








生物技术通报 ›› 2023, Vol. 39 ›› Issue (3): 278-289.doi: 10.13560/j.cnki.biotech.bull.1985.2022-0732
徐小文1( ), 李金仓2, 海都2, 查玉平1, 宋菲1, 王义勋1(
), 李金仓2, 海都2, 查玉平1, 宋菲1, 王义勋1( )
)
收稿日期:2022-06-17
出版日期:2023-03-26
发布日期:2023-04-10
通讯作者:
王义勋,博士,副研究员,研究方向:森林病虫害防治;E-mail: 594580124@qq.com作者简介:徐小文,博士,助理研究员,研究方向:森林病害防治;E-mail: xuxiaowen222@126.com
基金资助:
XU Xiao-wen1( ), LI Jin-cang2, HAI Du2, ZHA Yu-ping1, SONG Fei1, WANG Yi-xun1(
), LI Jin-cang2, HAI Du2, ZHA Yu-ping1, SONG Fei1, WANG Yi-xun1( )
)
Received:2022-06-17
Published:2023-03-26
Online:2023-04-10
摘要:
炭疽菌是核桃主要病害核桃炭疽病的病原,目前对核桃炭疽菌携带的病毒种类及病毒基因组序列信息了解较少。利用宏病毒组测序技术,对分离自我国3个不同省份的25株核桃炭疽菌所携带的病毒基因序列进行发掘、鉴定和分类。经同源比对分析,获得22种病毒基因组序列,其中19种为新病毒。在22种病毒中,有21种为正单链RNA病毒,1种为dsRNA病毒。正单链RNA分别隶属于裸露病毒科(Narnaviridae)、线粒体病毒科(Mitoviridae)和葡萄孢欧尔密病毒科(Botourmiaviridae),而dsRNA属于Alternaviridae病毒科。RT-PCR验证结果表明22种病毒都能在核桃炭疽菌株中被检测到,且25株炭疽菌的病毒携带率为100%,每个菌株都至少被1-11种病毒侵染。研究结果丰富了炭疽菌属所携带的病毒基因组信息,为后续深入分析炭疽菌真菌病毒的多样性和分子特性奠定基础。
徐小文, 李金仓, 海都, 查玉平, 宋菲, 王义勋. 核桃炭疽菌携带病毒种类鉴定及多样性分析[J]. 生物技术通报, 2023, 39(3): 278-289.
XU Xiao-wen, LI Jin-cang, HAI Du, ZHA Yu-ping, SONG Fei, WANG Yi-xun. Identification and Diversity Analysis of Mycoviruses from the Phytopathogenic Fungus Colletotrichum spp. of Walnut[J]. Biotechnology Bulletin, 2023, 39(3): 278-289.
| 种名Species | 菌株编号Strain No. | 采集地点Collecting location | 种名Species | 菌株编号Strain No. | 采集地点Collecting location | |
|---|---|---|---|---|---|---|
| C. gloeosporioides | SZ2-3 | Shangzhou, Shanxi, China | C. siamense | YX1-5 | Yunxi, Hubei, China | |
| C. gloeosporioides | DJ1-1 | Danjiang, Hubei, China | C. fioriniae | BK3-5 | Baokang, Hubei, China | |
| C. gloeosporioides | DJ2-1 | Danjiang, Hubei, China | C. fioriniae | CX3 | Chengxian, Gansu, China | |
| C. gloeosporioides | DJ2-4 | Danjiang, Hubei, China | C. fioriniae | BK2-3 | Baokang, Hubei, China | |
| C. gloeosporioides | YY1-11 | Yunyang, Hubei, China | C. fioriniae | CX1 | Chengxian, Gansu, China | |
| C. fructicola | XS6 | Xingshan, Hubei, China | C. fioriniae | CX7 | Chengxian, Gansu, China | |
| C. fructicola | FX1-3 | Fengxiang, Shanxi, China | C. nymphaeae | ZS3-13 | Zhushan, Hubei, China | |
| C. fructicola | CX6 | Chengxian, Gansu, China | C. nymphaeae | ZS3-8 | Zhushan, Hubei, China | |
| C. fructicola | YY1-28 | Yunyang, Hubei, China | C. nymphaeae | BK1-11 | Baokang, Hubei, China | |
| C. fructicola | BK2-1 | Baokang, Hubei, China | C. nymphaeae | ZS2-6 | Zhushan, Hubei, China | |
| C. siamense | YX1-7 | Yunxi, Hubei, China | C. nymphaeae | XS5 | Xingshan, Hubei, China | |
| C. siamense | ZS2-9 | Zhushan, Hubei, China | C. nymphaeae | BK4-4 | Baokang, Hubei, China | |
| C. siamense | DJ3-5 | Danjiang, Hubei, China |
表1 供试25株核桃炭疽菌株信息列表
Table 1 Information of the Colletotrichum species from 25 walnut plants for experiment
| 种名Species | 菌株编号Strain No. | 采集地点Collecting location | 种名Species | 菌株编号Strain No. | 采集地点Collecting location | |
|---|---|---|---|---|---|---|
| C. gloeosporioides | SZ2-3 | Shangzhou, Shanxi, China | C. siamense | YX1-5 | Yunxi, Hubei, China | |
| C. gloeosporioides | DJ1-1 | Danjiang, Hubei, China | C. fioriniae | BK3-5 | Baokang, Hubei, China | |
| C. gloeosporioides | DJ2-1 | Danjiang, Hubei, China | C. fioriniae | CX3 | Chengxian, Gansu, China | |
| C. gloeosporioides | DJ2-4 | Danjiang, Hubei, China | C. fioriniae | BK2-3 | Baokang, Hubei, China | |
| C. gloeosporioides | YY1-11 | Yunyang, Hubei, China | C. fioriniae | CX1 | Chengxian, Gansu, China | |
| C. fructicola | XS6 | Xingshan, Hubei, China | C. fioriniae | CX7 | Chengxian, Gansu, China | |
| C. fructicola | FX1-3 | Fengxiang, Shanxi, China | C. nymphaeae | ZS3-13 | Zhushan, Hubei, China | |
| C. fructicola | CX6 | Chengxian, Gansu, China | C. nymphaeae | ZS3-8 | Zhushan, Hubei, China | |
| C. fructicola | YY1-28 | Yunyang, Hubei, China | C. nymphaeae | BK1-11 | Baokang, Hubei, China | |
| C. fructicola | BK2-1 | Baokang, Hubei, China | C. nymphaeae | ZS2-6 | Zhushan, Hubei, China | |
| C. siamense | YX1-7 | Yunxi, Hubei, China | C. nymphaeae | XS5 | Xingshan, Hubei, China | |
| C. siamense | ZS2-9 | Zhushan, Hubei, China | C. nymphaeae | BK4-4 | Baokang, Hubei, China | |
| C. siamense | DJ3-5 | Danjiang, Hubei, China |
| Contig 编号 Contig code | 病毒名称 Name of putative viruses | 病毒名称缩写 Name abbreviation | 长度 Length | 序列相似度 Acid similarity/% | E值 E-value | 最佳匹配 Best match | 基因组类型 Genome type | 病毒科 Family | Contig 计数 Contig counts |
|---|---|---|---|---|---|---|---|---|---|
| Contig 22546 | Colletotrichum fioriniae alternavirus 1 | CfAV1 dsRNA2 | 2 463 bp | 70.4 | 3.50E-303 | Fusarium graminearum alternavirus 1(YP_009449446.1) | dsRNA | Alternaviridae | 1 |
| Contig11036 | Colletotrichum fioriniae alternavirus 1 | CfAV1 dsRNA1 | 3 534 bp | 73.3 | 0.00E+00 | Fusarium graminearum alternavirus 1(YP_009449439.1) | dsRNA | Alternaviridae | 1 |
| Contig22686 | Colletotrichum fioriniae alternavirus 1 | CfAV1 dsRNA3 | 2 454 bp | 72 | 3.00E-307 | Fusarium graminearum alternavirus 1(AUI80777.1) | dsRNA | Alternaviridae | 1 |
| Contig51809 | Colletotrichum fioriniae narnavirus 1 | CfNV1 | 1 136 nt | 59.28 | 3.00E-76 | Sclerotinia sclerotiorum narnavirus 2(QZE12028.1) | +ssRNA | Narnaviridae | 1 |
| Contig26097 | Colletotrichum siamense narnavirus 1 | CsNV1 | 2 239 nt | 65.3 | 0.00E+00 | Sclerotinia sclerotiorum narnavirus 2(QZE12027.1) | +ssRNA | Narnaviridae | 1 |
| Contig20122 | Colletotrichum siamense narnavirus 2 | CsNV2 | 2 420 nt | 33.2 | 8.00E-88 | Oidiodendron maius splipalmivirus 1(QNN89180.1) | +ssRNA | Narnaviridae | 1 |
| Contig11676 | Colletotrichum nymphaeae narnavirus 1 | CnNV1 | 3 090 nt | 65 | 0.00E+00 | Botryosphaeria dothidea narnavirus 3(QQD86177.1) | +ssRNA | Narnaviridae | 5 |
| Contig10410 | Colletotrichum fioriniae narnavirus 2 | CfNV2 | 3 619 nt | 55.3 | 0.00E+00 | Erysiphe necator associated narnavirus 15(QJT93747.1) | +ssRNA | Narnaviridae | 1 |
| Contig19559 | Colletotrichum nymphaeae narnavirus 2 | CnNV2 | 2 455 nt | 62.2 | 1.00E-278 | Erysiphe necator associated narnavirus 4(QHD64827.1) | +ssRNA | Narnaviridae | 2 |
| Contig4718 | Colletotrichum gloeosporioides narnavirus 1 | CgNV1 | 4 200 nt | 51.8 | 1.60E-300 | Monilinia narnavirus H (QED42934.1) | +ssRNA | Narnaviridae | 3 |
| Contig35776 | Colletotrichum fructicola narnavirus 1 | CfrNV1 | 1 689 nt | 60 | 1.00E-27 | Aspergillus lentulus narnavirus 1(BCH36644.1) | +ssRNA | Narnaviridae | 2 |
| Contig19474 | Colletotrichum fructicola botourmiavirus 1 | CfBV1 | 2 460 nt | 55.1 | 1.60E-178 | Pestalotiopsis botourmiavirus 3(QTH80197.1) | +ssRNA | Botourmiaviridae | 2 |
| Contig22684 | Colletotrichum fioriniae mitovirus 1 | CfMV1 | 2 455 nt | 51.7 | 4.00E-158 | Grapevine-associated mitovirus 10(QXN75363.1) | +ssRNA | Mitoviridae | 2 |
| Contig41113 | Colletotrichum fioriniae mitovirus 2 | CfMV2 | 1 493 nt | 78 | 9.10E-113 | Rhizoctonia solani mitovirus 33(QDW65423.1) | +ssRNA | Mitoviridae | 1 |
| Contig55502 | Colletotrichum fioriniae mitovirus 3 | CfMV3 | 1 012 nt | 68.2 | 1.50E-29 | Rhizoctonia solani mitovirus 33(QDW65423.1) | +ssRNA | Mitoviridae | 1 |
| Contig3026 | Colletotrichum gloeosporioides mitovirus 1 | CgMV1 | 4 742 nt | 91.4 | 4.20E-236 | Colletotrichum fructicola mitovirus 1(BBN51032.1) | +ssRNA | Mitoviridae | 1 |
| Contig18460 | Colletotrichum fructicola mitovirus 1 | CfrMV1 | 2 520 nt | 94.5 | 0.00E+00 | Colletotrichum fructicola mitovirus 1(BBN51032.1) | +ssRNA | Mitoviridae | 4 |
| Contig42866 | Colletotrichum siamense mitovirus 3 | CsMV3 | 1 462 nt | 93.1 | 4.20E-211 | Colletotrichum fructicola mitovirus 1(BBN51032.1) | +ssRNA | Mitoviridae | 5 |
| Contig46248 | Colletotrichum fioriniae mitovirus 5 | CfMV4 | 1 337 nt | 89.4 | 2.90E-227 | Colletotrichum fructicola mitovirus 1(BBN51032.1) | +ssRNA | Mitoviridae | 5 |
| Contig46871 | Colletotrichum gloeosporioides mitovirus 2 | CgMV2 | 1 313 nt | 92.4 | 1.30E-75 | Colletotrichum fructicola mitovirus 1(BBN51032.1) | +ssRNA | Mitoviridae | 5 |
| Contig50030 | Colletotrichum nymphaeae mitovirus 1 | CnMV1 | 1 197 nt | 94.9 | 3.90E-66 | Colletotrichum fructicola mitovirus 1(BBN51032.1) | +ssRNA | Mitoviridae | 5 |
| Contig23629 | Colletotrichum fioriniae mitovirus 6 | CfMV5 | 2 393 nt | 67.2 | 2.30E-259 | Colletotrichum falcatum mitovirus 1(AZT88621.1) | +ssRNA | Mitoviridae | 1 |
| Contig21451 | Colletotrichum siamense mitovirus 2 | CsMV2 | 2 539 nt | 65.2 | 1.20E-258 | Sclerotinia sclerotiorum mitovirus 32(AWY10990.1) | +ssRNA | Mitoviridae | 2 |
| Contig21504 | Colletotrichum siamense mitovirus 1 | CsMV1 | 2 536 nt | 65.5 | 1.5E-208 | Sclerotinia sclerotiorum mitovirus 6(QGY72611.1) | +ssRNA | Mitoviridae | 1 |
表2 测序得到的病毒序列信息
Table 2 Sequence information of sequenced viruses
| Contig 编号 Contig code | 病毒名称 Name of putative viruses | 病毒名称缩写 Name abbreviation | 长度 Length | 序列相似度 Acid similarity/% | E值 E-value | 最佳匹配 Best match | 基因组类型 Genome type | 病毒科 Family | Contig 计数 Contig counts |
|---|---|---|---|---|---|---|---|---|---|
| Contig 22546 | Colletotrichum fioriniae alternavirus 1 | CfAV1 dsRNA2 | 2 463 bp | 70.4 | 3.50E-303 | Fusarium graminearum alternavirus 1(YP_009449446.1) | dsRNA | Alternaviridae | 1 |
| Contig11036 | Colletotrichum fioriniae alternavirus 1 | CfAV1 dsRNA1 | 3 534 bp | 73.3 | 0.00E+00 | Fusarium graminearum alternavirus 1(YP_009449439.1) | dsRNA | Alternaviridae | 1 |
| Contig22686 | Colletotrichum fioriniae alternavirus 1 | CfAV1 dsRNA3 | 2 454 bp | 72 | 3.00E-307 | Fusarium graminearum alternavirus 1(AUI80777.1) | dsRNA | Alternaviridae | 1 |
| Contig51809 | Colletotrichum fioriniae narnavirus 1 | CfNV1 | 1 136 nt | 59.28 | 3.00E-76 | Sclerotinia sclerotiorum narnavirus 2(QZE12028.1) | +ssRNA | Narnaviridae | 1 |
| Contig26097 | Colletotrichum siamense narnavirus 1 | CsNV1 | 2 239 nt | 65.3 | 0.00E+00 | Sclerotinia sclerotiorum narnavirus 2(QZE12027.1) | +ssRNA | Narnaviridae | 1 |
| Contig20122 | Colletotrichum siamense narnavirus 2 | CsNV2 | 2 420 nt | 33.2 | 8.00E-88 | Oidiodendron maius splipalmivirus 1(QNN89180.1) | +ssRNA | Narnaviridae | 1 |
| Contig11676 | Colletotrichum nymphaeae narnavirus 1 | CnNV1 | 3 090 nt | 65 | 0.00E+00 | Botryosphaeria dothidea narnavirus 3(QQD86177.1) | +ssRNA | Narnaviridae | 5 |
| Contig10410 | Colletotrichum fioriniae narnavirus 2 | CfNV2 | 3 619 nt | 55.3 | 0.00E+00 | Erysiphe necator associated narnavirus 15(QJT93747.1) | +ssRNA | Narnaviridae | 1 |
| Contig19559 | Colletotrichum nymphaeae narnavirus 2 | CnNV2 | 2 455 nt | 62.2 | 1.00E-278 | Erysiphe necator associated narnavirus 4(QHD64827.1) | +ssRNA | Narnaviridae | 2 |
| Contig4718 | Colletotrichum gloeosporioides narnavirus 1 | CgNV1 | 4 200 nt | 51.8 | 1.60E-300 | Monilinia narnavirus H (QED42934.1) | +ssRNA | Narnaviridae | 3 |
| Contig35776 | Colletotrichum fructicola narnavirus 1 | CfrNV1 | 1 689 nt | 60 | 1.00E-27 | Aspergillus lentulus narnavirus 1(BCH36644.1) | +ssRNA | Narnaviridae | 2 |
| Contig19474 | Colletotrichum fructicola botourmiavirus 1 | CfBV1 | 2 460 nt | 55.1 | 1.60E-178 | Pestalotiopsis botourmiavirus 3(QTH80197.1) | +ssRNA | Botourmiaviridae | 2 |
| Contig22684 | Colletotrichum fioriniae mitovirus 1 | CfMV1 | 2 455 nt | 51.7 | 4.00E-158 | Grapevine-associated mitovirus 10(QXN75363.1) | +ssRNA | Mitoviridae | 2 |
| Contig41113 | Colletotrichum fioriniae mitovirus 2 | CfMV2 | 1 493 nt | 78 | 9.10E-113 | Rhizoctonia solani mitovirus 33(QDW65423.1) | +ssRNA | Mitoviridae | 1 |
| Contig55502 | Colletotrichum fioriniae mitovirus 3 | CfMV3 | 1 012 nt | 68.2 | 1.50E-29 | Rhizoctonia solani mitovirus 33(QDW65423.1) | +ssRNA | Mitoviridae | 1 |
| Contig3026 | Colletotrichum gloeosporioides mitovirus 1 | CgMV1 | 4 742 nt | 91.4 | 4.20E-236 | Colletotrichum fructicola mitovirus 1(BBN51032.1) | +ssRNA | Mitoviridae | 1 |
| Contig18460 | Colletotrichum fructicola mitovirus 1 | CfrMV1 | 2 520 nt | 94.5 | 0.00E+00 | Colletotrichum fructicola mitovirus 1(BBN51032.1) | +ssRNA | Mitoviridae | 4 |
| Contig42866 | Colletotrichum siamense mitovirus 3 | CsMV3 | 1 462 nt | 93.1 | 4.20E-211 | Colletotrichum fructicola mitovirus 1(BBN51032.1) | +ssRNA | Mitoviridae | 5 |
| Contig46248 | Colletotrichum fioriniae mitovirus 5 | CfMV4 | 1 337 nt | 89.4 | 2.90E-227 | Colletotrichum fructicola mitovirus 1(BBN51032.1) | +ssRNA | Mitoviridae | 5 |
| Contig46871 | Colletotrichum gloeosporioides mitovirus 2 | CgMV2 | 1 313 nt | 92.4 | 1.30E-75 | Colletotrichum fructicola mitovirus 1(BBN51032.1) | +ssRNA | Mitoviridae | 5 |
| Contig50030 | Colletotrichum nymphaeae mitovirus 1 | CnMV1 | 1 197 nt | 94.9 | 3.90E-66 | Colletotrichum fructicola mitovirus 1(BBN51032.1) | +ssRNA | Mitoviridae | 5 |
| Contig23629 | Colletotrichum fioriniae mitovirus 6 | CfMV5 | 2 393 nt | 67.2 | 2.30E-259 | Colletotrichum falcatum mitovirus 1(AZT88621.1) | +ssRNA | Mitoviridae | 1 |
| Contig21451 | Colletotrichum siamense mitovirus 2 | CsMV2 | 2 539 nt | 65.2 | 1.20E-258 | Sclerotinia sclerotiorum mitovirus 32(AWY10990.1) | +ssRNA | Mitoviridae | 2 |
| Contig21504 | Colletotrichum siamense mitovirus 1 | CsMV1 | 2 536 nt | 65.5 | 1.5E-208 | Sclerotinia sclerotiorum mitovirus 6(QGY72611.1) | +ssRNA | Mitoviridae | 1 |
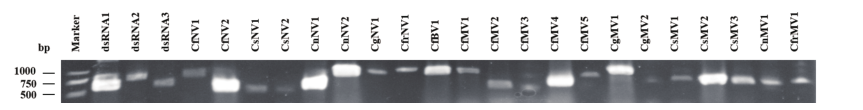
图1 病毒RNA RT-PCR扩增产物的琼脂糖凝胶电泳图 Marker为2 000 bp的DNA marker,产自大连的宝生物公司
Fig. 1 Agarose gel electrophoresis of the RT-PCR amplified products of viral RNAs M: 2 000 bp DNA marker(TaKaRa, Dalian, China)
| 菌株Strain | 种名Species | 携带病毒名称Contained mycoviruses |
|---|---|---|
| BK1-11 | C. nymphaeae | CfMV4,CnNV2,CsNV1, CfAV1,CfMV1,CfMV2,CnMV1 |
| BK2-1 | C. fructicola | CfMV4,CsNV1,CfBV1,CfMV1,CfMV2 |
| BK2-3 | C. fioriniae | CfMV2,CfMV3,CfMV4,CfMV5,CfNV1,CfMV1 |
| BK3-5 | C. fioriniae | CfMV2,CfMV3,CfNV2,CfBV1 |
| BK4-4 | C. nymphaeae | CfMV2,CsMV3,CgMV2,CfAV1,CnNV1,CfMV1 |
| CX1 | C. fioriniae | CfMV2 |
| CX3 | C. fioriniae | CfMV2,CfMV4,CnNV1,CfMV1 |
| CX6 | C. fructicola | CfrMV1,CfMV4,CsMV2,CnNV1,CfBV1,CfMV1,CsNV1 |
| CX7 | C. fioriniae | CfAV1,CfMV1,CfMV3,CfMV2 |
| DJ1-1 | C. gloeosporioides | CfMV2,CfMV3 |
| DJ2-1 | C. gloeosporioide | CfrMV1,CgNV1,CsNV1,CnNV1,CfBV1,CfMV1,CfMV3,CfMV2 |
| DJ2-4 | C. gloeosporioide | CgMV1,CfrMV1,CfMV4,CfMV1,CfMV3,CfMV2,CfNV1 |
| DJ3-5 | C. siamense | CsMV3,CfMV4,CfMV5,CsMV2,CsNV1,CfNV1,CfMV1,CfMV3,CfMV2 |
| FX1-3 | C. fructicola | CsMV3,CfMV4,CfMV5,CsMV2,CfMV3,CfNV2,CsNV1,CfNV1,CfAV1,CfMV1,CfMV2 |
| SZ2-3 | C. gloeosporioides | CgMV2,CsMV2,CgNV1,CsNV1,CfNV1,CnNV1,CfMV1,CfMV2 |
| XS5 | C. nymphaeae | CsMV3,CfMV1,CfrMV1,CsMV1,CfrNV1,CfMV2 |
| XS6 | C. fructicola | CsMV3,CfNV2,CfrNV1,CsNV1,CnNV1,CfMV1,CfMV2 |
| YX1-5 | C. siamense | CfMV2,CgMV1,CfrMV1,CfMV4,CfMV5,CsMV2,CfNV1,CnNV1,CfBV1,CfMV1 |
| YX1-7 | C. siamense | CfMV2,CfrMV1,CfMV4,CfrNV1,CsNV2,CsNV1,CfAV1,CnNV1,CfMV1,CfBV1 |
| YY1-11 | C. gloeosporioides | CfMV3,CfMV2,CfrMV1,CfMV4,CgMV2,CfMV5,CsMV1,CnNV2,CsNV1,CnNV1,CfMV1 |
| YY1-28 | C. fructicola | CfrMV1,CfMV1,CfMV3,CfMV2 |
| ZS3-8 | C. nymphaeae | CfrNV1,CfMV1,CfMV2 |
| ZS2-6 | C. nymphaeae | CgMV2,CsMV2,CnNV2,CfrNV1,CsNV1,CfMV2 |
| ZS2-9 | C. siamense | CgMV1,CfMV4,CsMV2,CsMV1,CnNV2,CsNV2,CfAV1,CnNV1,CfBV1,CfMV1 |
| ZS3-13 | C. nymphaeae | CsNV1,CfMV2 |
表3 22种病毒在核桃炭疽菌株中的具体定位信息列表
Table 3 List of identification results of 22 mycoviruses in Colletotrichum spp. strains
| 菌株Strain | 种名Species | 携带病毒名称Contained mycoviruses |
|---|---|---|
| BK1-11 | C. nymphaeae | CfMV4,CnNV2,CsNV1, CfAV1,CfMV1,CfMV2,CnMV1 |
| BK2-1 | C. fructicola | CfMV4,CsNV1,CfBV1,CfMV1,CfMV2 |
| BK2-3 | C. fioriniae | CfMV2,CfMV3,CfMV4,CfMV5,CfNV1,CfMV1 |
| BK3-5 | C. fioriniae | CfMV2,CfMV3,CfNV2,CfBV1 |
| BK4-4 | C. nymphaeae | CfMV2,CsMV3,CgMV2,CfAV1,CnNV1,CfMV1 |
| CX1 | C. fioriniae | CfMV2 |
| CX3 | C. fioriniae | CfMV2,CfMV4,CnNV1,CfMV1 |
| CX6 | C. fructicola | CfrMV1,CfMV4,CsMV2,CnNV1,CfBV1,CfMV1,CsNV1 |
| CX7 | C. fioriniae | CfAV1,CfMV1,CfMV3,CfMV2 |
| DJ1-1 | C. gloeosporioides | CfMV2,CfMV3 |
| DJ2-1 | C. gloeosporioide | CfrMV1,CgNV1,CsNV1,CnNV1,CfBV1,CfMV1,CfMV3,CfMV2 |
| DJ2-4 | C. gloeosporioide | CgMV1,CfrMV1,CfMV4,CfMV1,CfMV3,CfMV2,CfNV1 |
| DJ3-5 | C. siamense | CsMV3,CfMV4,CfMV5,CsMV2,CsNV1,CfNV1,CfMV1,CfMV3,CfMV2 |
| FX1-3 | C. fructicola | CsMV3,CfMV4,CfMV5,CsMV2,CfMV3,CfNV2,CsNV1,CfNV1,CfAV1,CfMV1,CfMV2 |
| SZ2-3 | C. gloeosporioides | CgMV2,CsMV2,CgNV1,CsNV1,CfNV1,CnNV1,CfMV1,CfMV2 |
| XS5 | C. nymphaeae | CsMV3,CfMV1,CfrMV1,CsMV1,CfrNV1,CfMV2 |
| XS6 | C. fructicola | CsMV3,CfNV2,CfrNV1,CsNV1,CnNV1,CfMV1,CfMV2 |
| YX1-5 | C. siamense | CfMV2,CgMV1,CfrMV1,CfMV4,CfMV5,CsMV2,CfNV1,CnNV1,CfBV1,CfMV1 |
| YX1-7 | C. siamense | CfMV2,CfrMV1,CfMV4,CfrNV1,CsNV2,CsNV1,CfAV1,CnNV1,CfMV1,CfBV1 |
| YY1-11 | C. gloeosporioides | CfMV3,CfMV2,CfrMV1,CfMV4,CgMV2,CfMV5,CsMV1,CnNV2,CsNV1,CnNV1,CfMV1 |
| YY1-28 | C. fructicola | CfrMV1,CfMV1,CfMV3,CfMV2 |
| ZS3-8 | C. nymphaeae | CfrNV1,CfMV1,CfMV2 |
| ZS2-6 | C. nymphaeae | CgMV2,CsMV2,CnNV2,CfrNV1,CsNV1,CfMV2 |
| ZS2-9 | C. siamense | CgMV1,CfMV4,CsMV2,CsMV1,CnNV2,CsNV2,CfAV1,CnNV1,CfBV1,CfMV1 |
| ZS3-13 | C. nymphaeae | CsNV1,CfMV2 |
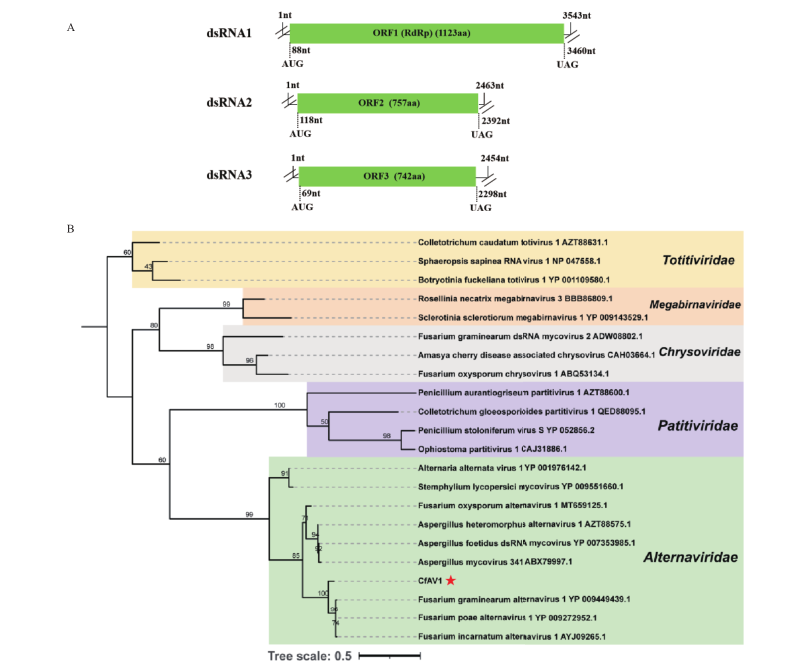
图2 CfAV1的3条dsRNA片段的基因组结构图和 Alternaviridae 及相关病毒的系统发育树 A:病毒CfAV1的3条dsRNA片段的基因组结构图;B:基于CfAV1和其相关病毒的RdRp序列,采用最大似然法构建的系统发育树。CfAV1 用红色五角星标示
Fig. 2 Genome organization of 3 dsRNA segments in CfAV1and phylogenetic analysis of viruses in family Alternaviridae and other related viruses A: Genome organization of 3 dsRNA segments in CfAV1. B: A maximum likelihood phylogenetic tree was constructed based on the RdRp sequence of CfAV1 and other related viruses. CfAV1is marked with red star
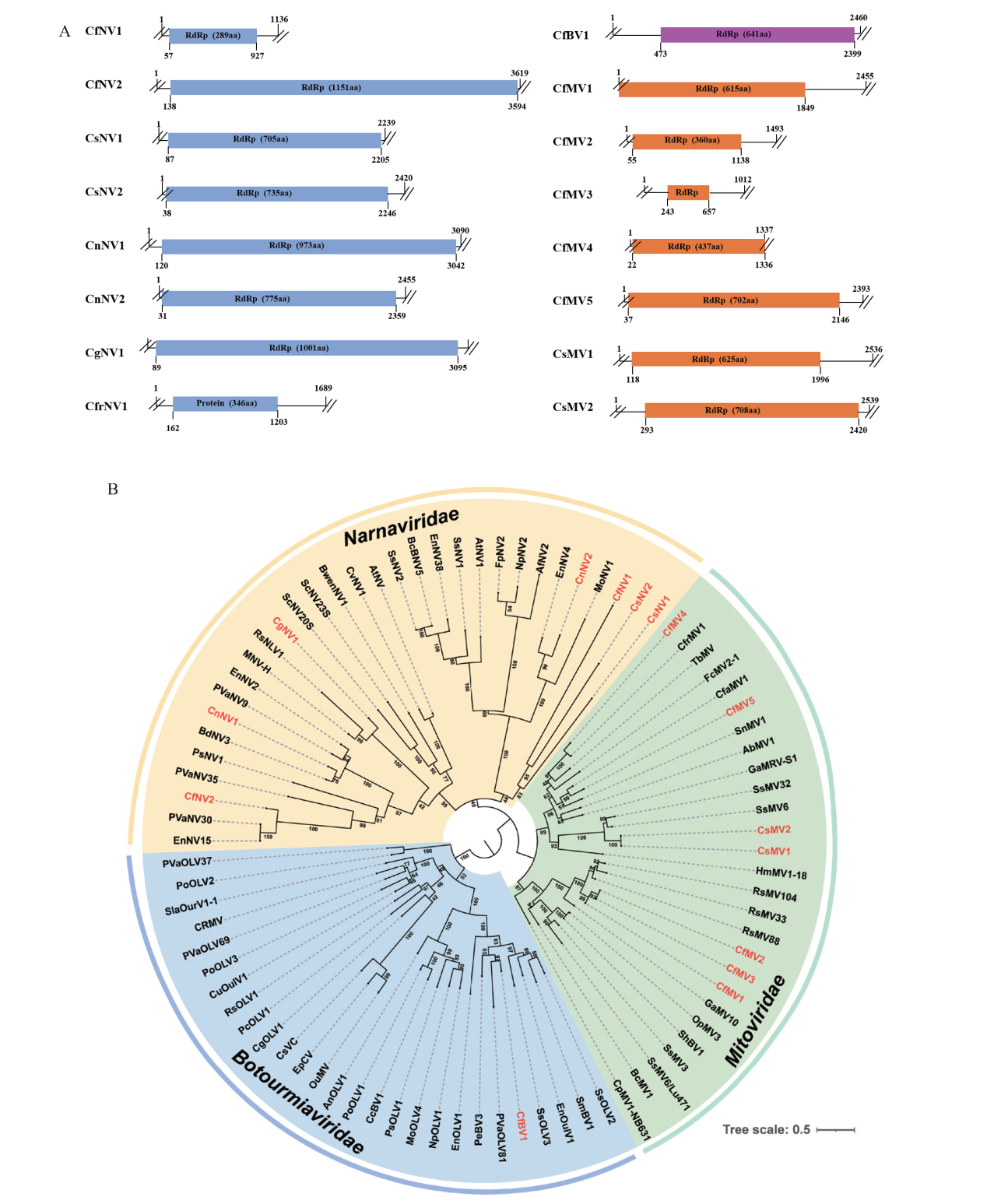
图3 隶属于裸露病毒科、线粒体病毒科和葡萄孢欧尔密病毒科的病毒基因组结构图和其系统发育树 A:隶属于裸露病毒科、线粒体病毒科和葡萄孢欧尔密病毒科的新病毒的基因组结构图;B:基于病毒的RdRp序列,采用最大似然法构建系统发育树。每个分支旁边的数字代表自展法进行 1 000 次重复的支持率。本研究鉴定出的病毒采用其缩写名称显示
Fig. 3 Genome organization and phylogenetic analysis of viruses in family Narnaviridae, Mitoviridae, and Botourmiaviridae A: Genome organization of novel viruses in family Narnaviridae, Mitoviridae, and Botourmiaviridae. B: A maximum likelihood phylogenetic tree constructed based on the amino acid sequences of viral RdRp. The numbers next to each branch refers to the bootstrap support based on 1 000 replicates. Viruses identified in this study are denoted by the abbreviated names
| [1] |
Ghabrial SA, Suzuki N. Viruses of plant pathogenic fungi[J]. Annu Rev Phytopathol, 2009, 47: 353-384.
doi: 10.1146/annurev-phyto-080508-081932 pmid: 19400634 |
| [2] |
Xie JT, Jiang DH. New insights into mycoviruses and exploration for the biological control of crop fungal diseases[J]. Annu Rev Phytopathol, 2014, 52: 45-68.
doi: 10.1146/annurev-phyto-102313-050222 pmid: 25001452 |
| [3] | Dolja VV, Koonin EV. Capsid-less RNA viruses[M]. In:eLS ed.ed. Chichester: John Wiley & Sons, 2012. |
| [4] |
Son M, Yu J, Kim KH. Five questions about mycoviruses[J]. PLoS Pathog, 2015, 11(11): e1005172.
doi: 10.1371/journal.ppat.1005172 URL |
| [5] |
Nuss DL. Hypovirulence: mycoviruses at the fungal-plant interface[J]. Nat Rev Microbiol, 2005, 3(8): 632-642.
pmid: 16064055 |
| [6] |
Anagnostakis SL. Biological control of chestnut blight[J]. Science, 1982, 215(4532): 466-471.
pmid: 17771259 |
| [7] |
Yu X, Li B, Fu Y, et al. A geminivirus-related DNA mycovirus that confers hypovirulence to a plant pathogenic fungus[J]. PNAS, 2010, 107(18): 8387-8392.
doi: 10.1073/pnas.0913535107 pmid: 20404139 |
| [8] |
Yu X, Li B, Fu Y, et al. Extracellular transmission of a DNA mycovirus and its use as a natural fungicide[J]. PNAS, 2013, 110(4): 1452-1457.
doi: 10.1073/pnas.1213755110 pmid: 23297222 |
| [9] |
Dean R, Kan JAV, Pretorius ZA, et al. The Top 10 fungal pathogens in molecular plant pathology[J]. Mol Plant Pathol, 2012, 13(4): 414-430.
doi: 10.1111/j.1364-3703.2011.00783.x pmid: 22471698 |
| [10] |
Zhong J, Chen D, Lei XH, et al. Detection and characterization of a novel Gammapartitivirus in the phytopathogenic fungus Colletotrichum acutatum strain HNZJ001[J]. Virus Res, 2014, 190: 104-109.
doi: 10.1016/j.virusres.2014.05.028 pmid: 25008759 |
| [11] |
Wang Y, Liu S, Zhu HJ, et al. Molecular characterization of a novel mycovirus from the plant pathogenic fungus Colletotrichum gloeosporioides[J]. Arch Virol, 2019, 164(11): 2859-2863.
doi: 10.1007/s00705-019-04354-2 |
| [12] |
Marzano SYL, Nelson BD, Ajayi-Oyetunde O, et al. Identification of diverse mycoviruses through metatranscriptomics characterization of the viromes of five major fungal plant pathogens[J]. J Virol, 2016, 90(15): 6846-6863.
doi: 10.1128/JVI.00357-16 URL |
| [13] |
Campo S, Gilbert KB, Carrington JC. Small RNA-based antiviral defense in the phytopathogenic fungus Colletotrichum higginsianum[J]. PLoS Pathog, 2016, 12(6): e1005640.
doi: 10.1371/journal.ppat.1005640 URL |
| [14] |
Zhong J, Pang XD, Zhu HJ, et al. Molecular characterization of a trisegmented mycovirus from the plant pathogenic fungus Colletotrichum gloeosporioides[J]. Viruses, 2016, 8(10): 268.
doi: 10.3390/v8100268 URL |
| [15] |
Zhai LF, Zhang MX, Hong N, et al. Identification and characterization of a novel hepta-segmented dsRNA virus from the phytopathogenic fungus Colletotrichum fructicola[J]. Front Microbiol, 2018, 9: 754.
doi: 10.3389/fmicb.2018.00754 URL |
| [16] |
Jia HX, Dong KL, Zhou LL, et al. A dsRNA virus with filamentous viral particles[J]. Nat Commun, 2017, 8(1): 168.
doi: 10.1038/s41467-017-00237-9 pmid: 28761042 |
| [17] |
Guo J, Zhu JZ, Zhou XY, et al. A novel ourmia-like mycovirus isolated from the plant pathogenic fungus Colletotrichum gloeosporioi-des[J]. Arch Virol, 2019, 164(10): 2631-2635.
doi: 10.1007/s00705-019-04346-2 |
| [18] |
Xu XW, Hai D, Li JC, et al. Molecular characterization of a novel penoulivirus from the phytopathogenic fungus Colletotrichum camelliae[J]. Arch Virol, 2022, 167(2): 641-644.
doi: 10.1007/s00705-021-05334-1 |
| [19] |
Liu H, Wang H, Lu X, et al. Molecular characterization of a novel single-stranded RNA virus, ChRV1, isolated from the plant pathogenic fungus Colletotrichum Higginsianum[J]. Archives of Virology, 2021, 166(6):1805-1809.
doi: 10.1007/s00705-021-05071-5 |
| [20] |
Wang H, Liu H, Zhou Q. The complete genome sequence of a new Mitovirus from the phytopathogenic fungus Colletotrichum higginsianum[J]. Arch Virol, 2021, 166(5): 1481-1484.
doi: 10.1007/s00705-021-04996-1 pmid: 33616726 |
| [21] |
Gilbert KB, Holcomb EE, Allscheid RL, et al. Hiding in plain sight: new virus genomes discovered via a systematic analysis of fungal public transcriptomes[J]. PLoS One, 2019, 14(7): e0219207.
doi: 10.1371/journal.pone.0219207 URL |
| [22] | Lima JS, Figueiredo JG, Gomes RG, et al. Genetic diversity of Colletotrichum spp. an endophytic fungi in a medicinal plant, Brazilian pepper tree[J]. ISRN Microbiol, 2012, 2012: 215716. |
| [23] |
Figueiredo LC, Figueiredo GS, Giancoli AC, et al. Detection of isometric, dsRNA-containing viral particles in Colletotrichum gloeosporioides isolated from cashew tree[J]. Tropical Plant Pathology, 2012, 37(2): 142-145.
doi: 10.1590/S1982-56762012000200007 URL |
| [24] |
Zhang HX, Xie JT, Fu YP, et al. A 2-kb mycovirus converts a pathogenic fungus into a beneficial endophyte for Brassica protection and yield enhancement[J]. Mol Plant, 2020, 13(10): 1420-1433.
doi: 10.1016/j.molp.2020.08.016 URL |
| [25] |
Bolger AM, Lohse M, Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data[J]. Bioinformatics, 2014, 30(15): 2114-2120.
doi: 10.1093/bioinformatics/btu170 pmid: 24695404 |
| [26] |
Kim D, Langmead B, Salzberg SL. HISAT: a fast spliced aligner with low memory requirements[J]. Nat Methods, 2015, 12(4): 357-360.
doi: 10.1038/nmeth.3317 pmid: 25751142 |
| [27] |
Li H, Handsaker B, Wysoker A, et al. The sequence alignment/map format and SAMtools[J]. Bioinformatics, 2009, 25(16): 2078-2079.
doi: 10.1093/bioinformatics/btp352 pmid: 19505943 |
| [28] |
Bankevich A, Nurk S, Antipov D, et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing[J]. J Comput Biol, 2012, 19(5): 455-477.
doi: 10.1089/cmb.2012.0021 pmid: 22506599 |
| [29] |
Fu LM, Niu BF, Zhu ZW, et al. CD-HIT: accelerated for clustering the next-generation sequencing data[J]. Bioinformatics, 2012, 28(23): 3150-3152.
doi: 10.1093/bioinformatics/bts565 pmid: 23060610 |
| [30] |
Buchfink B, Xie C, Huson DH. Fast and sensitive protein alignment using DIAMOND[J]. Nat Methods, 2015, 12(1): 59-60.
doi: 10.1038/nmeth.3176 pmid: 25402007 |
| [31] |
Nakamura T, Yamada KD, Tomii K, et al. Parallelization of MAFFT for large-scale multiple sequence alignments[J]. Bioinformatics, 2018, 34(14): 2490-2492.
doi: 10.1093/bioinformatics/bty121 pmid: 29506019 |
| [32] |
Kalyaanamoorthy S, Minh BQ, Wong TKF, et al. ModelFinder: fast model selection for accurate phylogenetic estimates[J]. Nat Methods, 2017, 14(6): 587-589.
doi: 10.1038/nmeth.4285 pmid: 28481363 |
| [33] |
Nguyen LT, Schmidt HA, von Haeseler A, et al. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies[J]. Mol Biol Evol, 2015, 32(1): 268-274.
doi: 10.1093/molbev/msu300 URL |
| [34] |
Sutela S, Forgia M, Vainio EJ, et al. The virome from a collection of endomycorrhizal fungi reveals new viral taxa with unprecedented genome organization[J]. Virus Evol, 2020, 6(2): veaa076.
doi: 10.1093/ve/veaa076 URL |
| [35] |
Poimala A, Parikka P, Hantula J, et al. Viral diversity in Phytophthora cactorum population infecting strawberry[J]. Environ Microbiol, 2021, 23(9): 5200-5221.
doi: 10.1111/1462-2920.15519 pmid: 33848054 |
| [36] |
Wen CY, Wan XR, Zhang YY, et al. Molecular characterization of the first alternavirus identified in Fusarium oxysporum[J]. Viruses, 2021, 13(10): 2026.
doi: 10.3390/v13102026 URL |
| [37] |
Aoki N, Moriyama H, Kodama M, et al. A novel mycovirus associated with four double-stranded RNAs affects host fungal growth in Alternaria alternata[J]. Virus Res, 2009, 140(1-2): 179-187.
doi: 10.1016/j.virusres.2008.12.003 URL |
| [38] |
Hillman BI, Cai GH. The family Narnaviridae: simplest of RNA viruses[J]. Adv Virus Res, 2013, 86: 149-176.
doi: 10.1016/B978-0-12-394315-6.00006-4 pmid: 23498906 |
| [39] | Chiba S. Fusariviruses[M]. In: Bamford DH and Zuckerman M eds. Encyclopedia of virology. Oxford: Academic Press, 2021: 577-581. |
| [40] | Ruiz-Padilla A, Rodríguez-Romero J, Gómez-Cid I, et al. Novel mycoviruses discovered in the mycovirome of a necrotrophic fungus[J]. mBio, 2021, 12(3): e03705-e03720. |
| [41] |
Turina M, Hillman BI, Izadpanah K, et al. ICTV virus taxonomy profile: Ourmiavirus[J]. The Journal of General Virology, 2017, 98(2):129-130.
doi: 10.1099/jgv.0.000725 URL |
| [42] |
Ayllón MA, Turina M, Xie J, et al. ICTV virus taxonomy profile: Botourmiaviridae[J]. J Gen Virol, 2020, 101(5): 454-455.
doi: 10.1099/jgv.0.001409 pmid: 32375992 |
| [1] | 余洋, 刘天海, 刘理旭, 唐杰, 彭卫红, 陈阳, 谭昊. 羊肚菌菌种生产车间气溶胶微生物群落研究[J]. 生物技术通报, 2023, 39(5): 267-275. |
| [2] | 李善家, 雷雨昕, 孙梦格, 刘海锋, 王兴敏. 种子内生细菌多样性与植物互馈作用研究进展[J]. 生物技术通报, 2023, 39(4): 166-175. |
| [3] | 郭文博, 路杨, 隋丽, 赵宇, 邹晓威, 张正坤, 李启云. 球孢白僵菌真菌病毒BbPmV-4外壳蛋白多克隆抗体制备及应用[J]. 生物技术通报, 2023, 39(10): 58-67. |
| [4] | 王子夜, 王志刚, 阎爱华. 不同树龄桑根际土壤原生生物群落组成多样性[J]. 生物技术通报, 2022, 38(8): 206-215. |
| [5] | 高小宁, 刘睿, 吴自林, 吴嘉云. 宿根矮化病抗感甘蔗品种茎部内生真菌和细菌群落特征分析[J]. 生物技术通报, 2022, 38(6): 166-173. |
| [6] | 徐扬, 张冠初, 丁红, 秦斐斐, 张智猛, 戴良香. 土壤类型对花生根际土壤细菌群落多样性和产量的影响[J]. 生物技术通报, 2022, 38(6): 221-234. |
| [7] | 钟辉, 刘亚军, 王滨花, 和梦洁, 吴兰. 分析方法对细菌群落16S rRNA基因扩增测序分析结果的影响[J]. 生物技术通报, 2022, 38(6): 81-92. |
| [8] | 周晓楠, 徐金青, 雷雨晴, 王海庆. 基于GBS-seq的青藏扁蓿豆SNP标记开发[J]. 生物技术通报, 2022, 38(4): 303-310. |
| [9] | 谢果珍, 唐圆, 宁晓妹, 邱集慧, 谭周进. 铁皮石斛多糖对高脂饮食小鼠肠黏膜结构及菌群的影响[J]. 生物技术通报, 2022, 38(2): 150-157. |
| [10] | 刘爽, 姚佳妮, 沈聪, 代金霞. 荒漠植物柠条根际土壤nifH基因荧光定量及固氮菌多样性分析[J]. 生物技术通报, 2022, 38(12): 252-262. |
| [11] | 陈宇捷, 郑华宝, 周昕彦. 改良高通量测序技术揭示除藻剂对藻类群落的影响[J]. 生物技术通报, 2022, 38(11): 70-79. |
| [12] | 李婷婷, 邓旭辉, 李若尘, 刘红军, 沈宗专, 李荣, 沈其荣. 番茄青枯病发生对土壤真菌群落多样性的影响[J]. 生物技术通报, 2022, 38(10): 195-203. |
| [13] | 颜珲璘, 芦光新, 邓晔, 顾松松, 颜程良, 马坤, 赵阳安, 张海娟, 王英成, 周学丽, 窦声云. 高寒地区根瘤菌拌种对禾/豆混播土壤微生物群落的影响[J]. 生物技术通报, 2022, 38(10): 204-215. |
| [14] | 张田田, 李永臻, 沈国平, 王嵘, 朱德锐, 邢江娃. 高盐盐湖可分离嗜盐耐盐菌的种群多样性及四氢嘧啶产量评价[J]. 生物技术通报, 2022, 38(1): 168-178. |
| [15] | 王志山, 黎妮, 王伟平, 刘洋. 水稻种子内生细菌研究进展[J]. 生物技术通报, 2022, 38(1): 236-246. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||