








Biotechnology Bulletin ›› 2023, Vol. 39 ›› Issue (11): 191-204.doi: 10.13560/j.cnki.biotech.bull.1985.2023-0723
Previous Articles Next Articles
ZHU Ying-xuan( ), LI Ke-jing, HE Min, ZHENG Dao-qiong(
), LI Ke-jing, HE Min, ZHENG Dao-qiong( )
)
Received:2023-07-30
Online:2023-11-26
Published:2023-12-20
Contact:
ZHENG Dao-qiong
E-mail:zhuyingxuan@zju.edu.cn;zhengdaoqiong@zju.edu.cn
ZHU Ying-xuan, LI Ke-jing, HE Min, ZHENG Dao-qiong. Research Progress in the Exploring Genomic Variations Driven by Stress Factors Using the Yeast Model[J]. Biotechnology Bulletin, 2023, 39(11): 191-204.
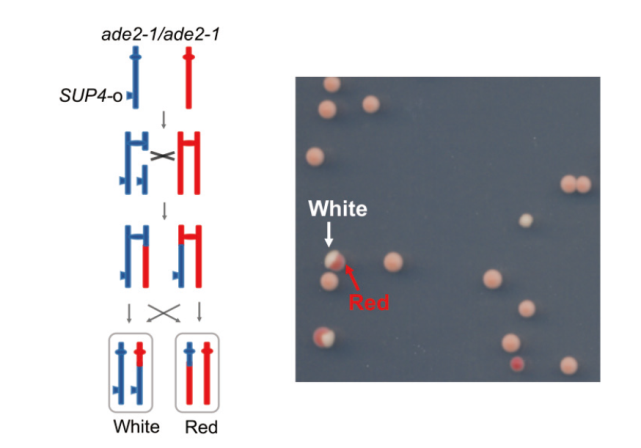
Fig. 1 Yeast white/red sectoring colony system to screen chromosomal crossover recombinants The homologous allele ade2-1 in the diploid yeast strain is ocher mutant. One copy of the SUP4-o gene can partially suppress this ochre mutation of ade2-1, making the colonies appear pink. Chromosomal crossover events can lead to one daughter cell having two copies of the SUP4-o gene, causing the colonies to appear white; another daughter cell without the SUP4-o gene would result in red colonies
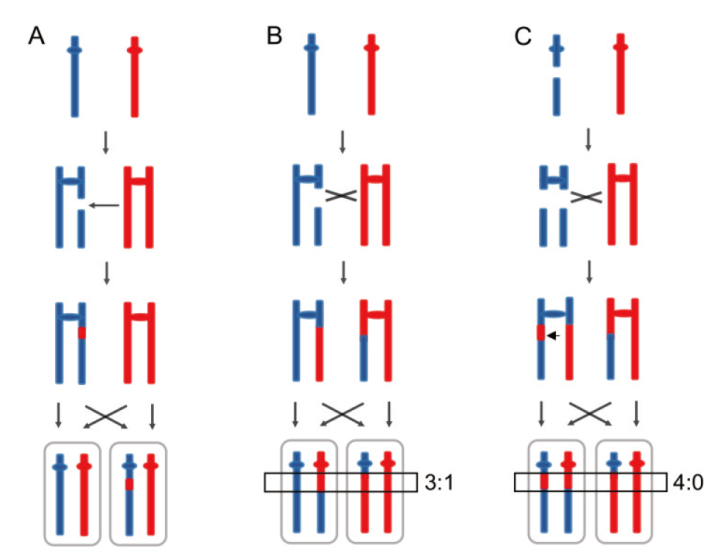
Fig. 2 LOH events induced by DNA homologous recombination in heterozygous diploid A: The interstitial LOH or gene conversion is resulted from repairing the DSB(double-strand DNA breaks)on a sister chromatid via homologous recombination. B: A 3∶1 pattern of the crossover associated with gene conversion tract(black rectangle)indicates repairing the DSB on one sister chromatid via homologous recombination. C: Chromosome breakage during the G1 phase results in DNA breaks on both sister chromosomes. A 4∶0 gene conversion district is formed at the breakpoint after this situation is repaired
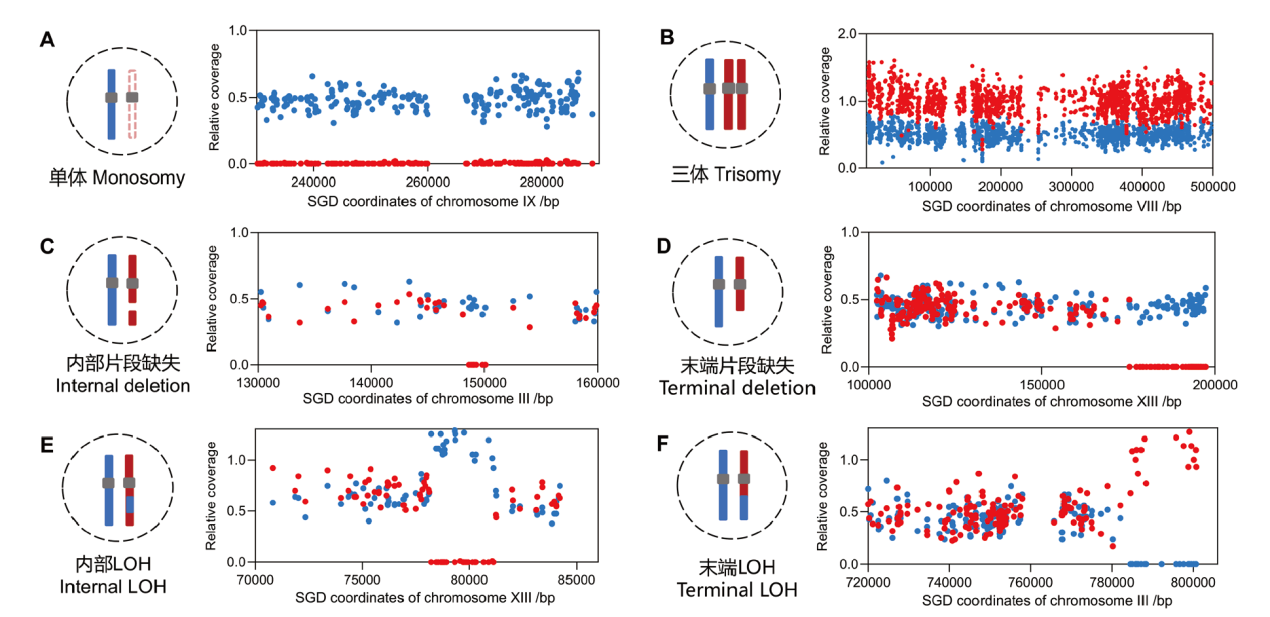
Fig. 3 Mapping genomic alterations by analyzing the relative sequencing coverages of SNP sites A: Chromosomal trisomy events. B: Chromosomal monosomy events. C: An internal deletion event in chromosome. D: A terminal deletion event in chromosome. E: Internal loss of heterozygosity(LOH)caused by gene conversion events. F: Crossover induced terminal LOH. The blue and red lines or points represent homologous chromosomes or SNP sites. The relative coverage value 0, 0.5, and 1 indicate zero, one, and two copies of DNA, respectively
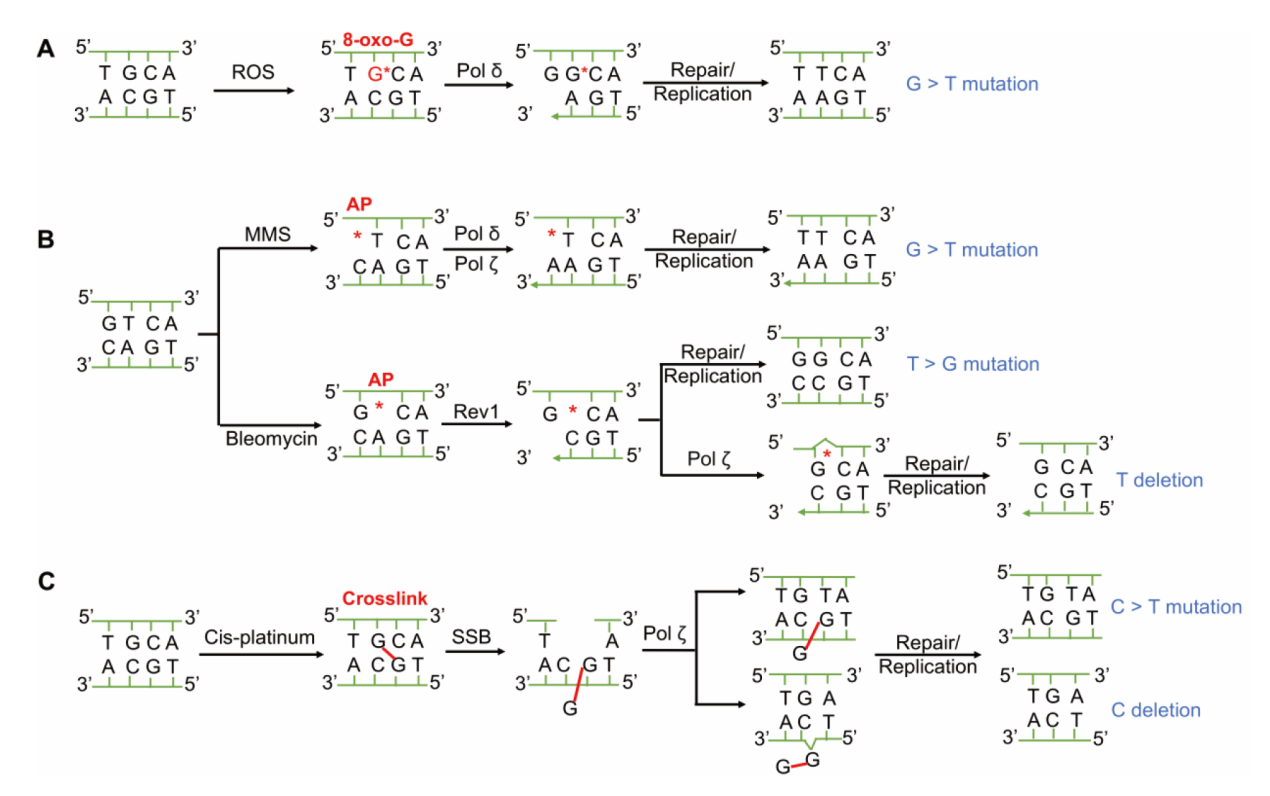
Fig. 4 Characteristic mutations and genetic mechanisms of yeast cells induced by different compounds A: Potassium bromate induces cells to produce ROS, which oxidizes base G to 8-oxo-G that prefers to pair with A, causing G > T. B: MMS and bleomycin treatment induces the generation of AP sites without base. DNA polymerase δ and Rev1 can recognize AP sites and prefer to insert A and C, respectively, causing G > T and T > G. The C inserted by Rev1 pairs with the G adjacent to the AP site when DNA polymerase ζ acts, this template slippage will cause one base T deletion. C: Cisplatin treatment induces C > T substitution or C deletion
| 表型效应 Phenotypic effect | 变异类型 Variation type | 遗传机理 Genetic mechanism | 参考文献 References |
|---|---|---|---|
| H2O2耐受性提高 | Ⅳ号染色体扩增 | 编码硫氧还蛋白过氧化物酶的TSP2基因拷贝数上升 | [ |
| VII号染色体上572-734 kb区域扩增 | 编码过氧化氢酶T的 CTT1基因拷贝数上升 | [ | |
| 热胁迫耐受性提高 | III号染色体扩增 | 该染色体上至少14个与耐热相关基因表达量上调 | [ |
| 博来霉素耐受性提高 | SKY1突变, XIII号染色体右臂缺失 | SKY1基因编码蛋白激酶,其缺失可抑制膜转运蛋白活性,减少药物进入细胞 | [ |
| 乙醇耐受性提高 | III号染色体扩增 | 未知 | [ |
| 对亚硫酸盐环境耐受性提高 | VIII和XVI号染色体易位,XV和XVI号染色体易位 | 染色体易位引起SSU1基因表达量提高,SSU1基因编码可泵出亚硫酸盐的膜蛋白 | [ |
| 氮限制环境耐受性提高 | 含有GAP1基因区域形成eccDNA | GAP1基因编码氨基酸透性酶 | [ |
| 尿囊素利用能力提高 | DAL4基因所在区域扩增 | DAL4基因编码尿囊素通透酶 | [ |
| 尿素利用能力提高 | DUR3基因所在区域扩增 | DUR3基因编码尿素通透酶 | [ |
| 冻融耐受性增强 | AQY2基因所在区域扩增 | AQY2基因介导水分子跨膜运输通道 | [ |
| 氟康唑耐受性增强 | ERG11基因所在区域扩增 | ERG11基因编码羊毛甾醇14α-去甲基化酶,可与氟康唑结合发挥耐药性作用 | [ |
| 雷帕霉素耐受性提高 | XII号染色体扩增 | 维持核糖体DNA拷贝数 | [ |
| NaCl耐受性提高 | Gcn20基因所在区域缺失 | 未知 | [ |
| 5-氟胞嘧啶耐受性提高 | FCY2基因所在区域扩增缺失 | FCY2基因编码胞嘧啶通透酶 | [ |
| 葡萄糖限制环境耐受性提高 | IV号染色体扩增 | 己糖转运蛋白编码基因HXT6和HXT7扩增 | [ |
Table 1 Examples of evolutionary adaptation induced by genomic alternations in yeast
| 表型效应 Phenotypic effect | 变异类型 Variation type | 遗传机理 Genetic mechanism | 参考文献 References |
|---|---|---|---|
| H2O2耐受性提高 | Ⅳ号染色体扩增 | 编码硫氧还蛋白过氧化物酶的TSP2基因拷贝数上升 | [ |
| VII号染色体上572-734 kb区域扩增 | 编码过氧化氢酶T的 CTT1基因拷贝数上升 | [ | |
| 热胁迫耐受性提高 | III号染色体扩增 | 该染色体上至少14个与耐热相关基因表达量上调 | [ |
| 博来霉素耐受性提高 | SKY1突变, XIII号染色体右臂缺失 | SKY1基因编码蛋白激酶,其缺失可抑制膜转运蛋白活性,减少药物进入细胞 | [ |
| 乙醇耐受性提高 | III号染色体扩增 | 未知 | [ |
| 对亚硫酸盐环境耐受性提高 | VIII和XVI号染色体易位,XV和XVI号染色体易位 | 染色体易位引起SSU1基因表达量提高,SSU1基因编码可泵出亚硫酸盐的膜蛋白 | [ |
| 氮限制环境耐受性提高 | 含有GAP1基因区域形成eccDNA | GAP1基因编码氨基酸透性酶 | [ |
| 尿囊素利用能力提高 | DAL4基因所在区域扩增 | DAL4基因编码尿囊素通透酶 | [ |
| 尿素利用能力提高 | DUR3基因所在区域扩增 | DUR3基因编码尿素通透酶 | [ |
| 冻融耐受性增强 | AQY2基因所在区域扩增 | AQY2基因介导水分子跨膜运输通道 | [ |
| 氟康唑耐受性增强 | ERG11基因所在区域扩增 | ERG11基因编码羊毛甾醇14α-去甲基化酶,可与氟康唑结合发挥耐药性作用 | [ |
| 雷帕霉素耐受性提高 | XII号染色体扩增 | 维持核糖体DNA拷贝数 | [ |
| NaCl耐受性提高 | Gcn20基因所在区域缺失 | 未知 | [ |
| 5-氟胞嘧啶耐受性提高 | FCY2基因所在区域扩增缺失 | FCY2基因编码胞嘧啶通透酶 | [ |
| 葡萄糖限制环境耐受性提高 | IV号染色体扩增 | 己糖转运蛋白编码基因HXT6和HXT7扩增 | [ |
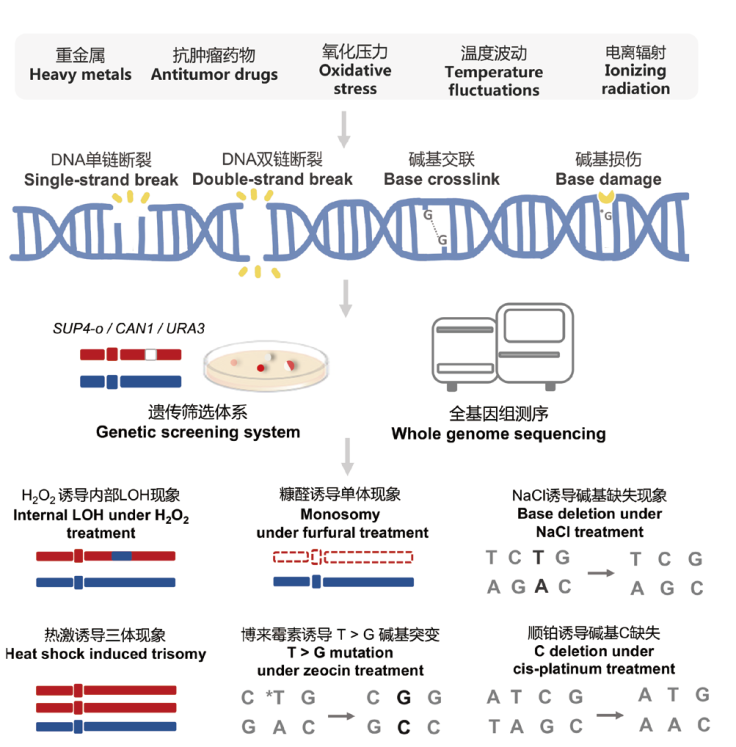
Fig. 5 Different environmental stressors result in differentiated and characteristic variations in genomes Various extracellular stressors can cause DNA damage, including DNA breaks, base damage, and cross-linking, thereby inducing different types of variations. Genetic screening systems and whole-genome sequencing are crucial technical methods for detecting environmental factors induced genomic alterations in yeast
| [1] |
Jensen RB, Rothenberg E. Preserving genome integrity in human cells via DNA double-strand break repair[J]. Mol Biol Cell, 2020, 31(9): 859-865.
doi: 10.1091/mbc.E18-10-0668 pmid: 32286930 |
| [2] |
Behringer MG, Hall DW. The repeatability of genome-wide mutation rate and spectrum estimates[J]. Curr Genet, 2016, 62(3): 507-512.
doi: 10.1007/s00294-016-0573-7 pmid: 26919990 |
| [3] |
Waterman DP, Haber JE, Smolka MB. Checkpoint responses to DNA double-strand breaks[J]. Annu Rev Biochem, 2020, 89: 103-133.
doi: 10.1146/annurev-biochem-011520-104722 pmid: 32176524 |
| [4] | Huang RX, Zhou PK. DNA damage repair: historical perspectives, mechanistic pathways and clinical translation for targeted cancer therapy[J]. Signal Transduct Target Ther, 2021, 6(1): 254. |
| [5] |
Jorgensen I, Rayamajhi M, Miao EA. Programmed cell death as a defence against infection[J]. Nat Rev Immunol, 2017, 17(3): 151-164.
doi: 10.1038/nri.2016.147 pmid: 28138137 |
| [6] |
Venkataram S, Dunn B, Li YP, et al. Development of a comprehensive genotype-to-fitness map of adaptation-driving mutations in yeast[J]. Cell, 2016, 166(6): 1585-1596.e22.
doi: S0092-8674(16)31010-8 pmid: 27594428 |
| [7] |
Kciuk M, Marciniak B, Mojzych M, et al. Focus on UV-induced DNA damage and repair-disease relevance and protective strategies[J]. Int J Mol Sci, 2020, 21(19): 7264.
doi: 10.3390/ijms21197264 URL |
| [8] |
Martin H, Shales M, Fernandez-Piñar P, et al. Differential genetic interactions of yeast stress response MAPK pathways[J]. Mol Syst Biol, 2015, 11(4): 800.
doi: 10.15252/msb.20145606 pmid: 25888283 |
| [9] |
Coelho M C, Pinto R M, Murray A W. Heterozygous mutations cause genetic instability in a yeast model of cancer evolution[J]. Nature, 2019, 566(7743): 275-278.
doi: 10.1038/s41586-019-0887-y |
| [10] |
Beaupere C, Dinatto L, Wasko BM, et al. Genetic screen identifies adaptive aneuploidy as a key mediator of ER stress resistance in yeast[J]. Proc Natl Acad Sci USA, 2018, 115(38): 9586-9591.
doi: 10.1073/pnas.1804264115 pmid: 30185560 |
| [11] |
Puddu F, Herzog M, Selivanova A, et al. Genome architecture and stability in the Saccharomyces cerevisiae knockout collection[J]. Nature, 2019, 573(7774): 416-420.
doi: 10.1038/s41586-019-1549-9 |
| [12] |
Ko N, Nishihama R, Pringle JR. Control of 5-FOA and 5-FU resistance by Saccharomyces cerevisiae YJL055W[J]. Yeast, 2008, 25(2): 155-160.
doi: 10.1002/yea.v25:2 URL |
| [13] |
Ahmad M, Bussey H. Yeast arginine permease: nucleotide sequence of the CAN1 gene[J]. Curr Genet, 1986, 10(8): 587-592.
pmid: 3327612 |
| [14] |
Ekwall K, Ruusala T. Budding yeast CAN1 gene as a selection marker in fission yeast[J]. Nucleic Acids Res, 1991, 19(5): 1150.
pmid: 2020549 |
| [15] |
Lang GI, Murray AW. Estimating the per-base-pair mutation rate in the yeast Saccharomyces cerevisiae[J]. Genetics, 2008, 178(1): 67-82.
doi: 10.1534/genetics.107.071506 URL |
| [16] |
Zheng DQ, Wang YT, Zhu YX, et al. Uncovering bleomycin-induced genomic alterations and underlying mechanisms in the yeast Saccharomyces cerevisiae[J]. Appl Environ Microbiol, 2022, 88(2): e0170321.
doi: 10.1128/AEM.01703-21 URL |
| [17] | Kiktev DA, Sheng ZW, Lobachev KS, et al. GC content elevates mutation and recombination rates in the yeast Saccharomyces cerevisiae[J]. Proc Natl Acad Sci USA, 2018, 115(30): E7109-E7118. |
| [18] |
Lippert MJ, Kim N, Cho JE, et al. Role for topoisomerase 1 in transcription-associated mutagenesis in yeast[J]. Proc Natl Acad Sci USA, 2011, 108(2): 698-703.
doi: 10.1073/pnas.1012363108 pmid: 21177427 |
| [19] | Putnam CD, Kolodner RD. Determination of gross chromosomal rearrangement rates[J]. Cold Spring Harb Protoc, 2010, 2010(9): pdb.prot5492. |
| [20] |
Shibata M, Keyamura K, Shioiri T, et al. Diploid-associated adaptation to chronic low-dose UV irradiation requires homologous recombination in Saccharomyces cerevisiae[J]. Genetics, 2022, 222(1): iyac115.
doi: 10.1093/genetics/iyac115 URL |
| [21] |
St Charles J, Petes TD. High-resolution mapping of spontaneous mitotic recombination hotspots on the 1.1 Mb arm of yeast chromosome IV[J]. PLoS Genet, 2013, 9(4): e1003434.
doi: 10.1371/journal.pgen.1003434 URL |
| [22] |
Yin Y, Petes TD. Genome-wide high-resolution mapping of UV-induced mitotic recombination events in Saccharomyces cerevisiae[J]. PLoS Genet, 2013, 9(10): e1003894.
doi: 10.1371/journal.pgen.1003894 URL |
| [23] |
Yin Y, Dominska M, Yim E, et al. High-resolution mapping of heteroduplex DNA formed during UV-induced and spontaneous mitotic recombination events in yeast[J]. eLife, 2017, 6: e28069.
doi: 10.7554/eLife.28069 URL |
| [24] |
Zhang K, Zheng DQ, Sui Y, et al. Genome-wide analysis of genomic alterations induced by oxidative DNA damage in yeast[J]. Nucleic Acids Res, 2019, 47(7): 3521-3535.
doi: 10.1093/nar/gkz027 pmid: 30668788 |
| [25] |
Sheng H, Qi L, Sui Y, et al. Mapping chromosomal instability induced by small-molecular therapeutics in a yeast model[J]. Appl Microbiol Biotechnol, 2019, 103(12): 4869-4880.
doi: 10.1007/s00253-019-09845-5 pmid: 31053912 |
| [26] | Qi L, Zhang K, Wang YT, et al. Global analysis of furfural-induced genomic instability using a yeast model[J]. Appl Environ Microbiol, 2019, 85(18): e01237-e01219. |
| [27] |
Qi L, Zhu YX, Wang YK, et al. Nonlethal furfural exposure causes genomic alterations and adaptability evolution in Saccharomyces cerevisiae[J]. Microbiol Spectr, 2023, 11(4): e0121623.
doi: 10.1128/spectrum.01216-23 URL |
| [28] |
Shen L, Wang YT, Tang XX, et al. Heat shock drives genomic instability and phenotypic variations in yeast[J]. AMB Express, 2020, 10(1): 146.
doi: 10.1186/s13568-020-01091-7 pmid: 32804300 |
| [29] |
St Charles J, Hazkani-Covo E, Yin Y, et al. High-resolution genome-wide analysis of irradiated(UV and γ-rays)diploid yeast cells reveals a high frequency of genomic loss of heterozygosity(LOH)events[J]. Genetics, 2012, 190(4): 1267-1284.
doi: 10.1534/genetics.111.137927 pmid: 22267500 |
| [30] |
Lynch M, Sung W, Morris K, et al. A genome-wide view of the spectrum of spontaneous mutations in yeast[J]. Proc Natl Acad Sci U S A, 2008, 105(27): 9272-9277.
doi: 10.1073/pnas.0803466105 URL |
| [31] |
Peter J, De Chiara M, Friedrich A, et al. Genome evolution across 1, 011 Saccharomyces cerevisiae isolates[J]. Nature, 2018, 556(7701): 339-344.
doi: 10.1038/s41586-018-0030-5 |
| [32] | Song W, Dominska M, Greenwell PW, et al. Genome-wide high-resolution mapping of chromosome fragile sites in Saccharomyces cerevisiae[J]. Proc Natl Acad Sci USA, 2014, 111(21): E2210-E2218. |
| [33] | Zhu YO, Siegal ML, Hall DW, et al. Precise estimates of mutation rate and spectrum in yeast[J]. Proc Natl Acad Sci USA, 2014, 111(22): E2310-E2318. |
| [34] |
Sui Y, Qi L, Wu JK, et al. Genome-wide mapping of spontaneous genetic alterations in diploid yeast cells[J]. Proc Natl Acad Sci USA, 2020, 117(45): 28191-28200.
doi: 10.1073/pnas.2018633117 pmid: 33106417 |
| [35] |
Rhoads A, Au KF. PacBio sequencing and its applications[J]. Genomics Proteomics Bioinformatics, 2015, 13(5): 278-289.
doi: 10.1016/j.gpb.2015.08.002 URL |
| [36] |
Hu TS, Chitnis N, Monos D, et al. Next-generation sequencing technologies: an overview[J]. Hum Immunol, 2021, 82(11): 801-811.
doi: 10.1016/j.humimm.2021.02.012 pmid: 33745759 |
| [37] |
O'Donnell S, Yue JX, Saada OA, et al. Telomere-to-telomere assemblies of 142 strains characterize the genome structural landscape in Saccharomyces cerevisiae[J]. Nat Genet, 2023, 55(8): 1390-1399.
doi: 10.1038/s41588-023-01459-y |
| [38] |
Qi L, Sui Y, Tang XX, et al. Shuffling the yeast genome using CRISPR/Cas9-generated DSBs that target the transposable Ty1 elements[J]. PLoS Genet, 2023, 19(1): e1010590.
doi: 10.1371/journal.pgen.1010590 URL |
| [39] |
Ogur M, Ogur S, St John R. Temperature dependence of the spontaneous mutation rate to respiration deficiency in Saccharomyces[J]. Genetics, 1960, 45(2): 189-194.
doi: 10.1093/genetics/45.2.189 pmid: 17247917 |
| [40] | Zhang K, Wu XC, Zheng DQ, et al. Effects of temperature on the meiotic recombination landscape of the yeast Saccharomyces cerevisiae[J]. mBio, 2017, 8(6): e02099-e02017. |
| [41] | Huang CJ, Lu MY, Chang YW, et al. Experimental evolution of yeast for high-temperature tolerance[J]. Mol Biol Evol, 2018, 35(8): 1823-1839. |
| [42] |
Finkel T, Holbrook NJ. Oxidants, oxidative stress and the biology of ageing[J]. Nature, 2000, 408(6809): 239-247.
doi: 10.1038/35041687 |
| [43] |
Hayashi M, Umezu K. Homologous recombination is required for recovery from oxidative DNA damage[J]. Genes Genet Syst, 2017, 92(2): 73-80.
doi: 10.1266/ggs.16-00066 pmid: 28381656 |
| [44] |
Poetsch AR. The genomics of oxidative DNA damage, repair, and resulting mutagenesis[J]. Comput Struct Biotechnol J, 2020, 18: 207-219.
doi: 10.1016/j.csbj.2019.12.013 URL |
| [45] |
Ikner A, Shiozaki K. Yeast signaling pathways in the oxidative stress response[J]. Mutat Res, 2005, 569(1-2): 13-27.
doi: 10.1016/j.mrfmmm.2004.09.006 URL |
| [46] |
Degtyareva NP, Heyburn L, Sterling J, et al. Oxidative stress-induced mutagenesis in single-strand DNA occurs primarily at cytosines and is DNA polymerase zeta-dependent only for adenines and guanines[J]. Nucleic Acids Res, 2013, 41(19): 8995-9005.
doi: 10.1093/nar/gkt671 pmid: 23925127 |
| [47] |
Degtyareva NP, Placentra VC, Gabel SA, et al. Changes in metabolic landscapes shape divergent but distinct mutational signatures and cytotoxic consequences of redox stress[J]. Nucleic Acids Res, 2023, 51(10): 5056-5072.
doi: 10.1093/nar/gkad305 URL |
| [48] |
Choi JE, Heo SH, Kim MJ, et al. Lack of superoxide dismutase in a rad51 mutant exacerbates genomic instability and oxidative stress-mediated cytotoxicity in Saccharomyces cerevisiae[J]. Free Radic Biol Med, 2018, 129: 97-106.
doi: 10.1016/j.freeradbiomed.2018.09.015 URL |
| [49] | Bjornsti MA, Knab AM, Benedetti P. Yeast Saccharomyces cerevisiae as a model system to study the cytotoxic activity of the antitumor drug camptothecin[J]. Cancer Chemother Pharmacol, 1994, 34 Suppl: S1-S5. |
| [50] |
Puddu F, Salguero I, Herzog M, et al. Chromatin determinants impart camptothecin sensitivity[J]. EMBO Rep, 2017, 18(6): 1000-1012.
doi: 10.15252/embr.201643560 pmid: 28389464 |
| [51] |
Pommier Y, Sun YL, Huang SY N, et al. Roles of eukaryotic topoisomerases in transcription, replication and genomic stability[J]. Nat Rev Mol Cell Biol, 2016, 17(11): 703-721.
doi: 10.1038/nrm.2016.111 |
| [52] |
Sloan R, Huang SY N, Pommier Y, et al. Effects of camptothecin or TOP1 overexpression on genetic stability in Saccharomyces cerevisiae[J]. DNA Repair, 2017, 59: 69-75.
doi: 10.1016/j.dnarep.2017.09.004 URL |
| [53] | Thomas AD, Johnson GE. DNA repair and its influence on points of departure for alkylating agent genotoxicity[M]// Thresholds of Genotoxic Carcinogens. Amsterdam: Elsevier, 2016: 67-82. |
| [54] |
Lundin C, North M, Erixon K, et al. Methyl methanesulfonate(MMS)produces heat-labile DNA damage but no detectable in vivo DNA double-strand breaks[J]. Nucleic Acids Res, 2005, 33(12): 3799-3811.
doi: 10.1093/nar/gki681 URL |
| [55] |
Haracska L, Unk I, Johnson RE, et al. Roles of yeast DNA polymerases delta and zeta and of Rev1 in the bypass of abasic sites[J]. Genes Dev, 2001, 15(8): 945-954.
doi: 10.1101/gad.882301 URL |
| [56] |
Dolling JA, Boreham DR, Brown DL, et al. Cisplatin-modification of DNA repair and ionizing radiation lethality in yeast, Saccharomyces cerevisiae[J]. Mutat Res, 1999, 433(2): 127-136.
doi: 10.1016/S0921-8777(98)00069-X URL |
| [57] |
Segovia R, Shen YQ, Lujan SA, et al. Hypermutation signature reveals a slippage and realignment model of translesion synthesis by Rev3 polymerase in cisplatin-treated yeast[J]. Proc Natl Acad Sci USA, 2017, 114(10): 2663-2668.
doi: 10.1073/pnas.1618555114 pmid: 28223526 |
| [58] |
Ward JF. The yield of DNA double-strand breaks produced intracellularly by ionizing radiation: a review[J]. Int J Radiat Biol, 1990, 57(6): 1141-1150.
doi: 10.1080/09553009014551251 pmid: 1971840 |
| [59] |
Brennan RJ, Schiestl RH. Persistent genomic instability in the yeast Saccharomyces cerevisiae induced by ionizing radiation and DNA-damaging agents[J]. Radiat Res, 2001, 155(6): 768-777.
pmid: 11352758 |
| [60] |
Argueso JL, Westmoreland J, Mieczkowski PA, et al. Double-strand breaks associated with repetitive DNA can reshape the genome[J]. Proc Natl Acad Sci USA, 2008, 105(33): 11845-11850.
doi: 10.1073/pnas.0804529105 pmid: 18701715 |
| [61] |
Franklin WA, Doetsch PW, Haseltine WA. Structural determination of the ultraviolet light-induced thymine-cytosine pyrimidine-pyrimidone(6-4)photoproduct[J]. Nucleic Acids Res, 1985, 13(14): 5317-5325.
pmid: 4022781 |
| [62] |
Bhavya G, Hiremath KY, Jogaiah S, et al. Heavy metal-induced oxidative stress and alteration in secretory proteins in yeast isolates[J]. Arch Microbiol, 2022, 204(3): 172.
doi: 10.1007/s00203-022-02756-6 pmid: 35165751 |
| [63] |
Diaconu M, Pavel LV, Hlihor RM, et al. Characterization of heavy metal toxicity in some plants and microorganisms-a preliminary approach for environmental bioremediation[J]. N Biotechnol, 2020, 56: 130-139.
doi: 10.1016/j.nbt.2020.01.003 URL |
| [64] |
Yu SS, Qin W, Zhuang GQ, et al. Monitoring oxidative stress and DNA damage induced by heavy metals in yeast expressing a redox-sensitive green fluorescent protein[J]. Curr Microbiol, 2009, 58(5): 504-510.
doi: 10.1007/s00284-008-9354-y pmid: 19184609 |
| [65] |
Wu K, Li HC, Wang YH, et al. Silver nanoparticles elevate mutagenesis of eukaryotic genomes[J]. G3, 2023, 13(3): jkad008.
doi: 10.1093/g3journal/jkad008 URL |
| [66] |
Liu HX, Zhang JZ. Yeast spontaneous mutation rate and spectrum vary with environment[J]. Curr Biol, 2019, 29(10): 1584-1591.e3.
doi: S0960-9822(19)30382-3 pmid: 31056389 |
| [67] |
Voordeckers K, Colding C, Grasso L, et al. Ethanol exposure increases mutation rate through error-prone polymerases[J]. Nat Commun, 2020, 11(1): 3664.
doi: 10.1038/s41467-020-17447-3 pmid: 32694532 |
| [68] |
Yona AH, Frumkin I, Pilpel Y. A relay race on the evolutionary adaptation spectrum[J]. Cell, 2015, 163(3): 549-559.
doi: 10.1016/j.cell.2015.10.005 pmid: 26496602 |
| [69] |
Zhang K, Fang YH, Gao KH, et al. Effects of genome duplication on phenotypes and industrial applications of Saccharomyces cerevisiae strains[J]. Appl Microbiol Biotechnol, 2017, 101(13): 5405-5414.
doi: 10.1007/s00253-017-8284-7 pmid: 28429058 |
| [70] |
Linder RA, Greco JP, Seidl F, et al. The stress-inducible peroxidase TSA2 underlies a conditionally beneficial chromosomal duplication in Saccharomyces cerevisiae[J]. G3, 2017, 7(9): 3177-3184.
doi: 10.1534/g3.117.300069 URL |
| [71] | Li J, Stenberg S, Yue JX, et al. Genome instability footprint under rapamycin and hydroxyurea treatments[J]. bioRxiv, 2023. DOI: 10.1101/2023.02.28.530484. |
| [72] |
Noer JB, Hørsdal OK, Xiang X, et al. Extrachromosomal circular DNA in cancer: history, current knowledge, and methods[J]. Trends Genet, 2022, 38(7): 766-781.
doi: 10.1016/j.tig.2022.02.007 pmid: 35277298 |
| [73] |
Gresham D, Usaite R, Germann SM, et al. Adaptation to diverse nitrogen-limited environments by deletion or extrachromosomal element formation of the GAP1 locus[J]. Proc Natl Acad Sci USA, 2010, 107(43): 18551-18556.
doi: 10.1073/pnas.1014023107 pmid: 20937885 |
| [74] |
Kang JP, Li JY, Guo Z, et al. Enhancement and mapping of tolerance to salt stress and 5-fluorocytosine in synthetic yeast strains via SCRaMbLE[J]. Synth Syst Biotechnol, 2022, 7(3): 869-877.
doi: 10.1016/j.synbio.2022.04.003 pmid: 35601823 |
| [75] |
Yona AH, Manor YS, Herbst RH, et al. Chromosomal duplication is a transient evolutionary solution to stress[J]. Proc Natl Acad Sci USA, 2012, 109(51): 21010-21015.
doi: 10.1073/pnas.1211150109 pmid: 23197825 |
| [76] |
Morard M, Macías LG, Adam AC, et al. Aneuploidy and ethanol tolerance in Saccharomyces cerevisiae[J]. Front Genet, 2019, 10: 82.
doi: 10.3389/fgene.2019.00082 URL |
| [77] |
García-Ríos E, Nuévalos M, Barrio E, et al. A new chromosomal rearrangement improves the adaptation of wine yeasts to sulfite[J]. Environ Microbiol, 2019, 21(5): 1771-1781.
doi: 10.1111/1462-2920.14586 pmid: 30859719 |
| [78] |
Hong J, Gresham D. Molecular specificity, convergence and constraint shape adaptive evolution in nutrient-poor environments[J]. PLoS Genet, 2014, 10(1): e1004041.
doi: 10.1371/journal.pgen.1004041 URL |
| [79] |
Hose J, Yong CM, Sardi M, et al. Dosage compensation can buffer copy-number variation in wild yeast[J]. eLife, 2015, 4: e05462.
doi: 10.7554/eLife.05462 URL |
| [80] |
Selmecki A, Forche A, Berman J. Aneuploidy and isochromosome formation in drug-resistant Candida albicans[J]. Science, 2006, 313(5785): 367-370.
pmid: 16857942 |
| [81] |
Dunham MJ, Badrane H, Ferea T, et al. Characteristic genome rearrangements in experimental evolution of Saccharomyces cerevisiae[J]. Proc Natl Acad Sci USA, 2002, 99(25): 16144-16149.
doi: 10.1073/pnas.242624799 URL |
| [1] | XU Fa-di, XU Kang, SUN Dong-ming, LI Meng-lei, ZHAO Jian-zhi, BAO Xiao-ming. Research Progress in Second-generation Fuel Ethanol Technology Based on Poplar(Populus sp.) [J]. Biotechnology Bulletin, 2023, 39(9): 27-39. |
| [2] | CHENG Ting, YUAN Shuai, ZHANG Xiao-yuan, LIN Liang-cai, LI Xin, ZHANG Cui-ying. Research Progress in the Regulation of Isobutanol Synthesis Pathway in Saccharomyces cerevisiae [J]. Biotechnology Bulletin, 2023, 39(7): 80-90. |
| [3] | WANG Chen-yu, ZHOU Chu-yuan, HE Di, FAN Zi-hao, WANG Meng-meng, YANG Liu-yan. Role and Mechanism of Polyphosphate in the Microbial Response to Environmental Stresses [J]. Biotechnology Bulletin, 2023, 39(11): 168-181. |
| [4] | SUN Yan-qiu, XIE Cai-yun, TANG Yue-qin. Construction and Mechanism Analysis of High-temperature Resistant Saccharomyces cerevisiae [J]. Biotechnology Bulletin, 2023, 39(11): 226-237. |
| [5] | WANG Wen-tao, FENG Qi, LIU Chen-guang, BAI Feng-wu, ZHAO Xin-qing. Redox-sensitive Genetic Parts Improve the Tolerance of Yeast to Lignocellulosic Hydrolysate Inhibitors [J]. Biotechnology Bulletin, 2023, 39(11): 360-372. |
| [6] | CUI Xin-gang, SUN Ya-xin, CUI Xiao-jing, DENG Yan-wen, SUN En-hao, WANG Jun-fang, CUI Hong-jing. Roles of Gene TAP42 in the Cell Wall Stress Response of Saccharomyces cerevisiae [J]. Biotechnology Bulletin, 2021, 37(10): 57-62. |
| [7] | CHEN Xue-lian, JIANG Gao-fei, ZHONG Zeng-tao. Research Progress of Horizontal Gene Transfer in Rhizobia Evolution [J]. Biotechnology Bulletin, 2019, 35(10): 18-24. |
| [8] | GUO Xue-wu, ZHANG Yu, GUAN Xiang-yu, NI Xiao-feng, WANG Qing, CHEN Ye-fu, XIAO Dong-guang. Transcriptomics Analysis of High-xylose-tolerance Klebsiella pneumonia Strain and Optimization of Fermentation Conditions for 2,3-butanediol Production [J]. Biotechnology Bulletin, 2018, 34(8): 159-169. |
| [9] | Li Hong. Molecular Mechanism of Genomic Stability and iPS Cells Reprogramming [J]. Biotechnology Bulletin, 2013, 0(12): 36-42. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||