








生物技术通报 ›› 2024, Vol. 40 ›› Issue (2): 1-8.doi: 10.13560/j.cnki.biotech.bull.1985.2023-0963
• 特约综述 • 下一篇
收稿日期:2023-10-18
出版日期:2024-02-26
发布日期:2024-03-13
作者简介:陈艳梅,博士,副教授,研究方向:植物蛋白质组学;E-mail: chenyanmei@cau.edu.cn
基金资助:Received:2023-10-18
Published:2024-02-26
Online:2024-03-13
摘要:
蛋白质的翻译后修饰在细胞信号转导过程中起核心调控作用。除了单一翻译后修饰的调控外,细胞中还存在由多个翻译后修饰共同调控一些关键的代谢或信号通路的分子机制。翻译后修饰是蛋白质之间的语言,通过不同修饰之间的相互作用和协同调控,把复杂的信号整合后传递到下游其他蛋白并形成信号网络,最终影响蛋白质的结构、亚细胞定位、相互作用以及生物学功能。本文概述磷酸化、泛素化和类泛素化三类重要的翻译后修饰在植物细胞信号通路中的串扰作用模式及协同调控机制。
陈艳梅. 蛋白质翻译后修饰之间的互作关系及其协同调控机理[J]. 生物技术通报, 2024, 40(2): 1-8.
CHEN Yan-mei. Crosstalk Between Different Post-translational Modifications and Its Regulatory Mechanisms in Plant Growth and Development[J]. Biotechnology Bulletin, 2024, 40(2): 1-8.
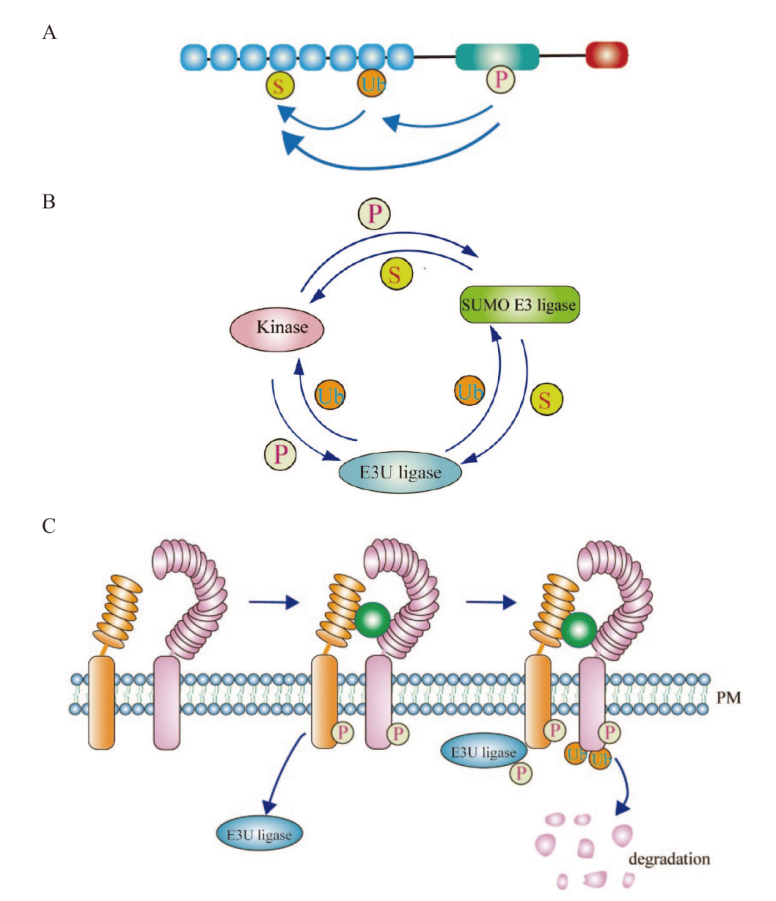
图1 “磷酸化-泛素化-类泛素化”之间的串扰调控机制 A:“磷酸化-泛素化-类泛素化”在同一个蛋白的不同氨基酸残基上发生修饰,不同修饰之间通过互相拮抗或促进的方式共同调控蛋白质的分子功能;B:“磷酸化-泛素化-类泛素化”三类修饰通过以不同蛋白或酶相互作为底物进行修饰,形成正调控或负反馈调控环而发挥作用;C:“磷酸化-泛素化”协同调控受体激酶信号通路。P:磷酸化;S:类泛素化;Ub:泛素化; PM:细胞质膜
Fig. 1 Crosstalk and interaction among “Phosphorylation-ubiquitination-sumoylation” A: “Phosphorylation-ubiquitination-sumoylation” function at different amino acid residues of the same protein. The molecular function of different modifications among proteins is jointly regulated through mutual antagonism or promotion. B: “Phosphorylation-ubiquitination-sumoylation” are the modifications conducted through the use of different proteins or enzymes as substrates. C: The signaling processes of receptor-like kinase are co-regulated by “phosphorylation-ubiquitination”. P: Phosphorylation; S: sumoylation; Ub: ubiquitination; PM: plasma membrane
| [1] |
Chen YM, Wang Y, Yang J, et al. Exploring the diversity of plant proteome[J]. J Integr Plant Biol, 2021, 63(7): 1197-1210.
doi: 10.1111/jipb.13087 |
| [2] |
Chen YM, Hoehenwarter W. Rapid and reproducible phosphopeptide enrichment by tandem metal oxide affinity chromatography: application to boron deficiency induced phosphoproteomics[J]. Plant J, 2019, 98(2): 370-384.
doi: 10.1111/tpj.2019.98.issue-2 URL |
| [3] |
Consortium TU. UniProt: the universal protein knowledgebase[J]. Nucleic Acids Res, 2017, 45(D1): D158-D169.
doi: 10.1093/nar/gkw1099 URL |
| [4] |
Willems P, Horne A, Van Parys T, et al. The Plant PTM Viewer, a central resource for exploring plant protein modifications[J]. Plant J, 2019, 99(4): 752-762.
doi: 10.1111/tpj.v99.4 URL |
| [5] |
Millar AH, Heazlewood JL, Giglione C, et al. The scope, functions, and dynamics of posttranslational protein modifications[J]. Annu Rev Plant Biol, 2019, 70: 119-151.
doi: 10.1146/annurev-arplant-050718-100211 pmid: 30786234 |
| [6] |
Chen YM, Wang Y. Mapping histone modification-dependent protein interactions with chemical proteomics[J]. Trends Biochem Sci, 2022, 47(3): 189-193.
doi: 10.1016/j.tibs.2021.11.002 URL |
| [7] |
Zhang Y, Zeng LR. Crosstalk between ubiquitination and other post-translational protein modifications in plant immunity[J]. Plant Commun, 2020, 1(4): 100041.
doi: 10.1016/j.xplc.2020.100041 URL |
| [8] |
Chen YM, Hoehenwarter W. Changes in the phosphoproteome and metabolome link early signaling events to rearrangement of photosynthesis and central metabolism in salinity and oxidative stress response in Arabidopsis[J]. Plant Physiol, 2015, 169(4): 3021-3033.
doi: 10.1104/pp.15.01486 URL |
| [9] |
Barbour H, Nkwe NS, Estavoyer B, et al. An inventory of crosstalk between ubiquitination and other post-translational modifications in orchestrating cellular processes[J]. iScience, 2023, 26(5): 106276.
doi: 10.1016/j.isci.2023.106276 URL |
| [10] |
Wang X, Ding YL, Li ZY, et al. PUB25 and PUB26 promote plant freezing tolerance by degrading the cold signaling negative regulator MYB15[J]. Dev Cell, 2019, 51(2): 222-235.e5.
doi: S1534-5807(19)30691-4 pmid: 31543444 |
| [11] |
Liu JH, Wu XY, Fang Y, et al. A plant RNA virus inhibits NPR1 sumoylation and subverts NPR1-mediated plant immunity[J]. Nat Commun, 2023, 14(1): 3580.
doi: 10.1038/s41467-023-39254-2 pmid: 37328517 |
| [12] | 高丰衣, 陈艳梅. 质谱技术探索蛋白磷酸化信号机制的研究进展[J]. 科学通报, 2021, 66(20): 2529-2541. |
|
Gao FY, Chen YM. Decoding protein phosphorylation signaling networks by mass spectrometry[J]. Chin Sci Bull, 2021, 66(20): 2529-2541.
doi: 10.1360/TB-2020-1088 URL |
|
| [13] |
Chen XX, Ding YL, Yang YQ, et al. Protein kinases in plant responses to drought, salt, and cold stress[J]. J Integr Plant Biol, 2021, 63(1): 53-78.
doi: 10.1111/jipb.13061 |
| [14] |
Bhaskara GB, Wong MM, Verslues PE. The flip side of phospho-signalling: regulation of protein dephosphorylation and the protein phosphatase 2Cs[J]. Plant Cell Environ, 2019, 42(10): 2913-2930.
doi: 10.1111/pce.13616 |
| [15] |
Linden KJ, Callis J. The ubiquitin system affects agronomic plant traits[J]. J Biol Chem, 2020, 295(40): 13940-13955.
doi: 10.1074/jbc.REV120.011303 URL |
| [16] |
Filipčík P, Curry JR, Mace PD. When worlds collide-mechanisms at the interface between phosphorylation and ubiquitination[J]. J Mol Biol, 2017, 429(8): 1097-1113.
doi: S0022-2836(17)30086-4 pmid: 28235544 |
| [17] |
Dubeaux G, Neveu J, Zelazny E, et al. Metal sensing by the IRT1 transporter-receptor orchestrates its own degradation and plant metal nutrition[J]. Mol Cell, 2018, 69(6): 953-964.e5.
doi: S1097-2765(18)30107-2 pmid: 29547723 |
| [18] |
Dong J, Ni WM, Yu RB, et al. Light-dependent degradation of PIF3 by SCFEBF1/2 promotes a photomorphogenic response in Arabidopsis[J]. Curr Biol, 2017, 27(16): 2420-2430.e6.
doi: S0960-9822(17)30799-6 pmid: 28736168 |
| [19] |
Yasuda S, Sato T, Maekawa S, et al. Phosphorylation of Arabidopsis ubiquitin ligase ATL31 is critical for plant carbon/nitrogen nutrient balance response and controls the stability of 14-3-3 proteins[J]. J Biol Chem, 2014, 289(22): 15179-15193.
doi: 10.1074/jbc.M113.533133 URL |
| [20] |
Sato T, Maekawa S, Yasuda S, et al. Identification of 14-3-3 proteins as a target of ATL31 ubiquitin ligase, a regulator of the C/N response in Arabidopsis[J]. Plant J, 2011, 68(1): 137-146.
doi: 10.1111/tpj.2011.68.issue-1 URL |
| [21] |
Yasuda S, Aoyama S, Hasegawa Y, et al. Arabidopsis CBL-interacting protein kinases regulate carbon/nitrogen-nutrient response by phosphorylating ubiquitin ligase ATL31[J]. Mol Plant, 2017, 10(4): 605-618.
doi: 10.1016/j.molp.2017.01.005 URL |
| [22] |
Nemoto K, Ramadan A, Arimura GI, et al. Tyrosine phosphorylation of the GARU E3 ubiquitin ligase promotes gibberellin signalling by preventing GID1 degradation[J]. Nat Commun, 2017, 8(1): 1004.
doi: 10.1038/s41467-017-01005-5 pmid: 29042542 |
| [23] |
Furlan G, Nakagami H, Eschen-Lippold L, et al. Changes in PUB22 ubiquitination modes triggered by MITOGEN-ACTIVATED PROTEIN KINASE3 dampen the immune response[J]. Plant Cell, 2017, 29(4): 726-745.
doi: 10.1105/tpc.16.00654 URL |
| [24] |
Wang J, Qu BY, Dou SJ, et al. The E3 ligase OsPUB15 interacts with the receptor-like kinase PID2 and regulates plant cell death and innate immunity[J]. BMC Plant Biol, 2015, 15: 49.
doi: 10.1186/s12870-015-0442-4 pmid: 25849162 |
| [25] |
Metzger MB, Pruneda JN, Klevit RE, et al. RING-type E3 ligases: master manipulators of E2 ubiquitin-conjugating enzymes and ubiquitination[J]. Biochim Biophys Acta, 2014, 1843(1): 47-60.
doi: 10.1016/j.bbamcr.2013.05.026 pmid: 23747565 |
| [26] | Wang ZB, Stockinger E. Arabidopsis homologs of transcription-domain-associated protein 1(Tra1)are essential for embryogenesis, growth and development[J]. Dev Biol, 2006, 295(1): 384. |
| [27] |
Liu HX, Stone SL. Abscisic acid increases Arabidopsis ABI5 transcription factor levels by promoting KEG E3 ligase self-ubiquitination and proteasomal degradation[J]. Plant Cell, 2010, 22(8): 2630-2641.
doi: 10.1105/tpc.110.076075 URL |
| [28] |
Lyzenga WJ, Sullivan V, Liu HX, et al. The kinase activity of calcineurin B-like interacting protein kinase 26(CIPK26)influences its own stability and that of the ABA-regulated ubiquitin ligase, keep on going(KEG)[J]. Front Plant Sci, 2017, 8: 502.
doi: 10.3389/fpls.2017.00502 pmid: 28443108 |
| [29] |
Lu DP, Lin WW, Gao XQ, et al. Direct ubiquitination of pattern recognition receptor FLS2 attenuates plant innate immunity[J]. Science, 2011, 332(6036): 1439-1442.
doi: 10.1126/science.1204903 pmid: 21680842 |
| [30] | Zhou JG, Liu DR, Wang P, et al. Regulation of Arabidopsis brassinosteroid receptor BRI1 endocytosis and degradation by plant U-box PUB12/PUB13-mediated ubiquitination[J]. Proc Natl Acad Sci USA, 2018, 115(8): E1906-E1915. |
| [31] |
Cheng CH, Wang ZJ, Ren ZY, et al. SCFAtPP2-B11 modulates ABA signaling by facilitating SnRK2.3 degradation in Arabidopsis thaliana[J]. PLoS Genet, 2017, 13(8): e1006947.
doi: 10.1371/journal.pgen.1006947 URL |
| [32] |
Niu D, Lin XL, Kong XX, et al. SIZ1-mediated SUMOylation of TPR1 suppresses plant immunity in Arabidopsis[J]. Mol Plant, 2019, 12(2): 215-228.
doi: S1674-2052(18)30368-X pmid: 30543996 |
| [33] |
Shahpasandzadeh H, Popova B, Kleinknecht A, et al. Interplay between sumoylation and phosphorylation for protection against α-synuclein inclusions[J]. J Biol Chem, 2014, 289(45): 31224-31240.
doi: 10.1074/jbc.M114.559237 pmid: 25231978 |
| [34] |
Saleh A, Withers J, Mohan R, et al. Posttranslational modifications of the master transcriptional regulator NPR1 enable dynamic but tight control of plant immune responses[J]. Cell Host Microbe, 2015, 18(2): 169-182.
doi: 10.1016/j.chom.2015.07.005 pmid: 26269953 |
| [35] |
Lee HJ, Park YJ, Seo PJ, et al. Systemic immunity requires SnRK2.8-mediated nuclear import of NPR1 in Arabidopsis[J]. Plant Cell, 2015, 27(12): 3425-3438.
doi: 10.1105/tpc.15.00371 URL |
| [36] |
Roy D, Sadanandom A. SUMO mediated regulation of transcription factors as a mechanism for transducing environmental cues into cellular signaling in plants[J]. Cell Mol Life Sci, 2021, 78(6): 2641-2664.
doi: 10.1007/s00018-020-03723-4 pmid: 33452901 |
| [37] |
Geoffroy MC, Hay RT. An additional role for SUMO in ubiquitin-mediated proteolysis[J]. Nat Rev Mol Cell Biol, 2009, 10(8): 564-568.
doi: 10.1038/nrm2707 |
| [38] |
Conti L, Nelis S, Zhang CJ, et al. Small Ubiquitin-like Modifier protein SUMO enables plants to control growth independently of the phytohormone gibberellin[J]. Dev Cell, 2014, 28(1): 102-110.
doi: 10.1016/j.devcel.2013.12.004 pmid: 24434138 |
| [39] |
Guo RK, Sun WN. Sumoylation stabilizes RACK1B and enhance its interaction with RAP2.6 in the abscisic acid response[J]. Sci Rep, 2017, 7: 44090.
doi: 10.1038/srep44090 pmid: 28272518 |
| [40] |
Praefcke GJK, Hofmann K, Dohmen RJ. SUMO playing tag with ubiquitin[J]. Trends Biochem Sci, 2012, 37(1): 23-31.
doi: 10.1016/j.tibs.2011.09.002 pmid: 22018829 |
| [41] |
McManus FP, Lamoliatte F, Thibault P. Identification of cross talk between SUMOylation and ubiquitylation using a sequential peptide immunopurification approach[J]. Nat Protoc, 2017, 12(11): 2342-2358.
doi: 10.1038/nprot.2017.105 pmid: 29048423 |
| [42] |
Tainer JA, Boddy MN. A SIM-ultaneous role for SUMO and ubiquitin[J]. Trends Biochem Sci, 2008, 33(5): 201-208.
doi: 10.1016/j.tibs.2008.02.001 pmid: 18403209 |
| [43] |
Elrouby N, Bonequi MV, Porri A, et al. Identification of Arabidopsis SUMO-interacting proteins that regulate chromatin activity and developmental transitions[J]. Proc Natl Acad Sci USA, 2013, 110(49): 19956-19961.
doi: 10.1073/pnas.1319985110 pmid: 24255109 |
| [44] |
Budhiraja R, Hermkes R, Müller S, et al. Substrates related to chromatin and to RNA-dependent processes are modified by Arabidopsis SUMO isoforms that differ in a conserved residue with influence on desumoylation[J]. Plant Physiol, 2009, 149(3): 1529-1540.
doi: 10.1104/pp.108.135053 pmid: 19151129 |
| [45] |
Lin XL, Niu D, Hu ZL, et al. An Arabidopsis SUMO E3 ligase, SIZ1, negatively regulates photomorphogenesis by promoting COP1 activity[J]. PLoS Genet, 2016, 12(4): e1006016.
doi: 10.1371/journal.pgen.1006016 URL |
| [46] |
Huang TT, Wuerzberger-Davis SM, Wu ZH, et al. Sequential modification of NEMO/IKKgamma by SUMO-1 and ubiquitin mediates NF-kappaB activation by genotoxic stress[J]. Cell, 2003, 115(5): 565-576.
doi: 10.1016/s0092-8674(03)00895-x pmid: 14651848 |
| [47] |
Sadanandom A, Ádám É, Orosa B, et al. SUMOylation of phytochrome-B negatively regulates light-induced signaling in Arabidopsis thaliana[J]. Proc Natl Acad Sci USA, 2015, 112(35): 11108-11113.
doi: 10.1073/pnas.1415260112 pmid: 26283376 |
| [48] |
Crozet P, Margalha L, Butowt R, et al. SUMOylation represses SnRK1 signaling in Arabidopsis[J]. Plant J, 2016, 85(1): 120-133.
doi: 10.1111/tpj.2016.85.issue-1 URL |
| [49] |
Kim JY, Park BS, Park SW, et al. Nitrate reductases are relocalized to the nucleus by AtSIZ1 and their levels are negatively regulated by COP1 and ammonium[J]. Int J Mol Sci, 2018, 19(4): 1202.
doi: 10.3390/ijms19041202 URL |
| [50] |
Lambeck IC, Fischer-Schrader K, Niks D, et al. Molecular mechanism of 14-3-3 protein-mediated inhibition of plant nitrate reductase[J]. J Biol Chem, 2012, 287(7): 4562-4571.
doi: 10.1074/jbc.M111.323113 pmid: 22170050 |
| [51] |
Heazlewood JL, Durek P, Hummel J, et al. PhosPhAt: a database of phosphorylation sites in Arabidopsis thaliana and a plant-specific phosphorylation site predictor[J]. Nucleic Acids Res, 2008, 36(Database issue): D1015-D1021.
doi: 10.1093/nar/gkm812 pmid: 17984086 |
| [52] |
Durek P, Schmidt R, Heazlewood JL, et al. PhosPhAt: the Arabidopsis thaliana phosphorylation site database. An update[J]. Nucleic Acids Res, 2010, 38(Database issue): D828-D834.
doi: 10.1093/nar/gkp810 URL |
| [53] |
Chen YM, Wang Y, Liang XL, et al. Mass spectrometric exploration of phytohormone profiles and signaling networks[J]. Trends Plant Sci, 2023, 28(4): 399-414.
doi: 10.1016/j.tplants.2022.12.006 URL |
| [54] |
Chen YM, Liu P. Proteome-wide chromatin interactomics to study plant epigenetics[J]. Trends Plant Sci, 2021, 26(7): 758-759.
doi: 10.1016/j.tplants.2021.04.002 URL |
| [1] | 杨艳, 胡洋, 刘霓如, 殷璐, 杨锐, 王鹏飞, 穆霄鹏, 张帅, 程春振, 张建成. ‘红满堂’苹果MbbZIP43基因的克隆与功能研究[J]. 生物技术通报, 2024, 40(2): 146-159. |
| [2] | 路喻丹, 刘晓驰, 冯新, 陈桂信, 陈义挺. 猕猴桃BBX基因家族成员鉴定与转录特征分析[J]. 生物技术通报, 2024, 40(2): 172-182. |
| [3] | 李亚男, 张豪杰, 梁梦静, 罗涛, 李旺宁, 张春辉, 季春丽, 李润植, 薛金爱, 崔红利. 雨生红球藻钙依赖蛋白激酶(CDPK)家族鉴定与表达分析[J]. 生物技术通报, 2024, 40(2): 300-312. |
| [4] | 邹修为, 岳佳妮, 李志宇, 戴良英, 李魏. 水稻热激转录因子HsfA2b调控非生物胁迫抗性的功能分析[J]. 生物技术通报, 2024, 40(2): 90-98. |
| [5] | 周会汶, 吴兰花, 韩德鹏, 郑伟, 余跑兰, 吴杨, 肖小军. 甘蓝型油菜种子硫苷含量全基因组关联分析[J]. 生物技术通报, 2024, 40(1): 222-230. |
| [6] | 吴圳, 张明英, 闫锋, 李依民, 高静, 颜永刚, 张岗. 掌叶大黄(Rheum palmatum L.)WRKY基因家族鉴定与分析[J]. 生物技术通报, 2024, 40(1): 250-261. |
| [7] | 王斌, 袁晓, 蒋园园, 王玉昆, 肖艳辉, 何金明. bHLH96的克隆及其在薄荷萜烯生物合成调控中的功能[J]. 生物技术通报, 2024, 40(1): 281-293. |
| [8] | 陈治民, 李翠, 韦继天, 李昕然, 刘峄, 郭强. 绿原酸生物合成调控及其应用研究进展[J]. 生物技术通报, 2024, 40(1): 57-71. |
| [9] | 黄小龙, 孙贵连, 马丹丹, 闫慧清. 水稻幼苗酵母单杂文库构建及LAZY1上游调控因子筛选[J]. 生物技术通报, 2023, 39(9): 126-135. |
| [10] | 韩浩章, 张丽华, 李素华, 赵荣, 王芳, 王晓立. 盐碱胁迫诱导的猴樟酵母cDNA文库构建及CbP5CS上游调控因子筛选[J]. 生物技术通报, 2023, 39(9): 236-245. |
| [11] | 丁丽, 都婷婷, 唐琼英, 高权新, 易少奎, 杨国梁. 罗氏沼虾蜕皮周期中内分泌调控和蜕皮信号通路相关基因的表达分析[J]. 生物技术通报, 2023, 39(9): 300-310. |
| [12] | 杨玉梅, 张坤晓. 应用CRISPR/Cas9技术建立ERK激酶相分离荧光探针定点整合的稳定细胞株[J]. 生物技术通报, 2023, 39(8): 159-164. |
| [13] | 吕秋谕, 孙培媛, 冉彬, 王佳蕊, 陈庆富, 李洪有. 苦荞转录因子基因FtbHLH3的克隆、亚细胞定位及表达分析[J]. 生物技术通报, 2023, 39(8): 194-203. |
| [14] | 徐靖, 朱红林, 林延慧, 唐力琼, 唐清杰, 王效宁. 甘薯IbHQT1启动子的克隆及上游调控因子的鉴定[J]. 生物技术通报, 2023, 39(8): 213-219. |
| [15] | 刘保财, 陈菁瑛, 张武君, 黄颖桢, 赵云青, 刘剑超, 危智诚. 多花黄精种子微根茎基因表达特征分析[J]. 生物技术通报, 2023, 39(8): 220-233. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||
