








Biotechnology Bulletin ›› 2025, Vol. 41 ›› Issue (4): 47-60.doi: 10.13560/j.cnki.biotech.bull.1985.2024-0982
Previous Articles Next Articles
LU Tian-yi1( ), LI Ai-peng1,2, FEI Qiang1,2(
), LI Ai-peng1,2, FEI Qiang1,2( )
)
Received:2024-10-08
Online:2025-04-26
Published:2025-04-25
Contact:
FEI Qiang
E-mail:luty6099@stu.xjtu.edu.cn;feiqiang@xjtu.edu.cn
LU Tian-yi, LI Ai-peng, FEI Qiang. Research Progress in the Biosynthesis of Polylactic Acid[J]. Biotechnology Bulletin, 2025, 41(4): 47-60.
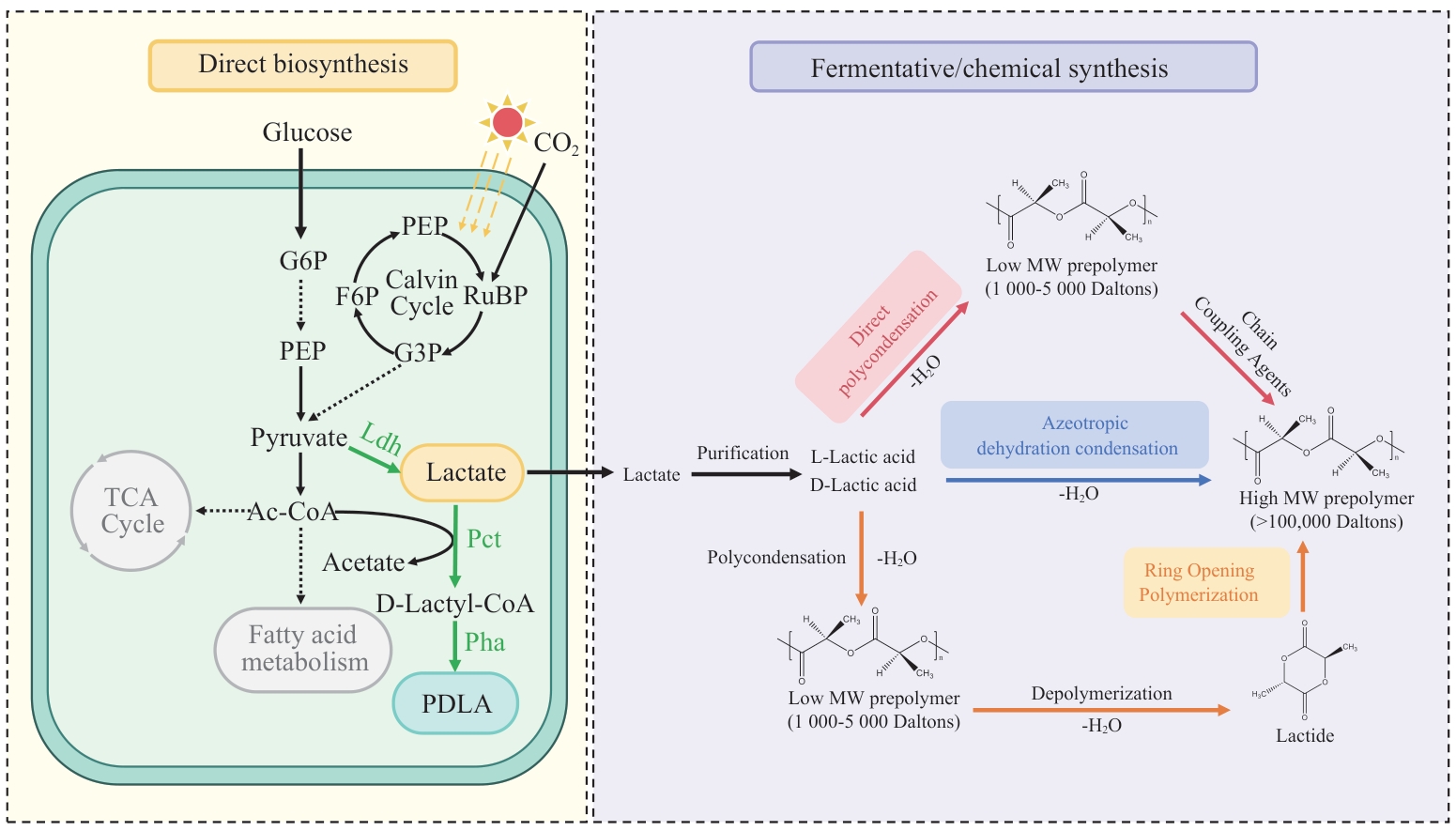
Fig. 3 Comparison of direct biosynthesis process and traditional biochemical hybrid process of PLAG6P: Glyceraldehyde 6-phosphate; PEP: phosphoenolpyruvate; Ac-CoA: acetyl-CoA; Ldh: lactate dehydrogenase; Pct: propionyl-CoA transferase; Pha: PHA synthase
| 方法 Method | 优点 Advantage | 缺点 Disadvantage |
|---|---|---|
直接缩合 Direct polycondensation | 1)操作简单且相对经济; 2)无需中间体的纯化,成本较低; 3)制得PLA的分子量低,降解速度快 | 1)易发生各种副反应; 2)低分子量PLA拉伸性能差,不能加工为塑料和纺织品; 3)易引入人体难以降解的杂质,不利于在医疗方面的应用[ |
共沸脱水缩合 Azeotropic dehydration condensation | 1)易除去反应中的水,使反应更易正向进行; 2)制得PLA的含水量低,分子量高 | 1)反应条件苛刻,设备和工艺复杂性较高; 2)消耗大量有机溶剂,成本高且安全性低; 3)易引入杂质和各种副反应; 4)产物提纯复杂; 5)制得的PLA易存在溶剂和催化剂残留,无法应用于医学领域[ |
开环聚合 Ring opening polymerization | 1)制得PLA的分子量高且集中; 2)制得PLA纯度高,力学性能好; 3)易调控PLA的化学结构,获得指定产物 | 1)丙交酯纯度要求高,纯化难度大; 2)合成工艺复杂、成本高、产率低 |
生物聚合 Biopolymerization | 1)反应条件温和,不易引入杂质和副反应; 2)PLA中不存在残留单体和毒性物质; 3)易生产优异对映纯度的聚合物; 4)易实现乳酸和其他单体的共聚 | 1)PLA产率低、分子量低; 2)生产成本和材料性能不可观; 3)尚处于实验室研究阶段,难以大规模应用 |
Table 1 Synthesis method of PLA and its advantages and disadvantages
| 方法 Method | 优点 Advantage | 缺点 Disadvantage |
|---|---|---|
直接缩合 Direct polycondensation | 1)操作简单且相对经济; 2)无需中间体的纯化,成本较低; 3)制得PLA的分子量低,降解速度快 | 1)易发生各种副反应; 2)低分子量PLA拉伸性能差,不能加工为塑料和纺织品; 3)易引入人体难以降解的杂质,不利于在医疗方面的应用[ |
共沸脱水缩合 Azeotropic dehydration condensation | 1)易除去反应中的水,使反应更易正向进行; 2)制得PLA的含水量低,分子量高 | 1)反应条件苛刻,设备和工艺复杂性较高; 2)消耗大量有机溶剂,成本高且安全性低; 3)易引入杂质和各种副反应; 4)产物提纯复杂; 5)制得的PLA易存在溶剂和催化剂残留,无法应用于医学领域[ |
开环聚合 Ring opening polymerization | 1)制得PLA的分子量高且集中; 2)制得PLA纯度高,力学性能好; 3)易调控PLA的化学结构,获得指定产物 | 1)丙交酯纯度要求高,纯化难度大; 2)合成工艺复杂、成本高、产率低 |
生物聚合 Biopolymerization | 1)反应条件温和,不易引入杂质和副反应; 2)PLA中不存在残留单体和毒性物质; 3)易生产优异对映纯度的聚合物; 4)易实现乳酸和其他单体的共聚 | 1)PLA产率低、分子量低; 2)生产成本和材料性能不可观; 3)尚处于实验室研究阶段,难以大规模应用 |
关键酶 Key enzyme | 来源菌株 Source strain | 宿主菌株 Host strain | 突变方式 Mutation method | 酶改造效果 Effect of enzyme modification | 参考文献 Reference | |
|---|---|---|---|---|---|---|
丙酰辅酶A转移酶 propionyl-CoA transferase | Clostridium ropionicum | E. coli XL1-Blue | V193A和4个沉默突变T78C、T669C、A1125G、T1158C(Pct540 Cp ) | 减缓酶表达对菌株的生长抑制,增强LA-CoA的合成,与野生型相比聚合物产量提高约5倍,聚合物中乳酸分数提高约8倍,酶体外特异性活性降低约39% | [ | |
| A243T和1个沉默突变A1200G(Pct532 Cp ) | 减缓酶表达对菌株的生长抑制,增强LA-CoA的合成,与野生型相比聚合物产量提高约4.5倍,聚合物中乳酸分数提高约9.5倍,酶体外特异性活性降低约52% | |||||
PHA合成酶 PHA synthetase | Ⅰ型 Type Ⅰ | Ralstonia eutropha | E. coli LS5218 | A510S | 赋予酶对LA-CoA的聚合能力,共聚物中乳酸分数最高可达26 mol%,且共聚物似为嵌段共聚物 | [ |
Ⅱ型 Type Ⅱ | Pseudomonas sp. 61-3 | E. coli JM109 | S325T/Q481K | 赋予酶对LA-CoA的聚合能力, 提高共聚物中乳酸分数至6 mol% | [ | |
| E. coli JW0885 | S325T/Q481K | 赋予酶对LA-CoA的聚合能力, 提高共聚物中乳酸分数至26 mol% | [ | |||
| S325T/Q481K/F392S | F392S的额外突变提高酶对LA-CoA的聚合能力,共聚物中乳酸分数提高约73%,聚合物含量提高约40.9% | |||||
| Pseudomonas sp. MBEL 6-19 | E. coli XL1-blue | E130D/Q481K | 赋予酶对LA-CoA的聚合能力, 提高共聚物中乳酸分数至35.3 mol% | [ | ||
| E130D/S477F/Q481K | 首次实现PLA均聚物(0.5 wt%)的合成,提高共聚物中乳酸分数至36.2 mol% | |||||
| E130D/S325T/Q481K | 提高酶对LA-CoA的聚合能力, 提高共聚物中乳酸分数至39.1 mol% | |||||
| E130D/S325T/S477R/Q481M | 提高酶对LA-CoA的聚合能力, 提高共聚物中乳酸分数至41.2 mol% | |||||
| E130D/S325T/S477F/Q481K | 提高酶对LA-CoA的聚合能力, 提高共聚物中乳酸分数至46 mol% | |||||
| E130D/S325T/S477G/Q481K | 提高酶对LA-CoA的聚合能力, 提高共聚物中乳酸分数至49 mol% | |||||
| Pseudomonas resinovorans | E. coli XL1-Blue | E130D/Q481K | 提高酶对LA-CoA的聚合能力,PLA产量提高至1.2 wt% | [ | ||
| E130D/S325T/S477G/Q481K | 提高酶对LA-CoA的聚合能力,PLA产量提高至7.3 wt% | |||||
嵌合酶 Chimerase | N端:Aeromonas cavia(Ⅱ型); C端:Ralstonia eutropha(Ⅰ型) | E. coli JM109 | N149D/F314H | 提高酶对LA-CoA的聚合能力, 共聚物中乳酸分数提高9.5倍 | [ | |
Table 2 Protein engineering strategies and effects of key enzymes in PLA biosynthesis
关键酶 Key enzyme | 来源菌株 Source strain | 宿主菌株 Host strain | 突变方式 Mutation method | 酶改造效果 Effect of enzyme modification | 参考文献 Reference | |
|---|---|---|---|---|---|---|
丙酰辅酶A转移酶 propionyl-CoA transferase | Clostridium ropionicum | E. coli XL1-Blue | V193A和4个沉默突变T78C、T669C、A1125G、T1158C(Pct540 Cp ) | 减缓酶表达对菌株的生长抑制,增强LA-CoA的合成,与野生型相比聚合物产量提高约5倍,聚合物中乳酸分数提高约8倍,酶体外特异性活性降低约39% | [ | |
| A243T和1个沉默突变A1200G(Pct532 Cp ) | 减缓酶表达对菌株的生长抑制,增强LA-CoA的合成,与野生型相比聚合物产量提高约4.5倍,聚合物中乳酸分数提高约9.5倍,酶体外特异性活性降低约52% | |||||
PHA合成酶 PHA synthetase | Ⅰ型 Type Ⅰ | Ralstonia eutropha | E. coli LS5218 | A510S | 赋予酶对LA-CoA的聚合能力,共聚物中乳酸分数最高可达26 mol%,且共聚物似为嵌段共聚物 | [ |
Ⅱ型 Type Ⅱ | Pseudomonas sp. 61-3 | E. coli JM109 | S325T/Q481K | 赋予酶对LA-CoA的聚合能力, 提高共聚物中乳酸分数至6 mol% | [ | |
| E. coli JW0885 | S325T/Q481K | 赋予酶对LA-CoA的聚合能力, 提高共聚物中乳酸分数至26 mol% | [ | |||
| S325T/Q481K/F392S | F392S的额外突变提高酶对LA-CoA的聚合能力,共聚物中乳酸分数提高约73%,聚合物含量提高约40.9% | |||||
| Pseudomonas sp. MBEL 6-19 | E. coli XL1-blue | E130D/Q481K | 赋予酶对LA-CoA的聚合能力, 提高共聚物中乳酸分数至35.3 mol% | [ | ||
| E130D/S477F/Q481K | 首次实现PLA均聚物(0.5 wt%)的合成,提高共聚物中乳酸分数至36.2 mol% | |||||
| E130D/S325T/Q481K | 提高酶对LA-CoA的聚合能力, 提高共聚物中乳酸分数至39.1 mol% | |||||
| E130D/S325T/S477R/Q481M | 提高酶对LA-CoA的聚合能力, 提高共聚物中乳酸分数至41.2 mol% | |||||
| E130D/S325T/S477F/Q481K | 提高酶对LA-CoA的聚合能力, 提高共聚物中乳酸分数至46 mol% | |||||
| E130D/S325T/S477G/Q481K | 提高酶对LA-CoA的聚合能力, 提高共聚物中乳酸分数至49 mol% | |||||
| Pseudomonas resinovorans | E. coli XL1-Blue | E130D/Q481K | 提高酶对LA-CoA的聚合能力,PLA产量提高至1.2 wt% | [ | ||
| E130D/S325T/S477G/Q481K | 提高酶对LA-CoA的聚合能力,PLA产量提高至7.3 wt% | |||||
嵌合酶 Chimerase | N端:Aeromonas cavia(Ⅱ型); C端:Ralstonia eutropha(Ⅰ型) | E. coli JM109 | N149D/F314H | 提高酶对LA-CoA的聚合能力, 共聚物中乳酸分数提高9.5倍 | [ | |
碳源 Carbon source | 宿主菌株 Host strain | 使用的关键酶 Key enzyme | 代谢工程调控策略 Metabolic engineering regulation strategies | 产量 Yield | 分子量 Molecular weight | 参考文献 Reference |
|---|---|---|---|---|---|---|
葡萄糖 Glucose | E. coli XL1-blue | Pct540 Cp PhaC Ps6-19E130D/S325T/S477R/Q481M | 1)乳酸供应强化:过表达ldhA;敲除ppc 2)乙酰辅酶A供应强化:敲除ackA、adhE;过表达acs | 11 wt% | N.D. | [ |
| E. coli XL1-blue | Pct540 Cp PhaC Ps6-19E130D/S477F/Q481K | 1)启动子改造:敲除lacI 2)乳酸供应强化:替换ldhA的 天然启动子为强启动子trc 3)乙酰辅酶A供应强化:过表达acs 4)副产物阻断:敲除pflB、frdABCD和adhE | 4.2 wt% | 21 000 Da | [ | |
| E. coli BL21(DE3) | Pct540 Cp PhaC Cs | 形态工程:过表达sulA | 2.23 wt% (955 mg/L) | 21 000 Da | [ | |
| Corynebacterium glutamicum ATCC13803 | Pct Me PhaC1 PsSTQK | 聚合途径强化:过表达PhaC1 PsSTQK | 1.4 wt% | Mw: 5.7 kD; Mn: 4.3 kD | [ | |
| Yarrowia lipolytica | Pct540 Cp PhaC PaE130D/S325T/S477R/Q481M | 1)乳酸供应强化:敲除YlDLD1 2)聚合途径强化:过表达Pct540 Cp 、PhaC PaE130D/S325T/S477R/Q481M 3)区室化工程:细胞质表达Pct540 Cp,过氧化物酶体表达PhaC PaE130D/S325T/S477R/Q481M | 26 mg/g DCW | 50.5 kD | [ | |
二氧化碳 Carbon dioxide | Synechococcus elongatus PCC7942 | Pct540 Cp PhaC Ps6-19 | 1)聚合途径强化:过表达Pct540 Cp 、PhaC Ps6-19 2)乙酰辅酶A供给强化:下调ackA,过表达acs 3)脂肪酸途径弱化:下调accABCD、fabH、fabF 4)高密度发酵和培养条件优化 | 23 mg/g DCW (108 mg/L) | Mw: 62.5 kD;Mn: 32.8 kD | [ |
Table 3 Examples of biosynthesis of polylactic acid and its metabolic engineering regulation strategies
碳源 Carbon source | 宿主菌株 Host strain | 使用的关键酶 Key enzyme | 代谢工程调控策略 Metabolic engineering regulation strategies | 产量 Yield | 分子量 Molecular weight | 参考文献 Reference |
|---|---|---|---|---|---|---|
葡萄糖 Glucose | E. coli XL1-blue | Pct540 Cp PhaC Ps6-19E130D/S325T/S477R/Q481M | 1)乳酸供应强化:过表达ldhA;敲除ppc 2)乙酰辅酶A供应强化:敲除ackA、adhE;过表达acs | 11 wt% | N.D. | [ |
| E. coli XL1-blue | Pct540 Cp PhaC Ps6-19E130D/S477F/Q481K | 1)启动子改造:敲除lacI 2)乳酸供应强化:替换ldhA的 天然启动子为强启动子trc 3)乙酰辅酶A供应强化:过表达acs 4)副产物阻断:敲除pflB、frdABCD和adhE | 4.2 wt% | 21 000 Da | [ | |
| E. coli BL21(DE3) | Pct540 Cp PhaC Cs | 形态工程:过表达sulA | 2.23 wt% (955 mg/L) | 21 000 Da | [ | |
| Corynebacterium glutamicum ATCC13803 | Pct Me PhaC1 PsSTQK | 聚合途径强化:过表达PhaC1 PsSTQK | 1.4 wt% | Mw: 5.7 kD; Mn: 4.3 kD | [ | |
| Yarrowia lipolytica | Pct540 Cp PhaC PaE130D/S325T/S477R/Q481M | 1)乳酸供应强化:敲除YlDLD1 2)聚合途径强化:过表达Pct540 Cp 、PhaC PaE130D/S325T/S477R/Q481M 3)区室化工程:细胞质表达Pct540 Cp,过氧化物酶体表达PhaC PaE130D/S325T/S477R/Q481M | 26 mg/g DCW | 50.5 kD | [ | |
二氧化碳 Carbon dioxide | Synechococcus elongatus PCC7942 | Pct540 Cp PhaC Ps6-19 | 1)聚合途径强化:过表达Pct540 Cp 、PhaC Ps6-19 2)乙酰辅酶A供给强化:下调ackA,过表达acs 3)脂肪酸途径弱化:下调accABCD、fabH、fabF 4)高密度发酵和培养条件优化 | 23 mg/g DCW (108 mg/L) | Mw: 62.5 kD;Mn: 32.8 kD | [ |
| 1 | Stubbins A, Law KL, Muñoz SE, et al. Plastics in the earth system [J]. Science, 2021, 373(6550): 51-55. |
| 2 | MacLeod M, Arp HPH, Tekman MB, et al. The global threat from plastic pollution [J]. Science, 2021, 373(6550): 61-65. |
| 3 | Lau WWY, Shiran Y, Bailey RM, et al. Evaluating scenarios toward zero plastic pollution [J]. Science, 2020, 369(6510): 1455-1461. |
| 4 | Soo XYD, Jia LR, Lim QF, et al. Hydrolytic degradation and biodegradation of polylactic acid electrospun fibers [J]. Chemosphere, 2024, 350: 141186. |
| 5 | Ma YX, Guo XY, Du MM, et al. Beyond biodegradation: upcycling of polylactic acid plastic waste into amino acids via cascade catalysis under mild conditions [J]. Green Chem, 2024, 26(7): 3995-4004. |
| 6 | Hao LT, Ren SL, Li JY, et al. Feasibility of biodegradable material polylactic acid as a substitute for polypropylene for disposable medical masks production verified by life cycle assessment [J]. J Clean Prod, 2024, 448: 141492. |
| 7 | Yang TH, Jung YK, Kang HO, et al. Tailor-made type II Pseudomonas PHA synthases and their use for the biosynthesis of polylactic acid and its copolymer in recombinant Escherichia coli [J]. Appl Microbiol Biotechnol, 2011, 90(2): 603-614. |
| 8 | Han X, Liu JQ, Tian S, et al. Microbial cell factories for bio-based biodegradable plastics production [J]. iScience, 2022, 25(11): 105462. |
| 9 | Park SJ, Lee SY, Kim TW, et al. Biosynthesis of lactate-containing polyesters by metabolically engineered bacteria [J]. Biotechnol J, 2012, 7(2): 199-212. |
| 10 | Van Wouwe P, Dusselier M, Vanleeuw E, et al. Lactide synthesis and chirality control for polylactic acid production [J]. ChemSusChem, 2016, 9(9): 907-921. |
| 11 | Yang WJ, Zhou QK, Pan WH, et al. Synthesis of vanillin-based porphyrin for remarkably enhancing the toughness, UV-resistance and self-extinguishing properties of polylactic acid [J]. Chem Eng J, 2023, 469: 143935. |
| 12 | Hu X, Wang BT, Guo ZH, et al. Roles of phosphoramide derivatives in flame retardancy, thermal degradation and crystallization behaviors of polylactic acid [J]. Int J Biol Macromol, 2022, 219: 558-570. |
| 13 | Makri SP, Xanthopoulou E, Valera MA, et al. Poly (lactic acid) composites with lignin and nanolignin synthesized by in situ reactive processing [J]. Polymers, 2023, 15(10): 2386. |
| 14 | Boarino A, Schreier A, Leterrier Y, et al. Uniformly dispersed poly (lactic acid)-grafted lignin nanoparticles enhance antioxidant activity and UV-barrier properties of poly (lactic acid) packaging films [J]. ACS Appl Polym Mater, 2022, 4(7): 4808-4817. |
| 15 | Huang SY, Xue YF, Yu B, et al. A review of the recent developments in the bioproduction of polylactic acid and its precursors optically pure lactic acids [J]. Molecules, 2021, 26(21): 6446. |
| 16 | Gao CT, Wang YW, Yang YL, et al. Poly (lactic acid) synthesized from non-food biomass feedstocks with tin-loaded ZA molecular sieve catalysts by direct melt polycondensation [J]. Polym Int, 2024, 73(4): 310-318. |
| 17 | Wang QM, Chen XY, Zeng SH, et al. In-situ polycondensate-coated cellulose nanofiber heterostructure for polylactic acid-based composites with superior mechanical and thermal properties [J]. Int J Biol Macromol, 2023, 240: 124515. |
| 18 | van den Berg SA, Zuilhof H, Wennekes T. Clickable polylactic acids by fast organocatalytic ring-opening polymerization in continuous flow [J]. Macromolecules, 2016, 49(6): 2054-2062. |
| 19 | Stopper A, Okuda J, Kol M. Ring-opening polymerization of lactide with Zr complexes of {ONSO} ligands: from heterotactically inclined to isotactically inclined poly (lactic acid) [J]. Macromolecules, 2012, 45(2): 698-704. |
| 20 | Taguchi S, Yamada M, Matsumoto K, et al. A microbial factory for lactate-based polyesters using a lactate-polymerizing enzyme [J]. Proc Natl Acad Sci U S A, 2008, 105(45): 17323-17327. |
| 21 | Madison LL, Huisman GW. Metabolic engineering of poly(3-hydroxyalkanoates): from DNA to plastic [J]. Microbiol Mol Biol Rev, 1999, 63(1): 21-53. |
| 22 | Yang TH, Kim TW, Kang HO, et al. Biosynthesis of polylactic acid and its copolymers using evolved propionate CoA transferase and PHA synthase [J]. Biotechnol Bioeng, 2010, 105(1): 150-160. |
| 23 | 谢彬, 白茸茸, 孙华山, 等. 聚乳酸塑料合成、生物降解及其废弃物处置的研究进展 [J]. 生物工程学报, 2023, 39(5): 1912-1929. |
| Xie B, Bai RR, Sun HS, et al. Synthesis, biodegradation and waste disposal of polylactic acid plastics: a review [J]. Chin J Biotechnol, 2023, 39(5): 1912-1929. | |
| 24 | Li G, Zhao MH, Xu F, et al. Synthesis and biological application of polylactic acid [J]. Molecules, 2020, 25(21): 5023. |
| 25 | Yu L, Zhao JB, Xu MM, et al. Metabolic engineering of Clostridium tyrobutyricum for n-butanol production: effects of CoA transferase [J]. Appl Microbiol Biotechnol, 2015, 99(11): 4917-4930. |
| 26 | Matsumoto K, Taguchi S. Biosynthetic polyesters consisting of 2-hydroxyalkanoic acids: current challenges and unresolved questions [J]. Appl Microbiol Biotechnol, 2013, 97(18): 8011-8021. |
| 27 | Volodina E, Schürmann M, Lindenkamp N, et al. Characterization of propionate CoA-transferase from Ralstonia eutropha H16 [J]. Appl Microbiol Biotechnol, 2014, 98(8): 3579-3589. |
| 28 | Matsumoto K, Taguchi S. Enzymatic and whole-cell synthesis of lactate-containing polyesters: toward the complete biological production of polylactate [J]. Appl Microbiol Biotechnol, 2010, 85(4): 921-932. |
| 29 | Lindenkamp N, Schürmann M, Steinbüchel A. A propionate CoA-transferase of Ralstonia eutropha H16 with broad substrate specificity catalyzing the CoA thioester formation of various carboxylic acids [J]. Appl Microbiol Biotechnol, 2013, 97(17): 7699-7709. |
| 30 | Park SJ, Kang KH, Lee H, et al. Propionyl-CoA dependent biosynthesis of 2-hydroxybutyrate containing polyhydroxyalkanoates in metabolically engineered Escherichia coli [J]. J Biotechnol, 2013, 165(2): 93-98. |
| 31 | Mezzolla V, D'Urso OF, Poltronieri P. Role of PhaC type I and type II enzymes during PHA biosynthesis [J]. Polymers, 2018, 10(8): 910. |
| 32 | Taguchi S. Current advances in microbial cell factories for lactate-based polyesters driven by lactate-polymerizing enzymes: towards the further creation of new LA-based polyesters [J]. Polym Degrad Stab, 2010, 95(8): 1421-1428. |
| 33 | Chek MF, Hiroe A, Hakoshima T, et al. PHA synthase (PhaC): interpreting the functions of bioplastic-producing enzyme from a structural perspective [J]. Appl Microbiol Biotechnol, 2019, 103(3): 1131-1141. |
| 34 | Zou HB, Shi MX, Zhang TT, et al. Natural and engineered polyhydroxyalkanoate (PHA) synthase: key enzyme in biopolyester production [J]. Appl Microbiol Biotechnol, 2017, 101(20): 7417-7426. |
| 35 | Tsuge T, Hyakutake M, Mizuno K. Class IV polyhydroxyalkanoate (PHA) synthases and PHA-producing Bacillus [J]. Appl Microbiol Biotechnol, 2015, 99(15): 6231-6240. |
| 36 | Rehm BHA. Polyester synthases: natural catalysts for plastics [J]. Biochem J, 2003, 376(Pt 1): 15-33. |
| 37 | Yuan W, Jia Y, Tian J, et al. Class I and III polyhydroxyalkanoate synthases from Ralstonia eutropha and Allochromatium vinosum: characterization and substrate specificity studies [J]. Arch Biochem Biophys, 2001, 394(1): 87-98. |
| 38 | Li AP, Cao XP, Fu RZ, et al. Biocatalysis of CO2 and CH4: key enzymes and challenges [J]. Biotechnol Adv, 2024, 72: 108347. |
| 39 | Liu Q, Xun GH, Feng Y. The state-of-the-art strategies of protein engineering for enzyme stabilization [J]. Biotechnol Adv, 2019, 37(4): 530-537. |
| 40 | Ochi A, Matsumoto K, Ooba T, et al. Engineering of class I lactate-polymerizing polyhydroxyalkanoate synthases from Ralstonia eutropha that synthesize lactate-based polyester with a block nature [J]. Appl Microbiol Biotechnol, 2013, 97(8): 3441-3447. |
| 41 | Yamada M, Matsumoto K, Shimizu K, et al. Adjustable mutations in lactate (LA)-polymerizing enzyme for the microbial production of LA-based polyesters with tailor-made monomer composition [J]. Biomacromolecules, 2010, 11(3): 815-819. |
| 42 | Phan HT, Furukawa S, Imai K, et al. Biosynthesis of high-molecular-weight poly(d-lactate)-containing block copolyesters using evolved sequence-regulating polyhydroxyalkanoate synthase PhaCAR [J]. ACS Sustainable Chem Eng, 2023, 11(30): 11123-11129. |
| 43 | Selmer T, Willanzheimer A, Hetzel M. Propionate CoA-transferase from Clostridium propionicum. Cloning of gene and identification of glutamate 324 at the active site [J]. Eur J Biochem, 2002, 269(1): 372-380. |
| 44 | Shi MX, Li MD, Yang AR, et al. Class I polyhydroxyalkanoate (PHA) synthase increased polylactic acid production in engineered Escherichia coli [J]. Front Bioeng Biotechnol, 2022, 10: 919969. |
| 45 | Matsumoto K, Hori C, Fujii R, et al. Dynamic changes of intracellular monomer levels regulate block sequence of polyhydroxyalkanoates in engineered Escherichia coli [J]. Biomacromolecules, 2018, 19(2): 662-671. |
| 46 | Jung YK, Kim TY, Park SJ, et al. Metabolic engineering of Escherichia coli for the production of polylactic acid and its copolymers [J]. Biotechnol Bioeng, 2010, 105(1): 161-171. |
| 47 | Jung YK, Lee SY. Efficient production of polylactic acid and its copolymers by metabolically engineered Escherichia coli [J]. J Biotechnol, 2011, 151(1): 94-101. |
| 48 | Matsumoto K, Tobitani K, Aoki S, et al. Improved production of poly(lactic acid)-like polyester based on metabolite analysis to address the rate-limiting step [J]. AMB Express, 2014, 4(1): 83. |
| 49 | Lajus S, Dusséaux S, Verbeke J, et al. Engineering the yeast Yarrowia lipolytica for production of polylactic acid homopolymer [J]. Front Bioeng Biotechnol, 2020, 8: 954. |
| 50 | Tan CL, Tao F, Xu P. Direct carbon capture for the production of high-performance biodegradable plastics by cyanobacterial cell factories [J]. Green Chem, 2022, 24(11): 4470-4483. |
| 51 | 邢敏钰, 冉淦侨, 谭丹. 酿酒酵母中萜类化合物的生物合成与代谢调控研究进展 [J]. 生物工程学报, 2024, 40(6): 1661-1693. |
| Xing MY, Ran GQ, Tan D. Advances in the biosynthesis and metabolic regulation of terpenoids in Saccharomyces cerevisiae [J]. Chin J Biotechnol, 2024, 40(6): 1661-1693. | |
| 52 | Krivoruchko A, Zhang YM, Siewers V, et al. Microbial acetyl-CoA metabolism and metabolic engineering [J]. Metab Eng, 2015, 28: 28-42. |
| 53 | 陈心宇, 李梦怡, 陈国强. 聚羟基脂肪酸酯PHA代谢工程研究30年 [J]. 生物工程学报, 2021, 37(5): 1794-1811. |
| Chen XY, Li MY, Chen GQ. Thirty years of metabolic engineering for biosynthesis of polyhydroxyalkanoates [J]. Chin J Biotechnol, 2021, 37(5): 1794-1811. | |
| 54 | Zheng YK, Cheng FY, Zheng B, et al. Enhancing single-cell hyaluronic acid biosynthesis by microbial morphology engineering [J]. Synth Syst Biotechnol, 2020, 5(4): 316-323. |
| 55 | Jordan A, Chandler J, MacCready JS, et al. Engineering cyanobacterial cell morphology for enhanced recovery and processing of biomass [J]. Appl Environ Microbiol, 2017, 83(9): e00053-17. |
| 56 | Wan L, Zhu YY, Ke JT, et al. Compartmentalization of pathway sequential enzymes into synthetic protein compartments for metabolic flux optimization in Escherichia coli [J]. Metab Eng, 2024, 85: 167-179. |
| 57 | Bertacchi S, Jayaprakash P, Morrissey JP, et al. Interdependence between lignocellulosic biomasses, enzymatic hydrolysis and yeast cell factories in biorefineries [J]. Microb Biotechnol, 2022, 15(3): 985-995. |
| 58 | Prangemeier T, Wildner C, Françani AO, et al. Yeast cell segmentation in microstructured environments with deep learning [J]. Biosystems, 2022, 211: 104557. |
| [1] | RAO Jun, ZHAO Chen, LI Duan-hua, LIAO Hao, HUANG Jia-yu, WANG Lu. Application of Auto-induction Strategy in Ergothioneine Biosynthesis [J]. Biotechnology Bulletin, 2025, 41(1): 333-346. |
| [2] | NIE Zhu-xin, GUO Jin, QIAO Zi-yang, LI Wei-wei, ZHANG Xue-yan, LIU Chun-yang, WANG Jing. Transcriptome Analysis of the Anthocyanin Biosynthesis in the Fruit Development Processes of Lycium ruthenicum Murr. [J]. Biotechnology Bulletin, 2024, 40(8): 106-117. |
| [3] | MA Xiao-xiang, MA Ze-yuan, LIU Ya-yue, ZHOU Long-jian, HE Yi-fan, ZHANG Yi. Effects of Simulated Mutational Biosynthetic Regulation on the Secondary Metabolites of Aspergillus terreus C23-3 [J]. Biotechnology Bulletin, 2024, 40(8): 275-287. |
| [4] | SHEN Zhen-hui, CAO Yao, YANG Lin-lei, LUO Xiang-ying, ZI Ling-shan, LU Qing-qing, LI Rong-chun. Cloning and Bioinformatics Analysis of the Ergothioneine Biosynthesis Genes in Naematelia aurantialba and Stereum hirsutum [J]. Biotechnology Bulletin, 2024, 40(7): 259-272. |
| [5] | HE Yu-bing, FU Zhen-hao, LI Ren-han, LIU Xiu-xia, LIU Chun-li, YANG Yan-kun, LI Ye, BAI Zhong-hu. Efficient Biosynthesis of 2-Naphthaleneethanol in Metabolically Engineered Saccharomyces cerevisiae [J]. Biotechnology Bulletin, 2024, 40(7): 99-107. |
| [6] | HU Jin-jin, LI Su-zhen, MA Xu-hui, LIU Xiao-qing, XIE Shan-shan, JIANG Hai-yang, CHEN Ru-mei. Regulation of Maize Anthocyanin Biosynthesis Metabolism [J]. Biotechnology Bulletin, 2024, 40(6): 34-44. |
| [7] | ZHANG Mei-yu, ZHAO Yu-bin, WANG Ling-yun, SONG Yuan-da, ZHAO Xin-he, REN Xiao-jie. Research Progress in the Production of Functional Fatty Acid DHA by Microalga Thraustochytrids [J]. Biotechnology Bulletin, 2024, 40(6): 81-94. |
| [8] | YANG Yan, HU Yang, LIU Ni-ru, YIN Lu, YANG Rui, WANG Peng-fei, MU Xiao-peng, ZHANG Shuai, CHENG Chun-zhen, ZHANG Jian-cheng. Cloning and Functional Analysis of MbbZIP43 Gene in ‘Hongmantang’ Red-flesh Apple [J]. Biotechnology Bulletin, 2024, 40(2): 146-159. |
| [9] | WANG Jun-fang, HUANG Qiu-bin, ZHANG Piao-dan, ZHANG Peng-pai. Structure and Biosynthesis of Surfactin as well as Its Role in Biological Control [J]. Biotechnology Bulletin, 2024, 40(1): 100-112. |
| [10] | CHEN Zhi-min, LI Cui, WEI Ji-tian, LI Xin-ran, LIU Yi, GUO Qiang. Research Progress in the Regulation of Chlorogenic Acid Biosynthesis and Its Application [J]. Biotechnology Bulletin, 2024, 40(1): 57-71. |
| [11] | HE Si-cheng, ZHANG Zi-yuan, HAN Yu-qing, MIAO Lin, ZHANG Cui-ying, YU Ai-qun. Research Progress in the Production of Polyunsaturated Fatty Acids by Yarrowia lipolytica Cell Factories [J]. Biotechnology Bulletin, 2024, 40(1): 72-85. |
| [12] | LI Liang, XU Shan-shan, JIANG Yan-jun. Research Progress in the Production of Ergothioneine by Biosynthesis [J]. Biotechnology Bulletin, 2024, 40(1): 86-99. |
| [13] | XUE Ning, WANG Jin, LI Shi-xin, LIU Ye, CHENG Hai-jiao, ZHANG Yue, MAO Yu-feng, WANG Meng. Construction of L-phenylalanine High-producing Corynebacterium glutamicum Engineered Strains via Multi-gene Simultaneous Regulation Combined with High-throughput Screening [J]. Biotechnology Bulletin, 2023, 39(9): 268-280. |
| [14] | CHENG Ya-nan, ZHANG Wen-cong, ZHOU Yuan, SUN Xue, LI Yu, LI Qing-gang. Synthetic Pathway Construction of Producing 2'-fucosyllactose by Lactococcus lactis and Optimization of Fermentation Medium [J]. Biotechnology Bulletin, 2023, 39(9): 84-96. |
| [15] | ZHAO Si-jia, WANG Xiao-lu, SUN Ji-lu, TIAN Jian, ZHANG Jie. Modification of Pichia pastoris for Erythritol Production by Metabolic Engineering [J]. Biotechnology Bulletin, 2023, 39(8): 137-147. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||