








生物技术通报 ›› 2022, Vol. 38 ›› Issue (4): 184-192.doi: 10.13560/j.cnki.biotech.bull.1985.2021-1198
肖艳华1,2( ), 邹智2(
), 邹智2( ), 赵永国3, 郭安平2, 张丽1(
), 赵永国3, 郭安平2, 张丽1( )
)
收稿日期:2021-09-16
出版日期:2022-04-26
发布日期:2022-05-06
通讯作者:
邹智,男,研究员,研究方向:基因组学与分子生物学;E-mail: zouzhi2008@126.com;作者简介:肖艳华,女,硕士研究生,研究方向:油莎豆分子育种;E-mail: 1740329672@qq.com
基金资助:
XIAO Yan-hua1,2( ), ZOU Zhi2(
), ZOU Zhi2( ), ZHAO Yong-guo3, GUO An-ping2, ZHANG Li1(
), ZHAO Yong-guo3, GUO An-ping2, ZHANG Li1( )
)
Received:2021-09-16
Published:2022-04-26
Online:2022-05-06
摘要:
乙酰乳酸合酶(ALS)在植物的除草剂抗性中发挥重要作用。为揭示油莎豆ALS编码基因的序列特征、进化关系及表达特性,研究基于转录组数据,应用RT-PCR技术对其进行了克隆。序列分析显示,CeALS预测编码645个氨基酸,理论分子量为69.94 kD、等电点为6.10,属于叶绿体定位的亲水性蛋白。保守结构域和同源分析显示蛋白含有TPP_enzyme_N、TPP_enzyme_M和TPP_enzyme_C三个完整的乙酰乳酸合酶功能域,与其他莎草科植物中同源蛋白的一致性都在90%以上。进化分析显示,莎草科与禾本科属禾本目的姊妹科。qRT-PCR分析显示,CeALS主要在叶片和块茎中表达;在叶片发育过程中呈现逐步递增的趋势,其在成熟和衰老叶片中的表达水平显著高于幼嫩叶片。序列比对和SNP分析显示,团队收集的56份油莎豆种质均不存在抗药性变异。这些结果为除草剂抗性分子育种及油莎豆的开发与利用奠定了坚实的基础。
肖艳华, 邹智, 赵永国, 郭安平, 张丽. 油莎豆乙酰乳酸合酶基因CeALS的克隆与分析[J]. 生物技术通报, 2022, 38(4): 184-192.
XIAO Yan-hua, ZOU Zhi, ZHAO Yong-guo, GUO An-ping, ZHANG Li. Molecular Cloning and Characterization of an Acetolactate Synthase Gene(CeALS)from Cyperus esculentus L.[J]. Biotechnology Bulletin, 2022, 38(4): 184-192.

图1 CeALS的全长cDNA及其推测编码的氨基酸 基因编码区用大写的碱基字母标识,5' UTR和3' UTR用小写的碱基字母标识,基因克隆引物CeALSF/R用单下划线标出,同源重组引物CeALSHF/R的基因序列部分用双下划线标出
Fig.1 Full-length cDNA sequence of CeALS and its dedu-ced coding amino acids The gene coding region is identified by uppercase base letters,the 5' UTR and 3' UTR are identified by lowercase bases,the cloning primer pair CeALSF/R are marked with single underlines,and the primer pair CeALSHF/R for homologous recombinant was marked by double underlines
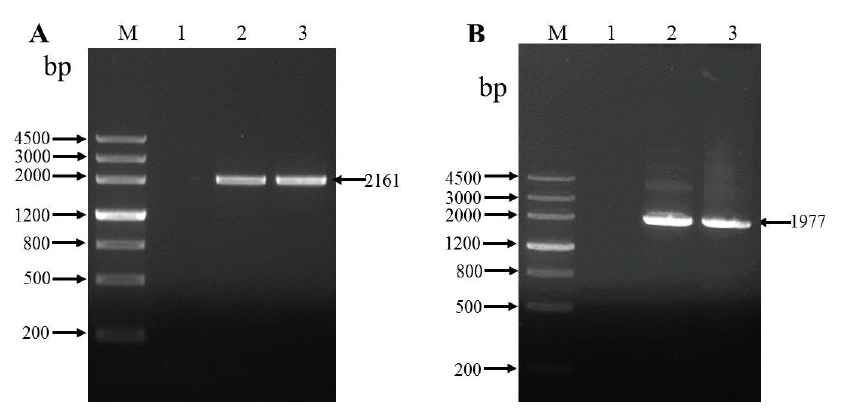
图2 油莎豆CeALS基因的PCR扩增 A:第一轮PCR扩增结果;B:第二轮PCR扩增结果;M:DNA marker III;1:空白对照;2和3:CeALS
Fig. 2 PCR amplification of CeALS in C. esculentus A:Results of the first round of PCR amplification. B:Results of the second round of PCR amplification. M:DNA marker III. 1:Blank control. 2 and 3:CeALS
| 基因 Gene | 内含子数 Number of intron | 蛋白 Protein/AA | 信号肽 Transit peptide/AA | 分子量 MW/kD | 等电点 pI | 总平均疏水指数 GRAVY | 脂肪族指数 AI | 不稳定系数 II |
|---|---|---|---|---|---|---|---|---|
| AtALS | 0 | 670 | 55 | 72.59 | 6.20 | -0.130 | 87.78 | 42.93 |
| PeALS | 0 | 657 | 58 | 71.51 | 6.78 | -0.126 | 95.16 | 40.20 |
| PdALS | 0 | 658 | 68 | 70.91 | 6.17 | -0.054 | 95.06 | 43.27 |
| EgALS | 0 | 658 | 68 | 70.89 | 6.27 | -0.063 | 94.92 | 41.18 |
| SpALS | 0 | 659 | 52 | 71.81 | 5.97 | -0.138 | 92.67 | 43.44 |
| AcALS | 0 | 655 | 59 | 70.44 | 6.37 | -0.061 | 93.89 | 45.85 |
| MaALS1 | 0 | 655 | 58 | 71.12 | 6.64 | -0.091 | 93.11 | 38.24 |
| MaALS2 | 0 | 660 | 54 | 71.84 | 6.43 | -0.103 | 93.03 | 37.67 |
| OsALS1 | 0 | 644 | 43 | 69.39 | 6.48 | -0.027 | 92.13 | 41.61 |
| OsALS2 | 0 | 663 | 79 | 71.18 | 6.30 | -0.111 | 87.59 | 36.30 |
| HvALS | 0 | 646 | 46 | 69.35 | 6.58 | -0.004 | 92.32 | 40.49 |
| BdALS1 | 0 | 646 | 46 | 69.45 | 6.55 | -0.015 | 92.46 | 43.17 |
| BdALS2 | 0 | 646 | 46 | 70.12 | 6.41 | -0.066 | 92.15 | 37.31 |
| SvALS | 0 | 643 | 41 | 69.39 | 6.45 | -0.053 | 88.96 | 42.81 |
| SiALS | 0 | 643 | 41 | 69.38 | 6.45 | -0.054 | 88.96 | 42.84 |
| SbALS | 0 | 641 | 40 | 69.05 | 6.42 | 0.000 | 92.14 | 40.01 |
| ZmALS1 | 0 | 638 | 39 | 68.93 | 6.69 | -0.060 | 91.16 | 40.26 |
| ZmALS2 | 0 | 638 | 39 | 68.99 | 6.77 | -0.036 | 91.33 | 40.15 |
| ClALS1 | 1 | 644 | 79 | 70.11 | 6.45 | -0.145 | 89.35 | 38.09 |
| ClALS2 | 1 | 644 | 79 | 70.34 | 6.80 | -0.163 | 88.59 | 37.94 |
| CbALS | 1 | 647 | 55 | 70.46 | 5.85 | -0.166 | 88.92 | 46.54 |
| CeALS | 1 | 645 | 54 | 69.94 | 6.10 | -0.156 | 89.36 | 44.93 |
| SwALS1 | 1 | 645 | 80 | 70.24 | 6.24 | -0.144 | 90.59 | 41.67 |
| SwALS2 | 1 | 645 | 77 | 70.26 | 6.45 | -0.163 | 89.98 | 41.97 |
| SjALS1 | 1 | 645 | 80 | 70.22 | 6.24 | -0.140 | 90.74 | 41.67 |
| SjALS2 | 1 | 645 | 80 | 70.26 | 6.45 | -0.163 | 89.98 | 41.97 |
| SmALS1 | 1 | 648 | 80 | 70.58 | 6.22 | -0.143 | 90.62 | 41.68 |
| SmALS2 | 1 | 645 | 77 | 70.21 | 6.45 | -0.157 | 90.28 | 41.37 |
表1 拟南芥和20种单子叶植物中ALS基因的结构及其编码蛋白的理化特性
Table 1 Gene structure and physicochemical properties of ALS genes in Arabidopsis and 20 monocots
| 基因 Gene | 内含子数 Number of intron | 蛋白 Protein/AA | 信号肽 Transit peptide/AA | 分子量 MW/kD | 等电点 pI | 总平均疏水指数 GRAVY | 脂肪族指数 AI | 不稳定系数 II |
|---|---|---|---|---|---|---|---|---|
| AtALS | 0 | 670 | 55 | 72.59 | 6.20 | -0.130 | 87.78 | 42.93 |
| PeALS | 0 | 657 | 58 | 71.51 | 6.78 | -0.126 | 95.16 | 40.20 |
| PdALS | 0 | 658 | 68 | 70.91 | 6.17 | -0.054 | 95.06 | 43.27 |
| EgALS | 0 | 658 | 68 | 70.89 | 6.27 | -0.063 | 94.92 | 41.18 |
| SpALS | 0 | 659 | 52 | 71.81 | 5.97 | -0.138 | 92.67 | 43.44 |
| AcALS | 0 | 655 | 59 | 70.44 | 6.37 | -0.061 | 93.89 | 45.85 |
| MaALS1 | 0 | 655 | 58 | 71.12 | 6.64 | -0.091 | 93.11 | 38.24 |
| MaALS2 | 0 | 660 | 54 | 71.84 | 6.43 | -0.103 | 93.03 | 37.67 |
| OsALS1 | 0 | 644 | 43 | 69.39 | 6.48 | -0.027 | 92.13 | 41.61 |
| OsALS2 | 0 | 663 | 79 | 71.18 | 6.30 | -0.111 | 87.59 | 36.30 |
| HvALS | 0 | 646 | 46 | 69.35 | 6.58 | -0.004 | 92.32 | 40.49 |
| BdALS1 | 0 | 646 | 46 | 69.45 | 6.55 | -0.015 | 92.46 | 43.17 |
| BdALS2 | 0 | 646 | 46 | 70.12 | 6.41 | -0.066 | 92.15 | 37.31 |
| SvALS | 0 | 643 | 41 | 69.39 | 6.45 | -0.053 | 88.96 | 42.81 |
| SiALS | 0 | 643 | 41 | 69.38 | 6.45 | -0.054 | 88.96 | 42.84 |
| SbALS | 0 | 641 | 40 | 69.05 | 6.42 | 0.000 | 92.14 | 40.01 |
| ZmALS1 | 0 | 638 | 39 | 68.93 | 6.69 | -0.060 | 91.16 | 40.26 |
| ZmALS2 | 0 | 638 | 39 | 68.99 | 6.77 | -0.036 | 91.33 | 40.15 |
| ClALS1 | 1 | 644 | 79 | 70.11 | 6.45 | -0.145 | 89.35 | 38.09 |
| ClALS2 | 1 | 644 | 79 | 70.34 | 6.80 | -0.163 | 88.59 | 37.94 |
| CbALS | 1 | 647 | 55 | 70.46 | 5.85 | -0.166 | 88.92 | 46.54 |
| CeALS | 1 | 645 | 54 | 69.94 | 6.10 | -0.156 | 89.36 | 44.93 |
| SwALS1 | 1 | 645 | 80 | 70.24 | 6.24 | -0.144 | 90.59 | 41.67 |
| SwALS2 | 1 | 645 | 77 | 70.26 | 6.45 | -0.163 | 89.98 | 41.97 |
| SjALS1 | 1 | 645 | 80 | 70.22 | 6.24 | -0.140 | 90.74 | 41.67 |
| SjALS2 | 1 | 645 | 80 | 70.26 | 6.45 | -0.163 | 89.98 | 41.97 |
| SmALS1 | 1 | 648 | 80 | 70.58 | 6.22 | -0.143 | 90.62 | 41.68 |
| SmALS2 | 1 | 645 | 77 | 70.21 | 6.45 | -0.157 | 90.28 | 41.37 |

图3 CeALS蛋白的多序列比对和序列特征 乙酰乳酸合酶结构域在其两端用实心箭头标识;叶绿体信号肽、TPP_enzyme_N、TPP_enzyme_M和TPP_enzyme_C用线框标识;可引起抗性改变的保守残基用空心箭头标识
Fig. 3 Multiple sequence alignment and sequence features of CeALS protein The acetolactate synthase domain is marked by solid arrows at both ends. Chloroplast transit peptide and domains of TPP_enzyme_N,TPP_enzyme_M,and TPP_enzyme_C are identified by wire frames,whereas conserved residues that can cause resistance changes are identified by hollow arrows
| 位点Site | CeALS | SNP | 位点Site | CeALS | SNP | 位点Site | CeALS | SNP | |||
|---|---|---|---|---|---|---|---|---|---|---|---|
| 120 | A | C | 435 | C | G | 1431 | T | G | |||
| 147 | C | T | 519 | C | T | 1512 | T | A | |||
| 237 | G | A | 631 | A | C | 1596 | T | C | |||
| 255 | G | A | 681 | T | C | 1612 | T | G | |||
| 405 | A | G | 685 | C | A | 1702 | A | G | |||
| 408 | T | C | 840 | C | T | 1749 | T | C | |||
| 411 | C | G | 945 | C | T | 1752 | T | C | |||
| 414 | C | T | 1128 | C | T | 1767 | T | C | |||
| 417 | C | T | 1146 | G | A | 1771 | G | T | |||
| 420 | C | A | 1404 | G | A | 1851 | G | T |
表2 SNP分布情况
Table 2 Distribution of SNPs
| 位点Site | CeALS | SNP | 位点Site | CeALS | SNP | 位点Site | CeALS | SNP | |||
|---|---|---|---|---|---|---|---|---|---|---|---|
| 120 | A | C | 435 | C | G | 1431 | T | G | |||
| 147 | C | T | 519 | C | T | 1512 | T | A | |||
| 237 | G | A | 631 | A | C | 1596 | T | C | |||
| 255 | G | A | 681 | T | C | 1612 | T | G | |||
| 405 | A | G | 685 | C | A | 1702 | A | G | |||
| 408 | T | C | 840 | C | T | 1749 | T | C | |||
| 411 | C | G | 945 | C | T | 1752 | T | C | |||
| 414 | C | T | 1128 | C | T | 1767 | T | C | |||
| 417 | C | T | 1146 | G | A | 1771 | G | T | |||
| 420 | C | A | 1404 | G | A | 1851 | G | T |
| [1] | 李杨, 马智宏, 李冰茹, 等. 我国主要作物中除草剂登记情况及存在问题[J]. 食品安全质量检测学报, 2018, 9(17):4483-4488. |
| Li Y, Ma ZH, Li BR, et al. Registration status and existing problems of herbicides in the major crops in China[J]. J Food Saf Qual, 2018, 9(17):4483-4488. | |
| [2] | 张玉池, 王晓蕾, 徐文蓉, 等. 国内外抗除草剂基因专利的分析[J]. 杂草学报, 2017, 35(2):1-22. |
| Zhang YC, Wang XL, Xu WR, et al. Analysis on the patents of herbicide resistance gene at home and abroad[J]. J Weed Sci, 2017, 35(2):1-22. | |
| [3] |
Liu YD, Li YY, Wang XY. Acetohydroxyacid synthases:evolution, structure, and function[J]. Appl Microbiol Biotechnol, 2016, 100(20):8633-8649.
doi: 10.1007/s00253-016-7809-9 URL |
| [4] |
Yu Q, Powles SB. Resistance to AHAS inhibitor herbicides:current understanding[J]. Pest Manag Sci, 2014, 70(9):1340-1350.
doi: 10.1002/ps.3710 URL |
| [5] | Heap I. The International Herbicide-Resistant Weed Database[EB/OL]. 2021 www.weedscience.org. |
| [6] | 张学昆. 我国油莎豆产业研发进展报告[J]. 中国农村科技, 2019(4):67-69. |
| Zhang XK. Research progress of the tigernut industry in China[J]. China Rural Sci Technol, 2019(4):67-69. | |
| [7] |
Turesson H, Marttila S, Gustavsson KE, et al. Characterization of oil and starch accumulation in tubers of Cyperus esculentus var. sativus(Cyperaceae):a novel model system to study oil reserves in nonseed tissues[J]. Am J Bot, 2010, 97(11):1884-1893.
doi: 10.3732/ajb.1000200 pmid: 21616827 |
| [8] | 邹智, 赵永国, 张丽, 等. 基于单分子实时测序的油莎豆全长转录组分析[J]. 中国油料作物学报, 2021, 43(2):229-235. |
| Zou Z, Zhao YG, Zhang L, et al. Single-molecule real-time(SMRT)-based full-length transcriptome analysis of tigernut(Cyperus esculentus L.)[J]. Chin J Oil Crop Sci, 2021, 43(2):229-235. | |
| [9] |
De Castro O, Gargiulo R, Del Guacchio E, et al. A molecular survey concerning the origin of Cyperus esculentus(Cyperaceae, Poales):two sides of the same coin(weed vs. crop)[J]. Ann Bot, 2015, 115(5):733-745.
doi: 10.1093/aob/mcv001 URL |
| [10] |
Tehranchian P, Norsworthy JK, Nandula V, et al. First report of resistance to acetolactate-synthase-inhibiting herbicides in yellow nutsedge(Cyperus esculentus):confirmation and characterization[J]. Pest Manag Sci, 2015, 71(9):1274-1280.
doi: 10.1002/ps.3922 pmid: 25307777 |
| [11] | Okuno J, Iwakami S, Uchino A, et al. Responses to halosulfuron-methyl and flazasulfuron and mutation of acetolactate synthase gene of Cyperus brebifolius survived in turf grass on golf course[J]. J Jpn Soc Turfgrass Sci, 2015, 43:159-162. |
| [12] |
Ntoanidou S, Kaloumenos N, Diamantidis G, et al. Molecular basis of Cyperus difformis cross-resistance to ALS-inhibiting herbicides[J]. Pestic Biochem Physiol, 2016, 127:38-45.
doi: 10.1016/j.pestbp.2015.09.004 pmid: 26821656 |
| [13] |
Goodstein DM, Shu S, Howson R, et al. Phytozome:a comparative platform for green plant genomics[J]. Nucleic Acids Res, 2012, 40(database issue):D1178-D1186.
doi: 10.1093/nar/gkr944 URL |
| [14] |
Benson DA, Karsch-Mizrachi I, Lipman DJ, et al. GenBank[J]. Nucleic Acids Res, 2010, 38(database issue):D46-D51.
doi: 10.1093/nar/gkp1024 URL |
| [15] | 邹智, 杨礼富. 巴西橡胶树铁硫簇支架蛋白cDNA的克隆与分析[J]. 热带作物学报, 2010, 31(10):1752-1756. |
| Zou Z, Yang LF. Cloning and analysis of a cDNA encoding a iron-sulfur cluster scaffold protein ISU1 from Hevea brasiliensis Müll. Arg[J]. Chin J Trop Crops, 2010, 31(10):1752-1756. | |
| [16] | 赵永国, 危文亮. 利用SRAP标记分析油莎豆遗传多样性[J]. 中国油料作物学报, 2011, 33(4):351-355. |
| Zhao YG, Wei WL. Genetic diversity analysis of tigernut(Cyperus esculentus)using SRAP markers[J]. Chin J Oil Crop Sci, 2011, 33(4):351-355. | |
| [17] |
Li H, Durbin R. Fast and accurate long-read alignment with Burrows-Wheeler transform[J]. Bioinformatics, 2010, 26(5):589-595.
doi: 10.1093/bioinformatics/btp698 URL |
| [18] |
McKenna A, Hanna M, Banks E, et al. The Genome Analysis Toolkit:a MapReduce framework for analyzing next-generation DNA sequencing data[J]. Genome Res, 2010, 20(9):1297-1303.
doi: 10.1101/gr.107524.110 pmid: 20644199 |
| [19] |
Zou Z, Xie GS, Yang LF. Papain-like cysteine protease encoding genes in rubber(Hevea brasiliensis):comparative genomics, phylogenetic, and transcriptional profiling analysis[J]. Planta, 2017, 246(5):999-1018.
doi: 10.1007/s00425-017-2739-z URL |
| [20] |
Zou Z, Gong J, An F, et al. Genome-wide identification of rubber tree(Hevea brasiliensis Muell. Arg. )aquaporin genes and their response to ethephon stimulation in the laticifer, a rubber-producing tissue[J]. BMC Genom, 2015, 16(1):1-18.
doi: 10.1186/1471-2164-16-1 URL |
| [21] |
Uchino A, Ogata S, Kohara H, et al. Molecular basis of diverse responses to acetolactate synthase-inhibiting herbicides in sulfonylurea-resistant biotypes of Schoenoplectus juncoides[J]. Weed Biol Manage, 2007, 7(2):89-96.
doi: 10.1111/j.1445-6664.2007.00240.x URL |
| [22] |
Scarabel L, Locascio A, Furini A, et al. Characterisation of ALS genes in the polyploid species Schoenoplectus mucronatus and implications for resistance management[J]. Pest Manag Sci, 2010, 66(3):337-344.
doi: 10.1002/ps.1883 pmid: 19921713 |
| [23] |
Sada Y, Ikeda H, Yamato S, et al. Characterization of sulfonylurea-resistant Schoenoplectus juncoides having a target-site Asp(376)Glu mutation in the acetolactate synthase[J]. Pestic Biochem Physiol, 2013, 107(1):106-111.
doi: 10.1016/j.pestbp.2013.05.013 URL |
| [24] |
Riar DS, Tehranchian P, Norsworthy JK, et al. Acetolactate synthase-inhibiting, herbicide-resistant rice flatsedge(Cyperus iria):cross-resistance and molecular mechanism of resistance[J]. Weed Sci, 2015, 63(4):748-757.
doi: 10.1614/WS-D-15-00014.1 URL |
| [25] |
McCullough PE, Yu JL, McElroy JS, et al. ALS-resistant annual sedge(Cyperus compressus)confirmed in turf grass[J]. Weed Sci, 2016, 64(1):33-41.
doi: 10.1614/WS-D-15-00094.1 URL |
| [26] |
Larridon I, Reynders M, Huygh W, et al. Affinities in C3 Cyperus li-neages(Cyperaceae)revealed using molecular phylogenetic data and carbon isotope analysis[J]. Bot J Linn Soc, 2011, 167(1):19-46.
doi: 10.1111/j.1095-8339.2011.01160.x URL |
| [1] | 陈海伟;张鲁华;陈德富;陈喜文;. 除草剂及抗除草剂作物的应用现状与展望[J]. , 2012, 0(10): 35-40. |
| [2] | 李思经;. 美国杜邦公司与先锋公司组建合资企业[J]. , 1997, 0(06): 47-48. |
| [3] | 吕蓓. 法院裁决Mycogen公司拥有Roundup技术[J]. , 1997, 0(02): 26-27. |
| [4] | 孙雷心. 加拿大转基因作物的田间试验[J]. , 1996, 0(03): 17-17. |
| [5] | 李思经. Calgene请求英国批准销售FLAVR SAVR[J]. , 1996, 0(02): 10-10. |
| [6] | 王颖. 英国环境释放政府顾问委员会批准除草剂抗性油菜[J]. , 1995, 0(03): 26-26. |
| [7] | 孙雷心. 美国批准首例rDNA除草剂抗性植物[J]. , 1994, 0(05): 18-19. |
| [8] | 陶冶;. Zeneca打算经销生物技术种子[J]. , 1993, 0(10): 13-14. |
| [9] | Jan Leemans;李思经;. 植物生物技术的十年[J]. , 1993, 0(09): 5-8. |
| [10] | . 植物遗传工程[J]. , 1993, 0(04): 68-78. |
| [11] | 李思经;. 美国花生品种基因转移成功[J]. , 1993, 0(01): 16-16. |
| [12] | . 植物遗传工程[J]. , 1992, 0(12): 67-76. |
| [13] | . 植物遗传工程[J]. , 1992, 0(11): 70-80. |
| [14] | 孙雷心;. 工程植物摄取除草剂并以之为养料[J]. , 1992, 0(10): 8-9. |
| [15] | 陶冶. USDA为田间试验遗传工程植物发放了8个许可证[J]. , 1992, 0(08): 8-8. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||