分析
Fig. 2
GO and KEGG enrichment analysis of differentially expressed proteins under ENR stress
A: Using DAVID online website and R language to analyze the biological processes of differentially expressed proteins. B: Molecular function. C: Enrichment analysis of KEGG metabolic pathway
2.4 KEGG富集分析
我们还利用David在线网站对差异蛋白的KEGG代谢通路途径进行了富集分析,并结合R语言的GOplot包进行了可视化分析。结果发现,ENR处理后嗜水气单胞菌的5个代谢通路显著富集(图2-C),主要包括不同环境下的微生物代谢(64个,上下调的差异蛋白比例为34∶30)、硫代谢(19个,上下调的差异蛋白比例为15∶4)、氧化磷酸化(19个,上下调的差异蛋白比例为14∶5)、代谢途径(185个,上下调的差异蛋白比例为108∶77)和碳代谢(28个,上下调的差异蛋白比例为20∶8)。其中至少改变了19个参与硫代谢的差异蛋白,说明该通路在ENR胁迫过程中发挥重要的作用。
2.5 差异蛋白中的核心蛋白网络
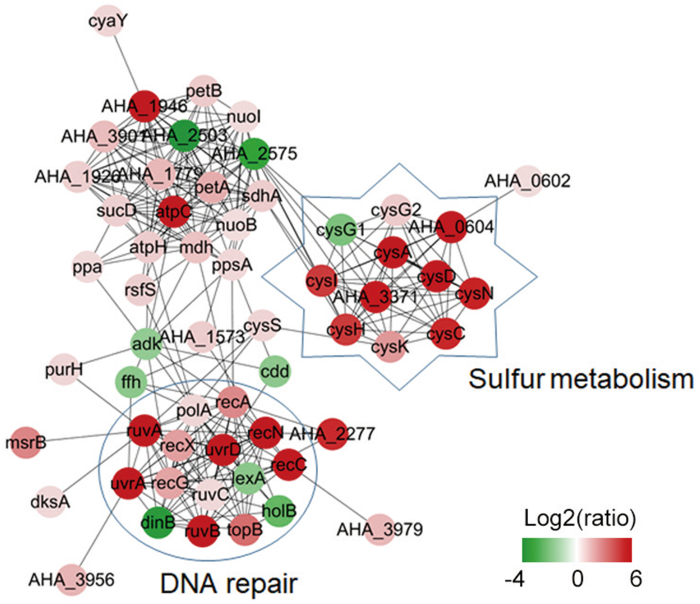
为了进一步研究蛋白质组学结果中的核心蛋白,本研究还利用Cytoscape软件预测了差异蛋白的蛋白与蛋白相互作用网络,并利用cytoHubba插件中的 MDNC方法鉴定了前10个核心蛋白及这些核心蛋白的扩展网络。结果如图3显示,这些蛋白跟DNA 修复和硫代谢等显著相关。这些结果表明,这些代谢通路可能在嗜水气单胞菌对ENR的抗性中起重要作用。
图3
图3
在ENR胁迫下差异蛋白的蛋白-蛋白相互作用预测(PPI)网络
核心蛋白扩展的网络。不同颜色表示差异蛋白的变化倍数(Log2)
Fig. 3
Protein-protein interaction prediction(PPI)net-work of differential proteins under ENR stress
The extended network of hub proteins. Different colors indicate the fold change of different proteins(Log2)
2.6 硫代谢在ENR抗生素耐药性中起着重要作用
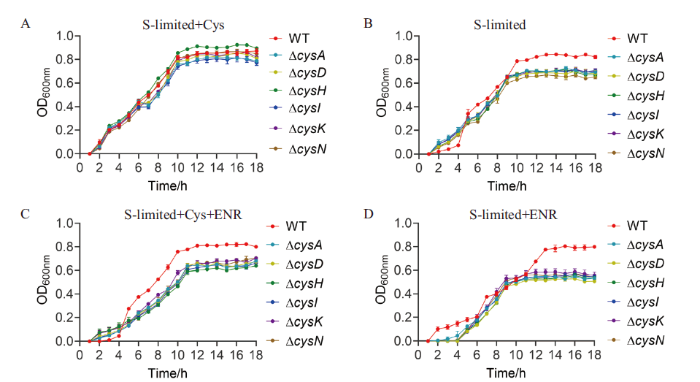
蛋白质组学数据显示,硫代谢在ENR胁迫下分子机制发生了变化。为了进一步验证硫代谢在治疗嗜水气单胞菌感染策略中的作用。本研究测定2 mmol/L半胱氨酸与0.78 µg/mL ENR处理下的敏感性。结果如图4所示,在硫限制培养基中加入2 mmol/L半胱氨酸,WT和缺失菌生长趋势大致相同;在硫限制性培养基中硫代谢相关基因的缺失相对于WT显著下降;在硫限制性培养基中加入2 mmol/L半胱氨酸和0.78 µg/mL ENR,硫代谢相关基因的缺失相对于WT显著下降;在硫限制性培养基中加入0.78 µg/mL ENR,硫代谢相关基因的缺失相对于WT更显著降低嗜水气单胞菌的存活率。这表明硫代谢参与了ENR的耐药,并且与抗生素联用可能是一个很好的治疗方向。
图4
图4
WT和硫代谢相关基因缺失菌株在硫限制培养基中分别添加2 mmol/L半胱氨酸和ENR后的生长情况
A: 硫限制培养基中加入2 mmol/L半胱氨酸;B: 硫限制培养基; C: 硫限制培养基中加入2 mmol/L半胱氨酸和0.78 µg/mL ENR;D: 硫限制培养基中加入ENR
Fig. 4
Growths of WT and sulfur metabolism-related gene deletion strains supplemented with 2 mmol/L cysteine and ENR in sulfur limiting medium, respectively
A: 2 mmol/L cysteine was added to the sulfur-limited medium. B: Sulfur-limited medium. C: Sulfur limited medium with 2 mmol/L cysteine and 0.78 µg/mL ENR. D: ENR was added to the sulfur-limited medium
2.7 组学数据验证
为了进一步验证蛋白质组学数据的准确性,我们选取了硫代谢途径中的6个基因(cysA、cysC、cysD、cysI、cysN、cysH、cysG2和cysK),采用qPCR方法比较有无ENR处理菌株的mRNA表达水平。如图5所示,与野生型相比,这些基因的转录水平均显著增加。这些结果与蛋白质组学结果一致,表明定量蛋白质组学数据具有一定的可靠性。
图5
图5
蛋白质组学数据的qPCR验证
采用qPCR方法验证嗜水气单胞菌在有无ENR胁迫下8个硫代谢相关基因在mRNA水平上的表达量
Fig. 5
qPCR validation of proteomic data
qPCR was used to verify the mRNA expression levels of 8 sulfur metabolism related genes in A. hydrophila under ENR stress or not
3 讨论
喹诺酮类抗生素是一种广谱型抗生素,对革兰氏阳性和革兰氏阴性细菌都能有效抑制[16]。然而,由于过度使用喹诺酮类抗生素,细菌已经对喹诺酮类药物产生了耐药性。现如今关于细菌对喹诺酮类抗生素的耐药性研究有很多[17]。根据以往的报道,喹诺酮类耐药菌主要采取以下3种抗生素耐药策略:一是通过破坏拓扑异构酶IV和DNA回旋酶来抑制细菌的核酸合成,并引起细菌染色体的断裂[18];二是外排泵引起的细胞内药物积累减少[19];三是喹诺酮类抗生素可诱导一些促进染色体突变的耐药基因(qnrA、qnrB、qnrS、qnrC和qnrD)表达上升,从而提高耐药水平[20]。细菌耐药性的提高使得在水产养殖等中治疗细菌引起的疾病变得越来越困难。并且,目前关于细菌对喹诺酮类抗生素的耐药分子机制还有很多未知之处,有待进一步研究。
本研究首先使用1/4 MIC浓度的ENR处理嗜水气单胞菌,并利用定量蛋白质组学的方法比较和分析,在鉴定到的2 838个蛋白中,有446个蛋白表达差异,其中233个蛋白表达上调和213个蛋白表达下调。在这些差异蛋白中,有一些蛋白参与细菌耐药的功能已有研究报道。例如Zhu等[8]研究发现,嗜水气单胞菌对恩诺沙星耐药与ABC转运体下调表达引起的胞内药物积累和拓扑异构酶IV表达增加密切相关。本研究发现在差异表达基因中,DNA修复相关基因在恩诺沙星胁迫下大部分均表达上升,这提示这些蛋白可能通过提高自身的DNA修复能力来减轻喹诺酮类抗生素对染色体的破坏。
此外,本研究还发现氧化磷酸化和碳代谢等重要代谢途径在恩诺沙星胁迫下发生了变化,提示这些代谢途径可能在细菌耐药过程中起着重要作用。研究表明,细菌耐药性的产生除了与关键耐药基因有关外,关键代谢途径的改变也会影响其耐药性。例如,Peng等[21]发现逆转三羧酸代谢途径,可以将耐药菌转化为敏感菌,提示控制耐药相关的关键代谢流,可以与抗生素协同抑制耐药细菌生长。细菌的硫代谢产生半胱氨酸,在恩诺沙星的耐药中起至关重要的作用,因此是决定细菌生存与否的重要代谢通路之一。本研究中,CysA、CysC、CysD、CysG2、CysH、CysI、CysK、CysN、AHA_0604和AHA_3371等硫代谢相关蛋白均显著上调表达,提示硫代谢可能参与细菌对ENR的耐药过程。本研究进一步通过测定cysA、cysD、cysH、cysI、cysK和cysN等缺失菌株在硫限制培养基、补充硫源和/或添加恩诺沙星下的生长情况发现,半胱氨酸与恩诺沙星联用显著影响硫代谢相关基因的生长,提示硫代谢通路在嗜水气单胞菌对喹诺酮类抗生素耐药中起重要的作用。综上所述,本研究通过对嗜水气单胞菌在恩诺沙星胁迫下的定量蛋白质组学比较,为细菌耐药机制研究以及今后新型抗生素的研发提供理论基础。
4 结论
嗜水气单胞菌ATCC7966在1/4 MIC恩诺沙星胁迫下,通过质谱鉴定共发现446个差异蛋白,其中233个蛋白表达上调,213个蛋白表达下调,这些蛋白调控多种代谢途径调节,特别是对DNA修复和硫代谢途径。本研究通过测定硫代谢相关缺失菌株的生长情况,发现半胱氨酸与恩诺沙星联用显著影响硫代谢相关基因的生长。
参考文献
Modern trends in Aeromonas hydrophila disease management with fish
[J].DOI:10.1080/10641260500320845 URL [本文引用: 1]
Fatal case of myonecrosis and septicaemia caused by Aeromonas hydrophila in Finland
[J].A 48-y-old female developed cellulitis, myonecrosis and sepsis after a prick wound in her hand while boning freshwater fish. Cultures revealed Aeromonas hydrophila, a Gram-negative bacillus. Despite prompt care the patient died 4 d after the incident. Our case shows that the occurrence of severe Aeromonas infections is not limited to tropical and subtropical areas of the world.
Antibiotic and heavy metal resistance genes in Aeromonas spp. isolated from marketed Manila Clam(Ruditapes philippinarum)in Korea
[J].
DOI:10.1111/jam.14355
PMID:31211903
[本文引用: 1]

Manila clam (Ruditapes philippinarum) is one of the most popular seafood in Korea, owing to their unique taste and nutritional value. This study aimed to disclose the antibiotic and heavy metal resistance characteristics of Aeromonas spp. isolated from marketed Manila clam in Korea.A total of 36 Aeromonas spp. strains were isolated and subjected to two tests: an antibiotic disk diffusion test to determine their resistance to antibiotics, and a broth dilution test to determine their resistance to heavy metals. PCR-based amplification was performed to detect the resistance genes. A high level of resistance to ampicillin (100%) and cephalothin (89%) was observed, while 42, 39, 36 and 36% of the isolates were resistant to oxytetracycline, imipenem, nalidixic acid and tetracycline respectively. In addition, among the tested heavy metals, cadmium (Cd) recorded the highest resistance rate (61%), followed by chromium (Cr) (50%), lead (Pb) (47%) and copper (Cu) (37%). However, mercury (Hg) resistance was not observed. PCRs revealed the occurrence of bla, bla, bla, qnrS, tetB, tetE, aac(6')-Ib, strA-strB and intI1 genes among 100, 31, 31, 78, 78, 89, 25, 50 and 72% of the isolates respectively. Moreover, heavy metal resistance genes, copA, merA and czcA were detected in 25, 47 and 61% of the isolates respectively.The results suggest the importance of multi-drug and heavy metal-resistant aeromonads in Manila clam to assess the consumer safety and public health.This study is the first to elaborate on the importance of multi-drug and heavy metal-resistant aeromonads in Manila clam. Particularly, the presence of extended-spectrum-β-lactamase genes and other antibiotic resistance genes intensifies the possible health risks and may complicate therapeutic treatments upon infection, while heavy metal resistance suggests possible heavy metal exposure.© 2019 The Society for Applied Microbiology.
Antimicrobial resistance of Aeromonas hydrophila isolated from different food sources: a mini-review
[J].DOI:10.1016/j.jiph.2015.10.006 URL [本文引用: 1]
Prevention of infection caused by enteropathogenic E. coli O157: H7 in intestinal cells using enrofloxacin entrapped in polymer based nanocarriers
[J].DOI:10.1016/j.jhazmat.2021.125454 URL [本文引用: 1]
Enrofloxacin
[J].DOI:10.1053/j.jepm.2005.11.011 URL [本文引用: 1]
Transcriptome responses in enrofloxacin-resistant strains of Aeromonas hydrophila exposed to Forsythia suspensa
[J].DOI:10.1016/j.aquaculture.2019.734809 URL [本文引用: 1]
Transcriptome differences between enrofloxacin-resistant and enrofloxacin-susceptible strains of Aeromonas hydrophila
[J].DOI:10.1371/journal.pone.0179549 URL [本文引用: 2]
In-situ removal of residual antibiotics(enrofloxacin)in recirculating aquaculture system: effect of ultraviolet photolysis plus biodegradation using immobilized microbial granules
[J].DOI:10.1016/j.jclepro.2021.130190 URL [本文引用: 1]
Comparison of detergent-based sample preparation workflows for LTQ-Orbitrap analysis of the Escherichia coli proteome
[J].DOI:10.1002/pmic.201200478 URL [本文引用: 1]
Proteomics analysis reveals the effect of Aeromonas hydrophila sirtuin CobB on biological functions
[J].DOI:10.1016/j.jprot.2020.103848 URL [本文引用: 1]
DAVID Bioinformatics Resources: expanded annotation database and novel algorithms to better extract biology from large gene lists
[J].
DOI:10.1093/nar/gkm415
PMID:17576678
[本文引用: 1]

All tools in the DAVID Bioinformatics Resources aim to provide functional interpretation of large lists of genes derived from genomic studies. The newly updated DAVID Bioinformatics Resources consists of the DAVID Knowledgebase and five integrated, web-based functional annotation tool suites: the DAVID Gene Functional Classification Tool, the DAVID Functional Annotation Tool, the DAVID Gene ID Conversion Tool, the DAVID Gene Name Viewer and the DAVID NIAID Pathogen Genome Browser. The expanded DAVID Knowledgebase now integrates almost all major and well-known public bioinformatics resources centralized by the DAVID Gene Concept, a single-linkage method to agglomerate tens of millions of diverse gene/protein identifiers and annotation terms from a variety of public bioinformatics databases. For any uploaded gene list, the DAVID Resources now provides not only the typical gene-term enrichment analysis, but also new tools and functions that allow users to condense large gene lists into gene functional groups, convert between gene/protein identifiers, visualize many-genes-to-many-terms relationships, cluster redundant and heterogeneous terms into groups, search for interesting and related genes or terms, dynamically view genes from their lists on bio-pathways and more. With DAVID (http://david.niaid.nih.gov), investigators gain more power to interpret the biological mechanisms associated with large gene lists.
Proteomics analysis reveals the importance of transcriptional regulator slyA in regulation of several physiological functions in Aeromonas hydrophila
[J].DOI:10.1016/j.jprot.2021.104275 URL [本文引用: 1]
Comparative transcriptomic analysis reveals the molecular mechanisms related to oxytetracycline- resistance in strains of Aeromonas hydrophila
[J].
SWATH based quantitative proteomics analysis reveals Hfq2 play an important role on pleiotropic physiological functions in Aeromonas hydrophila
[J].DOI:10.1016/j.jprot.2018.12.030 URL [本文引用: 1]
The effect of enrofloxacin on enteric Escherichia coli: fitting a mathematical model to in vivo data
[J].DOI:10.1371/journal.pone.0228138 URL [本文引用: 1]
Quinolones: mechanism, lethality and their contributions to antibiotic resistance
[J].
DOI:10.3390/molecules25235662
URL
[本文引用: 1]

Fluoroquinolones (FQs) are arguably among the most successful antibiotics of recent times. They have enjoyed over 30 years of clinical usage and become essential tools in the armoury of clinical treatments. FQs target the bacterial enzymes DNA gyrase and DNA topoisomerase IV, where they stabilise a covalent enzyme-DNA complex in which the DNA is cleaved in both strands. This leads to cell death and turns out to be a very effective way of killing bacteria. However, resistance to FQs is increasingly problematic, and alternative compounds are urgently needed. Here, we review the mechanisms of action of FQs and discuss the potential pathways leading to cell death. We also discuss quinolone resistance and how quinolone treatment can lead to resistance to non-quinolone antibiotics.
Quinolone antibiotics
[J].
DOI:10.1039/c9md00120d
PMID:31803393
[本文引用: 1]

The quinolone antibiotics arose in the early 1960s, with the first examples possessing a narrow-spectrum of activity with unfavorable pharmacokinetic properties. Over time, the development of new quinolone antibiotics has led to improved analogues with an expanded spectrum and high efficacy. Nowadays, quinolones are widely used for treating a variety of infections. Quinolones are broad-spectrum antibiotics that are active against both Gram-positive and Gram-negative bacteria, including mycobacteria, and anaerobes. They exert their actions by inhibiting bacterial nucleic acid synthesis through disrupting the enzymes topoisomerase IV and DNA gyrase, and by causing breakage of bacterial chromosomes. However, bacteria have acquired resistance to quinolones, similar to other antibacterial agents, due to the overuse of these drugs. Mechanisms contributing to quinolone resistance are mediated by chromosomal mutations and/or plasmid gene uptake that alter the topoisomerase targets, modify the quinolone, and/or reduce drug accumulation by either decreased uptake or increased efflux. This review discusses the development of this class of antibiotics in terms of potency, pharmacokinetics and toxicity, along with the resistance mechanisms which reduce the quinolones' activity against pathogens. Potential strategies for future generations of quinolone antibiotics with enhanced activity against resistant strains are suggested.This journal is © The Royal Society of Chemistry 2019.
Plasmid-mediated quinolone resistance genes and antibiotic residues in wastewater and soil adjacent to swine feedlots: potential transfer to agricultural lands
[J].DOI:10.1289/ehp.1104776 URL [本文引用: 1]
Investigating the impact of hospital antibiotic usage on aquatic environment and aquaculture systems: a molecular study of quinolone resistance in Escherichia coli
[J].DOI:10.1016/j.scitotenv.2020.141538 URL [本文引用: 1]
Exogenous alanine and/or glucose plus kanamycin kills antibiotic-resistant bacteria
[J].
DOI:S1550-4131(15)00009-1
PMID:25651179
[本文引用: 1]

Multidrug-resistant bacteria are an increasingly serious threat to human and animal health. However, novel drugs that can manage infections by multidrug-resistant bacteria have proved elusive. Here we show that glucose and alanine abundances are greatly suppressed in kanamycin-resistant Edwardsiella tarda by GC-MS-based metabolomics. Exogenous alanine or glucose restores susceptibility of multidrug-resistant E. tarda to killing by kanamycin, demonstrating an approach to killing multidrug-resistant bacteria. The mechanism underlying this approach is that exogenous glucose or alanine promotes the TCA cycle by substrate activation, which in turn increases production of NADH and proton motive force and stimulates uptake of antibiotic. Similar results are obtained with other Gram-negative bacteria (Vibrio parahaemolyticus, Klebsiella pneumoniae, Pseudomonas aeruginosa) and Gram-positive bacterium (Staphylococcus aureus), and the results are also reproduced in a mouse model for urinary tract infection. This study establishes a functional metabolomics-based strategy to manage infection by antibiotic-resistant bacteria.Copyright © 2015 Elsevier Inc. All rights reserved.