








生物技术通报 ›› 2022, Vol. 38 ›› Issue (2): 141-149.doi: 10.13560/j.cnki.biotech.bull.1985.2021-0241
叶鹏林( ), KwasiKyere-Yeboah, 高恶斌(
), KwasiKyere-Yeboah, 高恶斌( )
)
收稿日期:2021-03-03
出版日期:2022-02-26
发布日期:2022-03-09
作者简介:叶鹏林,男,硕士研究生,研究方向:蓝藻生物合成;E-mail: 基金资助:
YE Peng-lin( ), Kwasi Kyere-Yeboah, GAO E-bin(
), Kwasi Kyere-Yeboah, GAO E-bin( )
)
Received:2021-03-03
Published:2022-02-26
Online:2022-03-09
摘要:
为了增加工程集胞藻PCC 6803的乙醇合成产量,通过选用强启动子Pcpc560 驱动并提高外源乙醇合成基因(pdc,yqhD)的表达,从而促进乙醇的生产。具体方法利用同源双交换引入来源于运动型发酵单胞菌的丙酮酸脱羧酶基因(pdc)与来源于大肠杆菌的NADPH依赖型醛还原酶基因(yqhD)并选用不同的启动子来驱动其表达。通过逆转录定量PCR分析,比较在不同启动子驱动的情况下,外源乙醇合成基因(pdc,yqhD)的表达情况并检测相应突变株的乙醇产量。结果显示相较于中等启动子,铜离子诱导启动子PpetE,来源于集胞藻PCC 6803的光强启动子Pcpc560显著促进了外源乙醇合成基因(pdc,yqhD)的表达,并增加了工程菌株乙醇合成的产量。超强启动子Pcpc560搭配pdc,yqhD的组合表达,显著提高了工程菌株的乙醇合成产量。
叶鹏林, KwasiKyere-Yeboah, 高恶斌. 启动子PpetE与Pcpc560对集胞藻PCC 6803生物合成乙醇的影响[J]. 生物技术通报, 2022, 38(2): 141-149.
YE Peng-lin, Kwasi Kyere-Yeboah, GAO E-bin. Effects on the Biosynthesis of Ethanol by Promoters PpetE and Pcpc560 in Synechocystis sp. PCC 6803[J]. Biotechnology Bulletin, 2022, 38(2): 141-149.
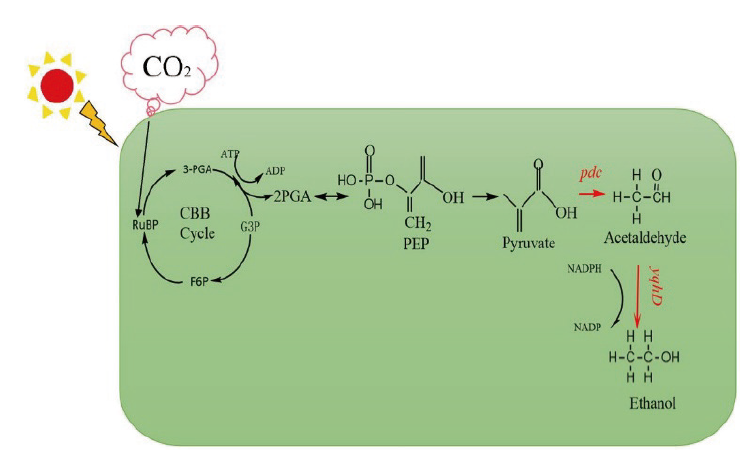
图1 集胞藻PCC 6803的乙醇合成路线 PEP:Phosphoenolpyruvate(磷酸烯醇丙酮酸),Pyruvate:丙酮酸,Acetal-dehyde:乙醛,Ethanol:乙醇,CBB cycle:卡尔文循环,RuBP:1,5-diphosphate ribulose(1,5-二磷酸核酮糖),G3P:glyceraldehyde-3-phosphate(三磷酸甘油醛),PGA:phosphoglyceric acid(磷酸甘油酸),F6P:fructose 6 Phosphate(磷酸果糖),NADP:烟酰胺腺嘌呤二核苷酸磷酸,NADPH:还原型烟酰胺腺嘌呤二核苷酸磷酸,pdc:丙酮酸脱羧酶基因,yqhD:NADPH依赖型醛还原酶基因
Fig.1 Ethanol synthesis route in Synechocystis sp. PCC 6803
| 名称 Name | 特点 Characteristic |
|---|---|
| pMD18-T | 氨苄霉素/AmpR |
| pMD0168 | 氨苄霉素/AmpR 将slr0168的上下游600 bp基因扩增并克隆到质粒pMD18-T的SphI/MluI和XbaI/KpnI位点 |
| pBE406 | 壮观霉素/ SpR 将壮观霉素基因扩增并克隆到所构建质粒pMD0168的XhoI/XbaI位点 |
| pBE407 | 壮观霉素/ SpR 将启动子PpetE扩增并克隆到所构建质粒pBE406的BamHI/SalI位点,将终止子TrbcL扩增并克隆到所构建质粒pBE406的SalI/HindIII位点 |
| pBE407-pdc | 壮观霉素/ SpR pdc基因被扩增并和质粒pBE407双酶切,通过NdeI/BamHI位点相连接获得质粒pBE407-pdc |
| pBE01 | 壮观霉素/ SpR yqhD 基因被扩增并和质粒pBE407-pdc双酶切,通过XbaI/KpnI位点相连接获得质粒pBE01 |
| pBE408 | 壮观霉素/ SpR 将启动子Pcpc560扩增并克隆到所构建质粒pBE406的BamHI/SalI位点,将终止子TrbcL扩增并克隆到所构建质粒pBE406的SalI/HindIII位点 |
| pBE408-pdc | 壮观霉素/ SpR pdc基因被扩增并和质粒pBE408双酶切,通过NdeI/BamHI位点相连接获得质粒pBE408-pdc |
| pBE02 | 壮观霉素/ SpR yqhD 基因被扩增并和质粒pBE408-pdc双酶切,通过XbaI/KpnI位点相连接获得质粒pBE02 |
表1 本文使用的质粒
Table 1 Plasmids used in this study
| 名称 Name | 特点 Characteristic |
|---|---|
| pMD18-T | 氨苄霉素/AmpR |
| pMD0168 | 氨苄霉素/AmpR 将slr0168的上下游600 bp基因扩增并克隆到质粒pMD18-T的SphI/MluI和XbaI/KpnI位点 |
| pBE406 | 壮观霉素/ SpR 将壮观霉素基因扩增并克隆到所构建质粒pMD0168的XhoI/XbaI位点 |
| pBE407 | 壮观霉素/ SpR 将启动子PpetE扩增并克隆到所构建质粒pBE406的BamHI/SalI位点,将终止子TrbcL扩增并克隆到所构建质粒pBE406的SalI/HindIII位点 |
| pBE407-pdc | 壮观霉素/ SpR pdc基因被扩增并和质粒pBE407双酶切,通过NdeI/BamHI位点相连接获得质粒pBE407-pdc |
| pBE01 | 壮观霉素/ SpR yqhD 基因被扩增并和质粒pBE407-pdc双酶切,通过XbaI/KpnI位点相连接获得质粒pBE01 |
| pBE408 | 壮观霉素/ SpR 将启动子Pcpc560扩增并克隆到所构建质粒pBE406的BamHI/SalI位点,将终止子TrbcL扩增并克隆到所构建质粒pBE406的SalI/HindIII位点 |
| pBE408-pdc | 壮观霉素/ SpR pdc基因被扩增并和质粒pBE408双酶切,通过NdeI/BamHI位点相连接获得质粒pBE408-pdc |
| pBE02 | 壮观霉素/ SpR yqhD 基因被扩增并和质粒pBE408-pdc双酶切,通过XbaI/KpnI位点相连接获得质粒pBE02 |
| 引物序列Sequence of a primer | 大小Size |
|---|---|
| SP-F:5'-CCACGCGTAAGCTTGGATCCGCTCACGCAACTGGTCCAGAA-3' SP-R:5'-CGGGAGCTCGAATTCTAGAGTGCTTAGTGCATCTAACGC-3' | 1.1 kb |
| Pcpc-F:5'-CGTCTAGAGGATCCCCTGTAGAGAAGAGTCCCTG-3' Pcpc-R:5'-TTTCTCCTCTTTTGAATTAATCTCCTACTTGACTTTATGAG-3' | 550 bp |
| PetE-F:5'-GCTCTAGACAAGGATTCATAGCGGTTGCCCAATC-3' PetE-R:5'-GCTGCCTAGGATTCTGGCGAAAGGGGGATGTG-3' | 200 bp |
| TrbcL-F:5'-CGCGTCGACCGGTGTTTGGATTGTCGGAGT-3' TrbcL-R:5'-CCGACGCGTAAGCTTCCGGTAATTGGTAAATTGCTGTC-3' | 250 bp |
| Pdc-F:5'-CCGAGATCTCATATGTCCTACACCGTGGGCACCT-3' Pdc-R:5'-CGCGGATCCTGCAGCTCGAGTCTAGATTACAACAATTTGTTCACGGGT-3' | 1.7 kb |
| YqhD-F:5'-CAAACTCGAGTCTAGATGAACAACTTTAACTTGCACACCCCCAC-3' YqhD-R:5'-CGGGGTACCTGCAGTTAGCGGGCGGCTTCGTATATACGGC-3' | 1.2 kb |
| slr0168Up-F:5'-GGCATGCCGAGCGGCACCACGGGGCACCACCGC-3' slr0168Up-R:5'-GACGCGTCGGCGCACAGCAGCGTGCGACGTGTG-3' | 600 bp |
| slr0168Dw-F:5'-CTCTAGAGTGCCACTACCTGGCGTGCCGCTACC-3' slr0168Dw-R:5'-GGGGTACCCCGCATGACCAGCTGCCGCCCCAGC-3' | 600 bp |
表2 本文使用的引物
Table 2 Primers used in this study
| 引物序列Sequence of a primer | 大小Size |
|---|---|
| SP-F:5'-CCACGCGTAAGCTTGGATCCGCTCACGCAACTGGTCCAGAA-3' SP-R:5'-CGGGAGCTCGAATTCTAGAGTGCTTAGTGCATCTAACGC-3' | 1.1 kb |
| Pcpc-F:5'-CGTCTAGAGGATCCCCTGTAGAGAAGAGTCCCTG-3' Pcpc-R:5'-TTTCTCCTCTTTTGAATTAATCTCCTACTTGACTTTATGAG-3' | 550 bp |
| PetE-F:5'-GCTCTAGACAAGGATTCATAGCGGTTGCCCAATC-3' PetE-R:5'-GCTGCCTAGGATTCTGGCGAAAGGGGGATGTG-3' | 200 bp |
| TrbcL-F:5'-CGCGTCGACCGGTGTTTGGATTGTCGGAGT-3' TrbcL-R:5'-CCGACGCGTAAGCTTCCGGTAATTGGTAAATTGCTGTC-3' | 250 bp |
| Pdc-F:5'-CCGAGATCTCATATGTCCTACACCGTGGGCACCT-3' Pdc-R:5'-CGCGGATCCTGCAGCTCGAGTCTAGATTACAACAATTTGTTCACGGGT-3' | 1.7 kb |
| YqhD-F:5'-CAAACTCGAGTCTAGATGAACAACTTTAACTTGCACACCCCCAC-3' YqhD-R:5'-CGGGGTACCTGCAGTTAGCGGGCGGCTTCGTATATACGGC-3' | 1.2 kb |
| slr0168Up-F:5'-GGCATGCCGAGCGGCACCACGGGGCACCACCGC-3' slr0168Up-R:5'-GACGCGTCGGCGCACAGCAGCGTGCGACGTGTG-3' | 600 bp |
| slr0168Dw-F:5'-CTCTAGAGTGCCACTACCTGGCGTGCCGCTACC-3' slr0168Dw-R:5'-GGGGTACCCCGCATGACCAGCTGCCGCCCCAGC-3' | 600 bp |
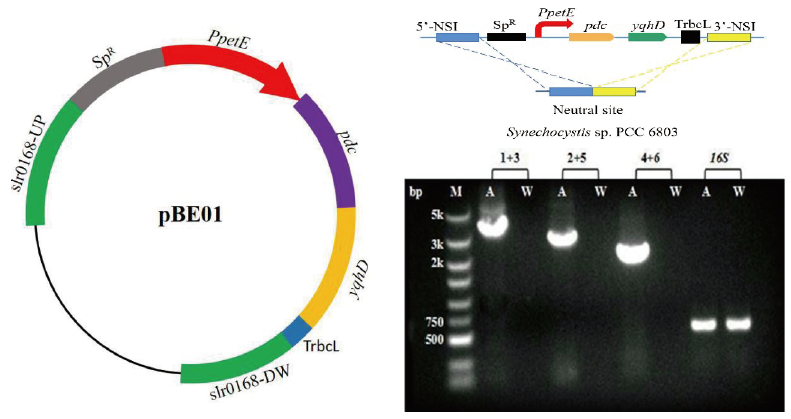
图2 SYN01菌株的构建示意图与PCR分析 M:DNA Marker,A:提取的转化子基因组DNA,W:提取的野生型集胞藻DNA,1:slr0168上游基因上游引物slr0168Up-F,2:pdc基因上游引物Pdc-F,3:pdc基因下游引物Pdc-R,4:yqhD基因上游引物YqhD-F,5:yqhD基因下游引物YqhD-R,6:slr0168下游基因下游引物slr0168DW-R,16S:扩增编码16S rRNA的内源基因(600 bp),1+3:使用引物1和引物3进行扩增(3.6 kb),2+5:使用引物2和引物5进行扩增(2.9 kb),4+6:使用引物4和引物6进行扩增(2 kb)
Fig.2 Construction diagram and PCR analysis of the SYN01 strain M:DNA marker. A:Extracted DNA from transformant genome. W:Extracted DNA from wild-type Synechocystis. 1:slr0168 upstream gene upstream primer slr0168Up-F. 2:pdc gene upstream primer Pdc-F. 3:pdc gene downstream primer Pdc-R. 4:yqhD gene upstream primer YqhD-F. 5:yqhD gene downstream primer YqhD-R. 6:slr0168 downstream gene downstream primer slr0168DW-R. 16S:Amplify the endogenous gene encoding 16S rRNA(600 bp). 1+3:Using primer 1 and primer 3 for amplification(3.6 kb). 2+5:Using primer 2 and primer 5 for amplification(2.9 kb). 4+6:Using primer 4 and primer 6 for amplification(2 kb)
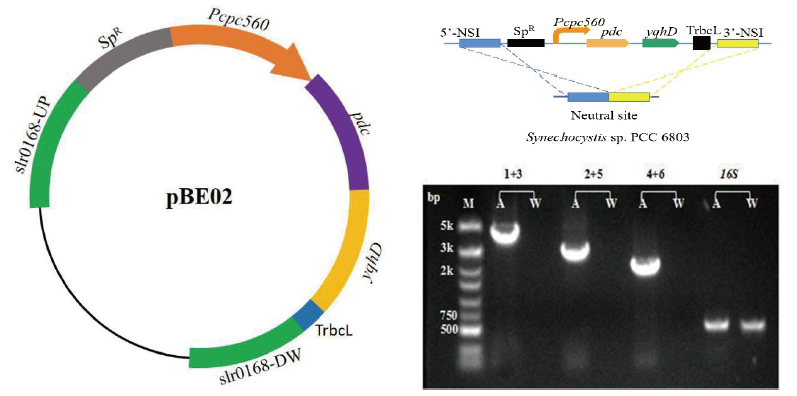
图3 SYN02菌株的构建示意图与PCR分析 M:DNA Marker,A:提取的转化子基因组DNA,W:提取的野生型集胞藻DNA,1:slr0168上游基因上游引物slr0168Up-F,2:pdc基因上游引物Pdc-F,3:pdc基因下游引物Pdc-R,4:yqhD基因上游引物YqhD-F,5:yqhD基因下游引物YqhD-R,6:slr0168下游基因下游引物slr0168DW-R,16S:扩增编码16S rRNA的内源基因(600 bp),1+3:使用引物1和引物3进行扩增(3.9 kb),2+5:使用引物2和引物5进行扩增(2.9 kb),4+6:使用引物4和引物6进行扩增(2 kb)
Fig.3 Construction diagram and PCR analysis of the SYN02 strain M:DNA marker. A:Extracted DNA from transformant genome. W:Extracted DNA from wild-type Synechocystis. 1:slr0168 upstream gene upstream primer slr0168Up-F. 2:pdc gene upstream primer Pdc-F. 3:pdc gene downstream primer Pdc-R. 4:yqhD gene upstream primer YqhD-F. 5:yqhD gene downstream primer YqhD-R. 6:slr0168 downstream gene downstream primer slr0168DW-R. 16S:Amplify the endogenous gene encoding 16S rRNA(600 bp). 1+3:Using primer 1 and primer 3 for amplification(3.9 kb). 2+5:Using primer 2 and primer 5 for amplification(2.9 kb). 4+6:Using primer 4 and primer 6 for amplification(2 kb)
| 基因Gene | 菌株Strain | 16S | CT1 | CT2 | CT3 | 平均 CT Average CT | ∆CT | 2-∆∆CT | 标准偏差SD |
|---|---|---|---|---|---|---|---|---|---|
| Pdc | SYN01 | 22.85 | 22.97 | 22.90 | 22.98 | 22.95 | 0.10 | 2.55 | 0.0252 |
| SYN02 | 22.85 | 21.60 | 21.57 | 21.63 | 21.60 | -1.25 | 0.0173 | ||
| YqhD | SYN01 | 22.85 | 23.76 | 23.72 | 24.04 | 23.84 | 0.99 | 2.31 | 0.1007 |
| SYN02 | 22.85 | 22.60 | 22.66 | 22.63 | 22.63 | -0.22 | 0.0173 |
表3 逆转录定量PCR的结果
Table 3 Reverse transcription of quantitative PCR results
| 基因Gene | 菌株Strain | 16S | CT1 | CT2 | CT3 | 平均 CT Average CT | ∆CT | 2-∆∆CT | 标准偏差SD |
|---|---|---|---|---|---|---|---|---|---|
| Pdc | SYN01 | 22.85 | 22.97 | 22.90 | 22.98 | 22.95 | 0.10 | 2.55 | 0.0252 |
| SYN02 | 22.85 | 21.60 | 21.57 | 21.63 | 21.60 | -1.25 | 0.0173 | ||
| YqhD | SYN01 | 22.85 | 23.76 | 23.72 | 24.04 | 23.84 | 0.99 | 2.31 | 0.1007 |
| SYN02 | 22.85 | 22.60 | 22.66 | 22.63 | 22.63 | -0.22 | 0.0173 |
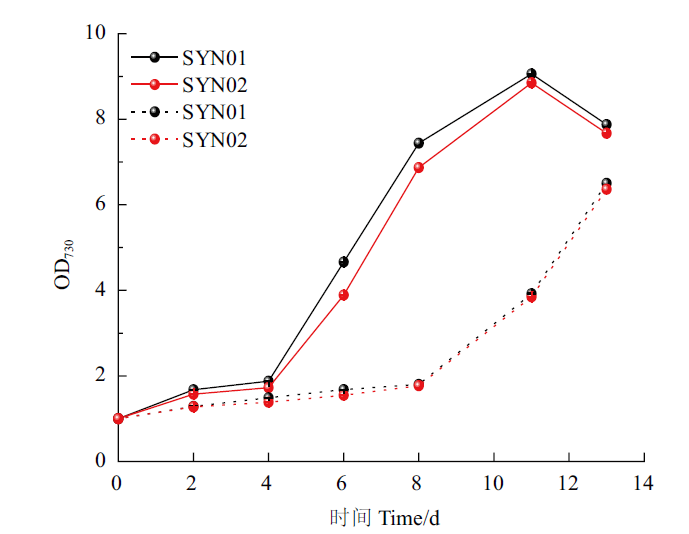
图8 优化培养下菌株的生长情况 实线为通入5% CO2+95%空气,虚线为通入100%空气,下同
Fig.8 Growth condition of the strains under the optimized culture The solid line refers to 5% CO2+95% air,and the dashed line refers to 100% air. The same below
| [1] |
Song CF, Liu QL, Deng S, et al. Cryogenic-based CO2 capture technologies:State-of-the-art developments and current challenges[J]. Renew Sustain Energy Rev, 2019, 101:265-278.
doi: 10.1016/j.rser.2018.11.018 URL |
| [2] |
Afgan NH, Gobaisi DA, Carvalho MG, et al. Sustainable energy development[J]. Renew Sustain Energy Rev, 1998, 2(3):235-286.
doi: 10.1016/S1364-0321(98)00002-1 URL |
| [3] |
Nemeth N. Environment and energy:Problems, resolutions, solutions[J]. Int J Hydrog Energy, 1990, 15(7):457-462.
doi: 10.1016/0360-3199(90)90102-5 URL |
| [4] |
Goldemberg J . Special issue on sustainability and energy:Perspectives:“Ethanol for a sustainable energy future”[J]. Science, 2007, 317(5843):1325-1325.
doi: 10.1126/science.317.5843.1325 URL |
| [5] |
Parmar A, Singh NK, Pandey A, et al. Cyanobacteria and microalgae:a positive prospect for biofuels[J]. Bioresour Technol, 2011, 102(22):10163-10172.
doi: 10.1016/j.biortech.2011.08.030 URL |
| [6] |
Diao J, Song X, Zhang L, et al. Tailoring cyanobacteria as a new platform for highly efficient synjournal of astaxanthin[J]. Metab Eng, 2020, 61:275-287.
doi: 10.1016/j.ymben.2020.07.003 URL |
| [7] | 高宏, 唐蜻, 徐旭东. 集胞藻PCC6803铜离子诱导表达平台的构建[J]. 水生生物学报, 2007, 31(2):240-244. |
| Gao H, Tang Q, Xu XD. Construction of copper-induced gene expression platform in Synechocystis sp pcc6803[J]. Acta Hydrobiol Sin, 2007, 31(2):240-244. | |
| [8] |
Ghassemian M, Wong B, Ferreira F, et al. Cloning, sequencing and transcriptional studies of the genes for cytochrome c-553 and plastocyanin from Anabaena sp. PCC 7120[J]. Microbiology, 1994, 140(5):1151-1159.
doi: 10.1099/13500872-140-5-1151 URL |
| [9] |
Gao ZX, Zhao H, Li ZM, et al. Photosynthetic production of ethanol from carbon dioxide in genetically engineered cyanobacteria[J]. Energy Environ Sci, 2012, 5(12):9857-9865.
doi: 10.1039/C2EE22675H URL |
| [10] |
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method[J]. Methods, 2001, 25(4):402-408.
doi: 10.1006/meth.2001.1262 pmid: 11846609 |
| [11] |
Woo JE, Jang YS. Metabolic engineering of microorganisms for the production of ethanol and butanol from oxides of carbon[J]. Appl Microbiol Biotechnol, 2019, 103(20):8283-8292.
doi: 10.1007/s00253-019-10072-1 URL |
| [12] |
Namakoshi K, Nakajima T, Yoshikawa K, et al. Combinatorial deletions of glgC and PhaCE enhance ethanol production in Synechocystis sp. PCC 6803[J]. J Biotechnol, 2016, 239:13-19.
doi: 10.1016/j.jbiotec.2016.09.016 URL |
| [13] |
Mock M, Schmid A, Bühler K. Photoautotrophic production of succinate via the oxidative branch of the tricarboxylic acid cycle influences glycogen accumulation in Synechocystis sp. PCC 6803[J]. Algal Res, 2019, 43:101645.
doi: 10.1016/j.algal.2019.101645 URL |
| [14] | Noreña-Caro D, Benton MG. Cyanobacteria as photoautotrophic biofactories of high-value chemicals[J]. J CO2 Util, 2018, 28:335-366. |
| [15] |
Angermayr SA, Gorchs Rovira A, Hellingwerf KJ. Metabolic engineering of cyanobacteria for the synjournal of commodity products[J]. Trends Biotechnol, 2015, 33(6):352-361.
doi: 10.1016/j.tibtech.2015.03.009 pmid: 25908503 |
| [16] |
Yoshida S, Tanaka H, Hirayama M, et al. Production of pyruvate from mannitol by mannitol-assimilating pyruvate decarboxylase-negative Saccharomyces cerevisiae[J]. Bioengineered, 2015, 6(6):347-350.
doi: 10.1080/21655979.2015.1112472 pmid: 26588105 |
| [17] |
Takahashi H, Uchimiya H, Hihara Y. Difference in metabolite levels between photoautotrophic and photomixotrophic cultures of Synechocystis sp. PCC 6803 examined by capillary electrophoresis electrospray ionization mass spectrometry[J]. J Exp Bot, 2008, 59(11):3009-3018.
doi: 10.1093/jxb/ern157 pmid: 18611912 |
| [18] |
Miao R, Liu X, Englund E, et al. Isobutanol production in Synechocystis PCC 6803 using heterologous and endogenous alcohol dehydrogenases[J]. Metab Eng Commun, 2017, 5:45-53.
doi: 10.1016/j.meteno.2017.07.003 pmid: 29188183 |
| [19] |
Dexter J, Fu PC. Metabolic engineering of cyanobacteria for ethanol production[J]. Energy Environ Sci, 2009, 2(8):857.
doi: 10.1039/b811937f URL |
| [20] |
Yoshikawa K, Hirasawa T, Shimizu H. Effect of malic enzyme on ethanol production by Synechocystis sp. PCC 6803[J]. J Biosci Bioeng, 2015, 119(1):82-84.
doi: 10.1016/j.jbiosc.2014.06.001 URL |
| [21] |
Gao EB, Kyere-Yeboah K, Wu JH, et al. Photoautotrophic production of p-Coumaric acid using genetically engineered Synechocystis sp. Pasteur Culture Collection 6803[J]. Algal Res, 2021, 54:102180.
doi: 10.1016/j.algal.2020.102180 URL |
| [22] |
Zhou J, Zhang H, Meng H, et al. Discovery of a super-strong promoter enables efficient production of heterologous proteins in cyanobacteria[J]. Sci Rep, 2014, 4:4500.
doi: 10.1038/srep04500 pmid: 24675756 |
| [23] | Castenholz RW. Culturing methods for cyanobacteria[M]// Methods in Enzymology. Amsterdam:Elsevier, 1988:68-93. |
| [24] | Moss NA, Leao T, Glukhov E, et al. Collection, culturing, and genome analyses of tropical marine filamentous benthic cyanobacteria[M]// Methods in Enzymology. Amsterdam:Elsevier, 2018:3-43. |
| [25] |
Velmurugan R, Incharoensakdi A. Co-cultivation of two engineered strains of Synechocystis sp. PCC 6803 results in improved bioethanol production[J]. Renew Energy, 2020, 146:1124-1133.
doi: 10.1016/j.renene.2019.07.025 URL |
| [1] | 刘玉玲, 王梦瑶, 孙琦, 马利花, 朱新霞. 启动子RD29A对转雪莲SikCDPK1基因烟草抗逆性的影响[J]. 生物技术通报, 2023, 39(9): 168-175. |
| [2] | 程亚楠, 张文聪, 周圆, 孙雪, 李玉, 李庆刚. 乳酸乳球菌生产2'-岩藻糖基乳糖的途径构建及发酵培养基优化[J]. 生物技术通报, 2023, 39(9): 84-96. |
| [3] | 李博, 刘合霞, 陈宇玲, 周兴文, 朱宇林. 金花茶CnbHLH79转录因子的克隆、亚细胞定位及表达分析[J]. 生物技术通报, 2023, 39(8): 241-250. |
| [4] | 叶云芳, 田清尹, 施婷婷, 王亮, 岳远征, 杨秀莲, 王良桂. 植物中β-紫罗兰酮生物合成及调控研究进展[J]. 生物技术通报, 2023, 39(8): 91-105. |
| [5] | 王玲, 卓燊, 付学森, 刘紫璇, 刘笑蓉, 王志辉, 周日宝, 刘湘丹. 莲生物碱生物合成途径及相关基因研究进展[J]. 生物技术通报, 2023, 39(7): 56-66. |
| [6] | 成婷, 苑帅, 张晓元, 林良才, 李欣, 张翠英. 酿酒酵母异丁醇合成途径调控的研究进展[J]. 生物技术通报, 2023, 39(7): 80-90. |
| [7] | 李帜奇, 袁月, 苗荣庆, 庞秋颖, 张爱琴. 盐胁迫盐芥和拟南芥褪黑素含量及合成相关基因表达模式分析[J]. 生物技术通报, 2023, 39(5): 142-151. |
| [8] | 姜晴春, 杜洁, 王嘉诚, 余知和, 王允, 柳忠玉. 虎杖转录因子PcMYB2的表达特性和功能分析[J]. 生物技术通报, 2023, 39(5): 217-223. |
| [9] | 周定定, 李辉虎, 汤兴涌, 余发新, 孔丹宇, 刘毅. 甘草酸和甘草苷生物合成与调控的研究进展[J]. 生物技术通报, 2023, 39(5): 44-53. |
| [10] | 郭三保, 宋美玲, 李灵心, 尧子钊, 桂明明, 黄胜和. 斑地锦查尔酮合酶基因及启动子的克隆与分析[J]. 生物技术通报, 2023, 39(4): 148-156. |
| [11] | 郁慧丽, 李爱涛. 细胞色素P450酶在香精香料绿色生物合成中的应用[J]. 生物技术通报, 2023, 39(4): 24-37. |
| [12] | 杨岚, 张晨曦, 樊学伟, 王阳光, 王春秀, 李文婷. 鸡 BMP15 基因克隆、表达模式及启动子活性分析[J]. 生物技术通报, 2023, 39(4): 304-312. |
| [13] | 姚晓文, 梁晓, 陈青, 伍春玲, 刘迎, 刘小强, 税军, 乔阳, 毛奕茗, 陈银华, 张银东. 二斑叶螨抗性木薯木质素合成途径基因表达特性研究[J]. 生物技术通报, 2023, 39(2): 161-171. |
| [14] | 王晓梅, 杨小薇, 李辉尚, 何微, 辛竹琳. 全球合成生物学发展现状及对我国的启示[J]. 生物技术通报, 2023, 39(2): 292-302. |
| [15] | 苗淑楠, 高宇, 李昕儒, 蔡桂萍, 张飞, 薛金爱, 季春丽, 李润植. 大豆GmPDAT1参与油脂合成和非生物胁迫应答的功能分析[J]. 生物技术通报, 2023, 39(2): 96-106. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||