








生物技术通报 ›› 2022, Vol. 38 ›› Issue (5): 112-122.doi: 10.13560/j.cnki.biotech.bull.1985.2021-0946
收稿日期:2021-07-24
出版日期:2022-05-26
发布日期:2022-06-10
作者简介:薛鲜丽,女,博士,讲师,研究方向:丝状真菌遗传改造;E-mail: 基金资助:
XUE Xian-li1( ), WANG Jing-ran1, BI Hang-hang1, WANG De-pei1,2(
), WANG Jing-ran1, BI Hang-hang1, WANG De-pei1,2( )
)
Received:2021-07-24
Published:2022-05-26
Online:2022-06-10
摘要:
SAGA复合体是一种多功能蛋白复合物,负责细胞内10%以上的基因转录。Spt7作为SAGA复合体的核心蛋白,维持SAGA复合体的稳定。除酵母之外的真菌中并未有Spt7相关的研究报道。本研究对不同黑曲霉、丝衣霉等丝状真菌来源的Spt7的氨基酸序列进行多序列比对及蛋白结构的分析,发现Spt7在不同物种一致性较低,但均存在保守型的Bromo结构域。以黑曲霉1062为出发菌株,通过农杆菌转化法将过表达spt7的质粒转入1062菌株中,获得黑曲霉OE spt7转化子。通过对OE spt7转化子及对照组在CM培养基生长形态进行观察分析发现,过表达Spt7有益于菌体的生长及分生孢子的生成,在72 h时OE spt7转化子和对照组的菌落直径分别达到2.9 cm和2.8 cm,且转化子的孢子数量达到(5.8-6.3)×107/cm2,较对照组(2.3×107/cm2)提高1.5-1.7倍,且转化子菌丝分支较对照组多。另外,与对照组相比,OE spt7转化子在15 mmol/L H2O2条件下孢子萌发更快,菌落直径是对照组4倍,且在15% NaCl高渗以及39℃高温的条件下较对照组生长更加鲜活。通过qRT-PCR分析参与菌体抗氧化胁迫的过氧化物酶及耐高温的热休克蛋白基因的转录水平,发现除了CatR转录水平下调3.56倍,其他的过氧化物酶SOD、CpeB、GPX分别上调了3.8、1.89、3.56倍,除Hsp40及Hsp70较对照组无显著差异,Hsp90的转录水平上调19倍。
薛鲜丽, 王静然, 毕杭杭, 王德培. 过表达Spt7对黑曲霉生长及抗逆性影响[J]. 生物技术通报, 2022, 38(5): 112-122.
XUE Xian-li, WANG Jing-ran, BI Hang-hang, WANG De-pei. Effect of Spt7 Overexpression of on the Growth and Stress Resistance of Aspergillus niger[J]. Biotechnology Bulletin, 2022, 38(5): 112-122.
| 名称Name | 引物序列Sequence of primers(5'-3') | 引物作用Functions of primers |
|---|---|---|
| Pgla-F | TCACTACAGATACAAGGATCCTGCCATTGGCGGAGGGGT | pOEspt7质粒构建引物(pgla-spt7通过重组酶连接到p66质粒) |
| Pgla-R | AGCGACATTGCTGAGGTGTAATGATGCTGG | |
| spt7-F | TACACCTCAGCAATGTCGCTCGGACACCACC | |
| spt7-R(含终止子) | TGTGATCCGCCTGGAGGATCCCAAGGCAAGTACACTGAACAAGGA | |
| YZ-Ku70LL-F | CCGTCCGCCATAGTAGAATGT | pOE spt7质粒验证引物 |
| YZ-spt7-5R | GTTCTTACGAACCTCGCTCAT | |
| YZ-spt7-3R | AAACAACAACGAGCCTTTGGTC | |
| YZ-ku70RR-R | TTGTTGGATTCGCTAGTGCGCT | |
| RT-actin-F | ACCACCGACTCCCTACTA | pOE spt7 qRT-PCR引物 |
| RT-actin-R | AGTCAAGAGAGAGATGGGAT | |
| RT-hsp90-F | TCTTCCCTCTCTCTCTACCTC | |
| RT-hsp90-R | AAAACGGTTTGAAAGGTGTG | |
| RT-hsp70-F | GTGGGTGTCTTCCGTGATGACCGCA | |
| RT-hsp70-R | GGACCTCAGCGTCGGCAAA | |
| RT-hsp40-F | TGTTCAAGAAACCCAACATCCT | |
| RT-hsp40-R | TGTGGGAGACGACGGCCATG | |
| RT-GPX-F | CGCCTTCTCAATCTCCTTCA | |
| RT-GPX-R | AGCGCATCAAGTGGAACTTT | |
| RT-SOD-F | TTATCGCTCACAATGGCTGCTT | |
| RT-SOD-R | GACAGGTCAGGGAGAGTAGC | |
| RT-catR-F | CTTGTCACCGAGTGCCCGTTT | |
| RT-catR-R | GTAATCCGGACCCTCCTGTTGGG | |
| RT-cpeB-F | AACGGTTCGCTCCTCTTAAC | |
| RT-cpeB-R | AGGAAATCTTTGCTCCGTACTT |
表1 试验中用到引物
Table 1 Primers used in the experiment
| 名称Name | 引物序列Sequence of primers(5'-3') | 引物作用Functions of primers |
|---|---|---|
| Pgla-F | TCACTACAGATACAAGGATCCTGCCATTGGCGGAGGGGT | pOEspt7质粒构建引物(pgla-spt7通过重组酶连接到p66质粒) |
| Pgla-R | AGCGACATTGCTGAGGTGTAATGATGCTGG | |
| spt7-F | TACACCTCAGCAATGTCGCTCGGACACCACC | |
| spt7-R(含终止子) | TGTGATCCGCCTGGAGGATCCCAAGGCAAGTACACTGAACAAGGA | |
| YZ-Ku70LL-F | CCGTCCGCCATAGTAGAATGT | pOE spt7质粒验证引物 |
| YZ-spt7-5R | GTTCTTACGAACCTCGCTCAT | |
| YZ-spt7-3R | AAACAACAACGAGCCTTTGGTC | |
| YZ-ku70RR-R | TTGTTGGATTCGCTAGTGCGCT | |
| RT-actin-F | ACCACCGACTCCCTACTA | pOE spt7 qRT-PCR引物 |
| RT-actin-R | AGTCAAGAGAGAGATGGGAT | |
| RT-hsp90-F | TCTTCCCTCTCTCTCTACCTC | |
| RT-hsp90-R | AAAACGGTTTGAAAGGTGTG | |
| RT-hsp70-F | GTGGGTGTCTTCCGTGATGACCGCA | |
| RT-hsp70-R | GGACCTCAGCGTCGGCAAA | |
| RT-hsp40-F | TGTTCAAGAAACCCAACATCCT | |
| RT-hsp40-R | TGTGGGAGACGACGGCCATG | |
| RT-GPX-F | CGCCTTCTCAATCTCCTTCA | |
| RT-GPX-R | AGCGCATCAAGTGGAACTTT | |
| RT-SOD-F | TTATCGCTCACAATGGCTGCTT | |
| RT-SOD-R | GACAGGTCAGGGAGAGTAGC | |
| RT-catR-F | CTTGTCACCGAGTGCCCGTTT | |
| RT-catR-R | GTAATCCGGACCCTCCTGTTGGG | |
| RT-cpeB-F | AACGGTTCGCTCCTCTTAAC | |
| RT-cpeB-R | AGGAAATCTTTGCTCCGTACTT |

图1 Spt7蛋白结构和氨基酸多序列比对及系统进化树分析 A:酵母及黑曲霉Spt7蛋白结构;B:酵母及黑曲霉中Spt7氨基酸多蛋白序列比对;C:丝状真菌Spt7的系统进化树
Fig.1 Structural and amino acid multiple sequence alignment of Spt7 protein and phylogenetic tree analysis A: Structure of Spt7 protein in the yeast and Aspergillus niger. B: Amino acid multiple sequence alignment of Spt7 protein in the yeast and Aspergillus niger. C: Phylogenetic tree of the filamentous fungus Spt7
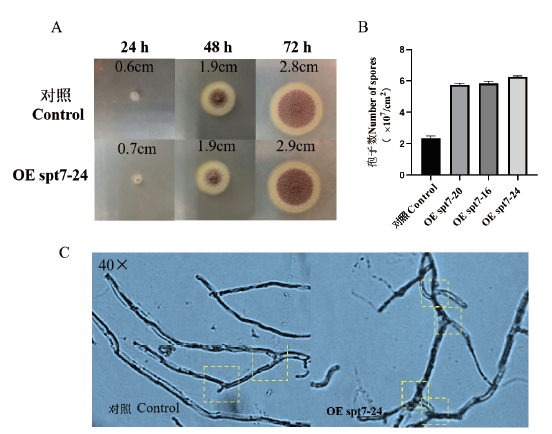
图2 30℃下对照组及转化子的生长形态观察及孢子计数 A:对照组及OE spt7菌落形态;B:孢子计数;C:40倍镜下对照组及OE spt7转化子菌丝形态
Fig. 2 Growth morphology observation and spore count of control and transformants at 30℃ A: Control group and OE spt7 colony morphology. B: Count of spores. C: Mycelial morphology of control and OE spt7 transformants under 40× microscope
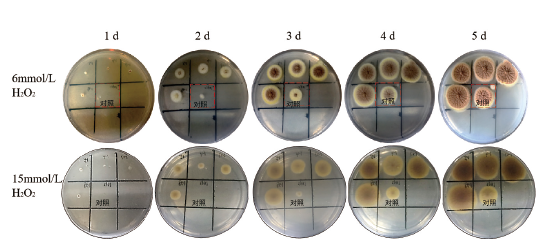
图3 对照组及OE spt7转化子在含有不同浓度H2O2 CM培养基的生长情况
Fig. 3 Growths of control group and OE spt7 transform-ants in CM medium containing different concen-trations of H2O2
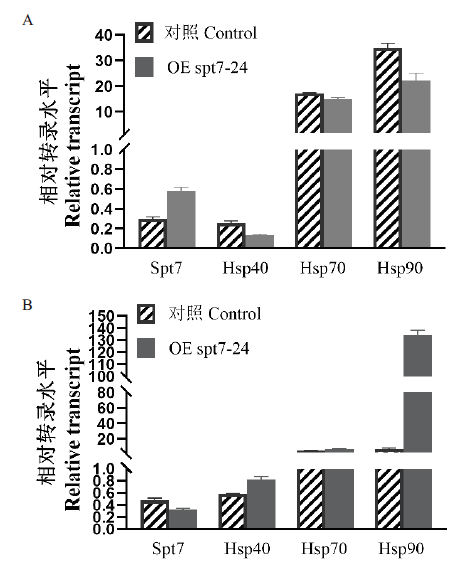
图7 对照组及OE spt7转化子在42℃热激条件下胞内热休克蛋白的转录水平 A:热激3 h后热休克蛋白转录水平;B:热激5 h后热休克蛋白转录水平
Fig. 7 Transcript levels of intracellular heat shock proteins in the control and OE spt7 transformants under heat stress conditions at 42℃ A:Transcript levels of heat shock protein after 3 h of heat stress. B:Transcript levels of heat shock protein after 5 h of heat stress
| [1] |
Soffers JHM, Workman JL. The SAGA chromatin-modifying complex:the sum of its parts is greater than the whole[J]. Genes Dev, 2020, 34(19/20):1287-1303.
doi: 10.1101/gad.341156.120 URL |
| [2] |
Hellman LM, Fried MG. Electrophoretic mobility shift assay(EMSA)for detecting protein-nucleic acid interactions[J]. Nat Protoc, 2007, 2(8):1849-1861.
pmid: 17703195 |
| [3] |
Prasad V, Voigt A. Quantification of protein aggregates using bimolecular fluorescence complementation[J]. Methods Mol Biol, 2019, 1873:183-193.
doi: 10.1007/978-1-4939-8820-4_11 pmid: 30341610 |
| [4] | Kurdistani SK, Grunstein M. Histone acetylation and deacetylation in yeast[J]. Nat Rev Mol Cell Biol, 2003, 4(4):276-284. |
| [5] |
Morgan MT, Haj-Yahya M, Ringel AE, et al. Structural basis for histone H2B deubiquitination by the SAGA DUB module[J]. Science, 2016, 351(6274):725-728.
doi: 10.1126/science.aac5681 URL |
| [6] |
Sung MK, Huh WK. In vivo quantification of protein-protein interactions in Saccharomyces cerevisiae using bimolecular fluorescence complementation assay[J]. J Microbiol Methods, 2010, 83(2):194-201.
doi: 10.1016/j.mimet.2010.08.021 URL |
| [7] |
Doncheva NT, Morris JH, Gorodkin J, et al. Cytoscape StringApp:network analysis and visualization of proteomics data[J]. J Proteome Res, 2019, 18(2):623-632.
doi: 10.1021/acs.jproteome.8b00702 URL |
| [8] |
VanHook AM. Linking light to development and metabolism[J]. Sci Signal, 2008, 1(24):ec222. DOI: 10.1126/scisignal.124ec222.
doi: 10.1126/scisignal.124ec222 |
| [9] | Moraga F, Aquea F. Composition of the SAGA complex in plants and its role in controlling gene expression in response to abiotic stresses[J]. Front Plant Sci, 2015, 6:865. |
| [10] |
Wu PY, Winston F. Analysis of Spt7 function in the Saccharomyces cerevisiae SAGA coactivator complex[J]. Mol Cell Biol, 2002, 22(15):5367-5379.
doi: 10.1128/MCB.22.15.5367-5379.2002 URL |
| [11] |
Winston F, Dollard C, Malone EA, et al. Three genes are required for trans-activation of Ty transcription in yeast[J]. Genetics, 1987, 115(4):649-656.
doi: 10.1093/genetics/115.4.649 pmid: 3034719 |
| [12] |
Gansheroff LJ, Dollard C, Tan P, et al. The Saccharomyces cerevisiae SPT7 gene encodes a very acidic protein important for transcription in vivo[J]. Genetics, 1995, 139(2):523-536.
doi: 10.1093/genetics/139.2.523 pmid: 7713415 |
| [13] |
Papai G, Frechard A, Kolesnikova O, et al. Structure of SAGA and mechanism of TBP deposition on gene promoters[J]. Nature, 2020, 577(7792):711-716.
doi: 10.1038/s41586-020-1944-2 URL |
| [14] |
Wang H, Dienemann C, Stützer A, et al. Structure of the transcription coactivator SAGA[J]. Nature, 2020, 577(7792):717-720.
doi: 10.1038/s41586-020-1933-5 URL |
| [15] |
Li X, Seidel CW, Szerszen LT, et al. Enzymatic modules of the SAGA chromatin-modifying complex play distinct roles in Drosophila gene expression and development[J]. Genes Dev, 2017, 31(15):1588-1600.
doi: 10.1101/gad.300988.117 URL |
| [16] | Torres-Zelada EF, Weake VM. The Gcn5 complexes in Drosophila as a model for Metazoa[J]. Biochim et Biophys Acta BBA Gene Regul Mech, 2021, 1864(2):194610. |
| [17] |
Grasser KD, Rubio V, Barneche F. Multifaceted activities of the plant SAGA complex[J]. Biochim Biophys Acta Gene Regul Mech, 2021, 1864(2):194613.
doi: 10.1016/j.bbagrm.2020.194613 URL |
| [18] |
Mustachio LM, Roszik J, Farria A, et al. Targeting the SAGA and ATAC transcriptional coactivator complexes in MYC-driven cancers[J]. Cancer Res, 2020, 80(10):1905-1911.
doi: 10.1158/0008-5472.CAN-19-3652 pmid: 32094302 |
| [19] |
Wang L, Dent SY. Functions of SAGA in development and disease[J]. Epigenomics, 2014, 6(3):329-339.
doi: 10.2217/epi.14.22 pmid: 25111486 |
| [20] |
Nguyen KT, Ho QN, Pham TH, et al. The construction and use of versatile binary vectors carrying pyrG auxotrophic marker and fluorescent reporter genes for Agrobacterium-mediated transformation of Aspergillus oryzae[J]. World J Microbiol Biotechnol, 2016, 32(12):1-9.
doi: 10.1007/s11274-015-1971-6 URL |
| [21] | 曹张磊, 王德培, 张岚. 提高根癌农杆菌介导黑曲霉转化效率的研究[J]. 天津科技大学学报, 2016, 31(2):20-25. |
| Cao ZL, Wang DP, Zhang L. Improvement of transformation efficiency of Aspergillus niger mediated by Agrobacterium tumefaciens[J]. J Tianjin Univ Sci Technol, 2016, 31(2):20-25. | |
| [22] | 周磊, 韩一帆, 段志强. 含溴结构域蛋白2结构与功能的研究进展[J]. 中国细胞生物学学报, 2021, 43(4):856-865. |
| Zhou L, Han YF, Duan ZQ. Advances in the structure and function of bromodomain-containing protein 2[J]. Chin J Cell Biol, 2021, 43(4):856-865. | |
| [23] | 魏松红, 刘伟, 王海宁, 等. 中国植物病理学会2017年学术年会论文集[C]. 泰安: 中国植物病理学会, 2017. |
| Wei SH, Liu W, Wang HN, et al Proceedings of the 2017 Annual Academic Conference of the Chinese Society of Phytopathology[C]. Tai’an: Chinese Society of Plant Pathology, 2017. | |
| [24] | 包鹏飞, 周敏芬, 汤璧嘉, 等. 黏膜上皮细胞对高渗盐水的耐受性研究[J]. 嘉兴学院学报, 2010, 22(6):29-33. |
| Bao PF, Zhou MF, Tang BJ, et al. On tolerance of epithelial cells in hypertonic saline[J]. J Jiaxing Univ, 2010, 22(6):29-33. | |
| [25] |
Jenkins DE, Schultz JE, Matin A. Starvation-induced cross protection against heat or H2O2 challenge in Escherichia coli[J]. J Bacteriol, 1988, 170(9):3910-3914.
pmid: 3045081 |
| [26] |
Jenkins DE, Auger EA, Matin A. Role of RpoH, a heat shock regulator protein, in Escherichia coli carbon starvation protein synthesis and survival[J]. J Bacteriol, 1991, 173(6):1992-1996.
pmid: 2002001 |
| [27] | Nystroem M, Nuortila-Jokinen J. Current Trends in Membrane Technology[J]. Kemia. Kemi, 1996, 23(8):. 663-672. |
| [28] |
Nyström T. To be or not to be:the ultimate decision of the growth-arrested bacterial cell[J]. FEMS Microbiol Rev, 1998, 21(4):283-290.
doi: 10.1016/S0168-6445(97)00060-0 URL |
| [29] |
Orr WC, Sohal RS. Extension of life-span by overexpression of superoxide dismutase and catalase in Drosophila melanogaster[J]. Science, 1994, 263(5150):1128-1130.
pmid: 8108730 |
| [30] |
Levine RL. Oxidative modification of glutamine synthetase. II. Characterization of the ascorbate model system[J]. J Biol Chem, 1983, 258(19):11828-11833.
pmid: 6137484 |
| [31] |
Kirkman HN, Gaetani GF. Mammalian catalase:a venerable enzyme with new mysteries[J]. Trends Biochem Sci, 2007, 32(1):44-50.
pmid: 17158050 |
| [32] |
Wang Y, Branicky R, Noë A, et al. Superoxide dismutases:Dual roles in controlling ROS damage and regulating ROS signaling[J]. J Cell Biol, 2018, 217(6):1915-1928.
doi: 10.1083/jcb.201708007 URL |
| [33] | Halliwell B, Gutteridge JMC. Redox chemistry:the essentials[M]// Free Radicals in Biology and Medicine. Oxford: Oxford University Press, 2015:30-76. |
| [34] |
Liu XZ, Huang BR. Heat stress injury in relation to membrane lipid peroxidation in creeping bentgrass[J]. Crop Sci, 2000, 40(2):503-510.
doi: 10.2135/cropsci2000.402503x URL |
| [35] |
Xu QZ, Huang BR. Morphological and physiological characteristics associated with heat tolerance in creeping bentgrass[J]. Crop Sci, 2001, 41(1):127-133.
doi: 10.2135/cropsci2001.411127x URL |
| [36] |
Zhang XZ, Ervin EH. Impact of seaweed extract-based cytokinins and Zeatin riboside on creeping bentgrass heat tolerance[J]. Crop Sci, 2008, 48(1):364-370.
doi: 10.2135/cropsci2007.05.0262 URL |
| [37] |
Ostankovitch M, Buchner J. The network of molecular chaperones:insights in the cellular proteostasis machinery[J]. J Mol Biol, 2015, 427(18):2899-2903.
doi: 10.1016/j.jmb.2015.08.010 pmid: 26363891 |
| [38] |
Hartl FU, Bracher A, Hayer-Hartl M. Molecular chaperones in protein folding and proteostasis[J]. Nature, 2011, 475(7356):324-332.
doi: 10.1038/nature10317 URL |
| [39] | Scharf KD, Berberich T, Ebersberger I, et al. The plant heat stress transcription factor(Hsf)family:structure, function and evolution[J]. Biochim Biophys Acta, 2012, 1819(2):104-119. |
| [40] |
Wang WX, Vinocur B, Shoseyov O, et al. Role of plant heat-shock proteins and molecular chaperones in the abiotic stress response[J]. Trends Plant Sci, 2004, 9(5):244-252.
doi: 10.1016/j.tplants.2004.03.006 URL |
| [41] | 蒲力群, 王逢会, 霍满鹏. 热休克蛋白的研究进展[J]. 延安大学学报:自然科学版, 2008, 27(1):72-75. |
| Pu LQ, Wang FH, Huo MP. The study progress of heat shock protein[J]. J Yanan Univ:Nat Sci Ed, 2008, 27(1):72-75. |
| [1] | 郭宇飞, 闫荣媚, 张小茹, 曹威, 刘浩. 代谢工程改造黑曲霉生产葡萄糖二酸[J]. 生物技术通报, 2022, 38(11): 227-237. |
| [2] | 李红叶, 陈立佼, 刘明丽, 郭天杰, 王道平, 潘映红, 赵明. 黑曲霉单宁酶基因Tan2克隆与表达[J]. 生物技术通报, 2021, 37(3): 44-52. |
| [3] | 孟晓建, 于建东, 郑小梅, 郑平, 李志敏, 孙际宾, 叶勤. 小分子代谢物对黑曲霉己糖激酶和丙酮酸激酶的酶活调控[J]. 生物技术通报, 2021, 37(12): 180-190. |
| [4] | 陈浩宇, 徐瑞涛, 程志翔, 高强, 张健. H2O2对黑曲霉氧化胁迫机理的研究[J]. 生物技术通报, 2018, 34(4): 201-207. |
| [5] | 魏姜勉, 鲁雷震, 焦国宝, 刘家扬, 陆隽鹤. 黑曲霉发酵菌渣对臧红T的吸附研究[J]. 生物技术通报, 2017, 33(10): 191-198. |
| [6] | 朱永瑞, 曾柏全, 曾磊, 刘辉. 黑曲霉C112纤维二糖酶基因的克隆与生物信息学分析[J]. 生物技术通报, 2016, 32(2): 116-122. |
| [7] | 张亮, 花尔并, 王华明. 粉红黏帚霉糖化酶基因的克隆、表达及酶学性质分析[J]. 生物技术通报, 2015, 31(7): 193-200. |
| [8] | 张谦, 王剑英, 林智, 贾佳, 郭宏涛. 华根霉脂肪酶在黑曲霉中的重组表达研究[J]. 生物技术通报, 2015, 31(3): 165-170. |
| [9] | 张晓立, 郑小梅, 满云, 罗虎, 于建东, 郑平, 刘浩, 孙际宾. 黑曲霉柠檬酸工业菌株原生质体制备与转化[J]. 生物技术通报, 2015, 31(3): 171-177. |
| [10] | 张谦,贾佳,林智,杨晓锋,郭宏涛,王剑英,Carol Sze Ki Lin. 产脂肪酶黑曲霉摇瓶发酵条件优化研究[J]. 生物技术通报, 2015, 31(12): 227-233. |
| [11] | 张欢, 曹焱鑫, 蒋林, 王晓明, 刘齐, 董晓莹, 寇巍. 黑曲霉(AS0006)产纤维素酶的纯化研究[J]. 生物技术通报, 2014, 0(6): 187-191. |
| [12] | 刘瑜,李丕武. 黑曲霉葡萄糖氧化酶高产基因工程菌研究进展[J]. 生物技术通报, 2013, 0(7): 12-19. |
| [13] | 张文丽,于寒松,朴春红,胡鑫,王舒婷,胡耀辉. 一株高产糖化酶生产菌的筛选与鉴定[J]. 生物技术通报, 2013, 0(7): 131-135. |
| [14] | 于海玲, 李树伟, 王华明. 葡萄穗霉中β-葡萄糖苷酶的基因克隆、表达及酶学性质分析 [J]. 生物技术通报, 2013, 0(6): 128-132. |
| [15] | 王红霞, 王华明, 张大龙, 罗成. 葡萄穗霉木聚糖酶XYA6205 在黑曲霉中的表达及酶学特性分析[J]. 生物技术通报, 2013, 0(2): 130-134. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||