








Biotechnology Bulletin ›› 2022, Vol. 38 ›› Issue (5): 149-158.doi: 10.13560/j.cnki.biotech.bull.1985.2021-1067
Previous Articles Next Articles
ZHAO Ming-ming1( ), TANG Yin1, GUO Lei-zhou1, HAN Jia-hui1, GE Jia-ming1, MENG Yong2, PING Shu-zhen1, ZHOU Zheng-fu1, WANG Jin1(
), TANG Yin1, GUO Lei-zhou1, HAN Jia-hui1, GE Jia-ming1, MENG Yong2, PING Shu-zhen1, ZHOU Zheng-fu1, WANG Jin1( )
)
Received:2021-08-09
Online:2022-05-26
Published:2022-06-10
Contact:
WANG Jin
E-mail:82101182064@caas.cn;wangjin@caas.cn
ZHAO Ming-ming, TANG Yin, GUO Lei-zhou, HAN Jia-hui, GE Jia-ming, MENG Yong, PING Shu-zhen, ZHOU Zheng-fu, WANG Jin. Function Analysis of Lon1 Protease Involved in High Temperature Stress and Cell Division of Deinococcus radiodurans R1[J]. Biotechnology Bulletin, 2022, 38(5): 149-158.
| 菌株及质粒Strains and plasmids | 特性Relevant characteristic | 来源Source | |
|---|---|---|---|
| 菌株 | Deinococcus radioduransR1 | 耐辐射异常球菌野生型 | 中科院微生物所菌种保藏中心 |
| △lon1 | 缺失lon1基因的突变株(Kanr) | 本研究构建 | |
| comlon1 | lon1回补株(Kanr+ Cmr) | 本研究构建 | |
| Z3-△lon1 | △lon1中导入质粒pRADZ3(Cmr) | 本研究构建 | |
| 质粒 | pRAD Z3 | 构建回补菌株所用质粒 | 本实验室保存 |
Table 1 Strains and Plasmids
| 菌株及质粒Strains and plasmids | 特性Relevant characteristic | 来源Source | |
|---|---|---|---|
| 菌株 | Deinococcus radioduransR1 | 耐辐射异常球菌野生型 | 中科院微生物所菌种保藏中心 |
| △lon1 | 缺失lon1基因的突变株(Kanr) | 本研究构建 | |
| comlon1 | lon1回补株(Kanr+ Cmr) | 本研究构建 | |
| Z3-△lon1 | △lon1中导入质粒pRADZ3(Cmr) | 本研究构建 | |
| 质粒 | pRAD Z3 | 构建回补菌株所用质粒 | 本实验室保存 |
| 扩增产物 Product for PCR amplification | 引物名称 Primer name | 序列Sequence(5'-3') | 片段长度Size/bp |
|---|---|---|---|
| lon1-Up | lon1-UP-F | AGCTGCTGACCCGTGACACCGAGCCATGGAGACCGAGGGCCC | 613 |
| lon1-up-R | GGGCCCTCGGTCTCCATGGCTCGGTGTCACGGGTCAGCAGCT | ||
| lon1-Kan | Kan-lon1-F | AGCTGCTGACCCGTGACACCGAGCCATGGAGACCGAGGGCCC | 965 |
| Kan-Down-R | TGGTCCACGCCAGGCCCTGCGTTAGAAAAACTCATCGAGCAT | ||
| lon1-Down | lon1-Down-F | ATGCTCGATGAGTTTTTCTAACGCAGGGCCTGGCGTGGACCA | 614 |
| lon1-Down-R | TTATGCGCCCGGCTGCACCGA | ||
| Kan内部序列 | YZ-Km-F | CGATTGTATGGGAAGCCCGAT | 591 |
| YZ-Km-R | CTCACCGAGGCAGTTCCATAG | ||
| 敲除的lon1序列 | YZ-lon1-F | ACCGTGGTTCGCAACTACAT | 400 |
| YZ-lon1-R | CGATCTCGACCTTCTCCTGC |
Table 2 Sequences of Primers
| 扩增产物 Product for PCR amplification | 引物名称 Primer name | 序列Sequence(5'-3') | 片段长度Size/bp |
|---|---|---|---|
| lon1-Up | lon1-UP-F | AGCTGCTGACCCGTGACACCGAGCCATGGAGACCGAGGGCCC | 613 |
| lon1-up-R | GGGCCCTCGGTCTCCATGGCTCGGTGTCACGGGTCAGCAGCT | ||
| lon1-Kan | Kan-lon1-F | AGCTGCTGACCCGTGACACCGAGCCATGGAGACCGAGGGCCC | 965 |
| Kan-Down-R | TGGTCCACGCCAGGCCCTGCGTTAGAAAAACTCATCGAGCAT | ||
| lon1-Down | lon1-Down-F | ATGCTCGATGAGTTTTTCTAACGCAGGGCCTGGCGTGGACCA | 614 |
| lon1-Down-R | TTATGCGCCCGGCTGCACCGA | ||
| Kan内部序列 | YZ-Km-F | CGATTGTATGGGAAGCCCGAT | 591 |
| YZ-Km-R | CTCACCGAGGCAGTTCCATAG | ||
| 敲除的lon1序列 | YZ-lon1-F | ACCGTGGTTCGCAACTACAT | 400 |
| YZ-lon1-R | CGATCTCGACCTTCTCCTGC |
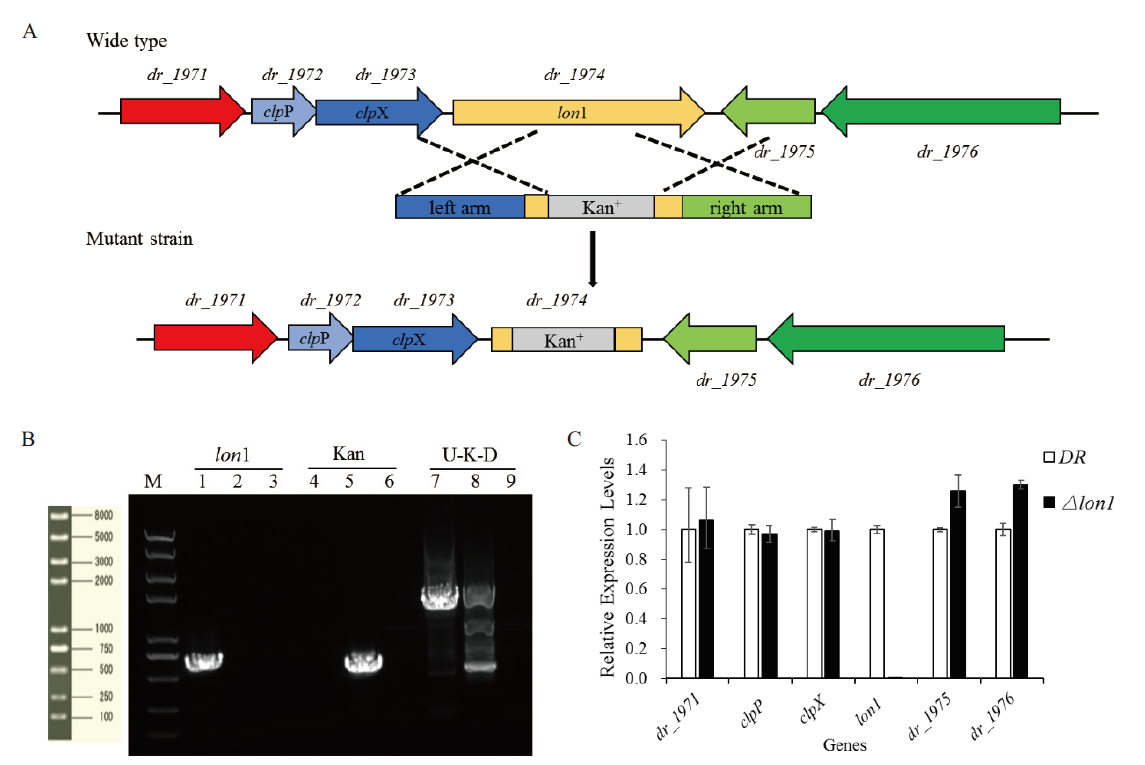
Fig. 3 Construction and identification of the lon1 mutant strain A:Schematic representation of the Δlon1mutant. B:Electrophoretogram of the lon1 mutant strain(M:Trans 2K PlusⅡ DNA marker. 1, 4, 7:PCR products amplified from wild type D. radiodurans. 2, 5, 8:PCR products amplified from the lon1mutant. 3, 6, 9:Negative control). C:Effects of lon1 deleted on adjacent genes in transcriptional level
| UniProtKB | Locus | Predicted function | Protein name | FC |
|---|---|---|---|---|
| Protein fate(heat shock responsive chaperones and protease) | ||||
| Q9RV58 | DR_1172 | Chaperone-like protein | DosH | 13.37 |
| Q9RWQ9 | DR_0607 | 60 kD chaperone | GroL | 7.08 |
| Q9RVI3 | DR_1046 | Chaperone protein ClpB | ClpB | 4.70 |
| Q9RXG4 | DR_0349 | Lon protease | Lon2 | 4.52 |
| Q9RWR0 | DR_0606 | 10 kD chaperone | GroS | 4.46 |
| Q9RY24 | DR_0128 | Chaperone protein GrpE | GrpE | 3.24 |
| DNA metabolism(repair and recombination) | ||||
| Q9RXI7 | DR_0326 | DNA damage response protein D | DdrD | 2.59 |
| Q9RS64 | DR_2263 | DNA protection during starvation protein 1 | Dps1 | 2.88 |
| Q9RY51 | DR_0099 | Single-stranded DNA-binding protein | Ssb | 3.25 |
| Q9RSJ6 | DR_2128 | DNA-directed RNA polymerase subunit alpha | RpoA | 2.06 |
| Q9RRE4 | DR_2548 | Transcriptional regulatory protein | 7.75 | |
| Membrane functions and transport | ||||
| Q9RWU0 | DR_0575 | Protein translocase subunit SecA | SecA | 2.18 |
| P56867 | DR_2508 | Hexagonally packed intermediate-layer surface protein | Hpi | 3.48 |
| Q9RWH3 | DR_0695 | V-type ATP synthase subunit | AtpI | 1.57 |
| Translation | ||||
| Q9RST0 | DR_2043 | 50S ribosomal protein L7/L12 | RplL | 0.14 |
| Central intermediary metabolism(synthesis of nitrogen,sulfur,polyamine compounds) | ||||
| Q9RR70 | DR_2627 | Fumarate hydratase class II | FumC | 7.59 |
| Q9RWB2 | DR_0757 | Citrate synthase | GltA | 3.06 |
| Q9RZ06 | DR_A0147 | Histidine ammonia-lyase | HutH | 2.66 |
| Q9RTN7 | DR_1720 | Aconitate hydratase A | Acn | 2.10 |
| P56861 | DR_A0014 | Adenylyl-sulfate kinase | CysC | 0.23 |
| Amino acid biosynthesis and energy metabolism | ||||
| Q9RRT0 | DR_2405 | Demethylmenaquinone methyltransferase | MenG | 0.51 |
| Q9RV69 | DR_1161 | 5-hydroxyisourate hydrolase | 0.33 | |
| Q9RV98 | DR_1131 | Ferrochelatase | HemH | 2.04 |
| Q9RRC4 | DR_2568 | Arginine--tRNA ligase | ArgS | 0.54 |
| Cell morphogenesis | ||||
| Q9RRJ4 | DR_2496 | UDP-N-acetylmuramoylalanine--D-glutamate ligase | MurD | 0.65 |
| Q9RWN8 | DR_0628 | UDP-N-acetylenolpyruvoylglucosamine reductase | MurB | 0.59 |
| Q9RXF1 | DR_0362 | D-alanine--D-alanine ligase | Ddl | 0.46 |
| Q9RWN9 | DR_0627 | UDP-N-acetylmuramate--L-alanine ligase | MurC | 0.42 |
| Q9RXL3 | DR_0297 | UDP-N-acetylmuramyl-tripeptide synthetase | MurE | 0.40 |
Table 3 Differentially expressed proteins associated with lon1 in response to high temperature stress
| UniProtKB | Locus | Predicted function | Protein name | FC |
|---|---|---|---|---|
| Protein fate(heat shock responsive chaperones and protease) | ||||
| Q9RV58 | DR_1172 | Chaperone-like protein | DosH | 13.37 |
| Q9RWQ9 | DR_0607 | 60 kD chaperone | GroL | 7.08 |
| Q9RVI3 | DR_1046 | Chaperone protein ClpB | ClpB | 4.70 |
| Q9RXG4 | DR_0349 | Lon protease | Lon2 | 4.52 |
| Q9RWR0 | DR_0606 | 10 kD chaperone | GroS | 4.46 |
| Q9RY24 | DR_0128 | Chaperone protein GrpE | GrpE | 3.24 |
| DNA metabolism(repair and recombination) | ||||
| Q9RXI7 | DR_0326 | DNA damage response protein D | DdrD | 2.59 |
| Q9RS64 | DR_2263 | DNA protection during starvation protein 1 | Dps1 | 2.88 |
| Q9RY51 | DR_0099 | Single-stranded DNA-binding protein | Ssb | 3.25 |
| Q9RSJ6 | DR_2128 | DNA-directed RNA polymerase subunit alpha | RpoA | 2.06 |
| Q9RRE4 | DR_2548 | Transcriptional regulatory protein | 7.75 | |
| Membrane functions and transport | ||||
| Q9RWU0 | DR_0575 | Protein translocase subunit SecA | SecA | 2.18 |
| P56867 | DR_2508 | Hexagonally packed intermediate-layer surface protein | Hpi | 3.48 |
| Q9RWH3 | DR_0695 | V-type ATP synthase subunit | AtpI | 1.57 |
| Translation | ||||
| Q9RST0 | DR_2043 | 50S ribosomal protein L7/L12 | RplL | 0.14 |
| Central intermediary metabolism(synthesis of nitrogen,sulfur,polyamine compounds) | ||||
| Q9RR70 | DR_2627 | Fumarate hydratase class II | FumC | 7.59 |
| Q9RWB2 | DR_0757 | Citrate synthase | GltA | 3.06 |
| Q9RZ06 | DR_A0147 | Histidine ammonia-lyase | HutH | 2.66 |
| Q9RTN7 | DR_1720 | Aconitate hydratase A | Acn | 2.10 |
| P56861 | DR_A0014 | Adenylyl-sulfate kinase | CysC | 0.23 |
| Amino acid biosynthesis and energy metabolism | ||||
| Q9RRT0 | DR_2405 | Demethylmenaquinone methyltransferase | MenG | 0.51 |
| Q9RV69 | DR_1161 | 5-hydroxyisourate hydrolase | 0.33 | |
| Q9RV98 | DR_1131 | Ferrochelatase | HemH | 2.04 |
| Q9RRC4 | DR_2568 | Arginine--tRNA ligase | ArgS | 0.54 |
| Cell morphogenesis | ||||
| Q9RRJ4 | DR_2496 | UDP-N-acetylmuramoylalanine--D-glutamate ligase | MurD | 0.65 |
| Q9RWN8 | DR_0628 | UDP-N-acetylenolpyruvoylglucosamine reductase | MurB | 0.59 |
| Q9RXF1 | DR_0362 | D-alanine--D-alanine ligase | Ddl | 0.46 |
| Q9RWN9 | DR_0627 | UDP-N-acetylmuramate--L-alanine ligase | MurC | 0.42 |
| Q9RXL3 | DR_0297 | UDP-N-acetylmuramyl-tripeptide synthetase | MurE | 0.40 |
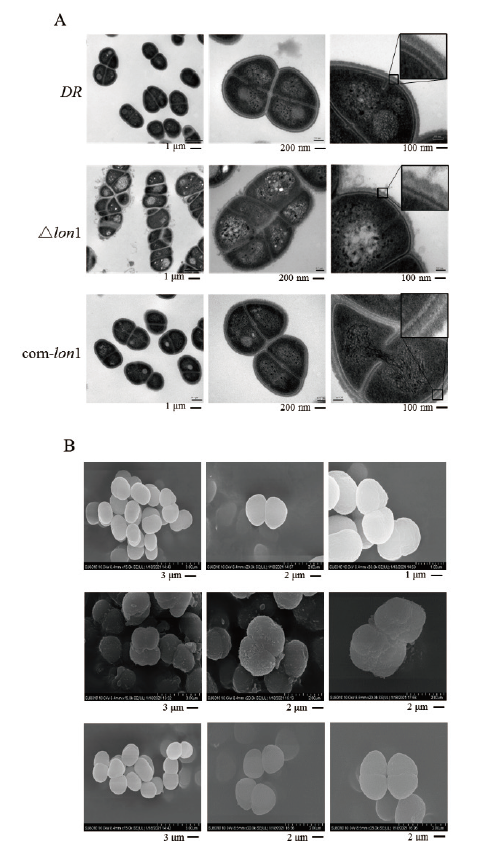
Fig. 6 Electron microscopy images of WT D. radiodurans DR,△lon1 and comlon1 A:Transmission electron microscopy(TEM). B:Scanning electron microscopy(SEM)
| [1] | Gur E, Ottofueling R, Dougan DA. Machines of destruction - AAA+ proteases and the adaptors that control them[M]// Subcellular Biochemistry. Dordrecht: Springer Netherlands, 2013:3-33. |
| [2] |
Mahmoud SA, Chien P. Regulated proteolysis in bacteria[J]. Annu Rev Biochem, 2018, 87:677-696.
doi: 10.1146/annurev-biochem-062917-012848 pmid: 29648875 |
| [3] | 刘郁夫, 董浩, 孙石静, 等. 细菌Lon蛋白酶研究进展[J]. 微生物学通报, 2019, 46(7):1706-1711. |
| Liu YF, Dong H, Sun SJ, et al. Research progress on bacterial Lon protease[J]. Microbiol China, 2019, 46(7):1706-1711. | |
| [4] | Gur E. The lon AAA+ protease[M]// Subcellular Biochemistry. Dordrecht: Springer Netherlands, 2013:35-51. |
| [5] |
Lin CC, Su SC, Su MY, et al. Structural insights into the allosteric operation of the lon AAA+ protease[J]. Structure, 2016, 24(5):667-675.
doi: 10.1016/j.str.2016.03.001 URL |
| [6] |
Tsilibaris V, Maenhaut-Michel G, van Melderen L. Biological roles of the Lon ATP-dependent protease[J]. Res Microbiol, 2006, 157(8):701-713.
pmid: 16854568 |
| [7] |
Jonas K, Liu J, Chien P, et al. Proteotoxic stress induces a cell-cycle arrest by stimulating Lon to degrade the replication initiator DnaA[J]. Cell, 2013, 154(3):623-636.
doi: 10.1016/j.cell.2013.06.034 URL |
| [8] |
Adler HI, Hardigree AA. Cell elongation in strains of Escherichia coli[J]. J Bacteriol, 1964, 87(5):1240-1242.
doi: 10.1128/jb.87.5.1240-1242.1964 pmid: 5334970 |
| [9] |
Howard-Flanders P, Simson E, Theriot L. A locus that controls filament formation and sensitivity to radiation in Escherichia coli k-12[J]. Genetics, 1964, 49:237-246.
doi: 10.1093/genetics/49.2.237 URL |
| [10] |
Fernández L, Breidenstein EBM, Taylor PK, et al. Interconnection of post-transcriptional regulation:The RNA-binding protein Hfq is a novel target of the Lon protease in Pseudomonas aeruginosa[J]. Sci Rep, 2016, 6:26811.
doi: 10.1038/srep26811 pmid: 27229357 |
| [11] |
Kim H, Lee H, Shin D. Lon-mediated proteolysis of the FeoC protein prevents Salmonella enterica from accumulating the Fe(II)transporter FeoB under high-oxygen conditions[J]. J Bacteriol, 2015, 197(1):92-98.
doi: 10.1128/JB.01826-14 URL |
| [12] |
Slade D, Radman M. Oxidative stress resistance in Deinococcus radiodurans[J]. Microbiol Mol Biol Rev, 2011, 75(1):133-191.
doi: 10.1128/MMBR.00015-10 URL |
| [13] |
Daly MJ. A new perspective on radiation resistance based on Deinococcus radiodurans[J]. Nat Rev Microbiol, 2009, 7(3):237-245.
doi: 10.1038/nrmicro2073 URL |
| [14] |
Daly MJ, Gaidamakova EK, Matrosova VY, et al. Small-molecule antioxidant proteome-shields in Deinococcus radiodurans[J]. PLoS One, 2010, 5(9):e12570.
doi: 10.1371/journal.pone.0012570 URL |
| [15] | 刘盈盈. 耐辐射异常球菌亲水蛋白DohL具有类分子伴侣和核酸内切酶功能并参与氧化胁迫保护[D]. 北京: 中国农业科学院, 2019. |
| Liu YY. Chaperone-like and catalytic functions in Deinococcus radiodurans of hydrophilic protein DohL involved in protective against oxidative stress[D]. Beijing: Chinese Academy of Agricultural Sciences, 2019. | |
| [16] | 陈晓楠. 耐辐射异常球菌黄嘌呤脱氢酶基因在氧化胁迫反应中的作用[D]. 北京: 中国农业科学院, 2020. |
| Chen XN. Role of xanthine dehydrogenase gene in oxidative stress response of Deinococcus radiodurans R1[D]. Beijing: Chinese Academy of Agricultural Sciences, 2020. | |
| [17] |
Karlin S, Mrazek J. Predicted highly expressed and putative alien genes of Deinococcus radiodurans and implications for resistance to ionizing radiation damage[J]. PNAS, 2001, 98(9):5240-5245.
pmid: 11296249 |
| [18] |
Heikkila JJ, Schultz GA, Iatrou K, et al. Expression of a set of fish genes following heat or metal ion exposure[J]. J Biol Chem, 1982, 257(20):12000-12005.
pmid: 7118927 |
| [19] |
Wickner S, Maurizi MR, Gottesman S. Posttranslational quality control:folding, refolding, and degrading proteins[J]. Science, 1999, 286(5446):1888-1893.
pmid: 10583944 |
| [20] |
Jacob P, Hirt H, Bendahmane A. The heat-shock protein/chaperone network and multiple stress resistance[J]. Plant Biotechnol J, 2017, 15(4):405-414.
doi: 10.1111/pbi.12659 URL |
| [21] |
Su SC, Stephens BB, Alexandre G, et al. Lon protease of the alpha-proteobacterium Agrobacterium tumefaciens is required for normal growth, cellular morphology and full virulence[J]. Microbiol Read Engl, 2006, 152(Pt 4):1197-1207.
doi: 10.1099/mic.0.28657-0 URL |
| [22] |
Breidenstein EB, Janot L, Strehmel J, et al. The Lon protease is essential for full virulence in Pseudomonas aeruginosa[J]. PLoS One, 2012, 7(11):e49123.
doi: 10.1371/journal.pone.0049123 URL |
| [23] |
Jukič M, Gobec S, Sova M. Reaching toward underexplored targets in antibacterial drug design[J]. Drug Dev Res, 2019, 80(1):6-10.
doi: 10.1002/ddr.21465 URL |
| [24] |
Gottesman S, Halpern E, Trisler P. Role of SulA and sulB in filamentation by lon mutants of Escherichia coli K-12[J]. J Bacteriol, 1981, 148(1):265-273.
doi: 10.1128/jb.148.1.265-273.1981 pmid: 7026534 |
| [25] |
Mizusawa S, Gottesman S. Protein degradation in Escherichia coli:the lon gene controls the stability of SulA protein[J]. PNAS, 1983, 80(2):358-362.
pmid: 6300834 |
| [1] | ZHOU Lu-qi, CUI Ting-ru, HAO Nan, ZHAO Yu-wei, ZHAO Bin, LIU Ying-chao. Application of Chemical Proteomics in Identifying the Molecular Targets of Natural Products [J]. Biotechnology Bulletin, 2023, 39(9): 12-26. |
| [2] | LYU Qiu-yu, SUN Pei-yuan, RAN Bin, WANG Jia-rui, CHEN Qing-fu, LI Hong-you. Cloning, Subcellular Localization and Expression Analysis of the Transcription Factor Gene FtbHLH3 in Fagopyrum tataricum [J]. Biotechnology Bulletin, 2023, 39(8): 194-203. |
| [3] | WANG Jia-rui, SUN Pei-yuan, KE Jin, RAN Bin, LI Hong-you. Cloning and Expression Analyses of C-glycosyltransferase Gene FtUGT143 in Fagopyrum tataricum [J]. Biotechnology Bulletin, 2023, 39(8): 204-212. |
| [4] | SUN Ming-hui, WU Qiong, LIU Dan-dan, JIAO Xiao-yu, WANG Wen-jie. Cloning and Expression Analysis of CsTMFs Gene in Tea Plant [J]. Biotechnology Bulletin, 2023, 39(7): 151-159. |
| [5] | ZHAO Xue-ting, GAO Li-yan, WANG Jun-gang, SHEN Qing-qing, ZHANG Shu-zhen, LI Fu-sheng. Cloning and Expression of AP2/ERF Transcription Factor Gene ShERF3 in Sugarcane and Subcellular Localization of Its Encoded Protein [J]. Biotechnology Bulletin, 2023, 39(6): 208-216. |
| [6] | LI Yuan-hong, GUO Yu-hao, CAO Yan, ZHU Zhen-zhou, WANG Fei-fei. Research Progress in the Microalgal Growth and Accumulation of Target Products Regulated by Exogenous Phytohormone [J]. Biotechnology Bulletin, 2023, 39(6): 61-72. |
| [7] | FENG Shan-shan, WANG Lu, ZHOU Yi, WANG You-ping, FANG Yu-jie. Research Progresses on WOX Family Genes in Regulating Plant Development and Abiotic Stress Response [J]. Biotechnology Bulletin, 2023, 39(5): 1-13. |
| [8] | LI Jing-rui, WANG Yu-bo, XIE Zi-wei, LI Chang, WU Xiao-lei, GONG Bin-bin, GAO Hong-bo. Identification and Expression Analysis of PIN Gene Family in Melon Under High Temperature Stress [J]. Biotechnology Bulletin, 2023, 39(5): 192-204. |
| [9] | ZHAI Ying, LI Ming-yang, ZHANG Jun, ZHAO Xu, YU Hai-wei, LI Shan-shan, ZHAO Yan, ZHANG Mei-juan, SUN Tian-guo. Heterologous Expression of Soybean Transcription Factor GmNF-YA19 Improves Drought Resistance of Transgenic Tobacco [J]. Biotechnology Bulletin, 2023, 39(5): 224-232. |
| [10] | GUO San-bao, SONG Mei-ling, LI Ling-xin, YAO Zi-zhao, GUI Ming-ming, HUANG Sheng-he. Cloning and Analysis of Chalcone Synthase Gene and Its Promoter from Euphorbia maculata [J]. Biotechnology Bulletin, 2023, 39(4): 148-156. |
| [11] | YANG Jun-zhao, ZHANG Xin-rui, ZHAO Guo-zhu, ZHENG Fei. Structure and Function Analysis of Novel GH5 Multi-domain Cellulase [J]. Biotechnology Bulletin, 2023, 39(4): 71-80. |
| [12] | SANG Tian, WANG Peng-cheng. Research Progress in Plant SUMOylation [J]. Biotechnology Bulletin, 2023, 39(3): 1-12. |
| [13] | GE Yan-rui, ZHAO Ran, XU Jing, LI Ruo-fan, HU Yun-tao, LI Rui-li. Advances in the Development and Regulation of Vascular Cambium [J]. Biotechnology Bulletin, 2023, 39(3): 13-25. |
| [14] | YANG Chun-hong, DONG Lu, CHEN Lin, SONG Li. Characterization of Soybean VAS1 Gene Family and Its Involvement in Lateral Root Development [J]. Biotechnology Bulletin, 2023, 39(3): 133-142. |
| [15] | LIU Si-jia, WANG Hao-nan, FU Yu-chen, YAN Wen-xin, HU Zeng-hui, LENG Ping-sheng. Cloning and Functional Analysis of LiCMK Gene in Lilium ‘Siberia’ [J]. Biotechnology Bulletin, 2023, 39(3): 196-205. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||