








生物技术通报 ›› 2025, Vol. 41 ›› Issue (3): 14-24.doi: 10.13560/j.cnki.biotech.bull.1985.2024-1038
收稿日期:2024-10-24
出版日期:2025-03-26
发布日期:2025-03-20
通讯作者:
马富强,男,博士,研究员,研究方向 :医药酶工程;E-mail: mafuqiang318@sibet.ac.cn作者简介:陆峰,男,博士,助理研究员,研究方向 :医药酶工程;E-mail: luf@sibet.ac.cn
基金资助:
LU Feng1( ), HUANG Yu-hong2, LIN Yan-na3, MA Fu-qiang1(
), HUANG Yu-hong2, LIN Yan-na3, MA Fu-qiang1( )
)
Received:2024-10-24
Published:2025-03-26
Online:2025-03-20
摘要:
随着全球对可持续能源转型和温室气体减排的迫切需求,CO₂的高效绿色转化成为能源、环境科学以及化学工程等多个领域的研究热点。特别是在应对气候变化的背景下,CO2的捕集和利用被视为实现碳中和的重要途径之一。甲酸脱氢酶(FDH)作为将CO₂还原为甲酸盐的重要生物催化剂,在绿色化学和生物能源转化中展现出巨大的应用潜力,然而其催化活性、热稳定性和辅酶特异性等方面仍存在一定局限性。近年来,随着蛋白质工程技术和分子生物学的不断发展,研究者们提出了多种改造FDH性能的策略,使得FDH的应用前景得到极大提升。例如通过定向突变、结构优化等酶工程手段提高酶的底物亲和力,增强酶的刚性和构象稳定性以及改变辅酶结合位点,拓宽其在绿色转化过程中的应用前景。本文将对近年来FDH催化CO2还原的研究进展进行全面总结,着重分析FDH在催化效率、热稳定性、辅酶特异性等方面的改进措施,探讨分子改造过程中所采取的具体策略,并总结这些策略的规律,以期为未来FDH的分子改造提供新的思路和方法,推动其在CO₂还原反应中的应用发展。此外,随着人工智能、机器学习和基因编辑等技术的发展,未来FDH的分子改造将更加高效和精确,这些新兴技术有望在较短的时间内筛选出具有优异性能的FDH突变体,为其在解决全球气候变化和能源危机方面提供可行的绿色解决方案。
陆峰, 黄玉红, 林燕娜, 马富强. CO2还原用甲酸脱氢酶分子改造的研究进展[J]. 生物技术通报, 2025, 41(3): 14-24.
LU Feng, HUANG Yu-hong, LIN Yan-na, MA Fu-qiang. Advances on Molecular Modifications of Formate Dehydrogenase for CO₂ Reduction[J]. Biotechnology Bulletin, 2025, 41(3): 14-24.

图1 多酶级联催化CO2合成甲醇流程图FDH:甲酸脱氢酶;FaldDH:甲醛脱氢酶;ADH:醇脱氢酶;NADH:还原型烟酰胺腺嘌呤二核苷酸;NAD+:氧化型烟酰胺腺嘌呤二核苷酸;W/Mo:钨/钼
Fig. 1 Flowchart of multi-enzyme cascade catalysis for CO2 to methanol synthesisFDH: Formate dehydrogenase; FaldDH: formaldehyde dehydrogenase; ADH: alcohol dehydrogenase; NADH: reduced nicotinamide adenine dinucleotide; NAD+: oxidized nicotinamide adenine dinucleotide; W/Mo: tungsten/molybdenum
| 类型 Types | 物种 Species | FDH | 分子量 Mw /kD | kcat /(s-1) | KM /(mmol·L-1) | kcat/KM / (mmol·L-1·s-1) | 参考文献 Reference |
|---|---|---|---|---|---|---|---|
| NAD+依赖型FDH | Candida boidinii | CbFDH | 41.0 | 0.015 0 | 31.3 | 0.000 400 | [ |
| Thiobacillus sp. KNK65MA | TsFDH | 45.0 | 0.318 | 9.23 | 0.034 0 | [ | |
| Thermochaetoides thermophila DSM 1495 | CtFDH | 45.0 | 0.023 0 | 0.320 | 0.069 0 | [ | |
| Candida methylica | CmFDH | 42.0 | 0.008 00 | 0.780 | 0.010 0 | [ | |
| Thermothelomyces thermophilus ATCC 42464 | MtFDH | 42.0 | 0.100 | 0.400 | 0.250 | [ | |
| Paracoccus sp. MKU1 | PsFDH | 44.0 | 0.073 0 | 0.928 | 0.079 0 | [ | |
| 金属依赖型FDH | Desulfovibrio desulfuricans | DdFDH | 135 | 46.6 | 0.015 7 | 2.97×103 | [ |
| Desulfovibrio vulgaris Hildenborough | DvFdhAB | 97.4 | 315 | 0.420 | 750 | [ | |
| Cupriavidus necator | FdsABG | 178 | 11.0 | 2.70 | 4.07 | [ | |
| Acetobacterium woodii | FdhF1/2 | 169 | 372 | 3.80 | 97.9 | [ | |
| Escherichia coli | EcFDH-H | 79.0 | 1.00 | 8.30 | 0.120 | [ | |
| Clostridium Ijungdahlii | ClFDH | 80.0 | 0.012 0 | 7.27 | 0.001 65 | [ | |
| Clostridium ljungdahlii | ClFDH | 75.0 | 5.66 | 66.2 | 0.085 5 | [ | |
| Clostridium autoethanogenum | CaFDH | 75.0 | 4.00 | 23.2 | 0.170 | [ | |
| Clostridium coskatii | CcFDH | 75.0 | 5.62 | 59.7 | 0.094 0 | [ | |
| Clostridium ragsdalei | CrFDH | 75.0 | 3.28 | 31.2 | 0.110 | [ | |
| Clostridium carboxidivorans P7T | FDHH_CloCa | 80.7 | 0.080 0 | 0.050 0 | 1.60 | [ | |
| Pseudomonas oxalaticus | Formate Dehydrogenase | 315 | 3.00 | 40.0 | 0.075 0 | [ | |
| Desulfosporosinus acididurans | DaFDH | 93.0 | 4.09 | 34.9 | 0.117 | [ | |
| Paraclostridium bifermentans | PbFDH | 93.0 | 4.45 | 30.6 | 0.145 | [ | |
| Clostridium scatologenes | CsFDH | 79.0 | 1.55 | 48.9 | 0.031 7 | [ |
表1 CO2还原用FDH的种类及其动力学参数
Table 1 Types and kinetic parameters of FDH for CO2 reduction
| 类型 Types | 物种 Species | FDH | 分子量 Mw /kD | kcat /(s-1) | KM /(mmol·L-1) | kcat/KM / (mmol·L-1·s-1) | 参考文献 Reference |
|---|---|---|---|---|---|---|---|
| NAD+依赖型FDH | Candida boidinii | CbFDH | 41.0 | 0.015 0 | 31.3 | 0.000 400 | [ |
| Thiobacillus sp. KNK65MA | TsFDH | 45.0 | 0.318 | 9.23 | 0.034 0 | [ | |
| Thermochaetoides thermophila DSM 1495 | CtFDH | 45.0 | 0.023 0 | 0.320 | 0.069 0 | [ | |
| Candida methylica | CmFDH | 42.0 | 0.008 00 | 0.780 | 0.010 0 | [ | |
| Thermothelomyces thermophilus ATCC 42464 | MtFDH | 42.0 | 0.100 | 0.400 | 0.250 | [ | |
| Paracoccus sp. MKU1 | PsFDH | 44.0 | 0.073 0 | 0.928 | 0.079 0 | [ | |
| 金属依赖型FDH | Desulfovibrio desulfuricans | DdFDH | 135 | 46.6 | 0.015 7 | 2.97×103 | [ |
| Desulfovibrio vulgaris Hildenborough | DvFdhAB | 97.4 | 315 | 0.420 | 750 | [ | |
| Cupriavidus necator | FdsABG | 178 | 11.0 | 2.70 | 4.07 | [ | |
| Acetobacterium woodii | FdhF1/2 | 169 | 372 | 3.80 | 97.9 | [ | |
| Escherichia coli | EcFDH-H | 79.0 | 1.00 | 8.30 | 0.120 | [ | |
| Clostridium Ijungdahlii | ClFDH | 80.0 | 0.012 0 | 7.27 | 0.001 65 | [ | |
| Clostridium ljungdahlii | ClFDH | 75.0 | 5.66 | 66.2 | 0.085 5 | [ | |
| Clostridium autoethanogenum | CaFDH | 75.0 | 4.00 | 23.2 | 0.170 | [ | |
| Clostridium coskatii | CcFDH | 75.0 | 5.62 | 59.7 | 0.094 0 | [ | |
| Clostridium ragsdalei | CrFDH | 75.0 | 3.28 | 31.2 | 0.110 | [ | |
| Clostridium carboxidivorans P7T | FDHH_CloCa | 80.7 | 0.080 0 | 0.050 0 | 1.60 | [ | |
| Pseudomonas oxalaticus | Formate Dehydrogenase | 315 | 3.00 | 40.0 | 0.075 0 | [ | |
| Desulfosporosinus acididurans | DaFDH | 93.0 | 4.09 | 34.9 | 0.117 | [ | |
| Paraclostridium bifermentans | PbFDH | 93.0 | 4.45 | 30.6 | 0.145 | [ | |
| Clostridium scatologenes | CsFDH | 79.0 | 1.55 | 48.9 | 0.031 7 | [ |

图2 CbFDH的结构及催化机制A:CbFDH三维结构(PDB:6D4C),红色和绿色分别为同源二聚体,橙色为NAD+;B:CbFDH的活性中心及与NAD+相近的氨基酸残基;C:NAD+依赖型FDH甲酸氧化(绿色)或CO2还原(粉色)的作用机制
Fig. 2 Structure and catalytic mechanism of CbFDHA: Three-dimensional structure of CbFDH (PDB: 6D4C). The two dimers are colored in red and green, while the NAD+ cofactor is colored in orange. B: Active center of CbFDH and amino acid residues closed to NAD+. C: Mechanism of action of NAD+-dependent FDH for formate oxidation (green arrows) or CO2 reduction (inverse sense, pink arrows)
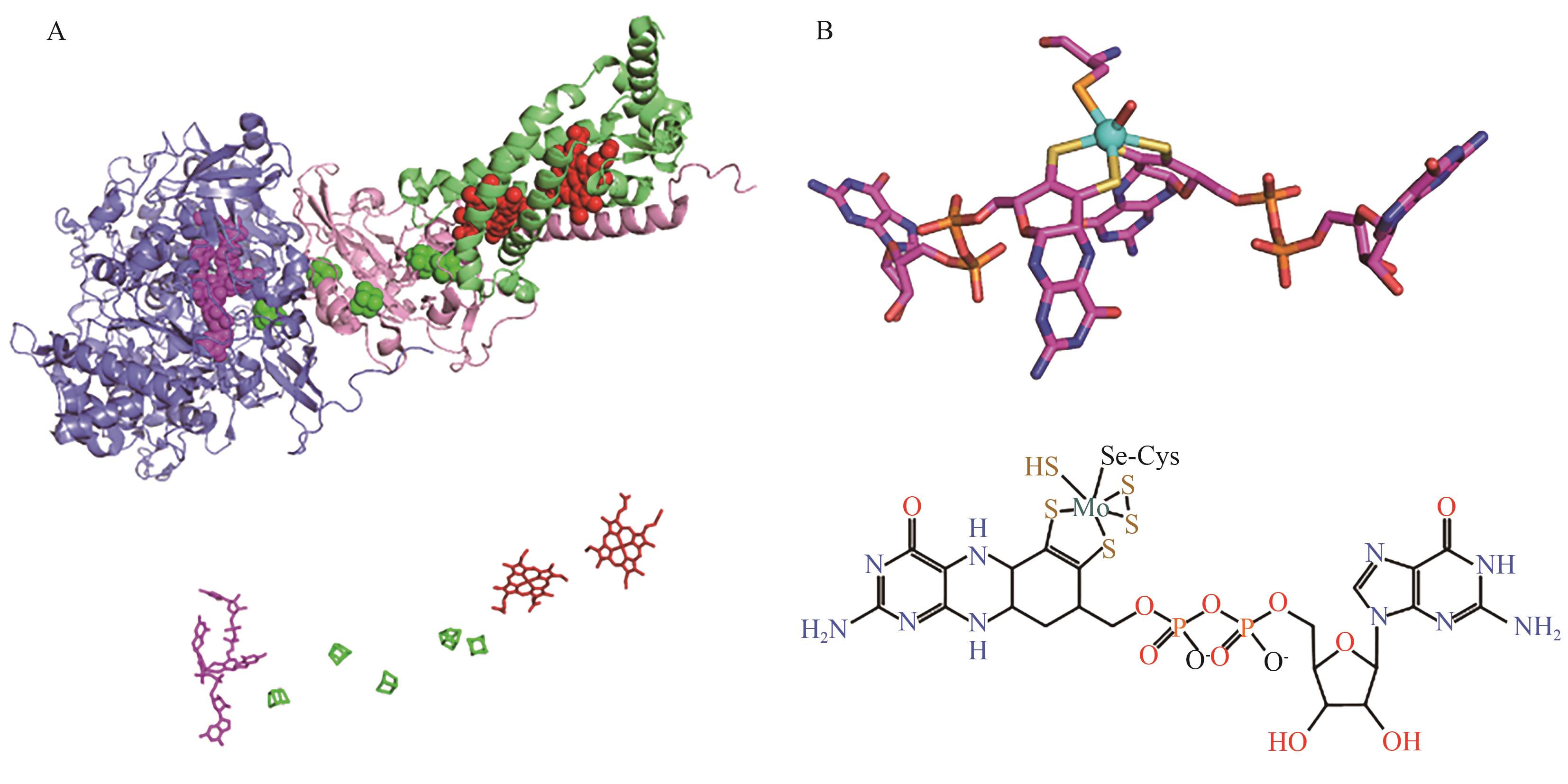
图3 大肠杆菌FDH N的三维结构及辅因子示意图A:大肠杆菌FDH N的三维结构(PDB:1KQF);3个多肽链分别用蓝色(链A)、粉色(链B)和绿色(链C)表示;钼喋呤(A链)、5个[4Fe-4S]簇(A链和B链)和2个血红素(C链)分别显示为粉色、绿色和红色; B:大肠杆菌FDH N中钼辅因子的图示及其化学式结构
Fig. 3 Three-dimensional structure and cofactor diagram of E. coli FDH NA: Three-dimensional structure of Escherichia coli FDH N (PDB: 1KQF). The three polypeptide chains are indicated by blue (chain A), pink (chain B) and green (chain C). The molybdopterin (in chain A), the five [4Fe-4S] clusters (in chain A and chain B) and the two heme (chain C) are displayed in pink, green and red, respectively. B: Graphic of the molybdenum cofactor and its chemical formula structure in E. coli FDH N
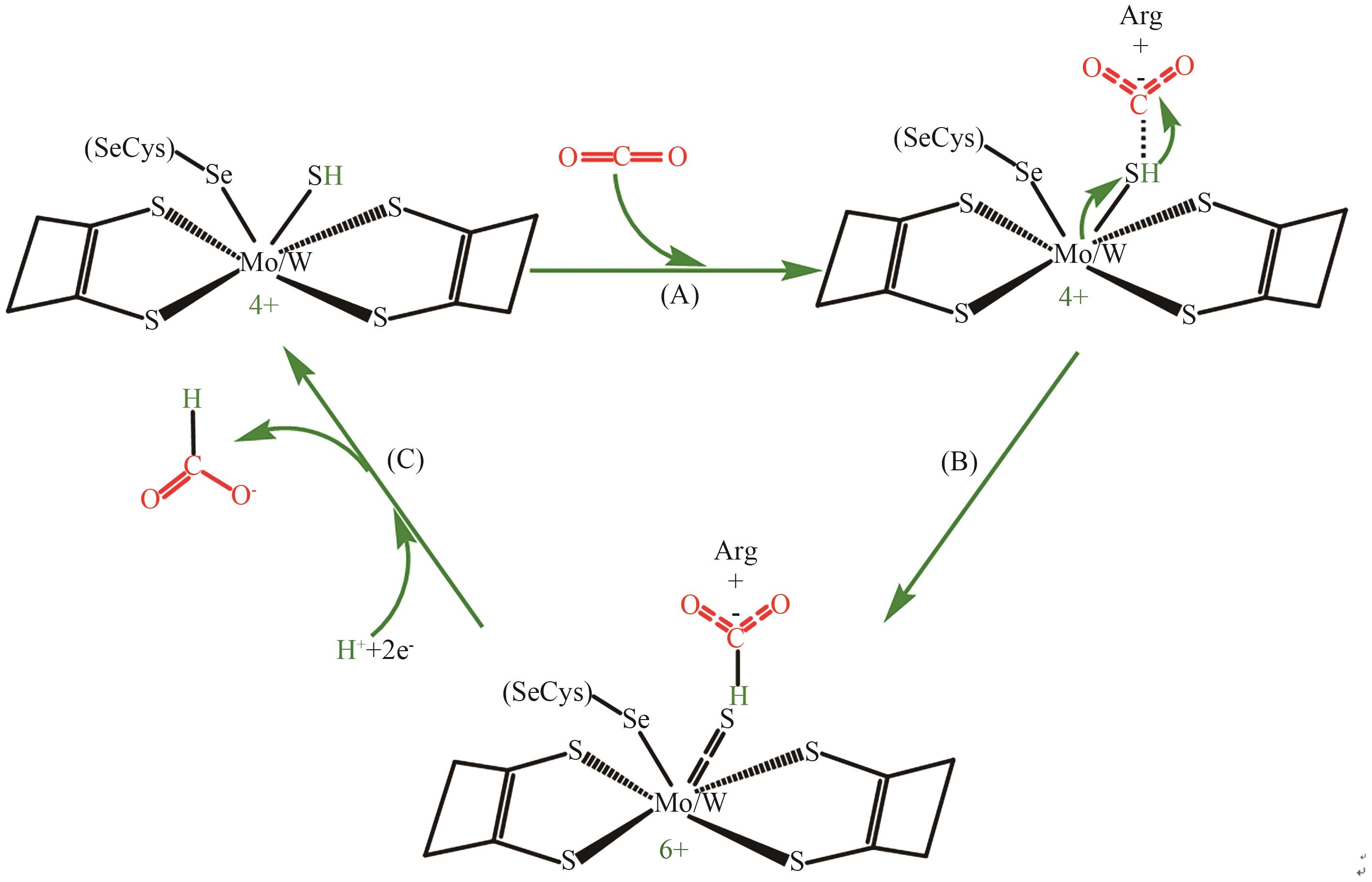
图4 金属依赖型FDH还原CO2成甲酸的可能机理A:FDH带正电的精氨酸残基静电捕获CO2; B:与金属Mo/W结合的硫醇氢攻击被精氨酸残基捕获的CO2,价态由四价氧化为六价;C:金属Mo/W在接受质子电子和释放甲酸的作用下,价态由六价又还原为四价,恢复初始状态
Fig. 4 Feasible mechanism of CO2 reduction to formic acid with metal-dependent FDHA: CO2 is electrostatically captured by positively charged arginine residues of FDH. B: Hydrogen mercaptan bound to metal Mo/W attacks CO2 captured by arginine residues, and the valence state is oxidized from four to six. C: Under the action of accepting proton electrons and releasing formic acid, the valence state of metal Mo/W is reduced from six to four, and the initial state is restored
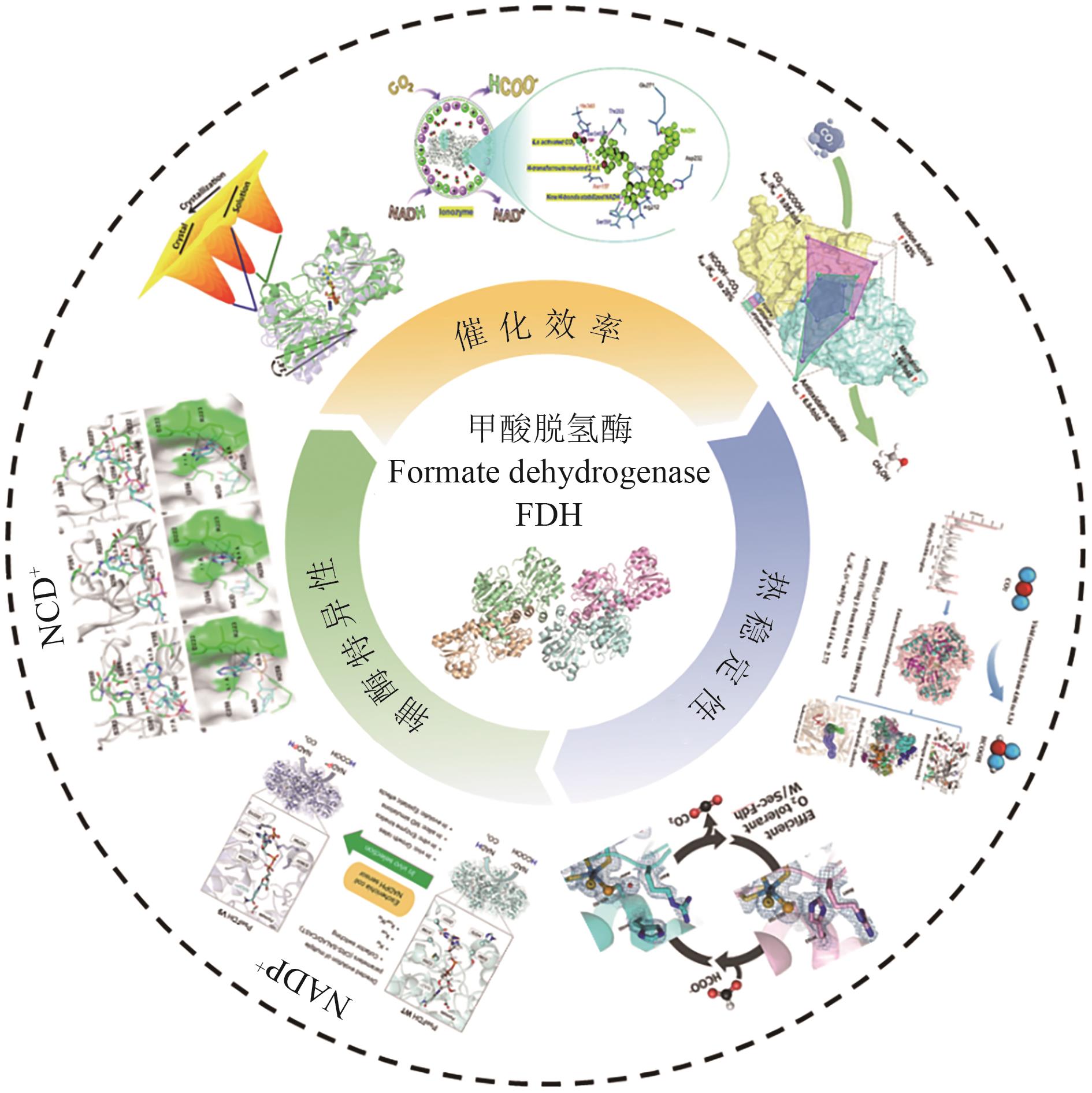
图5 分子改造提升FDH的催化活性、热稳定性以及辅酶偏好性示意图
Fig. 5 Schematic diagram of catalytic activity, thermal stability and coenzyme preference of FDH enhanced by molecular modification
| 35 | Alpdagtas S, Binay B. NADP+-dependent formate dehydrogenase: a review [J]. Biocatal Biotransform, 2021, 39(4): 260-268. |
| 36 | Alpdağtaş S, Turunen O, Valjakka J, et al. The challenges of using NAD+-dependent formate dehydrogenases for CO2 conversion [J]. Crit Rev Biotechnol, 2022, 42(6): 953-972. |
| 37 | Castillo R, Oliva M, Martí S, et al. A theoretical study of the catalytic mechanism of formate dehydrogenase [J]. J Phys Chem B, 2008, 112(32): 10012-10022. |
| 38 | Çakar MM, Ruupunen J, Mangas-Sanchez J, et al. Engineered formate dehydrogenase from Chaetomium thermophilum, a promising enzymatic solution for biotechnical CO2 fixation [J]. Biotechnol Lett, 2020, 42(11): 2251-2262. |
| 39 | Meneghello M, Oliveira AR, Jacq-Bailly A, et al. Formate dehydrogenases reduce CO2 rather than HCO3 -: an electrochemical demonstration [J]. Angew Chem Int Ed, 2021, 60(18): 9964-9967. |
| 40 | Pagano P, Guo Q, Ranasinghe C, et al. Oscillatory active-site motions correlate with kinetic isotope effects in formate dehydrogenase [J]. ACS Catal, 2019, 9(12): 11199-11206. |
| 41 | Cordas CM, Moura JJG. Molybdenum and tungsten enzymes redox properties-A brief overview [J]. Coord Chem Rev, 2019, 394: 53-64. |
| 42 | Kirk ML, Hille R. Spectroscopic studies of mononuclear molybdenum enzyme centers [J]. Molecules, 2022, 27(15): 4802. |
| 43 | Jormakka M, Törnroth S, Byrne B, et al. Molecular basis of proton motive force generation: structure of formate dehydrogenase-N [J]. Science, 2002, 295(5561): 1863-1868. |
| 44 | Maia LB, Moura I, Moura JJG. Molybdenum and tungsten-containing formate dehydrogenases: aiming to inspire a catalyst for carbon dioxide utilization [J]. Inorg Chim Acta, 2017, 455: 350-363. |
| 45 | Guo Q, Gakhar L, Wickersham K, et al. Structural and kinetic studies of formate dehydrogenase from Candida boidinii [J]. Biochemistry, 2016, 55(19): 2760-2771. |
| 46 | Sato R, Amao Y. Studies on the catalytic mechanism of formate dehydrogenase from Candida boidinii using isotope-labelled substrate and co-enzyme [J]. Catal Today, 2023, 411: 113796. |
| 47 | Jiang W, Lin P, Yang RN, et al. Identification of catalysis, substrate, and coenzyme binding sites and improvement catalytic efficiency of formate dehydrogenase from Candida boidinii [J]. Appl Microbiol Biotechnol, 2016, 100(19): 8425-8437. |
| 48 | Tishkov VI, Popov VO. Protein engineering of formate dehydrogenase [J]. Biomol Eng, 2006, 23(2/3): 89-110. |
| 49 | Pala U, Yelmazer B, Çorbacıoğlu M, et al. Functional effects of active site mutations in NAD+-dependent formate dehydrogenases on transformation of hydrogen carbonate to formate [J]. Protein Eng Des Sel, 2018, 31(9): 327-335. |
| 50 | Tülek A, Günay E, Servili B, et al. Sustainable production of formic acid from CO2 by a novel immobilized mutant formate dehydrogenase [J]. Sep Purif Technol, 2023, 309: 123090. |
| 51 | Kurt S, Ordu E. Effect of Met/Leu substitutions on the stability of NAD+-dependent formate dehydrogenases from Gossypium hirsutum [J]. Appl Microbiol Biotechnol, 2021, 105(7): 2787-2798. |
| 52 | Shi HL, Fu MR, Zhang TT, et al. Rational design of formate dehydrogenase for enhanced thermal stability and catalytic activity in bioelectrocatalysis [J]. J Agric Food Chem, 2024, 72(42): 23333-23344. |
| 53 | Schirwitz K, Schmidt A, Lamzin VS. High-resolution structures of formate dehydrogenase from Candida boidinii [J]. Protein Sci, 2007, 16(6): 1146-1156. |
| 54 | Alekseeva AA, Fedorchuk VV, Zarubina SA, et al. The role of Ala198 in the stability and coenzyme specificity of bacterial formate dehydrogenases [J]. Acta Naturae, 2015, 7(1): 60-69. |
| 55 | Calzadiaz-Ramirez L, Calvó-Tusell C, Stoffel GMM, et al. In vivo selection for formate dehydrogenases with high efficiency and specificity toward NADP+ [J]. ACS Catal, 2020, 10(14): 7512-7525. |
| 56 | Fogal S, Beneventi E, Cendron L, et al. Structural basis for double cofactor specificity in a new formate dehydrogenase from the Acidobacterium Granulicella mallensis MP5ACTX8 [J]. Appl Microbiol Biotechnol, 2015, 99(22): 9541-9554. |
| 57 | Robescu DMS, Rubini R, Beneventi DE, et al. From the amelioration of a NADP+-dependent formate dehydrogenase to the discovery of a new enzyme: round trip from theory to practice [J]. ChemCatChem, 2020, 12(9): 2478-2487. |
| 58 | Guo XJ, Wang XY, Liu YX, et al. Structure-guided design of formate dehydrogenase for regeneration of a non-natural redox cofactor [J]. Chemistry, 2020, 26(70): 16611-16615. |
| 1 | Jia ZC, Dang JN, Wen GB, et al. Constructing nanocaged enzymes for synergistic catalysis of CO2 reduction [J]. Adv Sci, 2023, 10(20): e2300752. |
| 2 | Chen H, Huang Y, Sha C, et al. Enzymatic carbon dioxide to formate: mechanisms, challenges and opportunities [J]. Renew Sustain Energy Rev, 2023, 178: 113271. |
| 3 | IEA. CO2 Emissions in 2023[EB/OL]. (2024-03-01) [2024-10-24]. . |
| 4 | Yuan SW, Fu MR, Xian YM, et al. Efficient heterologous expression of formate dehydrogenase and preliminary determination of the potential for conversion of carbon dioxide to formate [J]. Mol Catal, 2023, 548: 113455. |
| 5 | Li YX, Yan LH, Liu GH, et al. Enhanced electroenzymatic CO2 reduction by a multifunctional ZIF-8 layer on silica nanoflower with immobilized enzyme [J]. Chem Eng J, 2023, 466: 143198. |
| 6 | Cai T, Sun HB, Qiao J, et al. Cell-free chemoenzymatic starch synthesis from carbon dioxide [J]. Science, 2021, 373(6562): 1523-1527. |
| 7 | Ji XL, Guo H, Xue YJ, et al. Microenvironment: an efficient avenue for converting CO2 to high-value compounds [J]. Renew Sustain Energy Rev, 2023, 188: 113809. |
| 8 | Moon M, Park GW, Lee JP, et al. Recent progress in formate dehydrogenase (FDH) as a non-photosynthetic CO2 utilizing enzyme: a short review [J]. J CO2 Util, 2020, 42: 101353. |
| 9 | Villa R, Nieto S, Donaire A, et al. Direct biocatalytic processes for CO2 capture as a green tool to produce value-added chemicals [J]. Molecules, 2023, 28(14): 5520. |
| 10 | Kuwabata S, Tsuda R, Nishida K, et al. Electrochemical conversion of carbon dioxide to methanol with use of enzymes as biocatalysts [J]. Chem Lett, 1993, 22(9): 1631-1634. |
| 11 | Calzadiaz-Ramirez L, Meyer AS. Formate dehydrogenases for CO2 utilization [J]. Curr Opin Biotechnol, 2022, 73: 95-100. |
| 12 | Nielsen CF, Lange L, Meyer AS. Classification and enzyme kinetics of formate dehydrogenases for biomanufacturing via CO2 utilization [J]. Biotechnol Adv, 2019, 37(7): 107408. |
| 13 | Hille R, Hall J, Basu P. The mononuclear molybdenum enzymes [J]. Chem Rev, 2014, 114(7): 3963-4038. |
| 14 | Popov VO, Lamzin VS. NAD+-dependent formate dehydrogenase [J]. Biochem J, 1994, 301 (Pt 3)(Pt 3): 625-643. |
| 15 | Jollie DR, Lipscomb JD. Formate dehydrogenase from Methylosinus trichosporium OB3b. Purification and spectroscopic characterization of the cofactors [J]. J Biol Chem, 1991, 266(32): 21853-21863. |
| 16 | Amao Y. Formate dehydrogenase for CO2 utilization and its application [J]. J CO2 Util, 2018, 26: 623-641. |
| 17 | Oliveira AR, Mota C, Mourato C, et al. Toward the mechanistic understanding of enzymatic CO2 reduction [J]. ACS Catal, 2020, 10(6): 3844-3856. |
| 18 | Shi HL, Fu MR, Yuan SW, et al. Engineered Escherichia coli whole cell-mediated electro-biocatalysis for carbon dioxide to formic acid conversion [J]. ACS Sustainable Chem Eng, 2024, 12(14): 5544-5554. |
| 19 | Min K, Park YS, Park GW, et al. Elevated conversion of CO2 to versatile formate by a newly discovered formate dehydrogenase from Rhodobacter aestuarii [J]. Bioresour Technol, 2020, 305: 123155. |
| 20 | Choe H, Joo JC, Cho DH, et al. Efficient CO2-reducing activity of NAD-dependent formate dehydrogenase from Thiobacillus sp. KNK65MA for formate production from CO2 gas [J]. PLoS One, 2014, 9(7): e103111. |
| 21 | Aslan AS, Valjakka J, Ruupunen J, et al. Chaetomium thermophilum formate dehydrogenase has high activity in the reduction of hydrogen carbonate (HCO3 -) to formate [J]. Protein Eng Des Sel, 2017, 30(1): 47-55. |
| 22 | Altaş N, Aslan AS, Karataş E, et al. Heterologous production of extreme alkaline thermostable NAD+-dependent formate dehydrogenase with wide-range pH activity from Myceliophthora thermophila [J]. Process Biochem, 2017, 61: 110-118. |
| 23 | Xue YJ, Ji XL, Li Z, et al. NADH-dependent formate dehydrogenase mutants for efficient carbon dioxide fixation [J]. Bioresour Technol, 2024, 393: 130027. |
| 24 | Maia LB, Fonseca L, Moura I, et al. Reduction of carbon dioxide by a molybdenum-containing formate dehydrogenase: a kinetic and mechanistic study [J]. J Am Chem Soc, 2016, 138(28): 8834-8846. |
| 59 | Radon C, Mittelstädt G, Duffus BR, et al. Cryo-EM structures reveal intricate Fe-S cluster arrangement and charging in Rhodobacter capsulatus formate dehydrogenase [J]. Nat Commun, 2020, 11(1): 1912. |
| 25 | da Silva SM, Pimentel C, Valente FMA, et al. Tungsten and molybdenum regulation of formate dehydrogenase expression in Desulfovibrio vulgaris Hildenborough [J]. J Bacteriol, 2011, 193(12): 2909-2916. |
| 26 | Niks D, Duvvuru J, Escalona M, et al. Spectroscopic and kinetic properties of the molybdenum-containing, NAD+-dependent formate dehydrogenase from Ralstonia eutropha [J]. J Biol Chem, 2016, 291(3): 1162-1174. |
| 27 | Schuchmann K, Müller V. Direct and reversible hydrogenation of CO2 to formate by a bacterial carbon dioxide reductase [J]. Science, 2013, 342(6164): 1382-1385. |
| 28 | Bassegoda A, Madden C, Wakerley DW, et al. Reversible interconversion of CO2 and formate by a molybdenum-containing formate dehydrogenase [J]. J Am Chem Soc, 2014, 136(44): 15473-15476. |
| 29 | Mervan Çakar M, Mangas-Sanchez J, Birmingham WR, et al. Discovery of a new metal and NAD+-dependent formate dehydrogenase from Clostridium ljungdahlii [J]. Prep Biochem Biotechnol, 2018, 48(4): 327-334. |
| 30 | Moon M, Park GW, Lee JP, et al. Recombinant expression and characterization of formate dehydrogenase from Clostridium ljungdahlii (ClFDH) as CO2 reductase for converting CO2 to formate [J]. J CO2 Util, 2022, 57: 101876. |
| 31 | Min K, Moon M, Park GW, et al. Newly explored formate dehydrogenases from Clostridium species catalyze carbon dioxide to formate [J]. Bioresour Technol, 2022, 348: 126832. |
| 32 | Alissandratos A, Kim HK, Matthews H, et al. Clostridium carboxidivorans strain P7T recombinant formate dehydrogenase catalyzes reduction of CO2 to formate [J]. Appl Environ Microbiol, 2013, 79(2): 741-744. |
| 33 | Müller U, Willnow P, Ruschig U, et al. Formate dehydrogenase from Pseudomonas oxalaticus [J]. Eur J Biochem, 1978, 83(2): 485-498. |
| 34 | Ruschig U, Müller U, Willnow P, et al. CO2 reduction to formate by NADH catalysed by formate dehydrogenase from Pseudomonas oxalaticus [J]. Eur J Biochem, 1976, 70(2): 325-330. |
| [1] | 张阿娜, 韩雪, 谷天一, 辛凤姣, 王钰璐. 利用新型红酵母苯丙氨酸解氨酶制备低苯丙氨酸酪蛋白[J]. 生物技术通报, 2024, 40(8): 309-319. |
| [2] | 乔烨, 张楠, 杨建花, 张翠英, 朱蕾蕾. 糖磷酸酶的挖掘及其酶学性质研究[J]. 生物技术通报, 2024, 40(7): 299-306. |
| [3] | 韩雪, 张阿娜, 王海燕, 辛凤姣, 谷天一, 王钰璐. 基于计算设计的GH11家族木聚糖酶CDBFV的热稳定性改造及潜在机制研究[J]. 生物技术通报, 2024, 40(10): 305-314. |
| [4] | 韩惠, 张舰, 任宇红. 短链脱氢酶Lvchun的分子改造及其在氯霉胺合成中的应用[J]. 生物技术通报, 2023, 39(4): 81-92. |
| [5] | 张晨, 张佟佟, 刘海萍. 高活性和高热稳定性乙烯合成酶的筛选和鉴定[J]. 生物技术通报, 2022, 38(11): 269-276. |
| [6] | 张雪, 谭玉萌, 蒋海霞, 杨广宇. 基于单细胞超高通量筛选的α-1,2-岩藻糖基转移酶定向进化[J]. 生物技术通报, 2022, 38(1): 289-298. |
| [7] | 郝俊尧, 马富强, 杨广宇. 产碱杆菌Alcaligenes sp.KS-85来源肌酸酶活性中心的关键氨基酸功能研究[J]. 生物技术通报, 2021, 37(3): 75-83. |
| [8] | 陈春, 宿玲恰, 夏伟, 吴敬. 定向进化提高来源于Arthrobacter ramosus 的MTHase的热稳定性[J]. 生物技术通报, 2021, 37(3): 84-91. |
| [9] | 吴娇, 余桂珍, 袁航, 刘娴, 高艳秀, 龚明, 邹竹荣. 融合超嗜热菌Pyrococcus furiosus红素氧还蛋白可提高靶蛋白的热稳定性[J]. 生物技术通报, 2021, 37(10): 110-119. |
| [10] | 仲建锋, 李兴奎, 徐重新, 张霄, 刘贤金. Cry1B抗独特型单链抗体的定点突变及生物活性分析[J]. 生物技术通报, 2021, 37(10): 186-195. |
| [11] | 孙熙麟, 蒋振彦, 刘志屹, 戴璐, 孙非, 黄伟. 氨基酸定点突变提高灵芝蛋白LZ-8热稳定性的研究[J]. 生物技术通报, 2020, 36(1): 23-28. |
| [12] | 邱锦, 黄火清, 姚斌, 罗会颖. 解淀粉芽孢杆菌淀粉酶催化活力改良及其在枯草芽孢杆菌中的高效表达[J]. 生物技术通报, 2019, 35(9): 134-143. |
| [13] | 华晨, 李新新, 涂涛, 杨虹, 罗会颖, 陈家明, 姚斌, 柏映国, 彭书传. 基于酶热稳定性系统计算的乳酸氧化酶热稳定性改造[J]. 生物技术通报, 2018, 34(8): 144-150. |
| [14] | 袁林, 黄朝, 曾静, 郭建军, 张婷, 吕珺,. 植酸酶YiAPPA与生淀粉结合域SBD融合酶的构建及酶学性质分析[J]. 生物技术通报, 2018, 34(3): 200-207. |
| [15] | 任天雷, 杨海泉, 许菲. 基于分子结构与生物信息学等多维度特征的定向进化改造甲基对硫磷水解酶[J]. 生物技术通报, 2018, 34(10): 194-200. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||