








Biotechnology Bulletin ›› 2026, Vol. 42 ›› Issue (2): 207-217.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0789
Previous Articles Next Articles
FU Han( ), SUN Shu-hao, ZHANG Si-qing, AI Niu, YU Yang, YU Lian-wei, WANG Qiong-qiong, HAN Xiao-yu, SHI Yan, HAN Wei-li(
), SUN Shu-hao, ZHANG Si-qing, AI Niu, YU Yang, YU Lian-wei, WANG Qiong-qiong, HAN Xiao-yu, SHI Yan, HAN Wei-li( ), YANG Xue(
), YANG Xue( )
)
Received:2025-07-22
Online:2026-02-26
Published:2026-03-17
Contact:
HAN Wei-li, YANG Xue
E-mail:f2939735828@163.com;hanxing_zz@163.com;yangxuepphappy@126.com
FU Han, SUN Shu-hao, ZHANG Si-qing, AI Niu, YU Yang, YU Lian-wei, WANG Qiong-qiong, HAN Xiao-yu, SHI Yan, HAN Wei-li, YANG Xue. Genome-wide Identification and Expression Analysis of the BOI Gene Family in Nicotiana benthamiana[J]. Biotechnology Bulletin, 2026, 42(2): 207-217.
基因名称 Gene name | 上游引物 Forward primer (5'-3') | 下游引物 Reverse primer (5'-3') |
|---|---|---|
| NbBOI.1 | CGGGTGTAACGAGAGAGACG | AACACACAAATGGCGACACG |
| NbBOI.2 | CTAGTCAATCATCGGCTTTCCA | TAGGCGTGCCATTGTTTCCT |
| NbBOI.3 | TTGACTTAACCCAGCTCCAC | AGAATGATGTTGTTTTTGTTGGTGC |
| NbBRG1.2 | CAACAGCAGTTGCAAAAAGACAA | GCGTCAATCTCAATCGCTCC |
| NbBRG2.1 | TAGCTCCAGGTCCAGCTGAT | ACCCAAACGTCCATTATCCA |
| NbBRG2.3 | ACTATTACACCACCGCCACTT | GGCATCCAAATTGCAGCTGAG |
Table 1 Primers for quantitative fluorescence PCR
基因名称 Gene name | 上游引物 Forward primer (5'-3') | 下游引物 Reverse primer (5'-3') |
|---|---|---|
| NbBOI.1 | CGGGTGTAACGAGAGAGACG | AACACACAAATGGCGACACG |
| NbBOI.2 | CTAGTCAATCATCGGCTTTCCA | TAGGCGTGCCATTGTTTCCT |
| NbBOI.3 | TTGACTTAACCCAGCTCCAC | AGAATGATGTTGTTTTTGTTGGTGC |
| NbBRG1.2 | CAACAGCAGTTGCAAAAAGACAA | GCGTCAATCTCAATCGCTCC |
| NbBRG2.1 | TAGCTCCAGGTCCAGCTGAT | ACCCAAACGTCCATTATCCA |
| NbBRG2.3 | ACTATTACACCACCGCCACTT | GGCATCCAAATTGCAGCTGAG |
基因名称 Gene name | 上游引物 Forward primer (5'-3') | 下游引物 Reverse primer (5'-3') |
|---|---|---|
| NbBOI.1 | ATATGGCCATGGAGGCCAGTGAATTC ATGGCGGTTCAAGCTCAATACC | ATCTGCAGCTCGAGCTCGATGGATCC CTAACAAAGATAGACCTCAACGCTT |
| NbBOI.3 | ATATGGCCATGGAGGCCAGTGAATTC ATGGCCGTTCAAGCTCAATATGC | ATCTGCAGCTCGAGCTCGATGGATCC CTAACACAGAAAGCCCTCAACGC |
| NbBRG1.2 | ATATGGCCATGGAGGCCAGTGAATTCATGGCCATTCAAGCGCAGT | ATCTGCAGCTCGAGCTCGATGGATCC TCAAATCAAGGCCTCAATGGTAGC |
| NbBRG2.1 | ATATGGCCATGGAGGCCAGTGAATTC ATGGCTCTTCCTCATCACC | ATCTGCAGCTCGAGCTCGATGGATCC CTATATGTAAACTTCCATGCCTATATAC |
Table 2 Primers for Y2H vectors
基因名称 Gene name | 上游引物 Forward primer (5'-3') | 下游引物 Reverse primer (5'-3') |
|---|---|---|
| NbBOI.1 | ATATGGCCATGGAGGCCAGTGAATTC ATGGCGGTTCAAGCTCAATACC | ATCTGCAGCTCGAGCTCGATGGATCC CTAACAAAGATAGACCTCAACGCTT |
| NbBOI.3 | ATATGGCCATGGAGGCCAGTGAATTC ATGGCCGTTCAAGCTCAATATGC | ATCTGCAGCTCGAGCTCGATGGATCC CTAACACAGAAAGCCCTCAACGC |
| NbBRG1.2 | ATATGGCCATGGAGGCCAGTGAATTCATGGCCATTCAAGCGCAGT | ATCTGCAGCTCGAGCTCGATGGATCC TCAAATCAAGGCCTCAATGGTAGC |
| NbBRG2.1 | ATATGGCCATGGAGGCCAGTGAATTC ATGGCTCTTCCTCATCACC | ATCTGCAGCTCGAGCTCGATGGATCC CTATATGTAAACTTCCATGCCTATATAC |
基因名 Gene name | 基因ID Gene ID | 氨基酸数 Protein length (aa) | 分子量 Molecular weight (kD) | 等电点 pI | 不稳定指数Instability index | 总平均亲水系数Grand average of hydropathicity | 脂肪指数 Aliphatic index | 亚细胞定位 Subcellular location |
|---|---|---|---|---|---|---|---|---|
| NbBRG2.1 | Niben101Scf04995g03009.1 | 337 | 38 311.27 | 5.23 | 48.590 | -0.597 | 83.89 | Nucleus |
| NbBRG2.2 | Niben101Scf05551g00006.1 | 579 | 65 546.86 | 6.53 | 43.220 | -0.643 | 75.82 | Nucleus |
| NbBRG2.3 | Niben101Scf09599g00015.1 | 361 | 40 133.59 | 7.49 | 36.470 | -0.487 | 81.52 | Nucleus |
| NbBRG2.4 | Niben101Scf00468g01013.1 | 347 | 38 916.32 | 7.88 | 39.380 | -0.493 | 74.21 | Nucleus |
| NbBRG3.1 | Niben101Scf25562g00007.1 | 327 | 35 967.52 | 5.71 | 49.640 | -0.343 | 84.16 | Nucleus |
| NbBRG3.2 | Niben101Scf02156g11001.1 | 349 | 38 832.05 | 5.89 | 43.620 | -0.464 | 77.91 | Nucleus |
| NbBRG3.3 | Niben101Scf04499g00008.1 | 350 | 39 076.28 | 5.9 | 47.580 | -0.476 | 78.26 | Nucleus |
| NbBRG3.4 | Niben101Scf01455g00005.1 | 431 | 47 614.25 | 5.09 | 41.520 | -0.472 | 77.42 | Nucleus |
| NbBRG1.1 | Niben101Scf06106g01018.1 | 283 | 32 891.4 | 5.92 | 63.690 | -0.632 | 77.53 | Nucleus |
| NbBRG1.2 | Niben101Scf07614g02001.1 | 269 | 31 231.47 | 7.92 | 72.720 | -0.715 | 65.32 | Nucleus |
| NbBOI.1 | Niben101Scf04159g01016.1 | 316 | 34 681.7 | 6.52 | 68.610 | -0.671 | 66.2 | Nucleus |
| NbBOI.2 | Niben101Scf01526g12008.1 | 338 | 37 295.86 | 6.24 | 64.790 | -0.501 | 79.44 | Nucleus |
| NbBOI.3 | Niben101Scf06847g02001.1 | 339 | 37 640.29 | 7.2 | 67.220 | -0.561 | 79.23 | Nucleus |
Table 3 Physicochemical properties of the NbBOIs
基因名 Gene name | 基因ID Gene ID | 氨基酸数 Protein length (aa) | 分子量 Molecular weight (kD) | 等电点 pI | 不稳定指数Instability index | 总平均亲水系数Grand average of hydropathicity | 脂肪指数 Aliphatic index | 亚细胞定位 Subcellular location |
|---|---|---|---|---|---|---|---|---|
| NbBRG2.1 | Niben101Scf04995g03009.1 | 337 | 38 311.27 | 5.23 | 48.590 | -0.597 | 83.89 | Nucleus |
| NbBRG2.2 | Niben101Scf05551g00006.1 | 579 | 65 546.86 | 6.53 | 43.220 | -0.643 | 75.82 | Nucleus |
| NbBRG2.3 | Niben101Scf09599g00015.1 | 361 | 40 133.59 | 7.49 | 36.470 | -0.487 | 81.52 | Nucleus |
| NbBRG2.4 | Niben101Scf00468g01013.1 | 347 | 38 916.32 | 7.88 | 39.380 | -0.493 | 74.21 | Nucleus |
| NbBRG3.1 | Niben101Scf25562g00007.1 | 327 | 35 967.52 | 5.71 | 49.640 | -0.343 | 84.16 | Nucleus |
| NbBRG3.2 | Niben101Scf02156g11001.1 | 349 | 38 832.05 | 5.89 | 43.620 | -0.464 | 77.91 | Nucleus |
| NbBRG3.3 | Niben101Scf04499g00008.1 | 350 | 39 076.28 | 5.9 | 47.580 | -0.476 | 78.26 | Nucleus |
| NbBRG3.4 | Niben101Scf01455g00005.1 | 431 | 47 614.25 | 5.09 | 41.520 | -0.472 | 77.42 | Nucleus |
| NbBRG1.1 | Niben101Scf06106g01018.1 | 283 | 32 891.4 | 5.92 | 63.690 | -0.632 | 77.53 | Nucleus |
| NbBRG1.2 | Niben101Scf07614g02001.1 | 269 | 31 231.47 | 7.92 | 72.720 | -0.715 | 65.32 | Nucleus |
| NbBOI.1 | Niben101Scf04159g01016.1 | 316 | 34 681.7 | 6.52 | 68.610 | -0.671 | 66.2 | Nucleus |
| NbBOI.2 | Niben101Scf01526g12008.1 | 338 | 37 295.86 | 6.24 | 64.790 | -0.501 | 79.44 | Nucleus |
| NbBOI.3 | Niben101Scf06847g02001.1 | 339 | 37 640.29 | 7.2 | 67.220 | -0.561 | 79.23 | Nucleus |
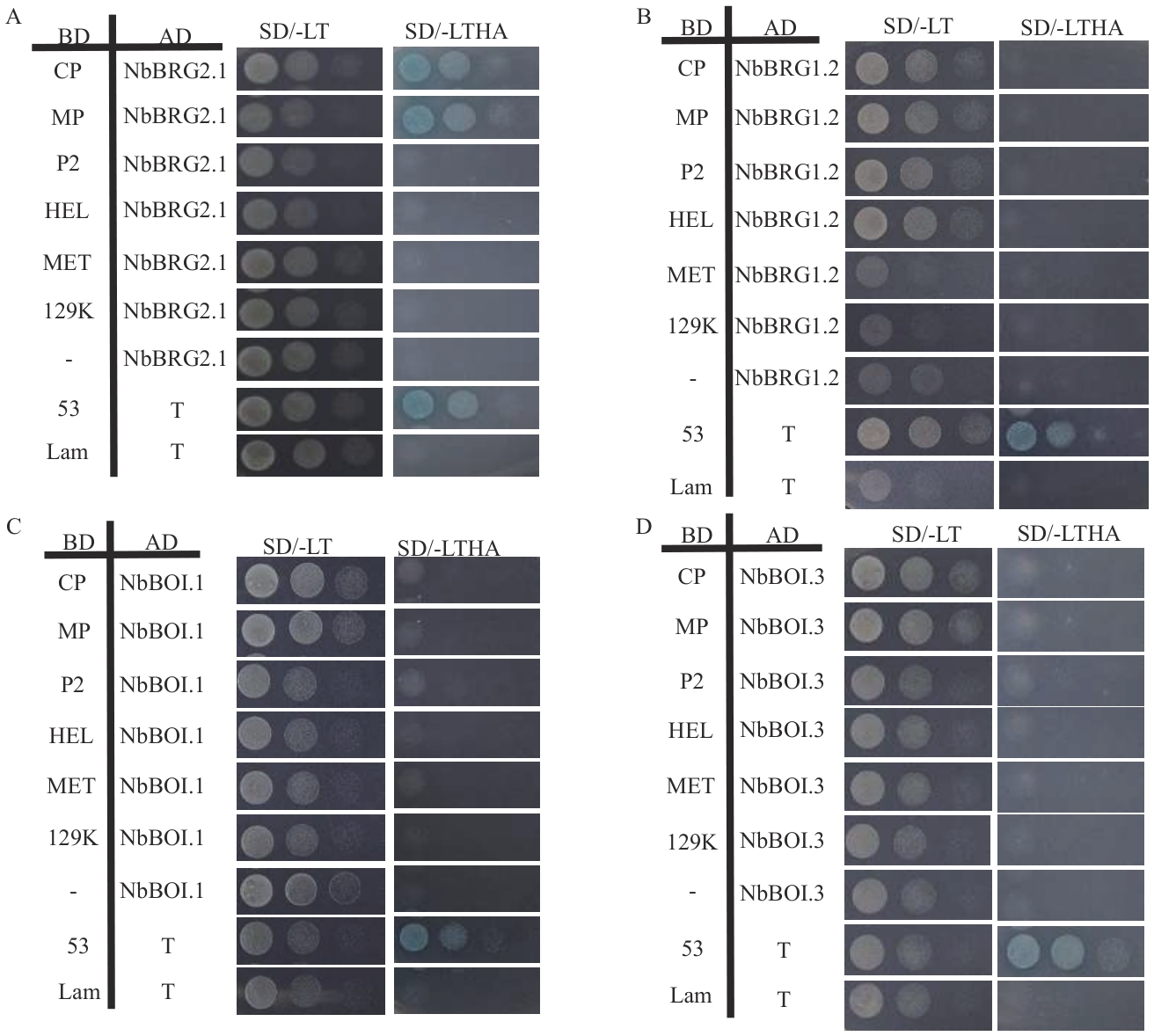
Fig. 8 Yeast two-hybrid assay for the interaction between NbBOIs and CGMMV encoded proteinsA: Verification of the interaction between NbBRG2.1 and CGMMV encoded proteins. B: Verification of the interaction between NbBRG1.2 and CGMMV encoded proteins. C: Verification of the interaction between NbBOI.1 and CGMMV encoded proteins. D: Verification of the interaction between NbBOI.3 and CGMMV encoded proteins
| [1] | Kawasaki T, Nam J, Boyes DC, et al. A duplicated pair of Arabidopsis RING-finger E3 ligases contribute to the RPM1- and RPS2-mediated hypersensitive response [J]. Plant J, 2005, 44(2): 258-270. |
| [2] | Li ZW, Huang JZ, Hu Y, et al. The RING-type E3 ligase BOI interacts with EXO70E2 and mediates its ubiquitination in Arabidopsis [J]. Life, 2024, 14(9): 1169. |
| [3] | Luo HL, Laluk K, Lai ZB, et al. The Arabidopsis botrytis Susceptible1 interactor defines a subclass of RING E3 ligases that regulate pathogen and stress responses [J]. Plant Physiol, 2010, 154(4): 1766-1782. |
| [4] | Sun C, Yao GF, Li LX, et al. E3 ligase BRG3 persulfidation delays tomato ripening by reducing ubiquitination of the repressor WRKY71 [J]. Plant Physiol, 2023, 192(1): 616-632. |
| [5] | 罗红丽, 宋凤鸣, Mengiste T. 拟南芥BOI基因在抗氧化胁迫和细胞死亡中的功能分析 [J]. 植物病理学报, 2012, 42(1): 65-72. |
| Luo HL, Song FM, Mengiste T. Functional analysis of Arabidopsis botrytis Susceptible 1 Interactor in oxidative stress and programmed cell death [J]. Acta Phytopathol Sin, 2012, 42(1): 65-72. | |
| [6] | 张萍. BnaBOI1与BnaPATL2互作调控油菜分枝数的分子机理研究 [D]. 重庆: 西南大学, 2022. |
| Zhang P. Molecular mechanism for regulating branch number in Brassica napus via interactions between BnaBOI1 and BnaPATL2 [D]. Chongqing: Southwest University, 2022. | |
| [7] | Huang JZ, Wu XQ, Gao ZY. The RING-type protein BOI negatively regulates the protein level of a CC-NBS-LRR in Arabidopsis [J]. Biochem Biophys Res Commun, 2021, 578: 104-109. |
| [8] | Nguyen KT, Park J, Park E, et al. The Arabidopsis RING domain protein BOI inhibits flowering via CO-dependent and CO-independent mechanisms [J]. Mol Plant, 2015, 8(12): 1725-1736. |
| [9] | Park J, Nguyen KT, Park E, et al. DELLA proteins and their interacting RING finger proteins repress gibberellin responses by binding to the promoters of a subset of gibberellin-responsive genes in Arabidopsis [J]. Plant Cell, 2013, 25(3): 927-943. |
| [10] | 陈智华, 乔振升, 李嘉其, 等. 滇杨TCP基因家族的全基因组鉴定与分析 [J]. 生物技术通报, 2024, 40(11): 214-226. |
| Chen ZH, Qiao ZS, Li JQ, et al. Genome-wide identification and analysis of the TCP gene family in Populus yunnanensis [J]. Biotechnol Bull, 2024, 40(11): 214-226. | |
| [11] | 吴浩, 董伟峰, 贺子天, 等. 水稻BXL基因家族的全基因组鉴定及表达分析 [J]. 生物技术通报, 2025(6): 87-98. |
| Wu H, Dong WF, He ZT, et al. Genome-wide identification and expression analysis of the Rice BXL Gene Family [J]. Biotechnology Bulletin, 2025(6): 87-98. | |
| [12] | Letunic I, Bork P. Interactive Tree of Life (iTOL) v6: recent updates to the phylogenetic tree display and annotation tool [J]. Nucleic Acids Res, 2024, 52(W1): W78-W82. |
| [13] | Ma WP, Liu X, Chen K, et al. Genome-wide re-identification and analysis of CrRLK1Ls in tomato [J]. Int J Mol Sci, 2023, 24(4): 3142. |
| [14] | 吴娅, 姚润, 杨含婷, 等. 凤梨薄荷SDR基因家族全基因组鉴定及表达分析 [J]. 生物技术通报, 2025, 41(5): 175-185. |
| Wu Y, Yao R, Yang HT, et al. Genome-wide identification and expression analysis of SDR gene family in Mentha suaveolens ‘variegata’ [J]. Biotechnol Bull, 2025, 41(5): 175-185. | |
| [15] | Rao SF, Wu XY, Zheng HY, et al. Genome-wide identification and analysis of Catharanthus roseus RLK1-like kinases in Nicotiana benthamiana [J]. BMC Plant Biol, 2021, 21(1): 425. |
| [16] | 于连伟, 姜兴林, 杨灵玲, 等. 转录因子NbERF RAP2-1在黄瓜绿斑驳花叶病毒侵染中的功能 [J]. 中国农业科学, 2023, 56(15): 2919-2928. |
| Yu LW, Jiang XL, Yang LL, et al. Function of transcription factor NbERF RAP2-1 in cucumber green mottle mosaic virus infection [J]. Sci Agric Sin, 2023, 56(15): 2919-2928. | |
| [17] | Yang X, Jiang XL, Fu H, et al. Cucumber green mottle mosaic virus coat protein hijacks mitochondrial ATPδ to promote viral infection [J]. Mol Plant Pathol, 2024, 25(11): e70034. |
| [18] | Huang JZ, Wu XQ, Gao ZY. A nucleocytoplasmic-localized E3 ligase affects the NLR receptor stability [J]. Biochem Biophys Res Commun, 2021, 583: 1-6. |
| [19] | Wang SS, Wang YJ. Harnessing hormone gibberellin knowledge for plant height regulation [J]. Plant Cell Rep, 2022, 41(10): 1945-1953. |
| [20] | Guo SQ, Chen YX, Ju YL, et al. Fine-tuning gibberellin improves rice alkali-thermal tolerance and yield [J]. Nature, 2025, 639(8053): 162-171. |
| [21] | 戴琪. 赤霉素结合2, 4-表油菜素内酯处理对缓解冷藏桃果实冷害的机制研究 [D]. 沈阳: 沈阳农业大学, 2023. |
| Dai Q. Research on the mechanism of gibberellin combined with 2,4 epibrassinolide treatment on alleviating cold damage of chilling peach fruits [D]. Shenyang: Shenyang Agricultural University, 2023. | |
| [22] | Huang JS, Xie B, Xian FJ, et al. Gibberellin signalling mediates nucleocytoplasmic trafficking of Sucrose Synthase 1 to regulate the drought tolerance in rice [J]. Plant Biotechnol J, 2025, 23(6): 1909-1926. |
| [23] | Bao SJ, Hua CM, Shen LS, et al. New insights into gibberellin signaling in regulating flowering in Arabidopsis [J]. J Integr Plant Biol, 2020, 62(1): 118-131. |
| [24] | Yimer HZ, Nahar K, Kyndt T, et al. Gibberellin antagonizes jasmonate-induced defense against Meloidogyne graminicola in rice [J]. New Phytol, 2018, 218(2): 646-660. |
| [25] | Li PB, Guo LM, Lang XY, et al. Geminivirus C4 proteins inhibit GA signaling via prevention of NbGAI degradation, to promote viral infection and symptom development in N. benthamiana [J]. PLoS Pathog, 2022, 18(4): e1010217. |
| [26] | Navarro L, Bari R, Achard P, et al. DELLAs control plant immune responses by modulating the balance of jasmonic acid and salicylic acid signaling [J]. Curr Biol, 2008, 18(9): 650-655. |
| [27] | 孔明悦. 柑橘木虱胁迫及喷施赤霉素和微肥对柑橘生理生化指标的影响 [D]. 佛山: 佛山科学技术学院, 2023. |
| Kong MY. Effects of Diaphorina citri,gibberellin and microfertilizer on hysiological and biochemical indices of citrus [D]. Foshan: Foshan University, 2023. | |
| [28] | Smalle J, Vierstra RD. The ubiquitin 26S proteasome proteolytic pathway [J]. Annu Rev Plant Biol, 2004, 55: 555-590. |
| [29] | Dong CH, Agarwal M, Zhang YY, et al. The negative regulator of plant cold responses, HOS1, is a RING E3 ligase that mediates the ubiquitination and degradation of ICE1 [J]. Proc Natl Acad Sci USA, 2006, 103(21): 8281-8286. |
| [30] | Ling KS, Li R, Zhang W. First report of Cucumber green mottle mosaic virus infecting greenhouse cucumber in Canada [J]. Plant Dis, 2014, 98(5): 701. |
| [31] | 姜兴林, 于连伟, 付涵, 等. 转录因子NbMYB1R1通过促进活性氧积累抑制病毒侵染 [J]. 中国农业科学, 2024, 57(8): 1490-1505. |
| Jiang XL, Yu LW, Fu H, et al. The transcription factor NbMYB1R1 inhibits viral infection by promoting ROS accumulation [J]. Sci Agric Sin, 2024, 57(8): 1490-1505. |
| [1] | LAI Shi-yu, LIANG Qiao-lan, WEI Lie-xin, NIU Er-bo, CHEN Ying-e, ZHOU Xin, YANG Si-zheng, WANG Bo. The Role of NbJAZ3 in the Infection of Nicotiana benthamiana by Alfalfa Mosaic Virus [J]. Biotechnology Bulletin, 2025, 41(8): 186-196. |
| [2] | HUANG Dan, PENG Bing-yang, ZHANG Pan-pan, JIAO Yue, LYU Jia-bin. Identification of HD-Zip Gene Family in Camellia oleifera and Analysis of Its Expression under Abiotic Stress [J]. Biotechnology Bulletin, 2025, 41(6): 191-207. |
| [3] | WANG Chen, LIU Guo-mei, CHEN Chang, ZHANG Jin-long, YAO Lin, SUN Xuan, DU Chun-fang. Genome-wide Identification and Expression Analysis of CCDs Family in Brassia rapa L. [J]. Biotechnology Bulletin, 2025, 41(3): 161-170. |
| [4] | CHEN Zhi-hua, QIAO Zhen-sheng, LI Jia-qi, ZHANG Xiao-lin, MA Shao-jie, HE Cheng-zhong, ZONG Dan. Genome-wide Identification and Analysis of the TCP Gene Family in Populus yunnanensis [J]. Biotechnology Bulletin, 2024, 40(11): 214-226. |
| [5] | SA Shi-juan, WU Han-yu, WEN Yuan, CHEN Xue-na, ZHENG Rui, YAO Xin-ling. Responses of Choloroplast Specific Protein Profile to Different Stomatal Densities in Nicotiana benthamiana [J]. Biotechnology Bulletin, 2023, 39(2): 193-202. |
| [6] | ZHANG Rui-ping, YANG Feng, CHEN Le-zhang, DENG Xing-guang, ZHANG Da-wei. Role of Alternative Respiratory Pathway in Brassinosteroids Inducing Heat Stress Response in Nicotiana benthamiana [J]. Biotechnology Bulletin, 2020, 36(10): 8-14. |
| [7] | HU Yu, WANG Ling, BI Wu, TAN Lin, XING Dan, ZUO Fu-yuan. Research Progress of miR-1246 in Diseases [J]. Biotechnology Bulletin, 2017, 33(3): 29-36. |
| [8] | Liu Rongdiao, Ruan Lingwei. PI3K-Akt Signaling Pathway and Viral Infection [J]. Biotechnology Bulletin, 2013, 0(6): 53-62. |
| [9] | Wang Linmei, Yue Dongmei, Li Shuying, Fan Qi, Ye Bo, Zhao Zhenjun, Zhang Bo. Study on Infection of Pupal Ovaries Cells of Antheraea pernyi with ApNPV [J]. Biotechnology Bulletin, 2013, 0(6): 172-176. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||