








Biotechnology Bulletin ›› 2022, Vol. 38 ›› Issue (5): 269-278.doi: 10.13560/j.cnki.biotech.bull.1985.2021-0892
Previous Articles Next Articles
Received:2021-07-12
Online:2022-05-26
Published:2022-06-10
Contact:
DUAN Xu-guo
E-mail:870129814@qq.com;xgduan@njfu.edu.cn
ZHU Qiu-yu, DUAN Xu-guo. Recombinant Expression and Site-directed Mutagenesis of L-aspartate-α-decarboxylase,and the Establishment of High-throughput Assay Method[J]. Biotechnology Bulletin, 2022, 38(5): 269-278.
| 引物Primer | 引物序列Primer sequence(5'-3') |
|---|---|
| BspanD-F | GAAGGAGATATACATATGTATCGAACAATGATGAGCGG |
| BspanD-R | GCTTGTCGACGGAGCTCCTACAAAATTGTACGGGCTGGT |
| I88M-F | GGTCATTATTATGTCCCACAAAATGATGTC |
| I88M-R | GACATCATTTTGTGGGACATAATAATGACC |
| I46V-F | GAAAAAGTACAAGTTGTGAATAATAATAATGGAG |
| I46V-R | CTCCATTATTATTATTCACAACTTGTACTTTTTC |
| K104S-F | CCATGAGCCGAGTGTGGCTGTTCT |
| K104S-R | AGAACAGCCACACTCGGCTCATGG |
| I126*-F | GAACCAGCCCGTACATAATTGTAGGAGC |
| I126*-R | GCTCCTACAATTATGTACGGGCTGGTTC |
Table 1 Primers used in this study
| 引物Primer | 引物序列Primer sequence(5'-3') |
|---|---|
| BspanD-F | GAAGGAGATATACATATGTATCGAACAATGATGAGCGG |
| BspanD-R | GCTTGTCGACGGAGCTCCTACAAAATTGTACGGGCTGGT |
| I88M-F | GGTCATTATTATGTCCCACAAAATGATGTC |
| I88M-R | GACATCATTTTGTGGGACATAATAATGACC |
| I46V-F | GAAAAAGTACAAGTTGTGAATAATAATAATGGAG |
| I46V-R | CTCCATTATTATTATTCACAACTTGTACTTTTTC |
| K104S-F | CCATGAGCCGAGTGTGGCTGTTCT |
| K104S-R | AGAACAGCCACACTCGGCTCATGG |
| I126*-F | GAACCAGCCCGTACATAATTGTAGGAGC |
| I126*-R | GCTCCTACAATTATGTACGGGCTGGTTC |
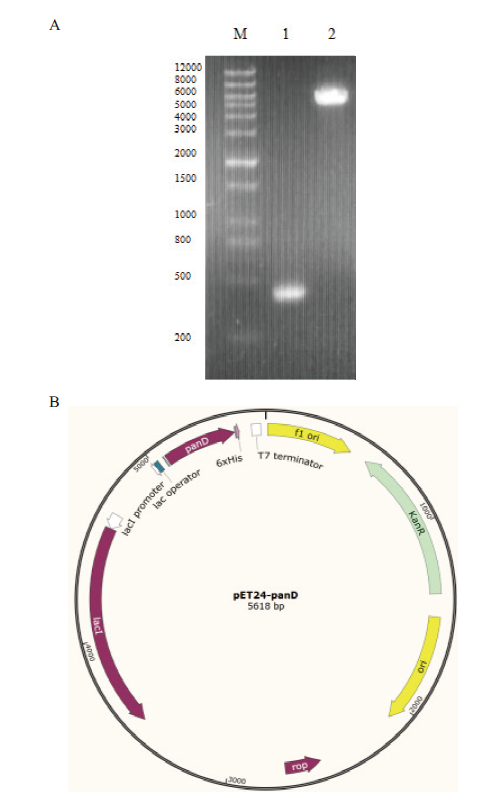
Fig. 1 Agarose gel electropherogram and structural sch-ematic diagram of expression vector A:PCR product of panDBs and plasmid enzyme digestion product(M:DNA marker;1:PCR product of panDBs;2:double restriction enzyme digestion product of plasmid pET24a(+)). B:Structural schematic diagram of recombinant expression vector pET24a(+)-panDBs
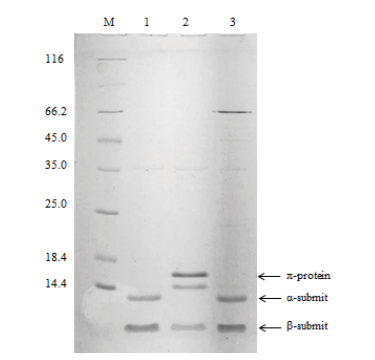
Fig. 2 Electropherogram of recombinant protein panDBs and its mutant proteins M:Protein marker. 1:Supernatant of panDBs. 2:Supernatant of mutant panDBs-1. 3:Supernatant of mutant panDBs-2
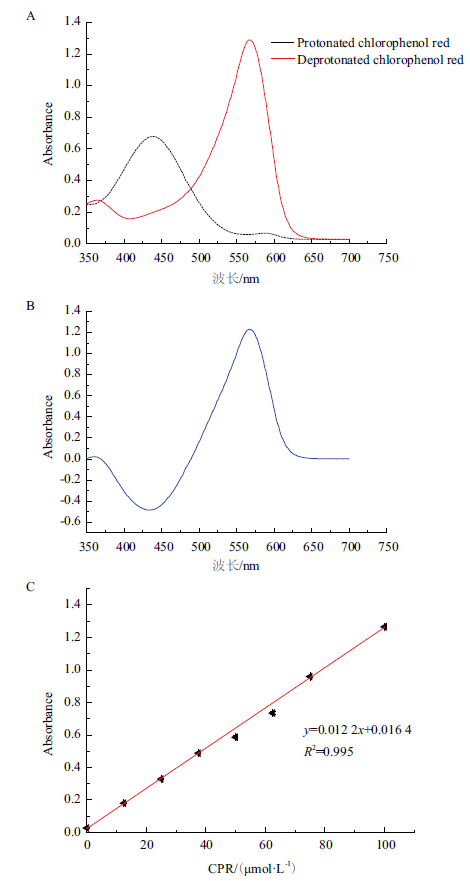
Fig. 4 Development of a colorimetric high-throughput scr-eening method for PanD assay A:Absorption spectra of protonated and deprotonated forms of CPR. B:Difference in absorption spectra between the two forms of CPR. C:Absorbance at 567 nm as a function of the concentration of CPR
| Mutation sites | Vmax/ (μmol·min-1·mg-1) | Km/ (mmol·L-1) | |
|---|---|---|---|
| panDBs | - | 0.61±0.06 | 2.78±0.09 |
| panDBs-1 | I46V/I88M/K104S/I126* | 0.68±0.03 | 2.98±0.16 |
| panDBs-2 | I88M | 1.11±0.11 | 1.49±0.05 |
Table 2 Partial kinetic parameters of recombinant enzyme and its mutants
| Mutation sites | Vmax/ (μmol·min-1·mg-1) | Km/ (mmol·L-1) | |
|---|---|---|---|
| panDBs | - | 0.61±0.06 | 2.78±0.09 |
| panDBs-1 | I46V/I88M/K104S/I126* | 0.68±0.03 | 2.98±0.16 |
| panDBs-2 | I88M | 1.11±0.11 | 1.49±0.05 |
| [1] |
Williamson JM, Brown GM. Purification and properties of L-Aspartate-alpha-decarboxylase, an enzyme that catalyzes the formation of beta-alanine in Escherichia coli[J]. J Biol Chem, 1979, 254(16):8074-8082.
pmid: 381298 |
| [2] | 范海洋. 重组大肠杆菌L-天冬氨酸-α-脱羧酶的制备及应用研究[D]. 上海: 华东理工大学, 2013. |
| Fan HY. Preparation and application of recombinant Escherichia coli L-aspartate-a-decdarboxylase[D]. Shanghai: East China University of Science and Technology, 2013. | |
| [3] | 赵连真, 张梁, 石贵阳. 谷氨酸棒杆菌L-天冬氨酸α-脱羧酶在大肠杆菌中的表达及酶转化生产β-丙氨酸[J]. 微生物学通报, 2013, 40(12):2161-2170. |
| Zhao LZ, Zhang L, Shi GY. Expression of L-aspartate α-decarboxylase from Corynebacterium glutamicum in Escherichia coli and its application in enzymatic synthesis of β-alanine[J]. Microbiol China, 2013, 40(12):2161-2170. | |
| [4] | 邓思颖, 张君丽, 蔡真, 等. 枯草芽胞杆菌L-天冬氨酸α-脱羧酶的酶学性质[J]. 生物工程学报, 2015, 31(8):1184-1193. |
| Deng SY, Zhang JL, Cai Z, et al. Characterization of L-aspartate-α-decarboxylase from Bacillus subtilis[J]. Chin J Biotechnol, 2015, 31(8):1184-1193. | |
| [5] | 陈涛, 徐世永, 冯炎. 结核杆菌L-天冬氨酸α-脱羧酶诱导表达条件研究[J]. 金陵科技学院学报, 2016, 32(3):80-83. |
| Chen T, Xu SY, Feng Y. Inducing conditions of recombined L-aspartate-α-decarboxylase in fermentor[J]. J Jinling Inst Technol, 2016, 32(3):80-83. | |
| [6] |
Kwon AR, Lee BI, Han BW, et al. Crystallization and preliminary X-ray crystallographic analysis of aspartate 1-decarboxylase from Helicobacter pylori[J]. Acta Crystallogr D Biol Crystallogr, 2002, 58(pt 5):861-863.
doi: 10.1107/S0907444902004833 URL |
| [7] |
Schmitzberger F, Kilkenny ML, Lobley CM, et al. Structural constraints on protein self-processing in L-aspartate-alpha-decarboxylase[J]. EMBO J, 2003, 22(23):6193-6204.
pmid: 14633979 |
| [8] | Li HH, Lu XL, Chen KQ, et al. β-alanine production using whole-cell biocatalysts in recombinant Escherichia coli[J]. Mol Catal, 2018, 449:93-98. |
| [9] |
Jang YS, Kim B, Shin JH, et al. Bio-based production of C2-C6 platform chemicals[J]. Biotechnol Bioeng, 2012, 109(10):2437-2459.
doi: 10.1002/bit.24599 URL |
| [10] | 张腾辉. L-天冬氨酸α-脱羧酶的表达、改造及全细胞制备β-丙氨酸[D]. 无锡: 江南大学, 2018. |
| Zhang TH. Expression and modification of L-aspartate α-decarboxylase for the whole-cell transformation of β-alanine[D]. Wuxi: Jiangnan University, 2018. | |
| [11] | 莫芹, 李由然, 石贵阳. 细菌L-天冬氨酸α-脱羧酶的分子机制及分子改造研究进展[J]. 微生物学通报, 2018, 45(7):1546-1554. |
| Mo Q, Li YR, Shi GY. Advances in molecular mechanism and modification of bacterial L-aspartate alpha-decarboxylase[J]. Microbiol China, 2018, 45(7):1546-1554. | |
| [12] | 莫芹. L-天冬氨酸α-脱羧酶催化失活相关分子机制的研究[D]. 无锡: 江南大学, 2019. |
| Mo Q. Molecular mechanism of the catalytic inactivation of L-aspartate alpha-decarboxylase[D]. Wuxi: Jiangnan University, 2019. | |
| [13] | 邢艳珑, 毛相朝, 王舒, 等. 应用荧光分析法检测酶的研究进展[J]. 生物工程学报, 2009, 25(12):1765-1769. |
| Xing YL, Mao N, et al. Recent advances in enzyme assays using fluoremetry[J]. Chin J Biotechnol, 2009, 25(12):1765-1769. | |
| [14] | 刘艳莉, 杨广宇, 王秋岩, 等. 脂肪酶和酯酶的定向进化及其应用[J]. 生物加工过程, 2006, 4(1):16-20, 26. |
| Liu YL, Yang GY, Wang QY, et al. Methods and application of directed evolution of lipase and esterase[J]. Chin J Bioprocess Eng, 2006, 4(1):16-20, 26. | |
| [15] |
Yu XJ, Huang CY, Xu XD, et al. Protein engineering of a pyridoxal-5'-phosphate-dependent l-aspartate-α-decarboxylase from Tribolium castaneum for β-alanine production[J]. Molecules, 2020, 25(6):1280.
doi: 10.3390/molecules25061280 URL |
| [16] | 陈虹. L-天冬氨酸-α-脱羧酶的蛋白质工程改造及其在β-丙氨酸生产中的应用[D]. 杭州: 浙江工业大学, 2019. |
| Chen H. Protein engineering of a L-aspartate-α-decarboxylase and its application in the β-alanine production[D]. Hangzhou: Zhejiang University of Technology, 2019. | |
| [17] | 王鹏. 氨基酸脱羧酶重组表达及应用研究[D]. 南京: 南京大学, 2015. |
| Wang P. Study on recombinant expression and application of amino acid decarboxylase[D]. Nanjing: Nanjing University, 2015. | |
| [18] |
Yu K, Hu S, Huang J, et al. A high-throughput colorimetric assay to measure the activity of glutamate decarboxylase[J]. Enzyme Microb Technol, 2011, 49(3):272-276.
doi: 10.1016/j.enzmictec.2011.06.007 URL |
| [19] |
Jiang H, Xia XX, Feng Y, et al. Development of a robust system for high-throughput colorimetric assay of diverse amino acid decarboxylases[J]. Process Biochem, 2017, 60:27-34.
doi: 10.1016/j.procbio.2017.05.028 URL |
| [20] |
Rosenberg RM, Herreid RM, et al. Indicator assay for amino acid decarboxylases[J]. Anal Biochem, 1989, 181(1):59-65.
pmid: 2817382 |
| [21] |
Gibbons BH, Edsall JT. Rate of hydration of carbon dioxide and dehydration of carbonic acid at 25 degrees[J]. J Biol Chem, 1963, 238:3502-3507.
doi: 10.1016/S0021-9258(18)48696-6 URL |
| [22] |
Persson M, Palcic MM. A high-throughput pH indicator assay for screening glycosyltransferase saturation mutagenesis libraries[J]. Anal Biochem, 2008, 378(1):1-7.
doi: 10.1016/j.ab.2008.03.006 URL |
| [23] |
Martínez-Martínez I, Montoro-García S, Lozada-Ramírez JD, et al. A colorimetric assay for the determination of acetyl xylan esterase or cephalosporin C acetyl esterase activities using 7-amino cephalosporanic acid, cephalosporin C, or acetylated xylan as substrate[J]. Anal Biochem, 2007, 369(2):210-217.
pmid: 17651681 |
| [24] |
Banerjee A, Kaul P, Sharma R, et al. A high-throughput amenable colorimetric assay for enantioselective screening of nitrilase-producing microorganisms using pH sensitive indicators[J]. J Biomol Screen, 2003, 8(5):559-565.
pmid: 14567783 |
| [25] |
Mo Q, Mao A, et al. Substrate inactivation of bacterial L-aspartate α-decarboxylase from Corynebacterium jeikeium K411 and improvement of molecular stability by saturation mutagenesis[J]. World J Microbiol Biotechnol, 2019, 35(4):62.
doi: 10.1007/s11274-019-2629-6 URL |
| [26] |
Pei WL, Zhang JL, Deng SY, et al. Molecular engineering of l-aspartate-α-decarboxylase for improved activity and catalytic stability[J]. Appl Microbiol Biotechnol, 2017, 101(15):6015-6021.
doi: 10.1007/s00253-017-8337-y URL |
| [27] |
Qian Y, Lu C, et al. Engineering protonation conformation of l-aspartate-α-decarboxylase to relieve mechanism-based inactivation[J]. Biotechnol Bioeng, 2020, 117(6):1607-1614.
doi: 10.1002/bit.27316 URL |
| [28] | Chapman E, Wong CH. A pH sensitive colorometric assay for the high-throughput screening of enzyme inhibitors and substrates:a case study using kinases[J]. Bioorg Med Chem, 2002, 10(3):551-555. |
| [29] |
Janes LE, Löwendahl AC, Kazlauskas RJ. Quantitative screening of hydrolase libraries using pH indicators:identifying active and enantioselective hydrolases[J]. Chem Eur J, 1998, 4(11):2324-2331.
doi: 10.1002/(SICI)1521-3765(19981102)4:11<2324::AID-CHEM2324>3.0.CO;2-I URL |
| [30] | 李晓涵, 郝建华, 等. 环糊精葡萄糖基转移酶高效异源表达研究进展[J]. 微生物学通报, 2020, 47(2):615-622. |
| Li XH, Hao JH, et al. Advance in high-level heterologous expression of cyclodextrin glycosyltransferase[J]. Microbiol China, 2020, 47(2):615-622. | |
| [31] | 石增秀, 崔文璟, 周丽, 等. 谷氨酸棒杆菌L-天冬氨酸α-脱羧酶基因的克隆及重组酶性质研究[J]. 生物技术通报, 2013(4):110-115. |
| Shi ZX, Cui WJ, Zhou L, et al. Cloning and Characterization of L-aspartate-α-decarboxylase from Corynebacterium glutamicum[J]. Biotechnol Bull, 2013(4):110-115. | |
| [32] | 田慧, 等. 酰胺酶高通量筛选方法研究进展[J]. 微生物学杂志, 2012, 32(3):66-71. |
| Tian H, et al. Advances in high throughput screening method for amidase[J]. J Microbiol, 2012, 32(3):66-71. |
| [1] | JIANG Bai-yang, BAI Wen-bin, ZHANG Jiang-hua, FAN Na, SHI Li-juan. Advances in Studies on Identification Methods and Molecular Biology of Drought Resistance in Sorghum [J]. Biotechnology Bulletin, 2021, 37(4): 260-272. |
| [2] | SUN Xi-lin, JIANG Zhen-yan, LIU Zhi-yi, DAI Lu, SUN Fei, HUANG Wei. Improvement of Thermal Stability of Ganoderma lucidum Protein LZ-8 by Site-directed Mutation of Amino Acids [J]. Biotechnology Bulletin, 2020, 36(1): 23-28. |
| [3] | XU You-fen, LI Zong, LIU Ru-yin, YU Zhi-sheng, ZHANG Hong-xun, HE Wei, LI Ye. Advances in Research and Application of Microbial Source Tracking Technology in Aquatic Environment [J]. Biotechnology Bulletin, 2019, 35(9): 35-44. |
| [4] | QIU Jin, HUANG Huo-qing, YAO Bin, LUO Hui-ying. Improvement of Catalytic Activity of Amylase from Bacillus amyloliquefaciens and Its High Expression in Bacillus subtilis [J]. Biotechnology Bulletin, 2019, 35(9): 134-143. |
| [5] | ZHANG Huan-wei, CHEN Biao, WEN Xin-yi, ZHANG Jie, WANG Xiao-dong, LI Ji-wei, XU Zi-cheng, HUANG Wu-xing. Effects of Exogenous Silicon on Growth,Leaf Photosynthesis and Physiological Indexes of Tobacco Seedlings Under Drought Stress [J]. Biotechnology Bulletin, 2019, 35(1): 17-26. |
| [6] | LIU Xiao-wei, YANG Xiu-yan, WU Hai-wen, ZHI Xiao-rong, ZHU Jian-feng, ZHANG Hua-xin. Effects of NaCl Stress on the Germination of Reaumuria soongorica and Evaluation of Salt Tolerance at Germination Stage [J]. Biotechnology Bulletin, 2019, 35(1): 27-34. |
| [7] | QIN Tian-yuan, SUN Chao, BI Zhen-zhen, WANG Han, LI Xin, ZENG Wen-jie, BAI Jiang-ping. Responses of PVC-pipe Seedlings and Their Root Tip Microstructures of Different Drought-Resistant Potato Varieties to Drought Stress [J]. Biotechnology Bulletin, 2018, 34(12): 102-109. |
| [8] | QIN Hai-bin, XIONG Tao, ZHANG Bo, NIU Kun. Site-directed Mutation of α-ketoglutorate Semialdehye Dehydrogenase and Its Enzymatic Properties [J]. Biotechnology Bulletin, 2017, 33(8): 180-185. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
