








• 研究报告 • 下一篇
郭苗( ), 许家佳, 孙天国, 蔡璨, 曹婉笛, 包纪星, 沙伟, 张梅娟, 彭疑芳, 马天意(
), 许家佳, 孙天国, 蔡璨, 曹婉笛, 包纪星, 沙伟, 张梅娟, 彭疑芳, 马天意( )
)
收稿日期:2025-09-28
出版日期:2026-03-09
通讯作者:
马天意,男,博士,副教授,研究方向 :植物逆境分子生物学;E-mail: tyma@qqhru.edu.cn作者简介:郭苗,女,硕士研究生,研究方向 :植物逆境分子生物学;E-mail: 1066602752@qq.com
基金资助:
GUO Miao( ), XU Jia-jia, SUN Tian-guo, CAI Can, CAO Wan-di, BAO Ji-xing, SHA Wei, ZHANG Mei-juan, PENG Yi-fang, MA Tian-yi(
), XU Jia-jia, SUN Tian-guo, CAI Can, CAO Wan-di, BAO Ji-xing, SHA Wei, ZHANG Mei-juan, PENG Yi-fang, MA Tian-yi( )
)
Received:2025-09-28
Published:2026-03-09
摘要:
目的 作为油体中含量最丰富的表面蛋白,油体蛋白(oleosin, OLEO, OLE)对植物油体形成及油脂储存至关重要,同时OLEO在植物应对逆境胁迫过程中也发挥重要作用。砂藓(Racomitrium canescens)是一种拥有强耐脱水性和耐高温性的典型旱生苔藓植物,通过验证砂藓中响应胁迫处理的油体蛋白基因RcOLEO1能否提高植物抗逆性,为进一步阐明砂藓耐旱及耐高温过程的分子机制,以及挖掘砂藓相关抗旱基因资源提供理论依据。 方法 利用实时荧光定量PCR(real-time quantitative PCR, RT-qPCR)检测复水、脱水和高温胁迫处理下砂藓中RcOLEO1的表达模式,对RcOLEO1的编码序列进行克隆并构建其过表达拟南芥(Arabidopsis thaliana)株系,对转基因拟南芥分别进行干旱及高温胁迫处理,观察表型,并对生理生化指标进行测定。 结果 RT-qPCR结果显示RcOLEO1响应砂藓的复水、脱水和高温胁迫处理;成功克隆得到RcOLEO1的编码序列,获得T2代过表达转基因拟南芥株系;对野生型及过表达转基因拟南芥进行高温及干旱胁迫处理,发现过表达砂藓RcOLEO1增强了拟南芥的耐旱性及耐高温性;对渗透调节物质含量、丙二醛含量、叶绿素含量等生理生化指标进行检测,初步阐释了过表达RcOLEO1增强拟南芥植株的耐旱性及耐高温性的生理响应方式。 结论 过表达RcOLEO1可能通过调控渗透调节物质积累与抗氧化酶活性,增强植物对干旱和高温胁迫的耐受性。
郭苗, 许家佳, 孙天国, 蔡璨, 曹婉笛, 包纪星, 沙伟, 张梅娟, 彭疑芳, 马天意. 过表达砂藓RcOLEO1基因增强拟南芥的耐旱性及耐高温性[J]. 生物技术通报, doi: 10.13560/j.cnki.biotech.bull.1985.2025-1043.
GUO Miao, XU Jia-jia, SUN Tian-guo, CAI Can, CAO Wan-di, BAO Ji-xing, SHA Wei, ZHANG Mei-juan, PENG Yi-fang, MA Tian-yi. Overexpression of RcOLEO1 Enhanced the Tolerance of Arabidopsis thaliana toDrought and High-temperature Stress[J]. Biotechnology Bulletin, doi: 10.13560/j.cnki.biotech.bull.1985.2025-1043.
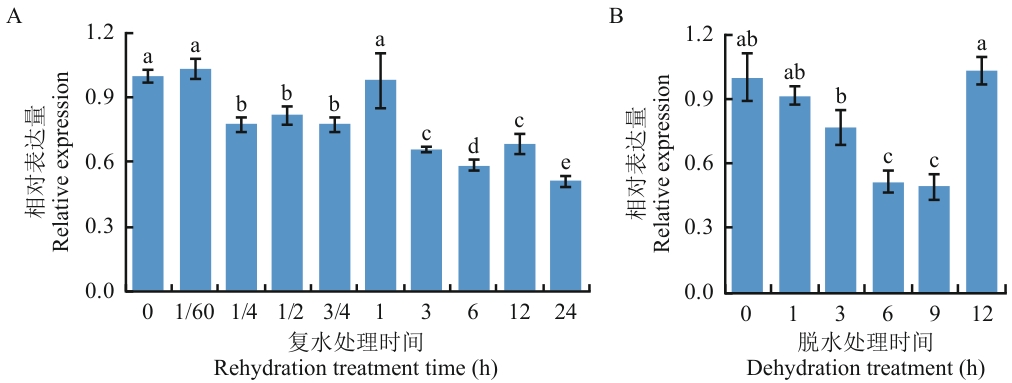
图1 砂藓复水及脱水处理过程中RcOLEO1的表达分析误差线表示标准偏差,误差线上字母表示不同的差异显著性(P<0.05),下同
Fig. 1 Expression analysis of RcOLEO1 during rehydration and dehydration treatments of Racomitrium canescensThe error bars indicates the standard deviation, and the letters on the error bars indicate different significance (P<0.05), the same below
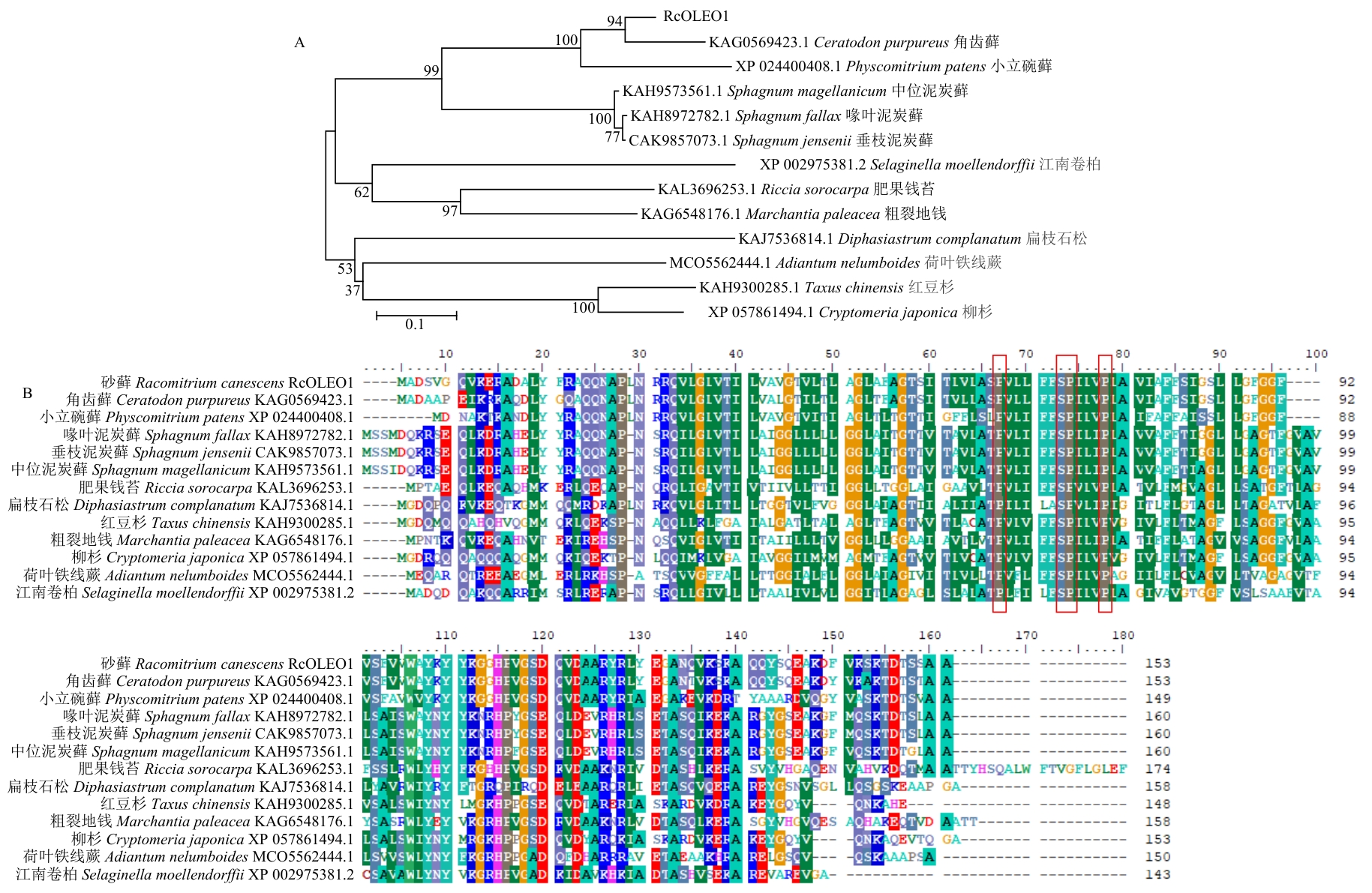
图2 RcOLEO1与近缘蛋白质的进化关系分析(A)及序列比对(B)红框内为脯氨酸结motif
Fig. 2 Phylogenic analysis (A) and sequence alignment (B) of RcOLEO1 and homologous proteinsThe red boxes indicate the conserved proline knot motif
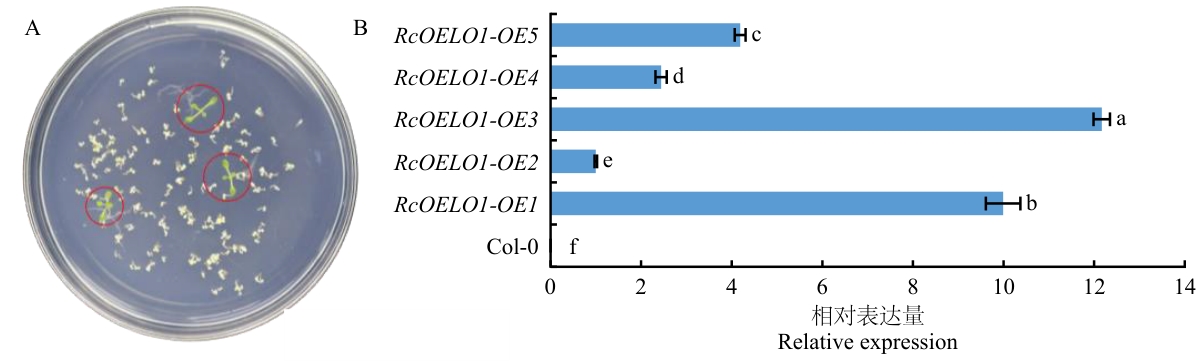
图3 RcOLEO1过表达转基因拟南芥的筛选及鉴定A:RcOLEO1过表达转基因拟南芥T1代植株的筛选;B:RcOLEO1过表达转基因拟南芥T2代植株的鉴定
Fig. 3 Screening and characterization of RcOLEO1 overexpression transgenic A. thalianaA: Screening of T1 generation plants of transgenic A. thaliana overexpressing RcOLEO1.B: Identification of T2RcOLEO1 overexpression transgenic A. thaliana
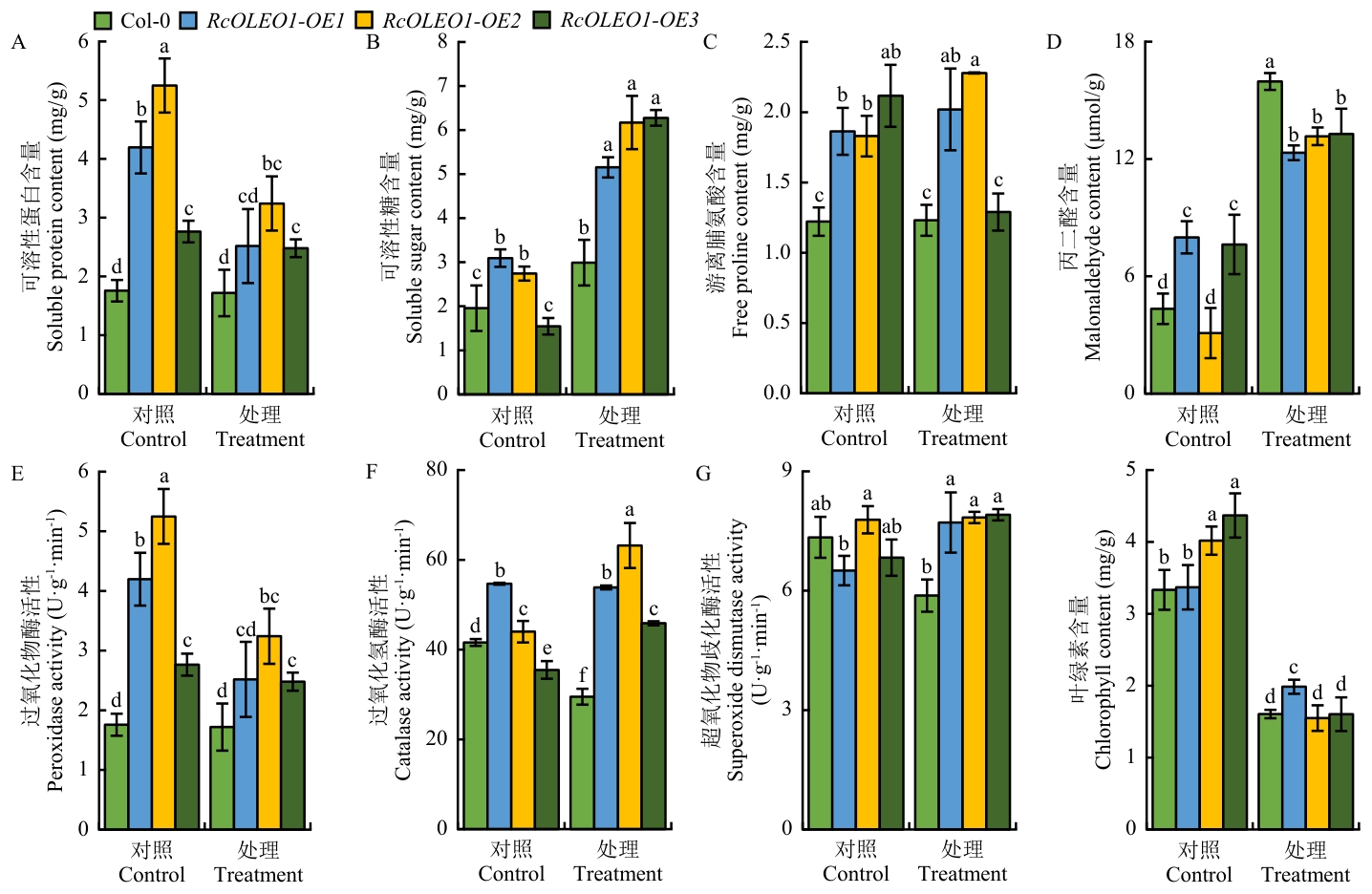
图5 干旱处理下野生型和RcOLEO1过表达转基因拟南芥植株生理生化指标的变化
Fig. 5 Changes in physiological and biochemical indexes of wild-type and RcOLEO1 overexpression transgenic A. thaliana under drought treatment
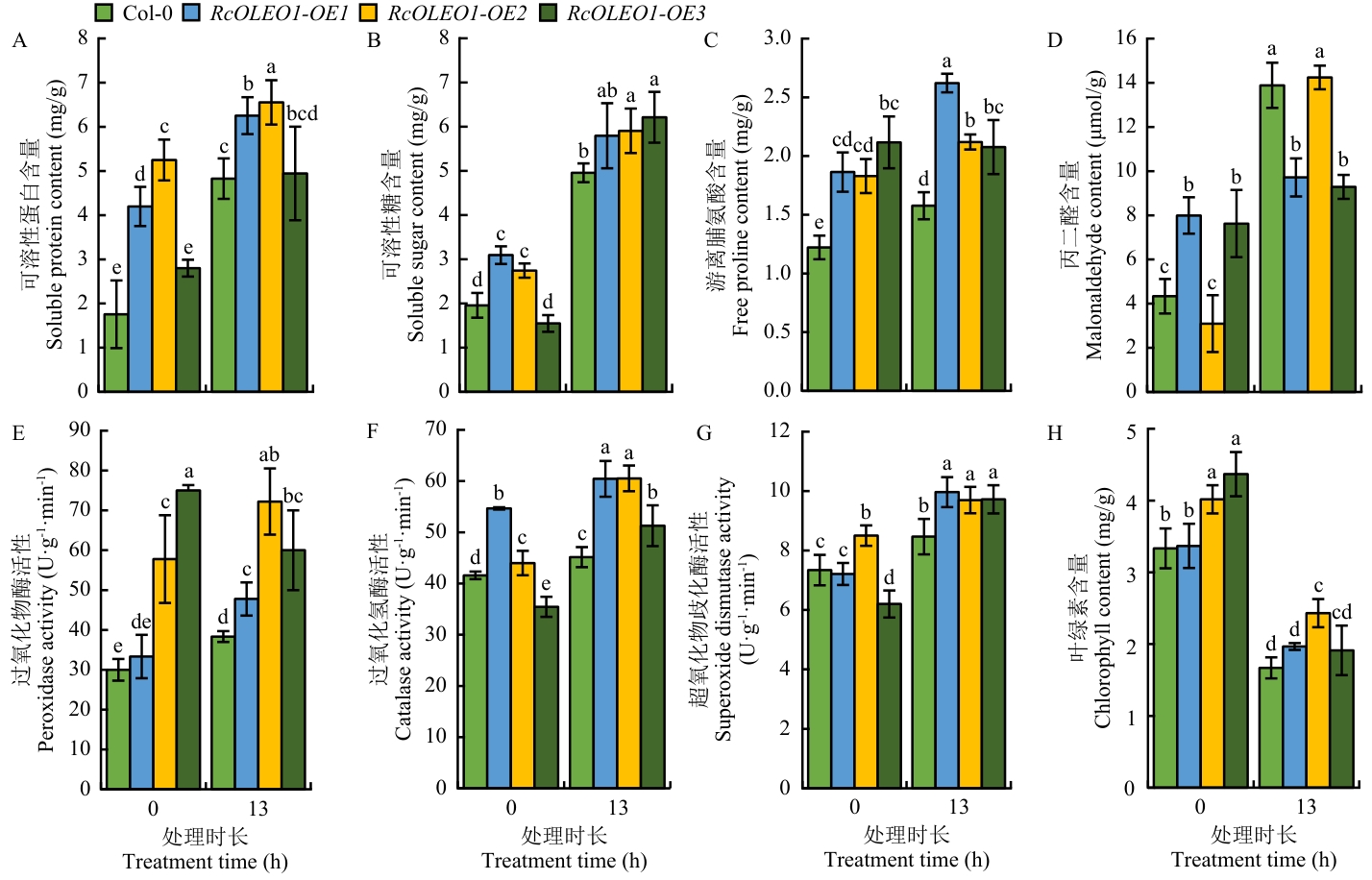
图7 45 ℃高温处理下野生型和RcOLEO1过表达转基因拟南芥植株生理生化指标变化
Fig. 7 Changes in physiological and biochemical indexes of wild-type and RcOLEO1 overexpression transgenic A. thaliana under 45 °C high-temperature treatment
| [1] | 祝亚丽, 刘洋, 孔祥慧, 等. 中国高温、干旱及其复合事件的研究进展和展望 [J]. 大气科学学报, 2025, 48(1): 26-36. |
| Zhu YL, Liu Y, Kong XH, et al. Research progress and prospect on the drought, heatwave, and compound drought and heatwave events in China [J]. Trans Atmos Sci, 2025, 48(1): 26-36. | |
| [2] | Wood AJ. The nature and distribution of vegetative desiccation-tolerance in hornworts, liverworts and mosses [J]. Bryologist, 2007, 110(2): 163-177. |
| [3] | 宋松泉, 刘军, 唐翠芳, 等. 种子耐脱水性的生理及分子机制研究进展 [J]. 中国农业科学, 2022, 55(6): 1047-1063. |
| Song SQ, Liu J, Tang CF, et al. Research progress on the physiology and its molecular mechanism of seed desiccation tolerance [J]. Sci Agric Sin, 2022, 55(6): 1047-1063. | |
| [4] | 李雯煜. 砂藓应答60 ℃高温胁迫的生理生化和蛋白质组学研究 [D]. 齐齐哈尔: 齐齐哈尔大学, 2019. |
| Li WY. Physiological, biochemical and proteomics research of Racomitrium canescens in response to 60 ℃ high temperature stresses [D]. Qiqihar: Qiqihar University, 2019. | |
| [5] | 赵浩强, 王小斐, 高少培. 植物油体蛋白基因家族研究进展 [J]. 遗传, 2022, 44(12): 1128-1140. |
| Zhao HQ, Wang XF, Gao SP. Progress on the functional role of oleosin gene family in plants [J]. Hereditas: Beijing, 2022, 44(12): 1128-1140. | |
| [6] | 胡佳, 刘春林. 植物油体研究进展 [J]. 植物学报, 2017, 52(5): 669-679. |
| Hu J, Liu CL. Research advances in plant oil body [J]. Chin Bull Bot, 2017, 52(5): 669-679. | |
| [7] | Huang AHC. Oleosins and oil bodies in seeds and other organs [J]. Plant Physiol, 1996, 110(4): 1055-1061. |
| [8] | Fang Y, Zhu RL, Mishler BD. Evolution of oleosin in land plants [J]. PLoS One, 2014, 9(8): e103806. |
| [9] | Tzen JT, Huang AH. Surface structure and properties of plant seed oil bodies [J]. J Cell Biol, 1992, 117(2): 327-335. |
| [10] | Shao Q, Liu XF, Su T, et al. New insights into the role of seed oil body proteins in metabolism and plant development [J]. Front Plant Sci, 2019, 10: 1568. |
| [11] | Huang AHC. Plant lipid droplets and their associated proteins: potential for rapid advances [J]. Plant Physiol, 2018, 176(3): 1894-1918. |
| [12] | Huang MD, Huang AHC. Bioinformatics reveal five lineages of oleosins and the mechanism of lineage evolution related to structure/function from green algae to seed plants [J]. Plant Physiol, 2015, 169(1): 453-470. |
| [13] | Zhang D, Zhang HY, Hu ZB, et al. Artificial selection on GmOLEO1 contributes to the increase in seed oil during soybean domestication [J]. PLoS Genet, 2019, 15(7): e1008267. |
| [14] | 陈晨, 王会伟, 李春鑫, 等. 油莎豆油体蛋白基因Oleosin的全基因组鉴定及功能分析 [J]. 植物遗传资源学报, 2025, 26(5): 1017-1030. |
| Chen C, Wang HW, Li CX, et al. Genome-wide identification and functional analysis of Oleosin genes in Cyperus esculentus L [J]. J Plant Genet Resour, 2025, 26(5): 1017-1030. | |
| [15] | Hu ML, Wu J, Xue XM, et al. Genome-wide identification and functional characterization of oleosin genes in peanut (Arachis hypogaea L.) [J]. Front Plant Sci, 2025, 16: 1623513. |
| [16] | 蒋茂双, 元香梅, 刘晓东, 等. 谷子Oleosin基因家族及其对干旱响应的分析 [J]. 山西农业大学学报: 自然科学版, 2018, 38(1): 16-20. |
| Jiang MS, Yuan XM, Liu XD, et al. Oleosin gene family of millet and its response to drought [J]. J Shanxi Agric Univ Nat Sci Ed, 2018, 38(1): 16-20. | |
| [17] | 郭新亚. 大豆油体蛋白基因GmOLE1和GmOLE2功能的初步研究 [D]. 南京: 南京农业大学, 2020. |
| Guo XY. Study on the functions of oleosin genes GmOLE1 and GmOLE2 from soybean [D]. Nanjing: Nanjing Agricultural University, 2020. | |
| [18] | 郭新亚, 章文华, 林峰. 大豆油体蛋白基因GmOLE2的功能分析 [J]. 南京农业大学学报, 2021, 44(3): 477-486. |
| Guo XY, Zhang WH, Lin F. Functional analysis of soybean oleosin gene GmOLE2 [J]. J Nanjing Agric Univ, 2021, 44(3): 477-486. | |
| [19] | 何汉琼, 姜洪进, 张文婧, 等. 蓖麻油体蛋白基因家族鉴定和盐碱胁迫响应分析 [J]. 种子, 2024, 43(8): 103-111. |
| He HQ, Jiang HJ, Zhang WJ, et al. Identification for oleosin gene family and analysis on its saline and alkaline stress response in Ricinus communis L [J]. Seed, 2024, 43(8): 103-111. | |
| [20] | Peng YF, Ma TY, Wang X, et al. Proteomic and transcriptomic responses of the desiccation-tolerant moss Racomitrium canescens in the rapid rehydration processes [J]. Genes, 2023, 14(2): 390. |
| [21] | 张梅娟, 沙伟. 东亚砂藓组织培养技术方法研究 [J]. 植物科学学报, 2013, 31(6): 616-622. |
| Zhang MJ, Sha W. Research on tissue culture technology of Racomitrium japonicum [J]. Plant Sci J, 2013, 31(6): 616-622. | |
| [22] | 沙伟, 王鑫, 张梅娟, 等. 砂藓抗高温相关基因RcDAPDC的克隆及表达分析 [J]. 分子植物育种, 2020, 18(24): 8092-8098. |
| Sha W, Wang X, Zhang MJ, et al. Cloning and expression analysis of high-temperature resistance related gene RcDAPDC in Racomitrium canescens [J]. Mol Plant Breed, 2020, 18(24): 8092-8098. | |
| [23] | 赵继发. 砂藓脱水胁迫响应查尔酮合成酶基因的克隆和抗逆性分析 [D]. 齐齐哈尔: 齐齐哈尔大学, 2023. |
| Zhao JF. Cloning and stress resistance analysis of RcCHS genes in response to dehydration stress in Racomitrium canescens [D]. Qiqihar: Qiqihar University, 2023. | |
| [24] | 唐宽宇. 砂藓耐脱水性相关基因RcTRX1、RcTRX2的克隆及抗逆性分析 [D]. 齐齐哈尔: 齐齐哈尔大学, 2023. |
| Tang KY. Cloning and stress resistance analysis of dehydration resistance related genes RcTRX1 and RcTRX2 of Racomitrium canescens [D]. Qiqihar: Qiqihar University, 2023. | |
| [25] | 马天意, 张时通, 朱巍巍, 等. 砂藓RcPLD基因的抗旱功能分析 [J]. 西北植物学报, 2021, 41(7): 1091-1100. |
| Ma TY, Zhang ST, Zhu WW, et al. Analysis of drought-resistance function of RcPLD gene from Racomitrium canescens [J]. Acta Bot Boreali Occidentalia Sin, 2021, 41(7): 1091-1100. | |
| [26] | 张春蕾, 沙伟, 张梅娟, 等. 砂藓生长素受体基因RcTIR1的克隆及表达分析 [J]. 基因组学与应用生物学, 2015, 34(1): 100-105. |
| Zhang CL, Sha W, Zhang MJ, et al. Cloning and expression analysis of auxin receptor gene RcTIR1 in Racomitrium canescens [J]. Genom Appl Biol, 2015, 34(1): 100-105. | |
| [27] | 马天意, 许家佳, 路文婧, 等. ‘金小童’大白菜BrcGASA3基因在盐碱胁迫下的表达分析及抗性鉴定 [J]. 生物技术通报, 2025, 41(2): 127-138. |
| Ma TY, Xu JJ, Lu WJ, et al. Expression analysis and resistance identification of BrcGASA3 in Chinese cabbage ‘Jinxiaotong’ cultivar under saline-alkali stress [J]. Biotechnol Bull, 2025, 41(2): 127-138. | |
| [28] | 王艺瑾. 油莎豆油体蛋白基因CeOle1和CeOle4克隆及功能研究 [D]. 长春: 吉林农业大学, 2023. |
| Wang YJ. Cloning and functional research of oleosin genes CeOle1 and CeOle4 in Cyperus esculentusis [D]. Changchun: Jilin Agricultural University, 2023. | |
| [29] | Abell BM, Holbrook LA, Abenes M, et al. Role of the proline knot motif in oleosin endoplasmic reticulum topology and oil body targeting [J]. Plant Cell, 1997, 9(8): 1481-1493. |
| [30] | 党瑗, 李维, 苗向, 等. 山杏油体蛋白基因PsOLE4克隆及其调控油脂累积功能分析 [J]. 生物技术通报, 2022, 38(11): 151-161. |
| Dang Y, Li W, Miao X, et al. Cloning of oleosin gene PsOLE4 from Prunus sibirica and its regulatory function analysis for oil accumulation [J]. Biotechnol Bull, 2022, 38(11): 151-161. | |
| [31] | 姜焕焕, 温思慧, 卢雨庭, 等. 二倍体野生种花生Oleosin基因家族全基因组鉴定及其表达分析 [J]. 中国油料作物学报, 2025, 47(1): 94-104. |
| Jiang HH, Wen SH, Lu YT, et al. Genome-wide analysis and stress-responsive expression profiling of the Oleosin gene family in diploid wild species Arachis duranensis and Arachis ipaensis [J]. Chin J Oil Crop Sci, 2025, 47(1): 94-104. | |
| [32] | Khan I, Awan SA, Rizwan M, et al. Silicon nanoparticles improved the osmolyte production, antioxidant defense system, and phytohormone regulation in Elymus sibiricus (L.) under drought and salt stress [J]. Environ Sci Pollut Res Int, 2024, 31(6): 8985-8999. |
| [33] | Wu YQ, Guo XY, Zhou XH, et al. Effects of drought stress on physiological properties of Gazania rigens L [J]. Adv Mater Res, 2014, 955-959: 217-221. |
| [34] | 谢亚军, 王兵, 梁新华, 等. 干旱胁迫对甘草幼苗活性氧代谢及保护酶活性的影响 [J]. 农业科学研究, 2008, 29(4): 19-22. |
| Xie YJ, Wang B, Liang XH, et al. Effect of drought stress on active oxygen metabolism and activities of protective enzymes of licorice seedlings [J]. J Agric Sci, 2008, 29(4): 19-22. | |
| [35] | 王亚凯. 多尺度叶绿素荧光与光合干旱胁迫响应机理及作物模型融合方法 [D]. 杨凌: 西北农林科技大学, 2023. |
| Wang YK. Multi-scale chlorophyll fluorescence and photosynthetic response mechanisms to drought stress and integration methods with crop model [D]. Yangling: Northwest A & F University, 2023. | |
| [36] | 张永超, 梁国玲, 秦燕, 等. 老芒麦衰老过程中叶片叶绿素和光合作用变化特征及对养分的响应 [J]. 草业学报, 2022, 31(1): 229-237. |
| Zhang YC, Liang GL, Qin Y, et al. Characteristics of chlorophyll and photosynthesis in leaves and their response to nutrients during aging of Elymus sibiricus [J]. Acta Prataculturae Sin, 2022, 31(1): 229-237. |
| [1] | 吴翠翠, 陈登科, 兰刚, 夏芝, 李朋波. 花生转录因子AhHDZ70的生物信息学分析及耐盐耐旱性研究[J]. 生物技术通报, 2026, 42(1): 198-207. |
| [2] | 倪莹, 李雷, 汪进萱, 马波, 孟昕, 冷平生, 吴静, 胡增辉. 紫丁香So4CL的克隆及功能分析[J]. 生物技术通报, 2026, 42(1): 139-149. |
| [3] | 陈静欢, 房国楠, 朱文豪, 叶广继, 苏旺, 贺苗苗, 杨生龙, 周云. 马铃薯种质资源淀粉表征及相关基因表达分析[J]. 生物技术通报, 2026, 42(1): 170-183. |
| [4] | 吕呈聪, 衡蒙, 陈思琪, 金雪花. 彩色马蹄莲花青素苷转运相关ZhGSTF的克隆及功能分析[J]. 生物技术通报, 2026, 42(1): 161-169. |
| [5] | 杨娟, 冯慧, 吉乃喆, 孙丽萍, 王赟, 张佳楠, 赵世伟. 月季AP2/ERF转录因子RcERF4和RcRAP2-12的克隆及功能分析[J]. 生物技术通报, 2026, 42(1): 150-160. |
| [6] | 陈强, 于璎霏, 张颖, 张冲. 茉莉酸甲酯对薄皮甜瓜‘绿宝石’采后冷害的调控[J]. 生物技术通报, 2025, 41(9): 105-114. |
| [7] | 程婷婷, 刘俊, 王利丽, 练从龙, 魏文君, 郭辉, 吴尧琳, 杨晶凡, 兰金旭, 陈随清. 杜仲查尔酮异构酶基因家族全基因组鉴定及其表达模式分析[J]. 生物技术通报, 2025, 41(9): 242-255. |
| [8] | 徐小萍, 杨成龙, 和兴, 郭文杰, 吴健, 方少忠. 百合LoAPS1克隆及其在休眠解除过程的功能分析[J]. 生物技术通报, 2025, 41(9): 195-206. |
| [9] | 董向向, 缪百灵, 许贺娟, 陈娟娟, 李亮杰, 龚守富, 朱庆松. 森林草莓FveBBX20基因的生物信息学分析及开花调控功能[J]. 生物技术通报, 2025, 41(9): 115-123. |
| [10] | 李珊, 马登辉, 马红义, 姚文孔, 尹晓. 葡萄SKP1基因家族鉴定与表达分析[J]. 生物技术通报, 2025, 41(9): 147-158. |
| [11] | 巩慧玲, 邢玉洁, 马俊贤, 蔡霞, 冯再平. 马铃薯LAC基因家族的鉴定及盐胁迫下表达分析[J]. 生物技术通报, 2025, 41(9): 82-93. |
| [12] | 赖诗雨, 梁巧兰, 魏列新, 牛二波, 陈应娥, 周鑫, 杨思正, 王博. NbJAZ3在苜蓿花叶病毒侵染本氏烟过程中的作用[J]. 生物技术通报, 2025, 41(8): 186-196. |
| [13] | 腊贵晓, 赵玉龙, 代丹丹, 余永亮, 郭红霞, 史贵霞, 贾慧, 杨铁钢. 红花质膜H+-ATPase基因家族成员鉴定及响应低氮低磷胁迫的表达分析[J]. 生物技术通报, 2025, 41(8): 220-233. |
| [14] | 任睿斌, 司二静, 万广有, 汪军成, 姚立蓉, 张宏, 马小乐, 李葆春, 王化俊, 孟亚雄. 大麦条纹病菌GH17基因家族的鉴定及表达分析[J]. 生物技术通报, 2025, 41(8): 146-154. |
| [15] | 程雪, 付颖, 柴晓娇, 王红艳, 邓欣. 谷子LHC基因家族鉴定及非生物胁迫表达分析[J]. 生物技术通报, 2025, 41(8): 102-114. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||