








生物技术通报 ›› 2026, Vol. 42 ›› Issue (2): 149-157.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0974
王晶1( ), 刘卓艳2, 张晓磊1, 刘宝海1, 关海涛1(
), 刘卓艳2, 张晓磊1, 刘宝海1, 关海涛1( ), 温洪涛1(
), 温洪涛1( )
)
收稿日期:2025-09-10
出版日期:2026-02-26
发布日期:2026-03-17
通讯作者:
温洪涛,男,博士,副研究员,研究方向 :转基因农作物产品成分检测技术和安全性评价;E-mail: wen0891@163.com作者简介:王晶,女,博士,助理研究员,研究方向 :生物技术产品检测技术;E-mail: buyijingjing@163.com基金资助:
WANG Jing1( ), LIU Zhuo-yan2, ZHANG Xiao-lei1, LIU Bao-hai1, GUAN Hai-tao1(
), LIU Zhuo-yan2, ZHANG Xiao-lei1, LIU Bao-hai1, GUAN Hai-tao1( ), WEN Hong-tao1(
), WEN Hong-tao1( )
)
Received:2025-09-10
Published:2026-02-26
Online:2026-03-17
摘要:
目的 转化体特异性检测是转基因生物安全监管中识别转化体合法性的重要技术手段,目前转化体特异性检测以实验室检测为主,急需开发能够现场快速检测的方法,满足转基因生物安全监管需求。 方法 以抗虫耐除草剂转基因玉米DBN9501为材料,利用环介导等温扩增(loop-mediated isothermal amplification, LAMP)技术结合CRISPR/Cas12a(clustered regularly interspaced short palindromic repeats,CRISPR)技术(LAMP-CRISPR/Cas12a),辅助试纸条显示,开发转化体特异性快速检测方法。 结果 本方法具有检测特异性,检出限为20拷贝,转化体含量为0.1%,检测时间<60 min。 结论 LAMP-CRISPR/Cas技术能够实现快速检测转基因玉米DBN9501转化体特异性,该方法将为转基因生物安全监管快速检测技术研究提供重要技术支撑。
王晶, 刘卓艳, 张晓磊, 刘宝海, 关海涛, 温洪涛. 基于LAMP-CRISPR/Cas12a技术快速检测转基因玉米DBN9501转化体特异性[J]. 生物技术通报, 2026, 42(2): 149-157.
WANG Jing, LIU Zhuo-yan, ZHANG Xiao-lei, LIU Bao-hai, GUAN Hai-tao, WEN Hong-tao. Rapid Detection of Transgenic Maize DBN9501 Event-specificity Based on LAMP-CRISPR-Cas12a Technology[J]. Biotechnology Bulletin, 2026, 42(2): 149-157.
| 名称 Name | 序列 Sequence (5′-3′) |
|---|---|
| DBN9501-LB-F3 | CCAGCCACTATTAAGTTTCATG |
| DBN9501-LB-B3 | AATCTGTACCTAGTTTAGCTAGT |
| DBN9501-LB-FIP | TGGCAAAAAGGCATCCTGTTTTAAAAAATCCAAAGAAACCATACG |
| DBN9501-LB-BIP | TTCAGGAAAAACAGGGGAAAAGTAGGACGTTAACAGGGACG |
| DBN9501-RB-F3 | TTAATACAAATCCACCTGCG |
| DBN9501-RB-B3 | GAACATTTCTTTATTCGGAAACA |
| DBN9501-RB-FIP | GAGTCACGTTATGACCCCCGTTTTGCGGTTCTGTCAGTTC |
| DBN9501-RB-BIP | AATTCTCCGCTCATGATCAGATTGTAATCCGGACTACTATACCATT |
| crRNA-LB | AAUUUCUACUGUUGUAGAUAAAAAAUCCAAAGAAACCAUACG |
| crRNA-RB | AAUUUCUACUGUUGUAGAUGCGGUUCUGUCAGUUCCAAACG |
| FB-reporter | 6FAM-TTTTTTTTTT-Biotin |
表1 引物、报告探针、crRNA序列信息
Table 1 Sequence information of primers, probe reporters, and CrRNA
| 名称 Name | 序列 Sequence (5′-3′) |
|---|---|
| DBN9501-LB-F3 | CCAGCCACTATTAAGTTTCATG |
| DBN9501-LB-B3 | AATCTGTACCTAGTTTAGCTAGT |
| DBN9501-LB-FIP | TGGCAAAAAGGCATCCTGTTTTAAAAAATCCAAAGAAACCATACG |
| DBN9501-LB-BIP | TTCAGGAAAAACAGGGGAAAAGTAGGACGTTAACAGGGACG |
| DBN9501-RB-F3 | TTAATACAAATCCACCTGCG |
| DBN9501-RB-B3 | GAACATTTCTTTATTCGGAAACA |
| DBN9501-RB-FIP | GAGTCACGTTATGACCCCCGTTTTGCGGTTCTGTCAGTTC |
| DBN9501-RB-BIP | AATTCTCCGCTCATGATCAGATTGTAATCCGGACTACTATACCATT |
| crRNA-LB | AAUUUCUACUGUUGUAGAUAAAAAAUCCAAAGAAACCAUACG |
| crRNA-RB | AAUUUCUACUGUUGUAGAUGCGGUUCUGUCAGUUCCAAACG |
| FB-reporter | 6FAM-TTTTTTTTTT-Biotin |
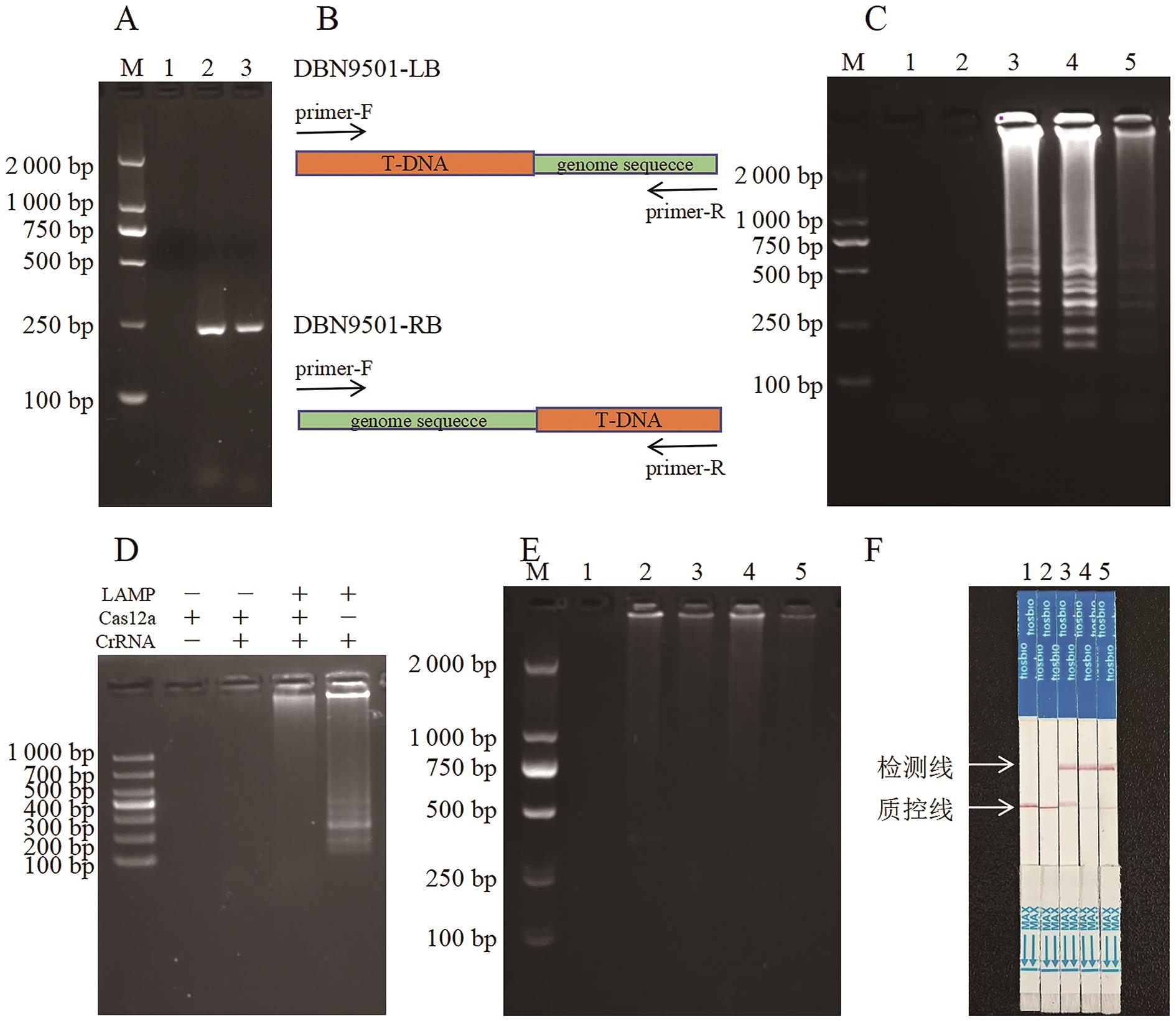
图1 DBN9501转化体特异性的快速检测流程A:LAMP-F3/B3扩增DBN9501转化体LB和RB序列的PCR产物(1:无模板对照;2:LB扩增产物;3:RB扩增产物);B:DBN9501-LB和DBN9501-RB质粒测序结果示意图;C:LAMP扩增结果;D:顺式酶切条件验证实验;E:顺式酶切实验完全切割结果;F:反式酶切试纸条检测结果。C-F图中,1:NTC;2:非转基因玉米;3:DBN9501 转化体;4:DBN9501-LB质粒;5:DBN9501-RB质粒
Fig. 1 Rapid detection workflow of DBN9501 event-specificityA: PCR products of LB and RB sequences of DBN9501 event amplified by LAMP-F3/B3 (1: No template control (NTC) ; 2: LB PCR product; 3:RB PCR product); B: Illustration of DBN9501-LB plasmid sequencing results; C: LAMP amplified results. D: Verification of cis-enzyme digestion condition. E: Trans-enzyme digestion results. F: Strip test assay. In Fig.C-F, 1: NTC; 2: Non-transgenic maize. 3: DBN9501 event. 4: DBN9501-LB plasmid. 5: DBN9501-RB plasmid
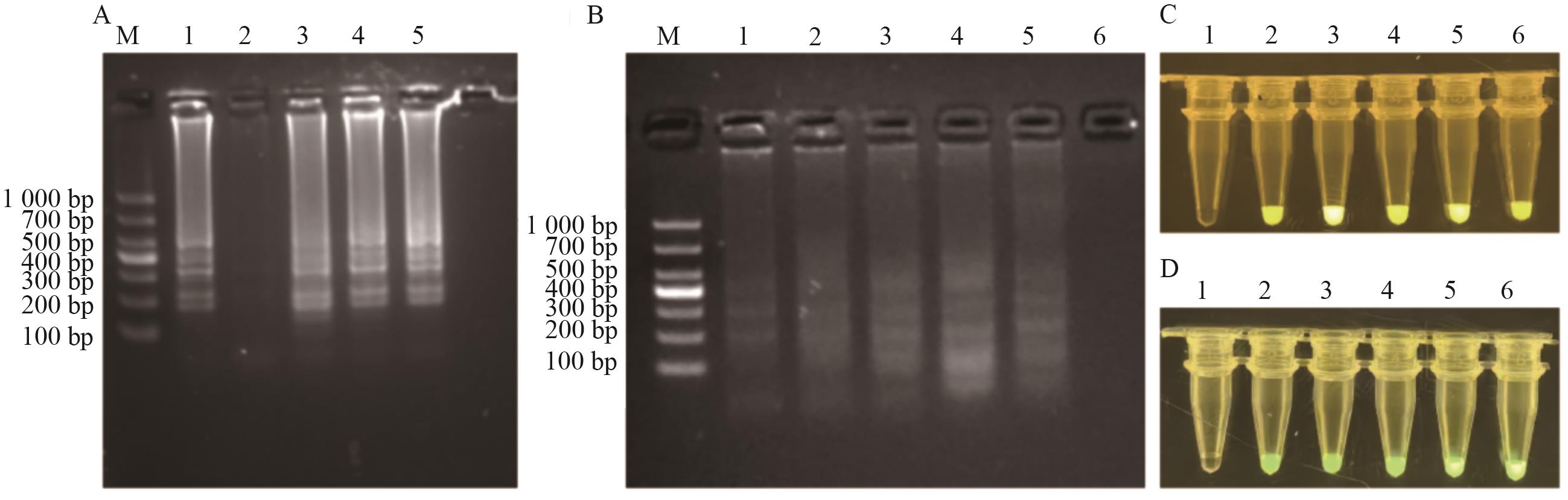
图2 LAMP-CRISPR/Cas12a反应条件优化结果A:LAMP引物FIP/BIP∶F3/B3浓度比例优化;B:LAMP反应温度优化;C:Cas12a/crRNA体积比例优化;D:FB reporter优化
Fig. 2 Optimized results of reaction conditions for the LAMP-CRISPR/Cas12aA: LAMP primer FIP/BIP∶F3/B3 Concentration optimization (1: Positive control (2∶1): 2: NTC; 3: 6∶1; 4: 8∶1; 5: 10∶1). B: Optimization of LAMP reaction temperature (1: 62 ℃; 2: 63 ℃; 3: 64 ℃; 4: 65 ℃; 5: 66 ℃; 6: NTC). C: Optimization of Cas12a/crRNA ratio (1: H2O; 2: 0.1∶0.5; 3: 0.2∶0.4; 4: 0.3∶0.3; 5: 0.4∶0.2; 6: 0.5∶0.1). D: Optimization of FB reporter (1: H2O; 2: 62.5; 3: 50.0; 4: 37.5; 5: 25.0; 6: 12.5 nmol/L)
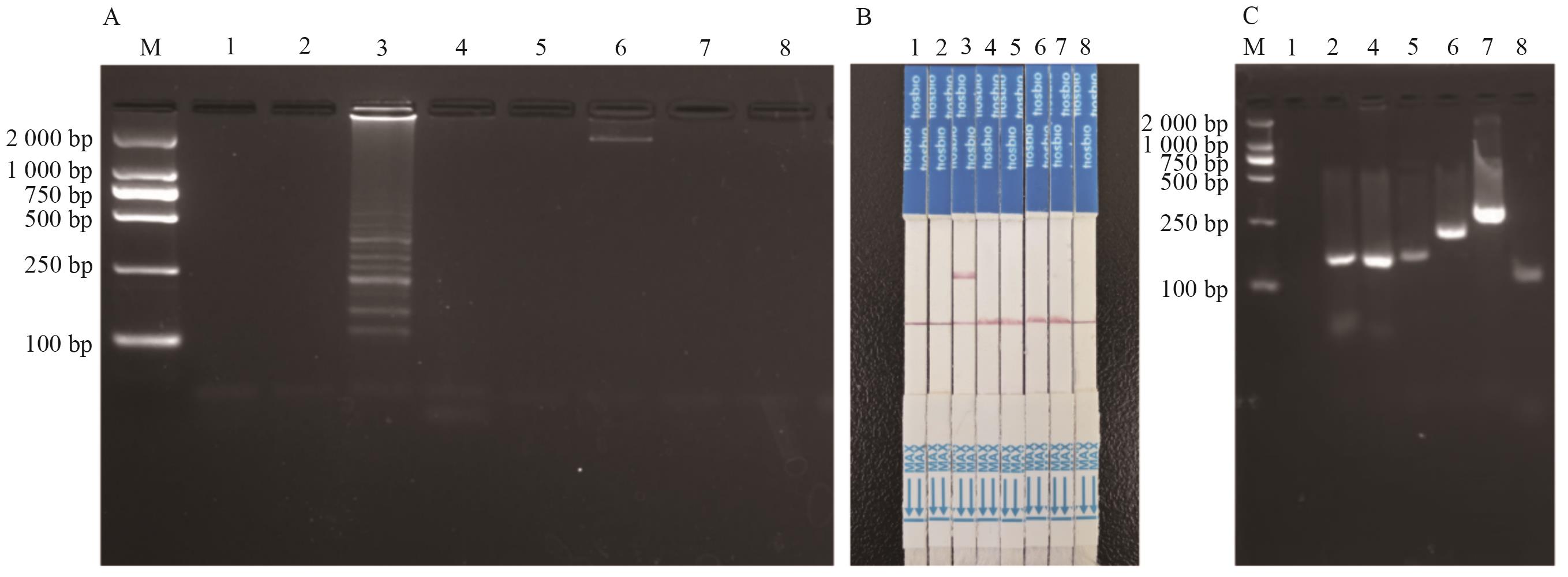
图3 LAMP-CRISPR/Cas12a方法特异性测试A:LAMP扩增结果;B:试纸条检测结果;C:内标准基因PCR扩增结果。1:空白对照;2:非转基因玉米;3:DBN9501转化体;4:转基因玉米(不含DBN9501);5:转基因油菜;6:转基因大豆;7:转基因水稻;8:转基因棉花
Fig. 3 Specificity detection for LAMP-CRISPR/Cas12a methodA: LAMP amplified results; B: strip test assay results; C: internal standard gene PCR amplified results. 1: Blank control; 2: non-transgenic maize; 3: DBN9501 event; 4: transgenic maize (excluding DBN9501); 5:transgenic rape; 6: transgenic soybean; 7: transgenic rice; 8: transgenic cotton
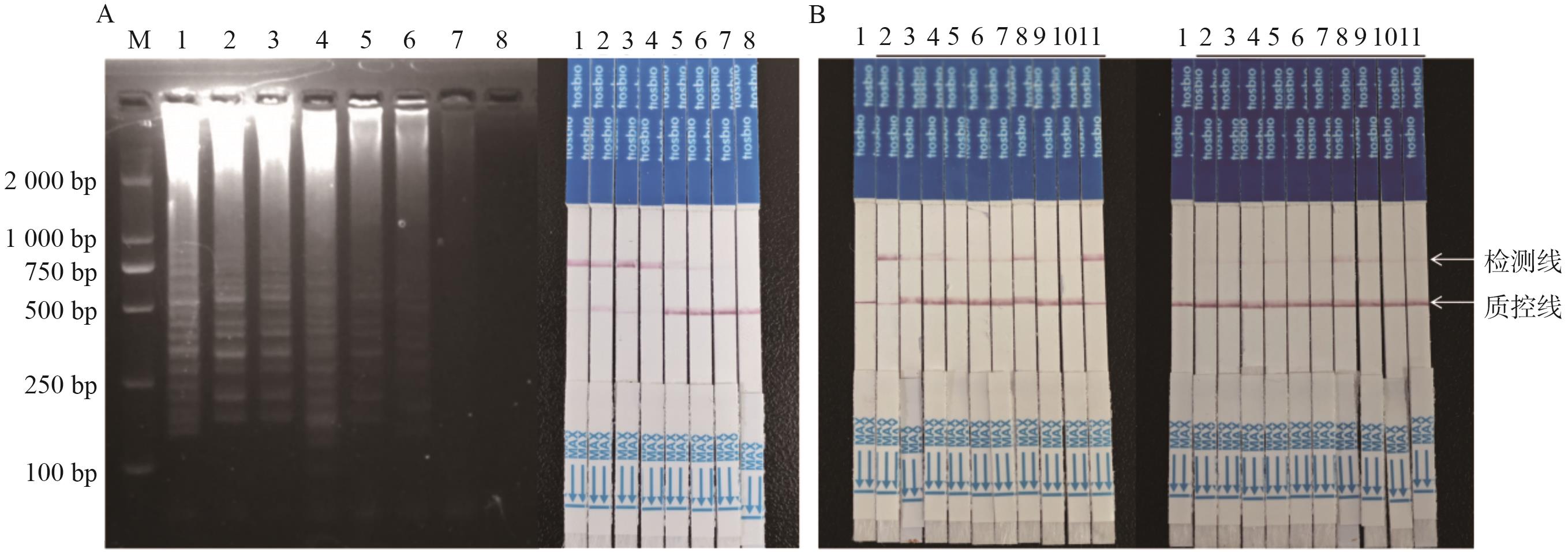

图4 LAMP-CRISPR/Cas12a方法灵敏度和检出限测试A:灵敏度测试,左图为LAMP扩增结果,右图为试纸条检测;B:检出限测试,左图为质粒检出限测试,右图为DBN9501转化体试纸条测试
Fig. 4 Assay of sensitivity and limit of detection for LAMP-CRISPR/Cas12a methodA: Sensitivity assay, the left picture is LAMP amplified results; the right picture is strip detection assay (1: 8.0×103; 2: 8.0×102; 3: 8.0×10; 4: 4.0×10; 5: 2.0×10; 7: 1.0×10; 7: 1.0; 8: NTC). B: Assay of detection limit, left picture is the assay of detection limit for plasmid, and right picture is DBN9501 event by strip test assay (1: NTC; 2-11: 20 copies DBN9501-LB plasmid)
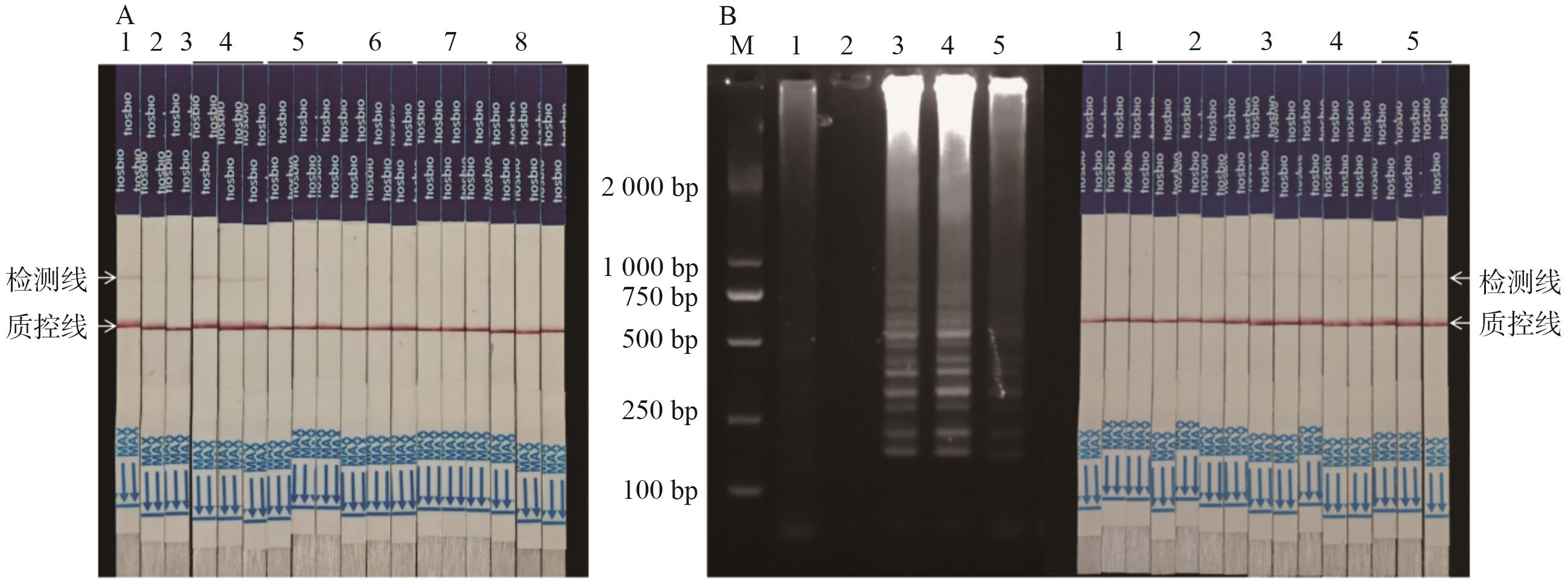
图5 盲样检测和方法应用A:盲样测试(1:DBN9501-LB;2:非转基因玉米;3:NTC;4:盲样(DBN9501);5:盲样(非转基因玉米);6:盲样(Bt11);7:盲样(DBN9936);8:盲样(MON810));B:检测方法应用,左图为LAMP扩增结果,右图为试纸条测试(M:marker;1:非转基因玉米;2:NTC;3:DBN9501-LB;4:DBN9501叶子;5:DBN9501种子,试纸条试验3次重复)
Fig. 5 Blind sample testing and method applicationA: Blind sample test (1: DBN9501-LB; 2: non-transgenic maize; 3: NTC; 4: blind sample (DBN9501). 5: Blind samples (non-transgenic maize); 6: blind sample (Bt11); 7: blind sample (DBN9936); 8: blind sample (MON810)). B: Application of assay method, the left is LAMP amplification results, the right is a test strip test (M: marker; 1: non-transgenic maize; 2: NTC; 3: DBN9501-LB; 4: DBN9501 leaves; 5: DBN9501 seeds, 3 replicates for strip test assay)
方法 Method | 操作步骤 Procedure | 检测对象 Test object | 灵敏度 Sensitivity | 检测时间 Testing time | 参考文献 References |
|---|---|---|---|---|---|
| RPA-Cas12a-FS | (1) RPA (2) CRISPR/Cas12a (3) 荧光/试纸条显示 | P-CaMV35S,T-nos | 10拷贝 | ~45 min | [ |
| MR-DCA | (1) 多重RPA (2) CRISPR/Cas13a或Cas12a+荧光/试纸条显示 | CaMV35S,NOS | 20拷贝 | ~35 min | [ |
| RPA-CRISPR/Cas12a | (1) RPA (2) CRISPR/Cas (3) G-四链体比色测定 | CaMV35 | 10 amol/L和0.01% | ~45 min | [ |
| RPA-CRISPR/Cas12a | (1) RPA (2) CRISPR/Cas12a (3) 荧光/试纸条显示 | CP4-EPSPS,Cry1Ab/Ac | 10 拷贝,0.5% | ~50 min | [ |
| RAA-CRISPR-Cas12a | (1) 重组酶辅助扩增(RAA); (2) CRISPR/Cas12a (3) 裸眼观察金纳米棒颜色变化 | NOS | 1 amol/L和0.1% | ~60 min | [ |
| LAMP-CRISPR/Cas12b | (1) LAMP (2) CRISPR/Cas12b (3) 荧光显示 | CaMV35S | 10拷贝,0.05% | ~40 min | [ |
| LAMP-CRISPR/Cas12a | (1) LAMP (2) CRISPR/Cas 12a (3) 试纸条显示 | DBN9501 | 20拷贝 | ~60 min | 本方法 |
表2 基于CRISPR/Cas技术的转基因检测时间比较
Table 2 Comparison of transgenic detecting time based on CRISPR/Cas technology
方法 Method | 操作步骤 Procedure | 检测对象 Test object | 灵敏度 Sensitivity | 检测时间 Testing time | 参考文献 References |
|---|---|---|---|---|---|
| RPA-Cas12a-FS | (1) RPA (2) CRISPR/Cas12a (3) 荧光/试纸条显示 | P-CaMV35S,T-nos | 10拷贝 | ~45 min | [ |
| MR-DCA | (1) 多重RPA (2) CRISPR/Cas13a或Cas12a+荧光/试纸条显示 | CaMV35S,NOS | 20拷贝 | ~35 min | [ |
| RPA-CRISPR/Cas12a | (1) RPA (2) CRISPR/Cas (3) G-四链体比色测定 | CaMV35 | 10 amol/L和0.01% | ~45 min | [ |
| RPA-CRISPR/Cas12a | (1) RPA (2) CRISPR/Cas12a (3) 荧光/试纸条显示 | CP4-EPSPS,Cry1Ab/Ac | 10 拷贝,0.5% | ~50 min | [ |
| RAA-CRISPR-Cas12a | (1) 重组酶辅助扩增(RAA); (2) CRISPR/Cas12a (3) 裸眼观察金纳米棒颜色变化 | NOS | 1 amol/L和0.1% | ~60 min | [ |
| LAMP-CRISPR/Cas12b | (1) LAMP (2) CRISPR/Cas12b (3) 荧光显示 | CaMV35S | 10拷贝,0.05% | ~40 min | [ |
| LAMP-CRISPR/Cas12a | (1) LAMP (2) CRISPR/Cas 12a (3) 试纸条显示 | DBN9501 | 20拷贝 | ~60 min | 本方法 |
| [1] | 王晶, 张晓磊, 白玉, 等. 不同玉米转化体通用PCR检测体系建立 [J]. 生物技术通报, 2024, 40(12): 34-44. |
| Wang J, Zhang XL, Bai Y, et al. Establishment of a universal PCR detection system for different maize events [J]. Biotechnol Bull, 2024, 40(12): 34-44. | |
| [2] | 王颢潜, 高鸿飞, 王梦雨, 等. 转基因生物成分快速检测技术研究进展 [J]. 中国油料作物学报, 2022, 44(3): 491-496. |
| Wang HQ, Gao HF, Wang MY, et al. Research progress on the rapid detection technologies for composition of genetically modified organisms [J]. Chin J Oil Crop Sci, 2022, 44(3): 491-496. | |
| [3] | Notomi T, Okayama H, Masubuchi H, et al. Loop-mediated isothermal amplification of DNA [J]. Nucleic Acids Res, 2000, 28(12): e63. |
| [4] | Han X, Lu MH, Zhang YR, et al. A thermostable Cas12b-powered bioassay coupled with loop-mediated isothermal amplification in a customized "one-pot" vessel for visual, rapid, sensitive, and on-site detection of genetically modified crops [J]. J Agric Food Chem, 2024, 72(19): 11195-11204. |
| [5] | Liang Y, He YH, Yang SF, et al. A novel cross-priming amplification technique combined with lateral flow strips for rapid and visual detection of zoonotic Toxoplasma gondii [J]. Vet Parasitol, 2025, 334: 110402. |
| [6] | Zhu XY, Yang H, Wang M, et al. Label-free detection of transgenic crops using an isothermal amplification reporting CRISPR/Cas12 assay [J]. ACS Synth Biol, 2022, 11(1): 317-324. |
| [7] | Ding L, Wang XF, Chen XY, et al. Development of a novel Cas13a/Cas12a-mediated ‘one-pot’ dual detection assay for genetically modified crops [J]. J Adv Res, 2025, 72: 97-106. |
| [8] | Park JW. Principles and applications of loop-mediated isothermal amplification to point-of-care tests [J]. Biosensors (Basel), 2022, 12(10): 857. |
| [9] | 中华人民共和国农业农村部. 转基因植物及其产品成分检测 环介导等温扩增方法制定指南: 农业农村部公告第323号-9—2020 [S]. 北京:中国农业出版社, 2020: 3. |
| Ministry of Agriculture and Rural Affairs of the People’s Republic of China. Detection of genetically modified plants and derived products-Guide for establishing loop-mediated isothermal amplification method: Announcement No. 323-9-2020 of the Ministry of Agriculture and Rural Affairs of the People’s Republic of China [S]. Beijing: China Agriculture Press, 2020: 3. | |
| [10] | Chylinski K, Makarova KS, Charpentier E, et al. Classification and evolution of type II CRISPR-Cas systems [J]. Nucleic Acids Res, 2014, 42(10): 6091-6105. |
| [11] | Zhu ZB, Li R, Zhang HW, et al. PAM-free loop-mediated isothermal amplification coupled with CRISPR/Cas12a cleavage (Cas-PfLAMP) for rapid detection of rice pathogens [J]. Biosens Bioelectron, 2022, 204: 114076. |
| [12] | 王渭霞, 朱廷恒, 赖凤香, 等. CRISPR/Cas系统在基因修饰植物及其产品检测应用中的原理和进展 [J]. 中国稻米, 2023, 29(6): 21-27. |
| Wang WX, Zhu TH, Lai FX, et al. Principle and progress of CRISPR/cas system in the detection of genetically modified plants and their products [J]. China Rice, 2023, 29(6): 21-27. | |
| [13] | Wang HM, Su AL, Chang JJ, et al. Sensitive detection of genetically modified maize based on a CRISPR/Cas12a system [J]. Analyst, 2024, 149(3): 836-845. |
| [14] | Li GY, Li SN, Li XH, et al. A novel electrochemical aptasensor based on NrGO-H-Mn3O4 NPs integrated CRISPR/Cas12a system for ultrasensitive low-density lipoprotein determination [J]. Microchim Acta, 2024, 191(9): 547. |
| [15] | Wang YH, Yang TM, Liu GF, et al. Application of CRISPR/Cas12a in the rapid detection of pathogens [J]. Clin Chim Acta, 2023, 548: 117520. |
| [16] | Xue PP, Peng YB, Wang RJ, et al. Advances, challenges, and opportunities for food safety analysis in the isothermal nucleic acid amplification/CRISPR-Cas12a era [J]. Crit Rev Food Sci Nutr, 2025, 65(13): 2473-2488. |
| [17] | Cao GH, Dong JB, Chen XL, et al. Simultaneous detection of CaMV35S and T-nos utilizing CRISPR/Cas12a and Cas13a with multiplex-PCR (MPT-Cas12a/13a) [J]. Chem Commun, 2022, 58(43): 6328-6331. |
| [18] | Huang D, Qian JJ, Shi ZW, et al. CRISPR-Cas12a-assisted multicolor biosensor for semiquantitative point-of-use testing of the nopaline synthase Terminator in genetically modified crops by unaided eyes [J]. ACS Synth Biol, 2020, 9(11): 3114-3123. |
| [19] | Wang Z, Huang CM, Wei S, et al. A CRISPR/Cas12a-mediated sensitive DNA detection system for gene-edited rice [J]. J AOAC Int, 2023, 106(3): 558-567. |
| [20] | Chen L, Wang C, Zhu ZB, et al. Field-deployable detection of genetically modified organisms with an integrated method of loop-mediated isothermal amplification and CRISPR/FnCas12a [J]. J Agric Food Chem, 2025, 73(9): 5625-5634. |
| [21] | Li R, Chen JW, Zhang XJ, et al. Mini-disk capillary array coupling with LAMP for visual detection of multiple nucleic acids using genetically modified organism analysis as an example [J]. J Agric Food Chem, 2020, 68(3): 899-906. |
| [22] | Wu H, He JS, Zhang F, et al. Contamination-free visual detection of CaMV35S promoter amplicon using CRISPR/Cas12a coupled with a designed reaction vessel: Rapid, specific and sensitive [J]. Anal Chim Acta, 2020, 1096: 130-137. |
| [23] | Wang J, Xiao B, Zhang RY, et al. Inter-laboratory validation by event-specific qPCR methods for the detection of genetically modified insect and herbicide-tolerant maize DBN9501 [J]. J Consum Prot Food Saf, 2023, 18(4): 451-460. |
| [24] | Li SY, Cheng QX, Liu JK, et al. CRISPR-Cas12a has both Cis- and trans-cleavage activities on single-stranded DNA [J]. Cell Res, 2018, 28(4): 491-493. |
| [25] | Liu H, Wang JB, Zeng HJ, et al. RPA-Cas12a-FS: a frontline nucleic acid rapid detection system for food safety based on CRISPR-Cas12a combined with recombinase polymerase amplification [J]. Food Chem, 2021, 334: 127608. |
| [26] | Li XH, Liu ML, Men DH, et al. Rapid, portable, and sensitive detection of CaMV35S by RPA-CRISPR/Cas12a-G4 colorimetric assays with high accuracy deep learning object recognition and classification [J]. Talanta, 2024, 278: 126441. |
| [27] | Wang JB, Luo JW, Liu H, et al. “Blue-red-purple” multicolored lateral flow immunoassay for simultaneous detection of GM crops utilizing RPA and CRISPR/Cas12a [J]. Talanta, 2025, 282: 127010. |
| [28] | Holst-Jensen A, Bertheau Y, de Loose M, et al. Detecting un-authorized genetically modified organisms (GMOs) and derived materials [J]. Biotechnol Adv, 2012, 30(6): 1318-1335. |
| [29] | Shehata HR, Ragupathy S, Shanmughanandhan D, et al. Guidelines for validation of qualitative real-time PCR methods for molecular diagnostic identification of probiotics [J]. J Aoac Int, 2019, 102(6): 1774-1778. |
| [30] | 麦晓东, 石磊, 常彦磊, 等. 环介导等温扩增技术(LAMP)在转基因大豆DBN9004品系快速检测中的应用 [J/OL]. 现代食品科技, 2025: 1-10. . |
| Mai XD, Shi L, Chang YL, et al. Application of Loop-mediated isothermal amplification (LAMP) in rapid identification of genetically modified soybean event DBN 9004. Modern food Sci Technol [J/OL]. Mod Food Sci Technol, 2025: 1-10. | |
| [31] | 咸若彤, 缪青梅, 彭城, 等. 转基因玉米WYN17132转化体特异性实时荧光PCR检测方法的建立与应用 [J]. 浙江农业学报, 2025, 37(7): 1397-1406. |
| Xian RT, Miao QM, Peng C, et al. Establishment and application of event-specific real-time PCR detection method of transgenic maize WYN17132 [J]. Acta Agric Zhejiangensis, 2025, 37(7): 1397-1406. |
| [1] | 李雅琦, 孙萌, 李秀丽, 魏静娜, 赵琳琳, 赵云平, 刘征辉, 苏蘩. 多种Cas12a蛋白普适性的高性能低成本荧光检测缓冲体系优化研究[J]. 生物技术通报, 2026, 42(4): 83-91. |
| [2] | 王婷婷, 何孟雅, 盛家顺, 高晨, 蔡含芳, 付彤, 孙宇, 高腾云, 张天留. 基于CRISPR/Cas9技术制备NCAPG基因敲除牛成纤维细胞系[J]. 生物技术通报, 2026, 42(2): 317-324. |
| [3] | 曾厅, 张兰, 罗睿. 转录因子MpR2R3-MYB17调控地钱胞芽发育的功能研究[J]. 生物技术通报, 2026, 42(1): 208-217. |
| [4] | 余永霞, 杜再慧, 朱龙佼, 许文涛. 基因编辑技术在牛种中的应用及研究进展[J]. 生物技术通报, 2025, 41(8): 34-41. |
| [5] | 刘华, 宋洁, 曾海娟, 王金斌, 钱韻芳. 单碱基突变检测方法及应用的研究进展[J]. 生物技术通报, 2025, 41(6): 61-70. |
| [6] | 霍贯中, 张欣濡, 田士军, 李君. CRISPR/Cas12a基因编辑技术在植物中的研究进展[J]. 生物技术通报, 2025, 41(6): 1-11. |
| [7] | 高畅, 庄添驰, 李宁, 刘云, 顾鹏飞, 赵昕怡, 季明辉. RPA-CRISPR/Cas12a结合重力驱动微流控芯片的MTB快检方法的建立[J]. 生物技术通报, 2025, 41(5): 62-69. |
| [8] | 文博霖, 万敏, 胡建军, 王克秀, 景晟林, 王心悦, 朱博, 唐铭霞, 李兵, 何卫, 曾子贤. 马铃薯川芋50遗传转化及基因编辑体系的建立[J]. 生物技术通报, 2025, 41(4): 88-97. |
| [9] | 卢勇杰, 夏海乾, 李永铃, 张文建, 余婧, 赵会纳, 王兵, 许本波, 雷波. 烟草AP2/ERF转录因子NtESR2的克隆及功能分析[J]. 生物技术通报, 2025, 41(4): 266-277. |
| [10] | 秦玉婷, 潘森涛, 陈渝萍. 非编码RNA的引导RNA的设计及其应用[J]. 生物技术通报, 2025, 41(3): 71-82. |
| [11] | 刘梓琦, 钟沛, 李琴, 郭成, 张艳梅, 张乃锋, 屠焰, 刁其玉, 毕研亮. CRISPR/Cas9技术在益生菌编辑中的应用与进展[J]. 生物技术通报, 2025, 41(11): 89-99. |
| [12] | 崔海洋, 谭淼, 全壮, 陈红利, 董艳敏, 唐立春. 利用Cas9TX实现非病毒TRAC定点整合制备T细胞[J]. 生物技术通报, 2024, 40(9): 190-197. |
| [13] | 侯文婷, 孙琳, 张艳军, 董合忠. 基因编辑技术在棉花种质创新和遗传改良中的应用[J]. 生物技术通报, 2024, 40(7): 68-77. |
| [14] | 陈墨岩, 祝诚. 基于CRISPR/Cas12a的生物传感平台的机制研究及应用[J]. 生物技术通报, 2024, 40(7): 90-98. |
| [15] | 朱恬仪, 孔桂美, 焦红梅, 郭停停, 乌日汗, 刘翠翠, 高成凤, 李国才. CRISPR/Cas9介导的adeG基因敲除大肠杆菌细菌模型的建立[J]. 生物技术通报, 2024, 40(2): 55-64. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||