








Biotechnology Bulletin ›› 2024, Vol. 40 ›› Issue (1): 194-206.doi: 10.13560/j.cnki.biotech.bull.1985.2023-0672
Previous Articles Next Articles
ZHANG Yi1,2( ), ZHANG Xin-ru1,2, ZHANG Jin-ke1,2, HU Li-zong1,2, SHANGGUAN Xin-xin1,2, ZHENG Xiao-hong1,2, HU Juan-juan1,2, ZHANG Cong-cong1,2, MU Gui-qing1,2, LI Cheng-wei3(
), ZHANG Xin-ru1,2, ZHANG Jin-ke1,2, HU Li-zong1,2, SHANGGUAN Xin-xin1,2, ZHENG Xiao-hong1,2, HU Juan-juan1,2, ZHANG Cong-cong1,2, MU Gui-qing1,2, LI Cheng-wei3( )
)
Received:2023-07-14
Online:2024-01-26
Published:2024-02-06
Contact:
LI Cheng-wei
E-mail:yizhang0401@sina.com;lichengweihist@163.com
ZHANG Yi, ZHANG Xin-ru, ZHANG Jin-ke, HU Li-zong, SHANGGUAN Xin-xin, ZHENG Xiao-hong, HU Juan-juan, ZHANG Cong-cong, MU Gui-qing, LI Cheng-wei. Functional Analysis of TaMYB1 Gene in Wheat Under Cadmium Stress[J]. Biotechnology Bulletin, 2024, 40(1): 194-206.
| 引物名称Primer name | 引物序列Primer sequence(5'-3') |
|---|---|
| TaMYB1-F | GTGAGTAAGGTTACCGAACTCCAAATGCAAATTGAG |
| TaMYB1-R | CGTGAGCTCGGTACCGGAGACGAGGCTAGCTCCGGC |
| TaMYB1-qPCR-F | AATTGAGGTCCAGAGACGACTG |
| TaMYB1-qPCR-R | GGTACTTCCCCTGGGCTTCG |
| TaActin-qPCR-F | CCAGGTATCGCTGACCGTAT |
| TaActin-qPCR-R | GCTGAGTGAGGCTAGGATGG |
Table 1 Primer sequences
| 引物名称Primer name | 引物序列Primer sequence(5'-3') |
|---|---|
| TaMYB1-F | GTGAGTAAGGTTACCGAACTCCAAATGCAAATTGAG |
| TaMYB1-R | CGTGAGCTCGGTACCGGAGACGAGGCTAGCTCCGGC |
| TaMYB1-qPCR-F | AATTGAGGTCCAGAGACGACTG |
| TaMYB1-qPCR-R | GGTACTTCCCCTGGGCTTCG |
| TaActin-qPCR-F | CCAGGTATCGCTGACCGTAT |
| TaActin-qPCR-R | GCTGAGTGAGGCTAGGATGG |

Fig. 1 Phenotype analysis of wheat plant under Cd stress A: Phenotype of wheat plants under Cd stress. B: Root length of wheat plants under Cd stress. C: Shoot length of wheat plants under Cd stress. D: Total biomass of wheat plants under Cd stress. ** indicates significant difference from control plants at P=0.01
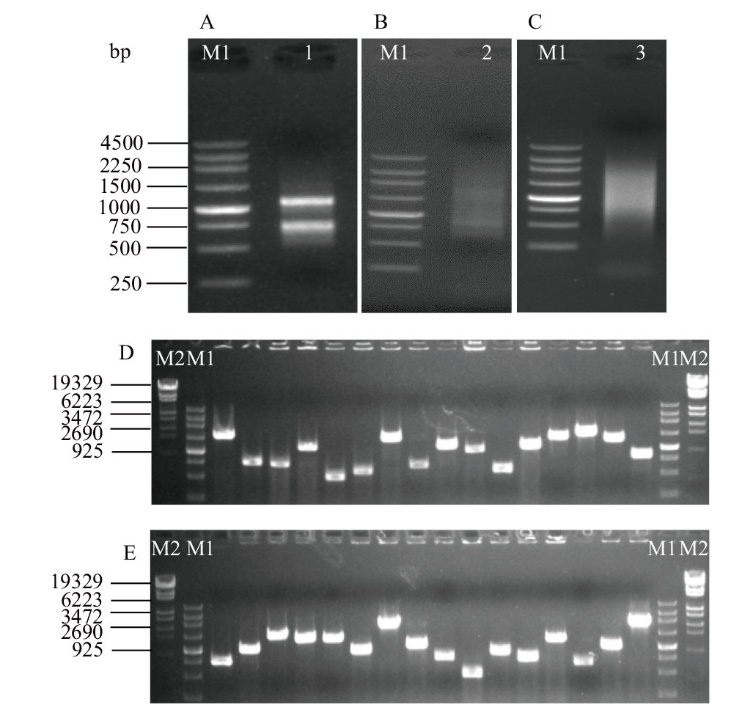
Fig. 2 Quality detection of yeast two-hybrid library of wheat roots under Cd stress A: Total RNA of wheat roots under Cd stress. B: High-quality cDNA generated using SMART cDNA synthesis technology. C: Purified cDNA with column chromatography. D, E: PCR analysis of inserted clones from the yeast library

Fig. 3 Transformants with the medium containing different cadmium concentration A-C: Refer to the screening results of yeast two-hybrid library in the yeast medium containing 0.002, 0.003 and 0.004 mol/L CdCl2
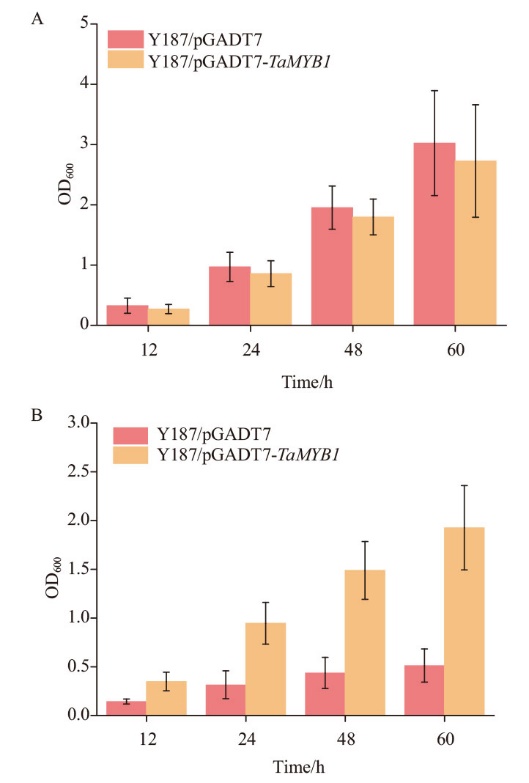
Fig. 4 Growth curves of yeast colonies in the SD-Leu medium added with CdCl2 A: Growth curves of yeast transformants growing in SD-Leu medium. B: Growth curves of yeast transformants growing in 0.004 mol/L CdCl2 medium
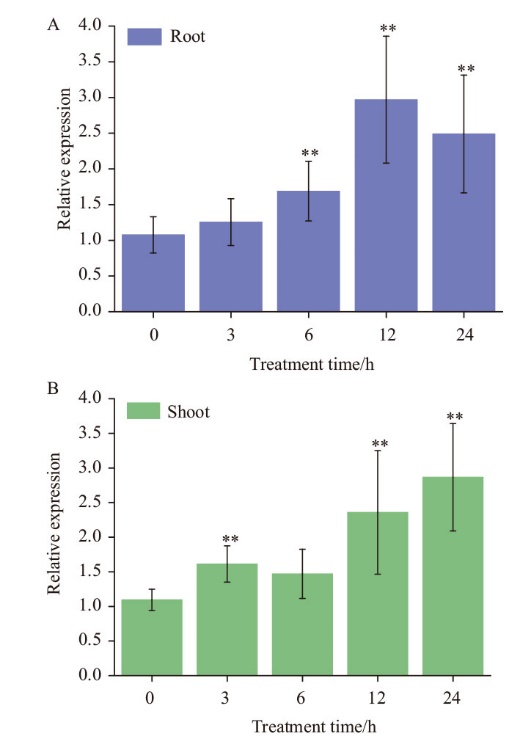
Fig. 5 Expression analysis of TaMYB1 under Cd stress A: Root. B: Shoot. ** indicates significant difference from control plants at P=0.01, the same below
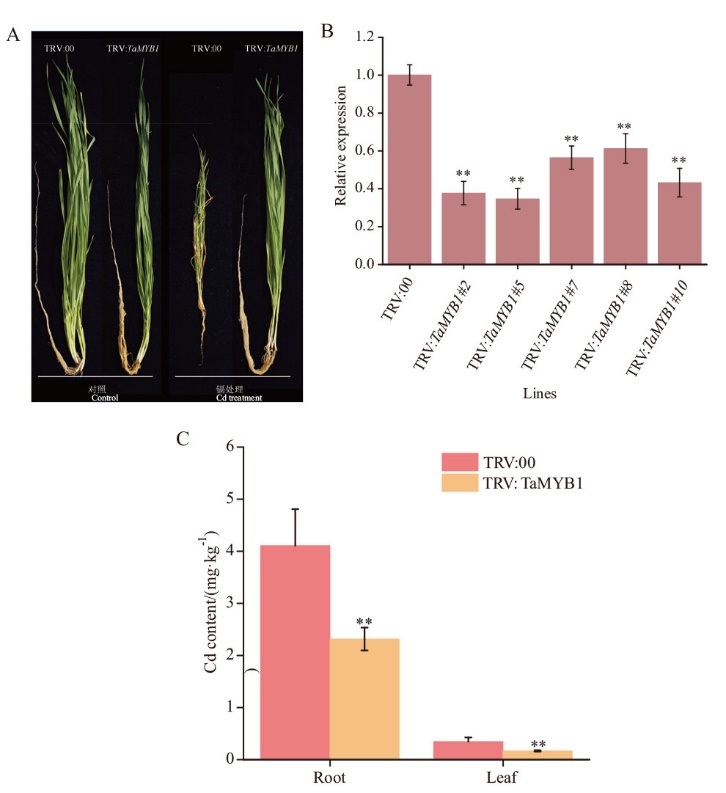
Fig. 6 Silencing of TaMYB1 enhanced wheat tolerance to cadmium A: Phenotypes of TaMYB1 gene-silenced and control plants under normal conditions and cadmium stress. B: Relative expression level of TaMYB1 gene in gene-silenced plants. C: Cadmium content of TaMYB1 gene-silenced and control plants after cadmium stress

Fig. 7 Physiological index detection of TaMYB1 gene-silenced and control plants under normal conditions and cadmium stress The data in A-D were all taken from wheat leaves

Fig. 9 Sequence features and phylogenetic relationships of TaMYB1 and its homologous genes in wheat A: Phylogenetic tree of TaMYB1 and its homologous proteins was analyzed. B: Conserved motifs of TaMYB1 and its homologous protein. C: Structures of TaMYB1 and its homologous genes. D: The sequence logo of the conserved motif 1. E: The sequence logo of the conserved motif 2
| [1] | 中华人民共和国环境保护部, 国土资源部. 全国土壤污染状况调查公报[R]. 北京: 中华人民共和国环境保护部, 国土资源部, 2014. |
| Ministry of Environmental Protection of PRC, Ministry of Land and Resources of PRC. Report on the national general survey of soil contamination[R]. Beijing: Ministry of Environmental Protection of PRC, Ministry of Land and Resources of PRC, 2014. | |
| [2] | 尚二萍, 许尔琪, 张红旗, 等. 中国粮食主产区耕地土壤重金属时空变化与污染源分析[J]. 环境科学, 2018, 39(10): 4670-4683. |
| Shang EP, Xu EQ, Zhang HQ, et al. Spatial-temporal trends and pollution source analysis for heavy metal contamination of cultivated soils in five major grain producing regions of China[J]. Environ Sci, 2018, 39(10): 4670-4683. | |
| [3] |
Raza A, Habib M, Kakavand SN, et al. Phytoremediation of cadmium: physiological, biochemical, and molecular mechanisms[J]. Biology, 2020, 9(7): 177.
doi: 10.3390/biology9070177 URL |
| [4] |
Rizwan M, Rubina Gilani S, Iqbal Durani A, et al. Materials diversity of hydrogel: synthesis, polymerization process and soil conditioning properties in agricultural field[J]. J Adv Res, 2021, 33: 15-40.
doi: 10.1016/j.jare.2021.03.007 pmid: 34603776 |
| [5] | 王怡雯, 芮玉奎, 李中阳, 等. 冬小麦吸收重金属特征及与影响因素的定量关系[J]. 环境科学, 2020, 41(3): 1482-1490. |
| Wang YW, Rui YK, Li ZY, et al. Characteristics of heavy metal absorption by winter wheat and its quantitative relationship with influencing factors[J]. Environ Sci, 2020, 41(3): 1482-1490. | |
| [6] | 赵多勇, 魏益民, 魏帅, 等. 区域小麦籽粒重金属分布及暴露评估[J]. 中国粮油学报, 2016, 31(7): 6-10, 18. |
| Zhao DY, Wei YM, Wei S, et al. Spatial distribution of heavy metal in wheat kernal and dietary exposure assessment of local residents in an industrial area[J]. J Chin Cereals Oils Assoc, 2016, 31(7): 6-10, 18. | |
| [7] | 肖冰, 薛培英, 韦亮, 等. 基于田块尺度的农田土壤和小麦籽粒镉砷铅污染特征及健康风险评价[J]. 环境科学, 2020, 41(6): 2869-2877. |
| Xiao B, Xue PY, Wei L, et al. Characteristics of Cd, As, and Pb in soil and wheat grains and health risk assessment of grain-Cd/As/Pb on the field scale[J]. Environ Sci, 2020, 41(6): 2869-2877. | |
| [8] |
Shi TR, Zhang YY, Gong YW, et al. Status of cadmium accumulation in agricultural soils across China(1975-2016): From temporal and spatial variations to risk assessment[J]. Chemosphere, 2019, 230: 136-143.
doi: 10.1016/j.chemosphere.2019.04.208 URL |
| [9] |
Hu YA, Cheng HF, Tao S. The challenges and solutions for cadmium-contaminated rice in China: A critical review[J]. Environ Int, 2016, 92/93: 515-532.
doi: 10.1016/j.envint.2016.04.042 URL |
| [10] | 王兴利, 王晨野, 吴晓晨, 等. 重金属污染土壤修复技术研究进展[J]. 化学与生物工程, 2019, 36(2): 1-7, 11. |
| Wang XL, Wang CY, Wu XC, et al. Research progress in remediation technology of heavy metal contaminated soil[J]. Chem Bioeng, 2019, 36(2): 1-7, 11. | |
| [11] |
Dinh N, van der Ent A, Mulligan DR, et al. Zinc and lead accumulation characteristics and in vivo distribution of Zn2+ in the hyperaccumulator Noccaea caerulescens elucidated with fluorescent probes and laser confocal microscopy[J]. Environ Exp Bot, 2018, 147: 1-12.
doi: 10.1016/j.envexpbot.2017.10.008 URL |
| [12] |
Wen X, Ding Y, Tan Z, et al. Identification and characterization of cadmium stress-related LncRNAs from Betula platyphylla[J]. Plant Sci, 2020, 299: 110601.
doi: 10.1016/j.plantsci.2020.110601 URL |
| [13] |
Chen L, Shi S, Jiang N, et al. Genome-wide analysis of long non-coding RNAs affecting roots development at an early stage in the rice response to cadmium stress[J]. BMC Genomics, 2018, 19(1): 460.
doi: 10.1186/s12864-018-4807-6 pmid: 29902991 |
| [14] |
Meng JG, Zhang XD, Tan SK, et al. Genome-wide identification of Cd-responsive NRAMP transporter genes and analyzing expression of NRAMP 1 mediated by miR167 in Brassica napus[J]. Biometals, 2017, 30(6): 917-931.
doi: 10.1007/s10534-017-0057-3 pmid: 28993932 |
| [15] |
Ding YF, Gong SH, Wang Y, et al. microRNA166 modulates cadmium tolerance and accumulation in rice[J]. Plant Physiol, 2018, 177(4): 1691-1703.
doi: 10.1104/pp.18.00485 pmid: 29925586 |
| [16] |
Wang NH, Zhou XY, Shi SH, et al. An miR156-regulated nucleobase-ascorbate transporter 2 confers cadmium tolerance via enhanced anti-oxidative capacity in barley[J]. J Adv Res, 2023, 44: 23-37.
doi: 10.1016/j.jare.2022.04.001 URL |
| [17] |
Han YY, Fan TT, Zhu XY, et al. WRKY12 represses GSH1 expression to negatively regulate cadmium tolerance in Arabidopsis[J]. Plant Mol Biol, 2019, 99(1): 149-159.
doi: 10.1007/s11103-018-0809-7 |
| [18] |
Li GZ, Zheng YX, Liu HT, et al. WRKY74 regulates cadmium tolerance through glutathione-dependent pathway in wheat[J]. Environ Sci Pollut Res Int, 2022, 29(45): 68191-68201.
doi: 10.1007/s11356-022-20672-6 |
| [19] |
Wang LJ, Lu WX, Ran LY, et al. R2R3-MYB transcription factor MYB6 promotes anthocyanin and proanthocyanidin biosynthesis but inhibits secondary cell wall formation in Populus tomentosa[J]. Plant J, 2019, 99(4): 733-751.
doi: 10.1111/tpj.v99.4 URL |
| [20] |
Pratyusha DS, Sarada DVL. MYB transcription factors-master regulators of phenylpropanoid biosynthesis and diverse developmental and stress responses[J]. Plant Cell Rep, 2022, 41(12): 2245-2260.
doi: 10.1007/s00299-022-02927-1 pmid: 36171500 |
| [21] |
Yang J, Chen Y, Xiao Z, et al. Multilevel regulation of anthocyanin-promoting R2R3-MYB transcription factors in plants[J]. Front Plant Sci, 2022, 13: 1008829.
doi: 10.3389/fpls.2022.1008829 URL |
| [22] |
Yin Y, Guo C, Shi HY, et al. Genome-wide comparative analysis of the R2R3-MYB gene family in five solanaceae species and identification of members regulating carotenoid biosynthesis in wolfberry[J]. Int J Mol Sci, 2022, 23(4): 2259.
doi: 10.3390/ijms23042259 URL |
| [23] |
Dubos C, Stracke R, Grotewold E et al. MYB transcription factors in Arabidopsis[J]. Trends Plant Sci, 2010, 15(10): 573-581.
doi: 10.1016/j.tplants.2010.06.005 URL |
| [24] |
Hu SB, Yu Y, Chen QH, et al. OsMYB45 plays an important role in rice resistance to cadmium stress[J]. Plant Sci, 2017, 264: 1-8.
doi: 10.1016/j.plantsci.2017.08.002 URL |
| [25] |
Zhang P, Wang RL, Ju Q, et al. The R2R3-MYB transcription factor MYB49 regulates cadmium accumulation[J]. Plant Physiol, 2019, 180(1): 529-542.
doi: 10.1104/pp.18.01380 pmid: 30782964 |
| [26] |
Zhu S, Shi W, Jie Y, et al. A MYB transcription factor, BnMYB2, cloned from ramie(Boehmeria nivea)is involved in cadmium tolerance and accumulation[J]. PLoS One, 2020, 15(5): e0233375.
doi: 10.1371/journal.pone.0233375 URL |
| [27] |
Tiwari P, Indoliya Y, Chauhan AS et al. Over-expression of rice R1-type MYB transcription factor confers different abiotic stress tolerance in transgenic Arabidopsis[J]. Ecotoxicol Environ Saf, 2020, 206: 111361.
doi: 10.1016/j.ecoenv.2020.111361 URL |
| [28] | 宋红改, 蒋晶, 乔桂荣, 等. 利用酵母建立植物抗逆基因快速筛选体系[J]. 浙江林学院学报, 2010, 27(6): 890-895. |
| Song HG, Jiang J, Qiao GR, et al. Plant stress-resistant gene cloning for Na and Cd using an INVSc1 yeast[J]. J Zhejiang for Coll, 2010, 27(6): 890-895. | |
| [29] | 张怡, 刘坤, 曹鹏, 等. 一种快速高效提取酵母质粒的方法[J]. 食品与生物技术学报, 2013, 32(11): 1194-1198. |
| Zhang Y, Liu K, Cao P, et al. A fast, efficient and economical method for isolating plasmid from yeast[J]. J Food Sci Biotechnol, 2013, 32(11): 1194-1198. | |
| [30] | Zhang Y, Hu L, Yu D, et al. Integrative analysis of the wheat PHT1 gene family reveals a novel member involved in arbuscular mycorrhizal phosphate transport and immunity[J]. Cells, 2019, 8(5): E490. |
| [31] | 刘萍, 李明军. 植物生理实验技术[M]. 第2版. 北京: 科学出版社, 2008. |
| Liu P, Li MJ. Plant physiology experimental technology[M]. 2nd ed. Beijing: Science Press, 2008. | |
| [32] |
Mistry J, Chuguransky S, Williams L, et al. Pfam: The protein families database in 2021[J]. Nucleic Acids Res, 2021, 49(D1): D412-D419.
doi: 10.1093/nar/gkaa913 pmid: 33125078 |
| [33] |
Li KB. ClustalW-MPI: ClustalW analysis using distributed and parallel computing[J]. Bioinformatics, 2003, 19(12): 1585-1586.
doi: 10.1093/bioinformatics/btg192 URL |
| [34] |
Kumar S, Stecher G, Li M, et al. MEGA X: molecular evolutionary genetics analysis across computing platforms[J]. Mol Biol Evol, 2018, 35(6): 1547-1549.
doi: 10.1093/molbev/msy096 pmid: 29722887 |
| [35] |
Letunic I, Bork JP. Interactive tree of life(iTOL)v5: an online tool for phylogenetic tree display and annotation[J]. Nucleic Acids Res, 2021, 49(W1): W293-W296.
doi: 10.1093/nar/gkab301 URL |
| [36] |
Hu B, Jin J, Guo AY, et al. GSDS 2.0: an upgraded gene feature visualization server[J]. Bioinformatics, 2015, 31(8): 1296-1297.
doi: 10.1093/bioinformatics/btu817 pmid: 25504850 |
| [37] |
Bailey TL, Boden M, Buske FA, et al. MEME SUITE: tools for motif discovery and searching[J]. Nucleic Acids Res, 2009, 37: W202-W208.
doi: 10.1093/nar/gkp335 URL |
| [38] |
Crooks GE, Hon G, Chandonia JM, et al. WebLogo: a sequence logo generator[J]. Genome Res, 2004, 14(6): 1188-1190.
doi: 10.1101/gr.849004 pmid: 15173120 |
| [39] |
Pingault L, Choulet F, Alberti A, et al. Deep transcriptome sequencing provides new insights into the structural and functional organization of the wheat genome[J]. Genome Biol, 2015, 16(1): 29.
doi: 10.1186/s13059-015-0601-9 URL |
| [40] |
Trapnell C, Roberts A, Goff L, et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks[J]. Nat Protoc, 2012, 7(3): 562-578.
doi: 10.1038/nprot.2012.016 pmid: 22383036 |
| [41] |
Ning WS, Wei YX, Gao LT, et al. HemI 2.0: an online service for heatmap illustration[J]. Nucleic Acids Res, 2022, 50(W1): W405-W411.
doi: 10.1093/nar/gkac480 pmid: 35670661 |
| [42] |
Zadoks JC, Chang TT, Konzak CF. A decimal code for the growth stages of cereals[J]. Weed Res, 1974, 14(6): 415-421.
doi: 10.1111/wre.1974.14.issue-6 URL |
| [43] |
Shan TL, Rong W, Xu HJ, et al. The wheat R2R3-MYB transcription factor TaRIM1 participates in resistance response against the pathogen Rhizoctonia cerealis infection through regulating defense genes[J]. Sci Rep, 2016, 6(1): 1-14.
doi: 10.1038/s41598-016-0001-8 |
| [44] |
Hawku MD, He FX, Bai XX, et al. A R2R3 MYB transcription factor, TaMYB391, is positively involved in wheat resistance to Puccinia striiformis f. sp. tritici[J]. Int J Mol Sci, 2022, 23(22): 14070.
doi: 10.3390/ijms232214070 URL |
| [45] |
Wei XN, Shan TL, Hong YT, et al. TaPIMP2, a pathogen-induced MYB protein in wheat, contributes to host resistance to common root rot caused by Bipolaris sorokiniana[J]. Sci Rep, 2017, 7(1): 1754.
doi: 10.1038/s41598-017-01918-7 |
| [46] | 何冠谛. 低积累Cd型马铃薯相关基因挖掘及耐镉分子机制研究[D]. 贵阳: 贵州大学, 2021. |
| He GD. Study on gene mining and molecular mechanism of cadmium tolerance in low accumulation cd potato[D]. Guiyang: Guizhou University, 2021. | |
| [47] | 汪怡文. 转录组和代谢组整合分析冬小麦镉胁迫响应的关键代谢通路[D]. 武汉: 华中农业大学, 2021. |
| Wang YW. transcriptome and metabolonmics integrated analysis of key metabolic pathways in response to cadmium stress in winter wheat[D]. Wuhan: Huazhong Agricultural University, 2021. | |
| [48] |
Niu M, Bao C, Zhan J, et al. Plasma membrane-localized protein BcHIPP16 promotes the uptake of copper and cadmium in planta[J]. Ecotoxicol Environ Saf, 2021, 227: 112920.
doi: 10.1016/j.ecoenv.2021.112920 URL |
| [49] |
Martel AB, Taylor AE, Qaderi MM. Individual and interactive effects of temperature and light intensity on canola growth, physiological characteristics and methane emissions[J]. Plant Physiol Biochem, 2020, 157: 160-168.
doi: 10.1016/j.plaphy.2020.10.016 URL |
| [50] | 葛依立, 陈心胜, 黄道友, 等. 湿地植物水蓼(Polygonum hydropiper L.)对镉的富集特征及生理响应[J]. 生态毒理学报, 2020, 15(2): 195-200. |
| Ge YL, Chen XS, Huang DY, et al. Accumulation characteristics and physiological responses of the wetland plant, Polygonum hydropiper L. to cadmium[J]. Asian J Ecotoxicol, 2020, 15(2): 190-200. | |
| [51] | 史怀宇. LcAPX基因对植物镉耐受的影响及水稻组织培养条件的优化[D]. 天津: 天津大学, 2018. |
| Shi HY. Effects of LcAPX on plant cadmium resistance and optimizing of rice callus culture condition[D]. Tianjin: Tianjin University, 2018. | |
| [52] | 黄云. 马铃薯WRKY6基因增强拟南芥镉耐受性功能验证[D]. 贵阳: 贵州大学, 2022. |
| Huang Y. Function validation of Arabidopsis thaliana cadmium tolerance enhanced by potato WRKY6[D]. Guiyang: Guizhou University, 2022. |
| [1] | HE Si-cheng, ZHANG Zi-yuan, HAN Yu-qing, MIAO Lin, ZHANG Cui-ying, YU Ai-qun. Research Progress in the Production of Polyunsaturated Fatty Acids by Yarrowia lipolytica Cell Factories [J]. Biotechnology Bulletin, 2024, 40(1): 72-85. |
| [2] | WANG Zi-ying, LONG Chen-jie, FAN Zhao-yu, ZHANG Lei. Screening of OsCRK5-interacted Proteins in Rice Using Yeast Two-hybrid System [J]. Biotechnology Bulletin, 2023, 39(9): 117-125. |
| [3] | HUANG Xiao-long, SUN Gui-lian, MA Dan-dan, YAN Hui-qing. Construction of Yeast One-hybrid Library and Screening of Factors Regulating LAZY1 Expression in Rice [J]. Biotechnology Bulletin, 2023, 39(9): 126-135. |
| [4] | WEN Xiao-lei, LI Jian-yuan, LI Na, ZHANG Na, YANG Wen-xiang. Construction and Utilization of Yeast Two-hybrid cDNA Library of Wheat Interacted by Puccinia triticina [J]. Biotechnology Bulletin, 2023, 39(9): 136-146. |
| [5] | HAN Hao-zhang, ZHANG Li-hua, LI Su-hua, ZHAO Rong, WANG Fang, WANG Xiao-li. Construction of cDNA Library of Cinnamomun bodinieri Induced by Saline-alkali Stress and Screening of CbP5CS Upstream Regulators [J]. Biotechnology Bulletin, 2023, 39(9): 236-245. |
| [6] | XU Jing, ZHU Hong-lin, LIN Yan-hui, TANG Li-qiong, TANG Qing-jie, WANG Xiao-ning. Cloning of IbHQT1 Promoter and Identification of Upstream Regulatory Factors in Sweet Potato [J]. Biotechnology Bulletin, 2023, 39(8): 213-219. |
| [7] | SONG Zhi-zhong, XU Wei-hua, XIAO Hui-lin, TANG Mei-ling, CHEN Jing-hui, GUAN Xue-qiang, LIU Wan-hao. Cloning, Expression and Function of Iron Regulated Transporter VvIRT1 in Wine Grape(Vitis vinifera L.) [J]. Biotechnology Bulletin, 2023, 39(8): 234-240. |
| [8] | LIU Zhen-yin, DUAN Zhi-zhen, PENG Ting, WANG Tong-xin, WANG Jian. Establishment and Optimization of Virus-induced Gene Silencing System in Bougainvillea peruviana ‘Thimma’ [J]. Biotechnology Bulletin, 2023, 39(7): 123-130. |
| [9] | YU Hui, WANG Jing, LIANG Xin-xin, XIN Ya-ping, ZHOU Jun, ZHAO Hui-jun. Isolation and Functional Verification of Genes Responding to Iron and Cadmium Stresses in Lycium barbarum [J]. Biotechnology Bulletin, 2023, 39(7): 195-205. |
| [10] | LI Yu-zhen, MEI Tian-xiu, LI Zhi-wen, WANG Qi, LI Jun, ZOU Yue, ZHAO Xin-qing. Advances in Genomic Studies and Metabolic Engineering of Red Yeasts [J]. Biotechnology Bulletin, 2023, 39(7): 67-79. |
| [11] | ZHAO Xue-ting, GAO Li-yan, WANG Jun-gang, SHEN Qing-qing, ZHANG Shu-zhen, LI Fu-sheng. Cloning and Expression of AP2/ERF Transcription Factor Gene ShERF3 in Sugarcane and Subcellular Localization of Its Encoded Protein [J]. Biotechnology Bulletin, 2023, 39(6): 208-216. |
| [12] | LI Yuan-hong, GUO Yu-hao, CAO Yan, ZHU Zhen-zhou, WANG Fei-fei. Research Progress in the Microalgal Growth and Accumulation of Target Products Regulated by Exogenous Phytohormone [J]. Biotechnology Bulletin, 2023, 39(6): 61-72. |
| [13] | FENG Shan-shan, WANG Lu, ZHOU Yi, WANG You-ping, FANG Yu-jie. Research Progresses on WOX Family Genes in Regulating Plant Development and Abiotic Stress Response [J]. Biotechnology Bulletin, 2023, 39(5): 1-13. |
| [14] | LIU Hui, LU Yang, YE Xi-miao, ZHOU Shuai, LI Jun, TANG Jian-bo, CHEN En-fa. Comparative Transcriptome Analysis of Cadmium Stress Response Induced by Exogenous Sulfur in Tartary Buckwheat [J]. Biotechnology Bulletin, 2023, 39(5): 177-191. |
| [15] | ZHAI Ying, LI Ming-yang, ZHANG Jun, ZHAO Xu, YU Hai-wei, LI Shan-shan, ZHAO Yan, ZHANG Mei-juan, SUN Tian-guo. Heterologous Expression of Soybean Transcription Factor GmNF-YA19 Improves Drought Resistance of Transgenic Tobacco [J]. Biotechnology Bulletin, 2023, 39(5): 224-232. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||