








生物技术通报 ›› 2025, Vol. 41 ›› Issue (1): 157-172.doi: 10.13560/j.cnki.biotech.bull.1985.2024-0656
何财林1,2( ), 卢晶1,2, 郭会会1,2, 李小安1,2, 吴琪1,2(
), 卢晶1,2, 郭会会1,2, 李小安1,2, 吴琪1,2( )
)
收稿日期:2024-07-10
出版日期:2025-01-26
发布日期:2025-01-22
通讯作者:
吴琪,男,博士,副研究员,研究方向 :藜麦分子遗传育种;E-mail: jerviswuqi@126.com作者简介:何财林,男,硕士研究生,研究方向:藜麦分子遗传育种;E-mail: 2737141093@qq.com
基金资助:
HE Cai-lin1,2( ), LU Jing1,2, GUO Hui-hui1,2, LI Xiao-an1,2, WU Qi1,2(
), LU Jing1,2, GUO Hui-hui1,2, LI Xiao-an1,2, WU Qi1,2( )
)
Received:2024-07-10
Published:2025-01-26
Online:2025-01-22
摘要:
【目的】开展藜麦CqMADS-box基因家族鉴定和表达分析,为藜麦产量品质的提高提供优异基因资源。【方法】基于拟南芥MADS-box蛋白序列,利用BLASTP和隐马尔可夫模型文件(Hidden Markov Model, HMM)鉴定CqMADS-box基因家族成员,采用最大似然法(maximum likelihood, ML)构建系统发育树,分析MADS-box基因在染色体的位置,利用MXSCan分析CqMADS-box基因家族成员在藜麦亚基因组A/B间,及其与甜菜二倍体近缘种C. pallidicaule(A)、C. suecicum(B),六倍体近缘种中国台湾红藜(Chenopodium formosanum)中MADS-box成员的共线性,利用WoLF PSORT、PlantCARE等在线网站分析CqMADS-box转录因子的理化性质、基因结构、启动子区顺式作用元件。基于前期发表的转录组数据,分析CqMADS-box在不同组织、6个花序发育时期、种子萌发的表达模式;利用转录组数据分析在长日照和短日照下,花期和花器官相关CqMADS-box的表达情况。【结果】鉴定得到117个藜麦MADS-box基因家族成员,分布在17条染色体上。I型CqMADS-box包含Mα、Mγ两个进化分支,II型包含CqPI(CqGLO)、CqAGL17等14个进化分支。不同分支CqMADS-box基因结构、蛋白Motif组成、转录因子结合位点类型和数量存在不同程度的差异。进化分析表明,串联复制是藜麦MADS-box家族扩张的主要途径,大部分复制的时间在四倍体化之前。表达模式结果表明,CqMADS-box基因在花序中表达量相对较高,而在花序发育前期(YP1或YP2),CqSEP3、CqAP1、CqGGM13类基因呈相对高表达;种子经过ABA处理后,CqSEP3类基因表达量显著上升;不同光周期下的表达模式表明,CqAP1、CqSOC1类基因在长日照和短日照下表达量均较高。【结论】CqMADS-box基因家族成员具有组织特异性表达模式,且CqSEP3类基因在藜麦花序发育和响应ABA激素处理的过程中发挥重要作用。
何财林, 卢晶, 郭会会, 李小安, 吴琪. 藜麦MADS-box基因家族的全基因组鉴定和表达分析[J]. 生物技术通报, 2025, 41(1): 157-172.
HE Cai-lin, LU Jing, GUO Hui-hui, LI Xiao-an, WU Qi. Genome-wide Identification and Expression Analysis of the MADS-box Gene Family in Quinoa[J]. Biotechnology Bulletin, 2025, 41(1): 157-172.
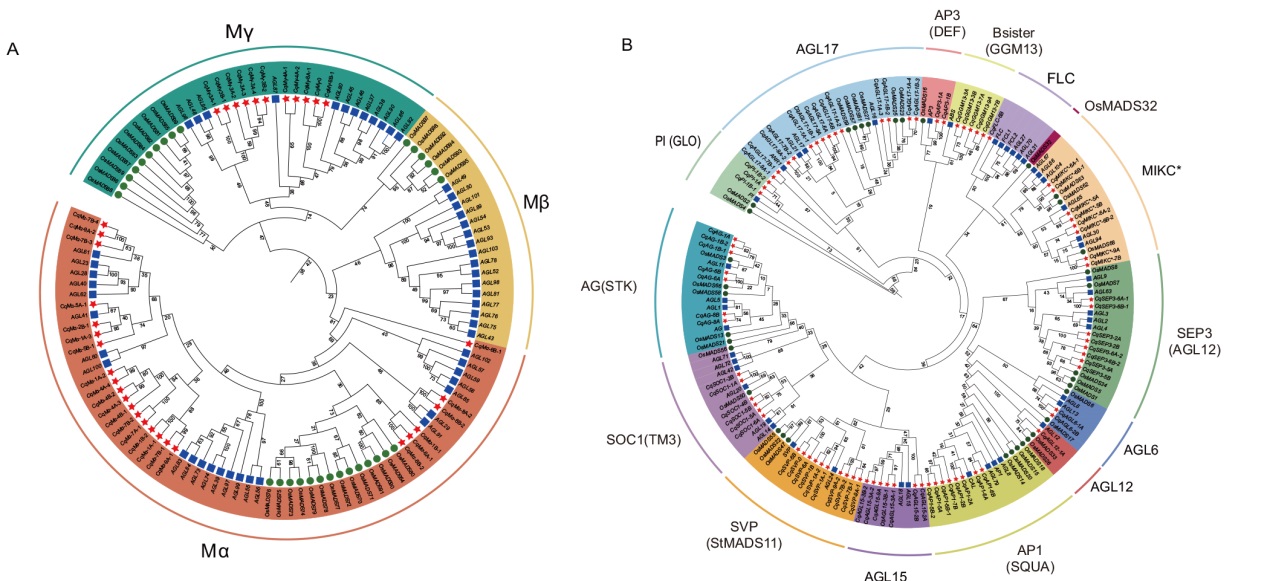
图1 藜麦、水稻和拟南芥MADS-box蛋白系统发育树 A:藜麦、拟南芥和水稻的I型MADS-box蛋白系统发育树;B:藜麦、拟南芥和水稻的II型MADS-box蛋白系统发育树;红色五角星代表藜麦蛋白,紫色方框代表拟南芥蛋白,绿色圆代表水稻蛋白
Fig. 1 MADS-box protein phylogenetic tree of quinoa, rice and Arabidopsis A: Phylogenetic tree of type I MADS-box proteins in quinoa(Chenopodium quinoa), Arabidopsis and rice(Oryza sativa). B: Phylogenetic tree of type II MADS-box proteins in quinoa, Arabidopsis and rice. The red five-pointed star indicates quinoa protein, the purple box indicates Arabidopsis protein, and the green circle indicates rice protein
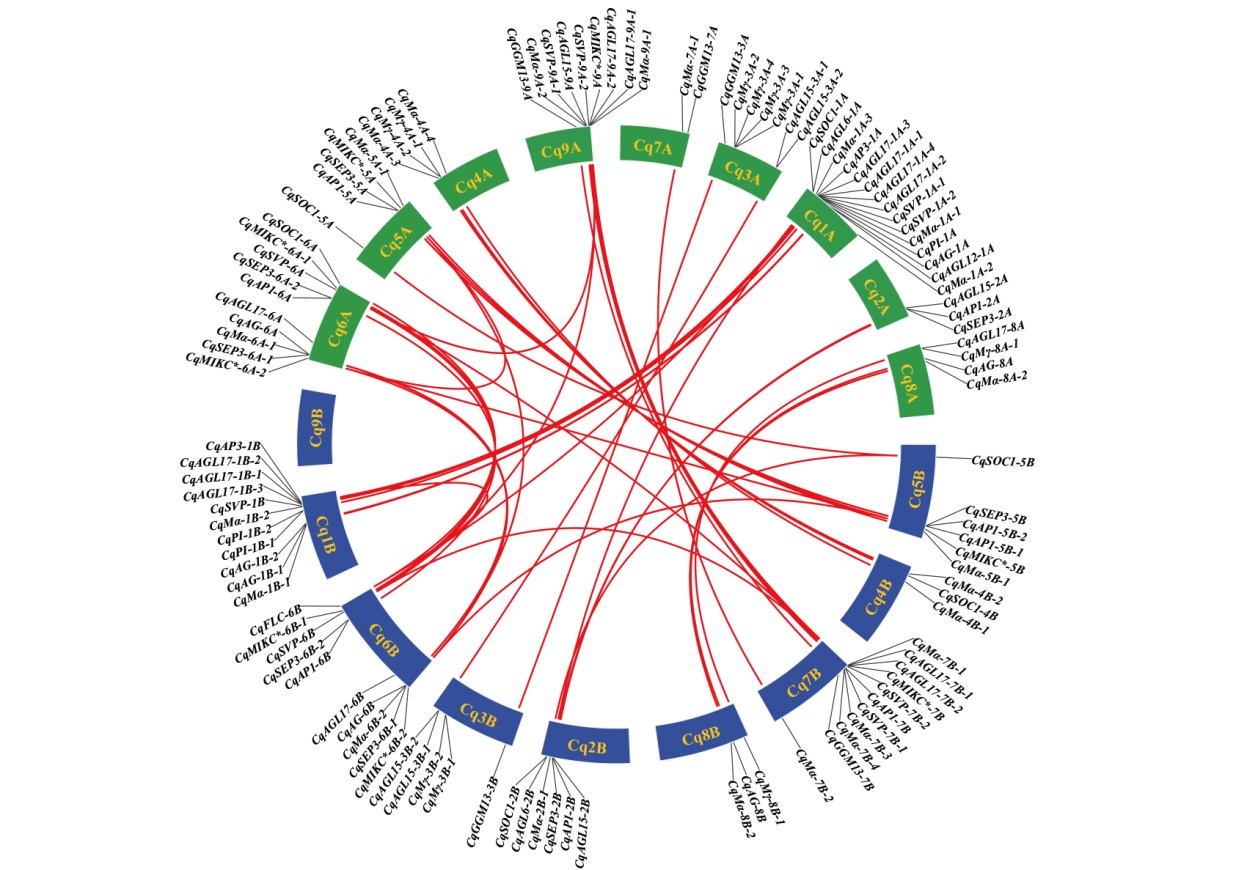
图4 藜麦亚基因组A和亚基因组B的MADS-box之间共线性 绿色表示藜麦A亚基因组,蓝色表示藜麦B亚基因组
Fig. 4 Collinearity between the MADS-box of quinoa sub-genome A and sub-genome B Green indicate quinoa A sub-genome, and blue represents quinoa B sub-genome

图5 藜麦与C. pallidicaule(A)和C. suecicum(B)的MADS-box共线性分析 绿色表示藜麦染色体,紫色表示C. pallidicaule染色体,蓝色表示C. suecicum染色体
Fig. 5 Collinearity analysis of quinoa MADS-box members with that in C. pallidicaule(A)and C. suecicum(B) Green indicates quinoa chromosome, purple indicates C. pallidicaule chromosome and blue indicates C. suecicum chromosome
| 基因名称 Gene name | 基因 ID Gene ID | 染色体 Chromosome | 起始位置 Start-end position(+/-strand) | 氨基酸数 Number of amino acids/aa | 类型 Type |
|---|---|---|---|---|---|
| CqSVP-1A-2 | CQ048095 | Cq1A | 1 174 6420-11 746 605(-) | 61 | 假基因-串联复制 |
| CqAGL17-1A-1 | CQ047997 | Cq1A | 10 659 725-10 659 946(-) | 73 | 假基因-串联复制 |
| CqAGL12-1A | CQ048800 | Cq1A | 22 590 418-22 590 639(-) | 73 | 假基因-节段复制 |
| CqAGL17-1A-2 | CQ048000 | Cq1A | 10 725 903-10 726 115(-) | 70 | 假基因-串联复制 |
| CqAG-1B-2 | CQ023885 | Cq1B | 49 507 013-49 507 303(-) | 96 | 假基因-串联复制 |
| CqMα-1B-1 | CQ023867 | Cq1B | 48 895 860-48 896 081(-) | 73 | 假基因-节段复制 |
| CqAGL17-1B-1 | CQ024731 | Cq1B | 64 342 030-64 342 293(+) | 87 | 假基因-串联复制 |
| CqAGL15-3B-2 | CQ019107 | Cq3B | 66 834 322-66 834 510(-) | 62 | 假基因-串联复制 |
| CqAGL15-3B-1 | CQ019105 | Cq3B | 66 809 225-66 812 527(-) | 99 | 假基因-节段复制 |
| CqSEP3-5B | CQ005895 | Cq5B | 67 053 435-67 053 671(+) | 78 | 假基因-串联复制 |
| CqAP1-5B-2 | CQ005897 | Cq5B | 67 131 352-67 131 611(+) | 64 | 假基因-串联复制 |
| CqMα-6A-1 | CQ025753 | Cq6A | 4 153 499-4 153 970(+) | 74 | 假基因-串联复制 |
| CqMα-6B-2 | CQ000301 | Cq6B | 3 428 220-3 429 042(+) | 75 | 假基因-节段复制 |
| CqSVP-7B-1 | CQ007326 | Cq7B | 3 310 550-3 310 735(-) | 61 | 假基因-串联复制 |
| CqMIKC*-7B | CQ007095 | Cq7B | 1 279 202-1 286 618(-) | 97 | 假基因-串联复制 |
| CqSVP-9A-1 | CQ052433 | Cq9A | 51 460 017-51 462 248(+) | 73 | 假基因-串联复制 |
| CqAGL17-9A-1 | CQ052648 | Cq9A | 53 340 285-53 346 018(+) | 77 | 假基因-串联复制 |
表1 藜麦MADS-box家族中假基因信息表
Table 1 Information of pseudogenes in the MADS-box family of quinoa
| 基因名称 Gene name | 基因 ID Gene ID | 染色体 Chromosome | 起始位置 Start-end position(+/-strand) | 氨基酸数 Number of amino acids/aa | 类型 Type |
|---|---|---|---|---|---|
| CqSVP-1A-2 | CQ048095 | Cq1A | 1 174 6420-11 746 605(-) | 61 | 假基因-串联复制 |
| CqAGL17-1A-1 | CQ047997 | Cq1A | 10 659 725-10 659 946(-) | 73 | 假基因-串联复制 |
| CqAGL12-1A | CQ048800 | Cq1A | 22 590 418-22 590 639(-) | 73 | 假基因-节段复制 |
| CqAGL17-1A-2 | CQ048000 | Cq1A | 10 725 903-10 726 115(-) | 70 | 假基因-串联复制 |
| CqAG-1B-2 | CQ023885 | Cq1B | 49 507 013-49 507 303(-) | 96 | 假基因-串联复制 |
| CqMα-1B-1 | CQ023867 | Cq1B | 48 895 860-48 896 081(-) | 73 | 假基因-节段复制 |
| CqAGL17-1B-1 | CQ024731 | Cq1B | 64 342 030-64 342 293(+) | 87 | 假基因-串联复制 |
| CqAGL15-3B-2 | CQ019107 | Cq3B | 66 834 322-66 834 510(-) | 62 | 假基因-串联复制 |
| CqAGL15-3B-1 | CQ019105 | Cq3B | 66 809 225-66 812 527(-) | 99 | 假基因-节段复制 |
| CqSEP3-5B | CQ005895 | Cq5B | 67 053 435-67 053 671(+) | 78 | 假基因-串联复制 |
| CqAP1-5B-2 | CQ005897 | Cq5B | 67 131 352-67 131 611(+) | 64 | 假基因-串联复制 |
| CqMα-6A-1 | CQ025753 | Cq6A | 4 153 499-4 153 970(+) | 74 | 假基因-串联复制 |
| CqMα-6B-2 | CQ000301 | Cq6B | 3 428 220-3 429 042(+) | 75 | 假基因-节段复制 |
| CqSVP-7B-1 | CQ007326 | Cq7B | 3 310 550-3 310 735(-) | 61 | 假基因-串联复制 |
| CqMIKC*-7B | CQ007095 | Cq7B | 1 279 202-1 286 618(-) | 97 | 假基因-串联复制 |
| CqSVP-9A-1 | CQ052433 | Cq9A | 51 460 017-51 462 248(+) | 73 | 假基因-串联复制 |
| CqAGL17-9A-1 | CQ052648 | Cq9A | 53 340 285-53 346 018(+) | 77 | 假基因-串联复制 |

图6 藜麦MADS-box家族成员启动子区域顺式作用元件预测 A:藜麦MADS-box家族成员启动子区域顺式作用元件分布图;B:藜麦MADS-box家族成员启动子区域结合的顺式作用元件数量统计
Fig. 6 Prediction of cis-acting elements in the promoter region of quinoa MADS-box family members A: The distribution map of cis-acting elements in the promoter region of quinoa MADS-box family members. B: Statistics on the number of cis-acting elements in the promoter region of quinoa MADS-box family members

图7 MADS-box在不同组织、六期花序和种子萌发中的表达分析 A:MADS-box在不同组织中的表达;表达值由FPKM值表示(下同);B:MADS-box在藜麦花序发育中的表达;YP1-YP4表示早期发育阶段的无分枝幼穗;P1和P2表示发育后期2个阶段的分枝穗;C:MADS-box在种子萌发过程中的表达变化;BL-A-5h/-15h和BL-5h/-15h分别表示ABA处理和未处理5 h和15 h的BL材料;每个表达值由3个重复生成;同一基因在6个阶段的表达水平逐行归一化为0-1
Fig. 7 Expression analysis of MADS-box in different tissues, six-stage inflorescences, and seed germination A: Expressions of MADS-box in different tissues. The expression value is represented by the FPKM values generated from the RNA-seq data(the same below). B: Expression of MADS-box in quinoa inflorescence development. YP1-YP4 indicates unbranched young spikelets in the early developmental stage. P1 and P2 indicate branched panicles at 2 stages of late development. C: Expression changes of MADS-box during seed germination. BL-A-5h/-15h and BL-5h/-15h indicate the ABA-treated and untreated BL materials at 5 h and 15 h after germination, respectively. Each expression value is generated from three replicates. The expressions of the same gene in the 6 stages were normalized to 0 to 1 by row
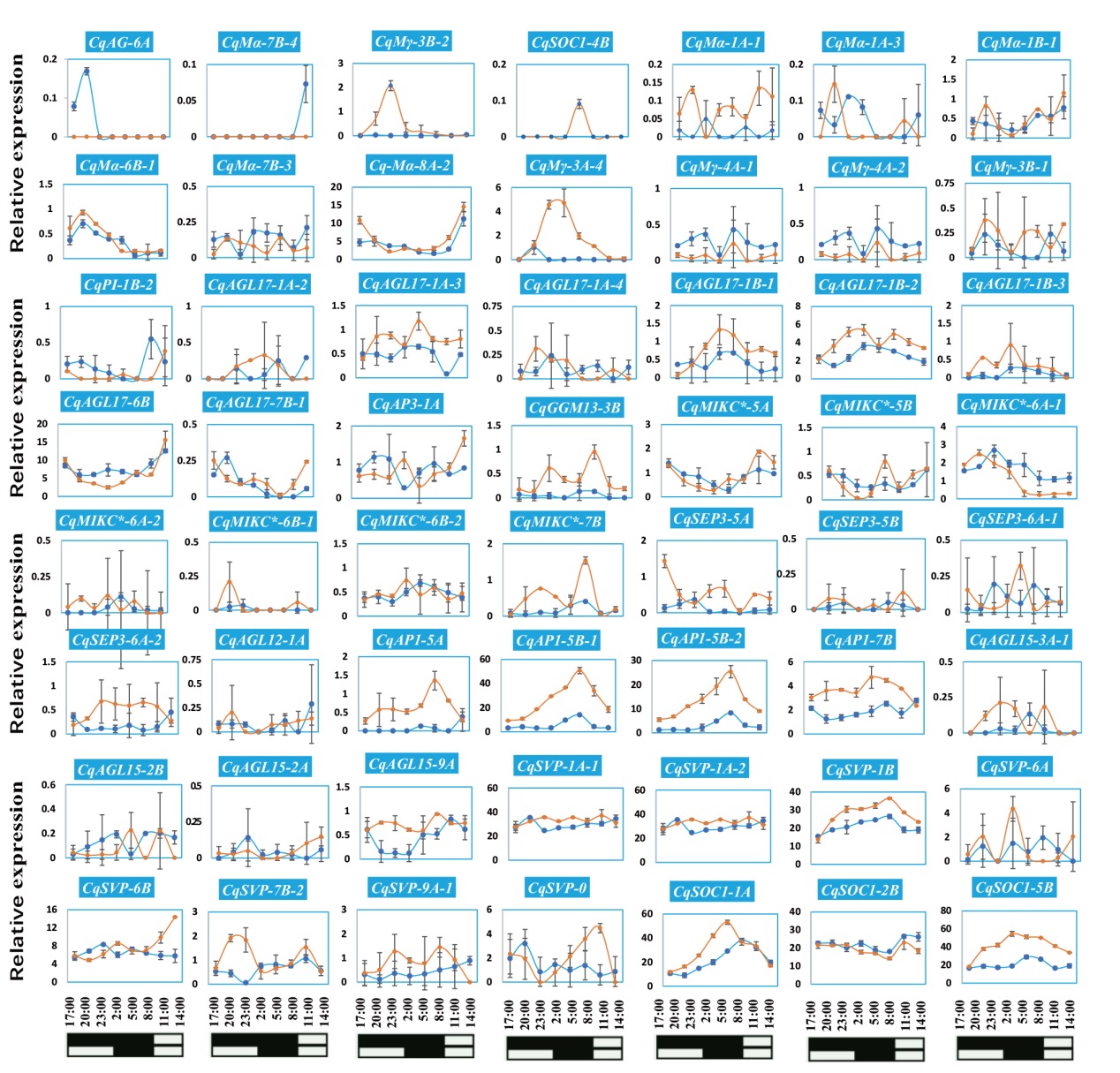
图8 短日照和长日照条件下CqMADS-box的昼夜表达模式 分别于17:00、20:00、23:00、02:00、05:00、08:00、11:00和14:00采集短日照和长日照条件下生长两周的藜麦幼苗叶片样本。采用RNA-seq分析CqMADS-box的表达模式。蓝色实线为LD下的基因表达曲线,橙色实线为SD下的基因表达曲线
Fig. 8 Diurnal expression patterns of CqMADS-box under short- and long-day conditions Leaf samples of quinoa seedlings grown for two weeks under short-day and long-day conditions were collected at 17:00, 20:00, 23:00, 02:00, 05:00, 08:00, 11:00 and 14:00, respectively. RNA-seq is used to analyze the expression pattern of CqMADS-box. The solid blue line is the gene expression curve under LD, and the solid orange line is the gene expression curve under SD
| [1] |
López-Marqués RL, Nørrevang AF, Ache P, et al. Prospects for the accelerated improvement of the resilient crop quinoa[J]. J Exp Bot, 2020, 71(18): 5333-5347.
doi: 10.1093/jxb/eraa285 pmid: 32643753 |
| [2] | Patiranage DSR, Asare E, Maldonado-Taipe N, et al. Haplotype variations of major flowering time genes in quinoa unveil their role in the adaptation to different environmental conditions[J]. Plant Cell Environ, 2021, 44(8): 2565-2579. |
| [3] |
Benlloch R, Berbel A, Ali L, et al. Genetic control of inflorescence architecture in legumes[J]. Front Plant Sci, 2015, 6: 543.
doi: 10.3389/fpls.2015.00543 pmid: 26257753 |
| [4] |
Schilling S, Pan SR, Kennedy A, et al. MADS-box genes and crop domestication: the jack of all traits[J]. J Exp Bot, 2018, 69(7): 1447-1469.
doi: 10.1093/jxb/erx479 pmid: 29474735 |
| [5] | Alvarez-Buylla ER, Pelaz S, Liljegren SJ, et al. An ancestral MADS-box gene duplication occurred before the divergence of plants and animals[J]. Proc Natl Acad Sci U S A, 2000, 97(10): 5328-5333. |
| [6] |
Kaufmann K, Melzer R, Theissen G. MIKC-type MADS-domain proteins: structural modularity, protein interactions and network evolution in land plants[J]. Gene, 2005, 347(2): 183-198.
doi: 10.1016/j.gene.2004.12.014 pmid: 15777618 |
| [7] |
Riechmann JL, Wang M, Meyerowitz EM. DNA-binding properties of Arabidopsis MADS domain homeotic proteins APETALA1, APETALA3, PISTILLATA and AGAMOUS[J]. Nucleic Acids Res, 1996, 24(16): 3134-3141.
pmid: 8774892 |
| [8] | Melzer R, Verelst W, Theiβen G. The class E floral homeotic protein SEPALLATA3 is sufficient to loop DNA in ‘floral quartet’-like complexes in vitro[J]. Nucleic Acids Res, 2009, 37(1): 144-157. |
| [9] | Honma T, Goto K. Complexes of MADS-box proteins are sufficient to convert leaves into floral organs[J]. Nature, 2001, 409(6819): 525-529. |
| [10] |
Wang YY, Jiang H, Wang GF. PHERES1 controls endosperm gene imprinting and seed development[J]. Trends Plant Sci, 2020, 25(6): 517-519.
doi: S1360-1385(20)30086-8 pmid: 32407690 |
| [11] | Paul P, Dhatt BK, Miller M, et al. MADS78 and MADS79 are essential regulators of early seed development in rice[J]. Plant Physiol, 2020, 182(2): 933-948. |
| [12] | Zhang JL, Dong TT, Hu ZL, et al. A SEPALLATA MADS-box transcription factor, SlMBP21, functions as a negative regulator of flower number and fruit yields in tomato[J]. Plants, 2024, 13(10): 1421. |
| [13] |
Amasino RM, Michaels SD. The timing of flowering[J]. Plant Physiol, 2010, 154(2): 516-520.
doi: 10.1104/pp.110.161653 pmid: 20921176 |
| [14] | Liu Y, Guan CN, Chen YY, et al. Evolutionary analysis of MADS-box genes in buckwheat species and functional study of FdMADS28 in flavonoid metabolism[J]. Plant Physiol Biochem, 2024, 210: 108637. |
| [15] |
Rey E, Maughan PJ, Maumus F, et al. A chromosome-scale assembly of the quinoa genome provides insights into the structure and dynamics of its subgenomes[J]. Commun Biol, 2023, 6(1): 1263.
doi: 10.1038/s42003-023-05613-4 pmid: 38092895 |
| [16] | Wu Q, Bai X, Wu XY, et al. Transcriptome profiling identifies transcription factors and key homologs involved in seed dormancy and germination regulation of Chenopodium quinoa[J]. Plant Physiol Biochem, 2020, 151: 443-456. |
| [17] | Wu Q, Bai X, Zhao W, et al. Investigation into the underlying regulatory mechanisms shaping inflorescence architecture in Chenopo-dium quinoa[J]. BMC Genomics, 2019, 20(1): 658. |
| [18] | Wu Q, Luo YM, Wu XY, et al. Identification of the specific long-noncoding RNAs involved in night-break mediated flowering retardation in Chenopodium quinoa[J]. BMC Genomics, 2021, 22(1): 284. |
| [19] |
Albert VA, Krabbenhoft TJ. Navigating the CoGe online software suite for polyploidy research[J]. Methods Mol Biol, 2023, 2545: 19-45.
doi: 10.1007/978-1-0716-2561-3_2 pmid: 36720806 |
| [20] | Garcia-Hernandez M, Berardini TZ, Chen GH, et al. TAIR: a resource for integrated Arabidopsis data[J]. Funct Integr Genomics, 2002, 2(6): 239-253. |
| [21] |
Parenicová L, de Folter S, Kieffer M, et al. Molecular and phylogenetic analyses of the complete MADS-box transcription factor family in Arabidopsis: new openings to the MADS world[J]. Plant Cell, 2003, 15(7): 1538-1551.
doi: 10.1105/tpc.011544 pmid: 12837945 |
| [22] |
Chen CJ, Chen H, Zhang Y, et al. TBtools: an integrative toolkit developed for interactive analyses of big biological data[J]. Mol Plant, 2020, 13(8): 1194-1202.
doi: S1674-2052(20)30187-8 pmid: 32585190 |
| [23] | Mistry J, Chuguransky S, Williams L, et al. Pfam: the protein families database in 2021[J]. Nucleic Acids Res, 2021, 49(D1): D412-D419. |
| [24] | Paysan-Lafosse T, Blum M, Chuguransky S, et al. InterPro in 2022[J]. Nucleic Acids Res, 2023, 51(D1): D418-D427. |
| [25] | Horton P, Park KJ, Obayashi T, et al. WoLF PSORT: protein localization predictor[J]. Nucleic Acids Res, 2007, 35(Web Server issue): W585-W587. |
| [26] |
Arora R, Agarwal P, Ray S, et al. MADS-box gene family in rice: genome-wide identification, organization and expression profiling during reproductive development and stress[J]. BMC Genomics, 2007, 8: 242.
pmid: 17640358 |
| [27] | Nguyen LT, Schmidt HA, von Haeseler A, et al. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies[J]. Mol Biol Evol, 2015, 32(1): 268-274. |
| [28] |
Rozas J, Sánchez-DelBarrio JC, Messeguer X, et al. DnaSP, DNA polymorphism analyses by the coalescent and other methods[J]. Bioinformatics, 2003, 19(18): 2496-2497.
doi: 10.1093/bioinformatics/btg359 pmid: 14668244 |
| [29] | Xie JB, Chen SS, Xu WJ, et al. Origination and function of plant Pseudogenes[J]. Plant Signal Behav, 2019, 14(8): 1625698. |
| [30] |
Zou CS, Chen AJ, Xiao LH, et al. A high-quality genome assembly of quinoa provides insights into the molecular basis of salt bladder-based salinity tolerance and the exceptional nutritional value[J]. Cell Res, 2017, 27(11): 1327-1340.
doi: 10.1038/cr.2017.124 pmid: 28994416 |
| [31] | Wu Q, Bai X, Nie MP, et al. Genome-wide identification and expression analysis disclose the pivotal PHOSPHATIDYLETHAN-OLAMINE BINDING PROTEIN members that may be utilized for yield improvement of Chenopodium quinoa[J]. Front Plant Sci, 2022, 13: 1119049. |
| [32] | Mirzaghaderi G. Genome-wide analysis of MADS-box transcription factor gene family in wild emmer wheat(Triticum turgidum subsp. dicoccoides)[J]. PLoS One, 2024, 19(3): e0300159. |
| [33] | Yang XY, Zhang MJ, Xi DX, et al. Genome-wide identification and expression analysis of the MADS gene family in sweet orange(Citrus sinensis)infested with pathogenic bacteria[J]. PeerJ, 2024, 12: e17001. |
| [34] |
Abdullah-Zawawi MR, Ahmad-Nizammuddin NF, Govender N, et al. Comparative genome-wide analysis of WRKY, MADS-box and MYB transcription factor families in Arabidopsis and rice[J]. Sci Rep, 2021, 11(1): 19678.
doi: 10.1038/s41598-021-99206-y pmid: 34608238 |
| [35] | Qiu YC, Li Z, Walther D, et al. Updated phylogeny and protein structure predictions revise the hypothesis on the origin of MADS-box transcription factors in land plants[J]. Mol Biol Evol, 2023, 40(9): msad194. |
| [36] | Jarvis DE, Sproul JS, Navarro-Domínguez B, et al. Chromosome-scale genome assembly of the hexaploid taiwanese goosefoot “djulis”(Chenopodium formosanum)[J]. Genome Biol Evol, 2022, 14(8): evac120. |
| [37] | Li L, Nie MP, Lu J, et al. Genome-wide identification, expression analysis and gene duplication analysis of Dof transcription factors between quinoa and its ancestral diploid sub-genome species reveal key CDF homologs involved in photoperiodic flowering regulation[J]. S Afr N J Bot, 2023, 162(6): 545-558. |
| [38] | Chai SY, Li KX, Deng XX, et al. Genome-wide analysis of the MADS-box gene family and expression analysis during anther development in Salvia miltiorrhiza[J]. Int J Mol Sci, 2023, 24(13): 10937. |
| [39] | Bai G, Yang DH, Cao PJ, et al. Genome-wide identification, gene structure and expression analysis of the MADS-box gene family indicate their function in the development of tobacco(Nicotiana tabacum L.)[J]. Int J Mol Sci, 2019, 20(20): 5043. |
| [40] | Zeng F, Zheng CM, Ge WX, et al. Regulatory function of the endogenous hormone in the germination process of quinoa seeds[J]. Front Plant Sci, 2023, 14: 1322986. |
| [1] | 崔原瑗, 王昭懿, 白双宇, 任毓昭, 豆飞飞, 刘彩霞, 刘凤楼, 王掌军, 李清峰. 大麦非特异性磷脂酶C基因家族全基因组鉴定及苗期胁迫表达分析[J]. 生物技术通报, 2024, 40(8): 74-82. |
| [2] | 杨巍, 赵丽芬, 唐兵, 周麟笔, 杨娟, 莫传园, 张宝会, 李飞, 阮松林, 邓英. 芥菜SRO基因家族全基因组鉴定与表达分析[J]. 生物技术通报, 2024, 40(8): 129-141. |
| [3] | 林彤, 袁程, 董陈文华, 曾孟琼, 杨燕, 毛自朝, 林春. 藜麦配子发育相关基因CqSTK的筛选及功能分析[J]. 生物技术通报, 2024, 40(8): 83-94. |
| [4] | 臧文蕊, 马明, 砗根, 哈斯阿古拉. 甜瓜BZR转录因子家族基因的全基因组鉴定及表达模式分析[J]. 生物技术通报, 2024, 40(7): 163-171. |
| [5] | 孙慧琼, 张春来, 王锡亮, 徐宏申, 窦苗苗, 杨博慧, 柴文婷, 赵珊珊, 姜晓东. 藜麦FLS基因家族的鉴定、表达及DNA变异分析[J]. 生物技术通报, 2024, 40(7): 172-182. |
| [6] | 阿丽亚·外力, 陈永坤, 克拉热木·克里木江, 王宝庆, 陈凌娜. 核桃SPL基因家族的系统进化和表达分析[J]. 生物技术通报, 2024, 40(6): 180-189. |
| [7] | 龚丽丽, 余花, 杨杰, 陈天池, 赵双滢, 吴月燕. 葡萄CYP707A基因家族的鉴定及对果实成熟的功能验证[J]. 生物技术通报, 2024, 40(2): 160-171. |
| [8] | 路喻丹, 刘晓驰, 冯新, 陈桂信, 陈义挺. 猕猴桃BBX基因家族成员鉴定与转录特征分析[J]. 生物技术通报, 2024, 40(2): 172-182. |
| [9] | 姜宇舢, 兰倩, 王芳, 姜亮, 裴成成. 一个影响酪氨酸代谢藜麦突变体的鉴定[J]. 生物技术通报, 2024, 40(10): 253-261. |
| [10] | 牛德, 何跃辉. 季节性因素调控小麦开花时间的分子与表观遗传机制[J]. 生物技术通报, 2024, 40(10): 30-40. |
| [11] | 张路阳, 韩文龙, 徐晓雯, 姚健, 李芳芳, 田效园, 张智强. 烟草TCP基因家族的鉴定及表达分析[J]. 生物技术通报, 2023, 39(6): 248-258. |
| [12] | 赖瑞联, 冯新, 高敏霞, 路喻丹, 刘晓驰, 吴如健, 陈义挺. 猕猴桃过氧化氢酶基因家族全基因组鉴定与表达分析[J]. 生物技术通报, 2023, 39(4): 136-147. |
| [13] | 段敏杰, 李怡斐, 杨小苗, 王春萍, 黄启中, 黄任中, 张世才. 辣椒锌指蛋白DnaJ-Like基因家族鉴定及对高温胁迫的表达响应[J]. 生物技术通报, 2023, 39(1): 187-198. |
| [14] | 袁星, 郭彩华, 刘金明, 亢超, 全绍文, 牛建新. 核桃CONSTANS-Like基因家族全基因组鉴定及表达分析[J]. 生物技术通报, 2022, 38(9): 167-179. |
| [15] | 冯建英, 李立芹, 鲁黎明. 马铃薯bHLH转录因子家族全基因组鉴定与表达分析[J]. 生物技术通报, 2022, 38(2): 21-33. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||