








YANG Yue1( ), LI Chang-ning1, SUN Xiao-lu1, LI Ya-qian1, DU Wei-jun1, WANG Li-xiang1, WANG Min1(
), LI Chang-ning1, SUN Xiao-lu1, LI Ya-qian1, DU Wei-jun1, WANG Li-xiang1, WANG Min1( ), JI Yue-mei2
), JI Yue-mei2
Received:2025-04-02
Online:2025-12-11
Contact:
WANG Min
E-mail:1297720641@qq.com;wangmin3502@126.com
YANG Yue, LI Chang-ning, SUN Xiao-lu, LI Ya-qian, DU Wei-jun, WANG Li-xiang, WANG Min, JI Yue-mei. Cloning and Identification of Salt Tolerance Function of Soybean Transcription Factor GmMYB069[J]. Biotechnology Bulletin, doi: 10.13560/j.cnki.biotech.bull.1985.2025-0348.
引物名称 Primer name | 引物序列 Primer sequence(5′-3′) |
|---|---|
| GmMYB069-F | ATGGGAAGGTCACCGTGTTG |
| GmMYB069-R | CCACAAAATTGGTGACGCAGAA |
| qRT-GmCYP2-F | CGGGACCAGTGTGCTTCTTCA |
| qRT-GmCYP2-R | CCCCTCCACTACAAAGGCTCG |
| qRT-GmMYB069-F | GCCTCATGTGGTGTTGAGTG |
| qRT-GmMYB069-R | TACAGCCACTGTCCATTTGGT |
| GmMYB069-pUBi-F | TTGATGTGATTACAGTCTAGAATGGGAAGGTCACCGTGTTG |
| GmMYB069-pUBi-R | GTTAATTAACCCGCTGGTACCCTACCACAAAATTGGTGACGCA |
Table 1 GmMYB069 primer sequence
引物名称 Primer name | 引物序列 Primer sequence(5′-3′) |
|---|---|
| GmMYB069-F | ATGGGAAGGTCACCGTGTTG |
| GmMYB069-R | CCACAAAATTGGTGACGCAGAA |
| qRT-GmCYP2-F | CGGGACCAGTGTGCTTCTTCA |
| qRT-GmCYP2-R | CCCCTCCACTACAAAGGCTCG |
| qRT-GmMYB069-F | GCCTCATGTGGTGTTGAGTG |
| qRT-GmMYB069-R | TACAGCCACTGTCCATTTGGT |
| GmMYB069-pUBi-F | TTGATGTGATTACAGTCTAGAATGGGAAGGTCACCGTGTTG |
| GmMYB069-pUBi-R | GTTAATTAACCCGCTGGTACCCTACCACAAAATTGGTGACGCA |
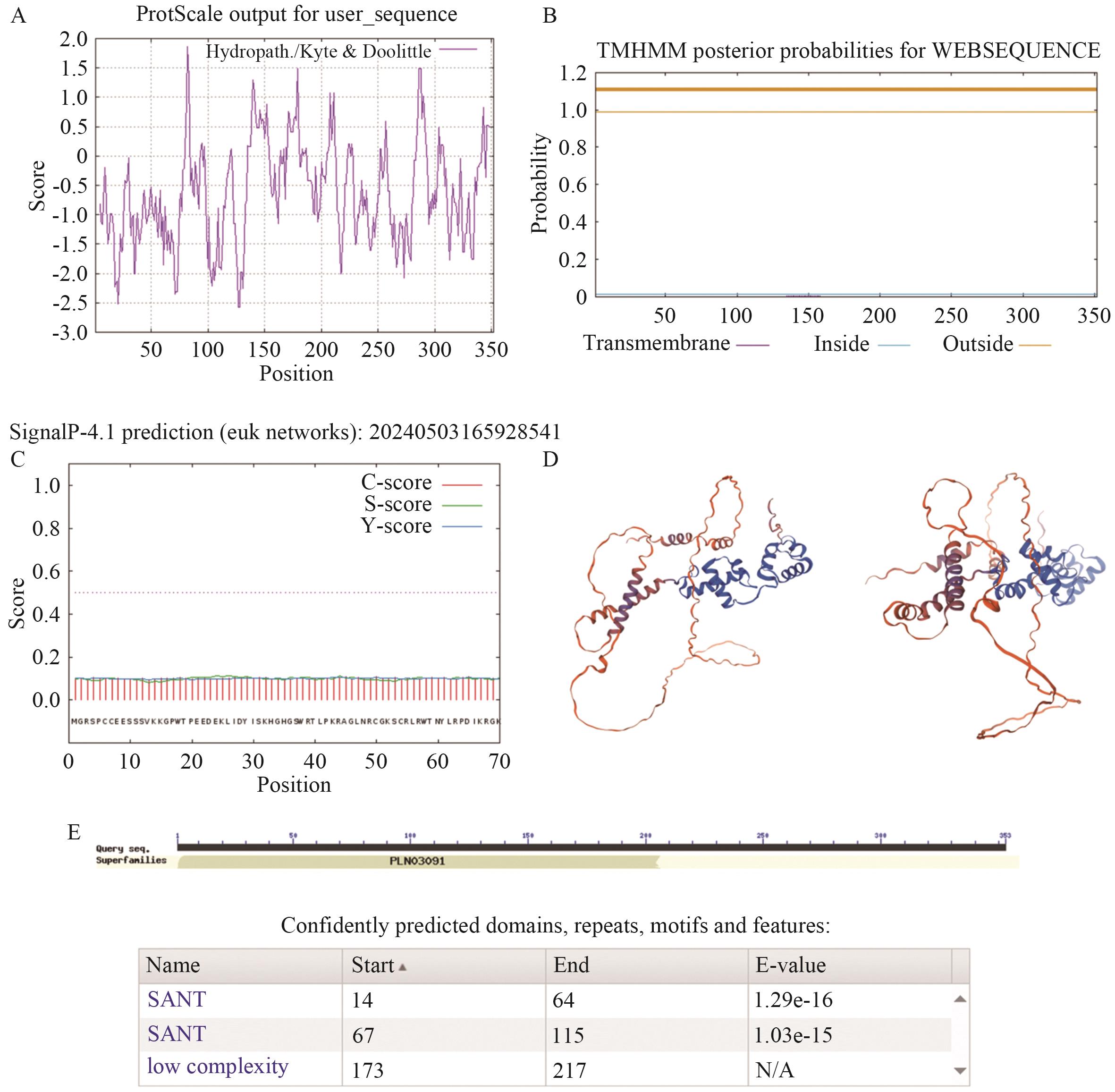
Fig. 5 Characterization of GmMYB069 proteinA. GmMYB069 protein hydrophobicity prediction map. B. GmMYB069 protein transmembrane domain prediction diagram. C. Signal peptide analysis of GmMYB069 protein (The horizontal coordinate indicates the position of the amino acid, and the vertical coordinate indicates the probability of the signal peptide). D. Conservative domains of GmMYB069 protein. E. Tertiary structure of GmMYB069 protein
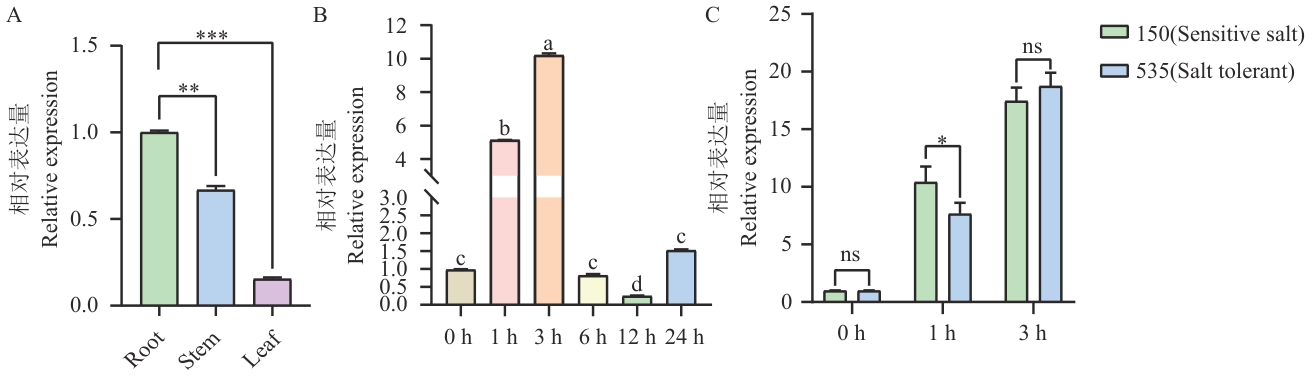
Fig. 7 Temporal and spatial expression pattern diagram of soybeanA. Expressions of GmMYB069 gene in different tissues of soybean. B. The expressions of GmMYB069 gene in soybean W82 under salt stress at different time periods. C. The expressions of GmMYB069 gene at different time periods in material 150 and 535. The data shown is mean±variance, and significant differences were analyzed using t-test. ns indicates no significant difference, *P<0.05, **P<0.01, and ***P<0.001, the same below
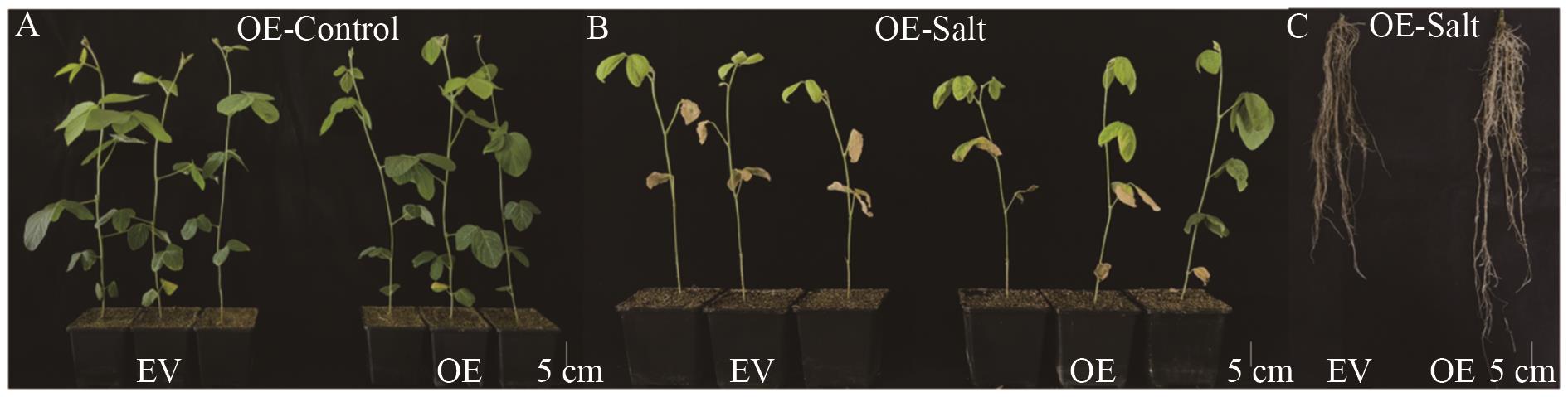

Fig. 9 Phenotypic analysis of GmMYB069 OverexpressionA. Phenotypes of GmMYB069-OE and control EV under water treatment (OE-Control: Overexpressed chimeric plants under water treatment). B. Phenotypes of GmMYB069-OE and control EV under salt treatment (OE-Salt: Overexpressed chimeric plants under salt treatment). C. Subterranean phenotype of GmMYB069-OE and control EV under salt stress (OE-Salt: Overexpressed chimeric plants under salt treatment)
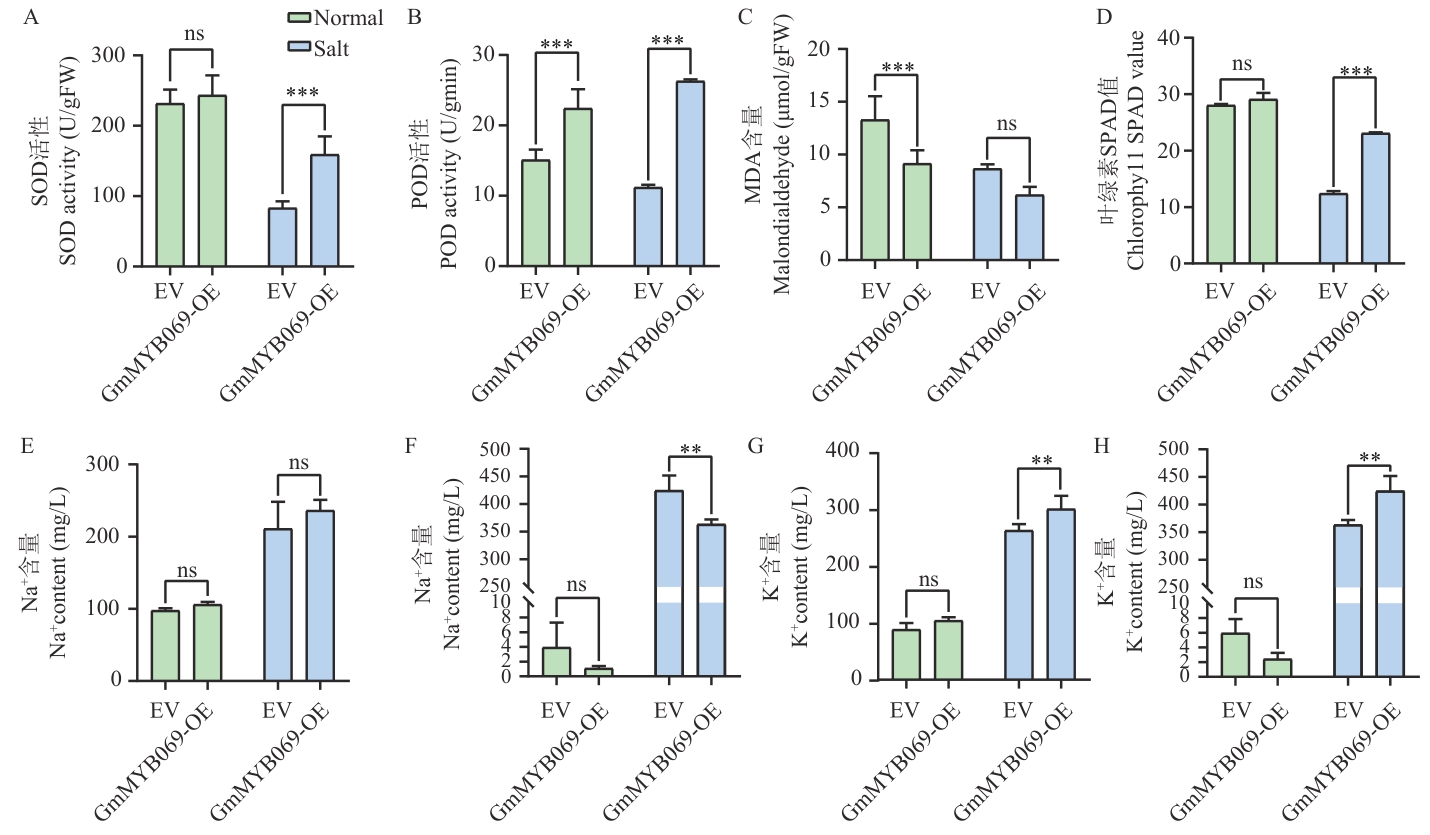
Fig. 10 Physiological indicators and Na+, K+ contents of GmMYB069 overexpressing chimeric plants under salt stressA: SOD activity in GmMYB069-OE leaves. B: POD activity in GmMYB069-OE leaves. C: MDA content in GmMYB069-OE leaves. D: SPAD value of GmMYB069-OE leaves. E: Na+ content in the roots of GmMYB069-OE plants. F: Na+ content in the leaves of GmMYB069-OE plants. G: K+ content in the roots of GmMYB069-OE plants. H: K+ content in the leaves of GmMYB069-OE plants
| [1] | 朱子坤, 马超, 王锦辉, 等. 大豆GmNARK基因家族鉴定及分析 [J]. 大豆科技, 2023(2): 10-20. |
| Zhu ZK, Ma C, Wang JH, et al. Identification and functional analysis of GmNARK gene family in soybean [J]. Soybean Sci & Technol, 2023(2): 10-20. | |
| [2] | Cheng Q, Gan ZR, Wang YP, et al. The soybean gene J contributes to salt stress tolerance by up-regulating salt-responsive genes [J]. Front Plant Sci, 2020, 11: 272. |
| [3] | 王燕. 樱桃李(Prunus cerasifera Ehrh .)果实主要花色苷组分及相关特性分析 [D]. 泰安: 山东农业大学, 2013. |
| Wang Y. Analysis of the main anthocyanin components and related characteristics of cherry plum(Prunus cerasifera Ehrh .)fruit [D]. Taian: Shandong Agricultural University, 2013. | |
| [4] | Aoyagi LN, Lopes-Caitar VS, de Carvalho MCCG, et al. Genomic and transcriptomic characterization of the transcription factor family R2R3-MYB in soybean and its involvement in the resistance responses to Phakopsora pachyrhizi [J]. Plant Sci, 2014, 229: 32-42. |
| [5] | Liao Y, Zou HF, Wang HW, et al. Soybean GmMYB76, GmMYB92, and GmMYB177 genes confer stress tolerance in transgenic Arabidopsis plants [J]. Cell Res, 2008, 18(10): 1047-1060. |
| [6] | Niu CF, Wei W, Zhou QY, et al. Wheat WRKY genes TaWRKY2 and TaWRKY19 regulate abiotic stress tolerance in transgenic Arabidopsis plants [J]. Plant Cell Environ, 2012, 35(6): 1156-1170. |
| [7] | Hao YJ, Wei W, Song QX, et al. Soybean NAC transcription factors promote abiotic stress tolerance and lateral root formation in transgenic plants [J]. Plant J, 2011, 68(2): 302-313. |
| [8] | Seo YJ, Park JB, Cho YJ, et al. Overexpression of the ethylene-responsive factor gene BrERF4 from Brassica rapa increases tolerance to salt and drought in Arabidopsis plants [J]. Mol Cells, 2010, 30(3): 271-278. |
| [9] | Jeong JS, Kim YS, Baek KH, et al. Root-specific expression of OsNAC10 improves drought tolerance and grain yield in rice under field drought conditions [J]. Plant Physiol, 2010, 153(1): 185-197. |
| [10] | Zou XL, Neuman D, Shen QJ. Interactions of two transcriptional repressors and two transcriptional activators in modulating gibberellin signaling in aleurone cells [J]. Plant Physiol, 2008, 148(1): 176-186. |
| [11] | Liao Y, Zou HF, Wei W, et al. Soybean GmbZIP44, GmbZIP62 and GmbZIP78 genes function as negative regulator of ABA signaling and confer salt and freezing tolerance in transgenic Arabidopsis [J]. Planta, 2008, 228(2): 225-240. |
| [12] | Nakashima K, Tran LP, Van Nguyen D, et al. Functional analysis of a NAC-type transcription factor OsNAC6 involved in abiotic and biotic stress-responsive gene expression in rice [J]. Plant J, 2007, 51(4): 617-630. |
| [13] | Hu HH, Dai MQ, Yao JL, et al. Overexpressing a NAM, ATAF, and CUC (NAC) transcription factor enhances drought resistance and salt tolerance in rice [J]. Proc Natl Acad Sci U S A, 2006, 103(35): 12987-12992. |
| [14] | Tran LP, Nakashima K, Sakuma Y, et al. Isolation and functional analysis of Arabidopsis stress-Inducible NAC transcription factors that bind to a drought-responsivecis-element in theearly responsive to dehydration stress 1 promoter W [J]. Plant Cell, 2004, 16(9): 2481-2498. |
| [15] | DuanMu HZ, Wang Y, Bai X, et al. Wild soybean roots depend on specific transcription factors and oxidation reduction related genesin response to alkaline stress [J]. Funct Integr Genom, 2015, 15(6): 651-660. |
| [16] | Gonzalez A, Zhao MZ, Leavitt JM, et al. Regulation of the anthocyanin biosynthetic pathway by the TTG1/bHLH/Myb transcriptional complex in Arabidopsis seedlings [J]. Plant J, 2008, 53(5): 814-827. |
| [17] | Rabinowicz PD, Braun EL, Wolfe AD, et al. Maize R2R3 myb genes: sequence analysis reveals amplification in the higher plants [J]. Genetics, 1999, 153(1): 427-444. |
| [18] | Du H, Yang SS, Liang Z, et al. Genome-wide analysis of the MYB transcription factor superfamily in soybean [J]. BMC Plant Biol, 2012, 12(1): 106. |
| [19] | 赵盼盼. 番茄R2R3MYB转录因子家族鉴定及SlMYB41和SlMYB64基因功能研究 [D]. 泰安: 山东农业大学, 2017. |
| Zhao PP. Identification of tomato R2R3MYB transcription factor family and functional study of SlMYB41 and SlMYB64 genes [D]. Tai’an: Shandong Agricultural University, 2017. | |
| [20] | 王欣怡, 王晓倩, 刘亚玲, 等. 紫花苜蓿MsMYB33基因克隆、亚细胞定位及转录自激活检测 [J]. 草地学报, 2023, 31(5): 1331-1337. |
| Wang XY, Wang XQ, Liu YL, et al. Cloning, subcellular localization and autonomous transcriptional activation testing of MsMYB33 gene from Medicago sativa [J]. Acta Agrestia Sin, 2023, 31(5): 1331-1337. | |
| [21] | Stracke R, Werber M, Weisshaar B. The R2R3-MYB gene family in Arabidopsis thaliana [J]. Curr Opin Plant Biol, 2001, 4(5): 447-456. |
| [22] | Dubos C, Stracke R, Grotewold E, et al. MYB transcription factors in Arabidopsis [J]. Trends Plant Sci, 2010, 15(10): 573-581. |
| [23] | Zhang LC, Zhao GY, Jia JZ, et al. Molecular characterization of 60 isolated wheat MYB genes and analysis of their expression during abiotic stress [J]. J Exp Bot, 2012, 63(1): 203-214. |
| [24] | 牛义岭, 姜秀明, 许向阳. 植物转录因子MYB基因家族的研究进展 [J]. 分子植物育种, 2016, 14(8): 2050-2059. |
| Niu YL, Jiang XM, Xu XY. Reaserch advances on transcription factor MYB gene family in plant [J]. Mol Plant Breed, 2016, 14(8): 2050-2059. | |
| [25] | Wang XP, Niu YL, Zheng Y. Multiple functions of MYB transcription factors in abiotic stress responses [J]. Int J Mol Sci, 2021, 22(11): 6125. |
| [26] | Zhang P, Wang RL, Yang XP, et al. The R2R3-MYB transcription factor AtMYB49 modulates salt tolerance in Arabidopsis by modulating the cuticle formation and antioxidant defence [J]. Plant Cell Environ, 2020, 43(8): 1925-1943. |
| [27] | Tang YH, Bao XX, Zhi YL, et al. Overexpression of a MYB family gene, OsMYB6, increases drought and salinity stress tolerance in transgenic rice [J]. Front Plant Sci, 2019, 10: 168. |
| [28] | 石广成, 杨万明, 杜维俊, 等. 大豆耐盐种质的筛选及其耐盐生理特性分析 [J]. 生物技术通报, 2022, 38(4): 174-183. |
| Shi GC, Yang WM, Du WJ, et al. Screening of salt-tolerant soybean germplasm and physiological characteristics analysis of its salt tolerance [J]. Biotechnol Bull, 2022, 38(4): 174-183. | |
| [29] | 孙群, 胡景江. 植物生理学研究技术 [M]. 杨凌: 西北农林科技大学出版社, 2006. |
| Sun Q, Hu JJ. Research technology of plant physiology [M]. Yangling: Northwest A&F University Press, 2006. | |
| [30] | Matsushita N, Matoh T. Characterization of Na+ exclusion mechanisms of salt-tolerant reed plants in comparison with salt-sensitive rice plants [J]. Physiol Plant, 1991, 83(1): 170-176. |
| [31] | 赵团结, 盖钧镒. 栽培大豆起源与演化研究进展 [J]. 中国农业科学, 2004, 37(7): 954-962. |
| Zhao TJ, Gai JY. The origin and evolution of cultivated soybean [Glycine max (L.) merr. [J]. Sci Agric Sin, 2004, 37(7): 954-962. | |
| [32] | Qiu PC, Du YC, Song MG, et al. Genetic overlap between salt and low-temperature tolerance loci at germination stage of soybean [J]. Eng Sci, 2013, 11(5): 37-40. |
| [33] | Rossi M, Borromeo I, Capo C, et al. PGPB improve photosynthetic activity and tolerance to oxidative stress in Brassica napus grown on salinized soils [J]. Applied Sciences, 2021, 11(23): 11442. |
| [34] | Lotkowska ME, Tohge T, Fernie AR, et al. The Arabidopsis transcription factor MYB112 promotes anthocyanin formation during salinity and under high light stress [J]. Plant Physiol, 2015, 169(3): 1862-1880. |
| [35] | 王春荣, 王庆杰, 李晓玲, 等. 葡萄中盐诱导的R2R3-MYB基因的筛选与表达分析 [J]. 园艺学报, 2014, 41(3): 529-535. |
| Wang CR, Wang QJ, Li XL, et al. Screening and expression analysis of salt-induced R2R3-MYB genes in grapes [J]. Acta Horticulturae Sinica, 2014, 41(3): 529-535. | |
| [36] | Wang L, Qiu TQ, Yue JR, et al. Arabidopsis ADF1 is regulated by MYB73 and is involved in response to salt stress affecting actin filament organization [J]. Plant Cell Physiol, 2021, 62(9): 1387-1395. |
| [37] | He YX, Dong YS, Yang XD, et al. Functional activation of a novel R2R3-MYB protein gene, GmMYB68, confers salt-alkali resistance in soybean (Glycine max L.) [J]. Genome, 2020, 63(1): 13-26. |
| [38] | Zhang WX, Wang N, Yang JT, et al. The salt-induced transcription factor GmMYB84 confers salinity tolerance in soybean [J]. Plant Sci, 2020, 291: 110326. |
| [39] | Du YT, Zhao MJ, Wang CT, et al. Identification and characterization of GmMYB118 responses to drought and salt stress [J]. BMC Plant Biol, 2018, 18(1): 320. |
| [40] | Pi EX, Xu J, Li HH, et al. Enhanced salt tolerance of rhizobia-inoculated soybean correlates with decreased phosphorylation of the transcription factor GmMYB183 and altered flavonoid biosynthesis [J]. Mol Cell Proteom, 2019, 18(11): 2225-2243. |
| [41] | Pi EX, Zhu CM, Fan W, et al. Quantitative phosphoproteomic and metabolomic analyses reveal GmMYB173 optimizes flavonoid metabolism in soybean under salt stress [J]. Mol Cell Proteom, 2018, 17(6): 1209-1224. |
| [42] | Wang C, Wang LJ, Lei J, et al. IbMYB308, a sweet potato R2R3-MYB gene, improves salt stress tolerance in transgenic tobacco [J]. Genes, 2022, 13(8): 1476. |
| [43] | Zhang X, Chen LC, Shi QH, et al. SlMYB102, an R2R3-type MYB gene, confers salt tolerance in transgenic tomato [J]. Plant Sci, 2020, 291: 110356. |
| [44] | Ullah A, Ul Qamar MT, Nisar M, et al. Characterization of a novel cotton MYB gene, GhMYB108-like responsive to abiotic stresses [J]. Mol Biol Rep, 2020, 47(3): 1573-1581. |
| [45] | 刘佳欣, 刘慧子, 石晶静, 等. 白桦MYB基因响应激素及盐旱处理的表达研究 [J]. 植物研究, 2020, 40(5): 743-750. |
| Liu JX, Liu HZ, Shi JJ, et al. Expression of MYB genes of birch in response to hormones, salt and drought [J]. Bull Bot Res, 2020, 40(5): 743-750. | |
| [46] | Roy SJ, Negrão S, Tester M. Salt resistant crop plants [J]. Curr Opin Biotechnol, 2014, 26: 115-124. |
| [47] | Deinlein U, Stephan AB, Horie T, et al. Plant salt-tolerance mechanisms [J]. Trends Plant Sci, 2014, 19(6): 371-379. |
| [48] | Kar RK. Plant responses to water stress: Role of reactive oxygen species [J]. Plant Signal Behav, 2011, 6(11): 1741-1745. |
| [49] | You J, Chan ZL. ROS regulation during abiotic stress responses in crop plants [J]. Front Plant Sci, 2015, 6: 1092. |
| [50] | Xu ZY, Wang CY, Xue F, et al. Wheat NAC transcription factor TaNAC29 is involved in response to salt stress [J]. Plant Physiol Biochem, 2015, 96: 356-363. |
| [51] | 李妍. 盐和PEG胁迫对丝瓜幼苗抗氧化酶活性及丙二醛含量的影响 [J]. 干旱地区农业研究, 2009, 27(2): 159-162, 178. |
| Li Y. Effect of salt and PEG to antioxidant enzymes activity and MDA concentration of Luffa cylindrical Roem [J]. Agric Res Arid Areas, 2009, 27(2): 159-162, 178. |
| [1] | DONG Xiang-xiang, MIAO Bai-ling, XU He-juan, CHEN Juan-juan, LI Liang-jie, GONG Shou-fu, ZHU Qing-song. Bioinformatics Analysis and Flowering Regulation Function of FveBBX20 Gene in Woodland Strawberry [J]. Biotechnology Bulletin, 2025, 41(9): 115-123. |
| [2] | LI Shan, MA Deng-hui, MA Hong-yi, YAO Wen-kong, YIN Xiao. Identification and Expression Analysis of SKP1 Gene Family in Grapevine (Vitis vinifera L.) [J]. Biotechnology Bulletin, 2025, 41(9): 147-158. |
| [3] | HUANG Guo-dong, DENG Yu-xing, CHENG Hong-wei, DAN Yan-nan, ZHOU Hui-wen, WU Lan-hua. Genome-wide Identification and Expression Analysis of the ZIP Gene Family in Soybean [J]. Biotechnology Bulletin, 2025, 41(9): 71-81. |
| [4] | GONG Hui-ling, XING Yu-jie, MA Jun-xian, CAI Xia, FENG Zai-ping. Identification of Laccase (LAC) Gene Family in Potato (Solanum tuberosum L.) and Its Expression Analysis under Salt Stresses [J]. Biotechnology Bulletin, 2025, 41(9): 82-93. |
| [5] | GUAN Zhi-hao, SHAN Zhi-yi, XIONG He, ZHAO Rui-xue. Computational Literature-based Knowledge Discovery for Soybean Coupling Traits [J]. Biotechnology Bulletin, 2025, 41(9): 345-356. |
| [6] | CHENG Ting-ting, LIU Jun, WANG Li-li, LIAN Cong-long, WEI Wen-jun, GUO Hui, WU Yao-lin, YANG Jing-fan, LAN Jin-xu, CHEN Sui-qing. Genome-wide Identification of the Chalcone Isomerase Gene Family in Eucommia ulmoides and Analysis of Their Expression Patterns [J]. Biotechnology Bulletin, 2025, 41(9): 242-255. |
| [7] | XU Xiao-ping, YANG Cheng-long, HE Xing, GUO Wen-jie, WU Jian, FANG Shao-zhong. Cloning of the LoAPS1 and Its Function Analysis during the Process of Dormancy Release in Lilium [J]. Biotechnology Bulletin, 2025, 41(9): 195-206. |
| [8] | ZHANG Yong-yan, GUO Si-jian, LI Jing, HAO Si-yi, LI Rui-de, LIU Jia-peng, CHENG Chun-zhen. Gene Cloning and Functional Analysis of the Anthocyanin-related VcGSTF19 Gene in Blueberry (Vaccinium corymbosum L.) [J]. Biotechnology Bulletin, 2025, 41(9): 139-146. |
| [9] | LI Ya-tao, ZHANG Zhi-peng, ZHAO Meng-yao, LYU Zhen, GAN Tian, WEI Hao, WU Shu-feng, MA Yu-chao. Whole Genome Analysis of Bradyrhizobium sp. Bd1 and the Negative Regulating Function of TetR3 during Cell Growth and Nodulation [J]. Biotechnology Bulletin, 2025, 41(9): 289-301. |
| [10] | CHENG Xue, FU Ying, CHAI Xiao-jiao, WANG Hong-yan, DENG Xin. Identification of LHC Gene Family in Setaria italica and Expression Analysis under Abiotic Stresses [J]. Biotechnology Bulletin, 2025, 41(8): 102-114. |
| [11] | LA Gui-xiao, ZHAO Yu-long, DAI Dan-dan, YU Yong-liang, GUO Hong-xia, SHI Gui-xia, JIA Hui, YANG Tie-gang. Identification of Plasma Membrane H+-ATPase Gene Family in Safflower and Expression Analysis in Response to Low Nitrogen and Low Phosphorus Stress [J]. Biotechnology Bulletin, 2025, 41(8): 220-233. |
| [12] | LAI Shi-yu, LIANG Qiao-lan, WEI Lie-xin, NIU Er-bo, CHEN Ying-e, ZHOU Xin, YANG Si-zheng, WANG Bo. The Role of NbJAZ3 in the Infection of Nicotiana benthamiana by Alfalfa Mosaic Virus [J]. Biotechnology Bulletin, 2025, 41(8): 186-196. |
| [13] | KANG Qin, WANG Xia, SHEN Ming-yang, XU Jing-tian, CHEN Shi-lan, LIAO Ping-yang, XU Wen-zhi, WU Wei, XU Dong-bei. Cloning and Expression Analysis of the UV-B Receptor Gene McUVR8 in Mentha canadensis L. [J]. Biotechnology Bulletin, 2025, 41(8): 255-266. |
| [14] | ZHU Li-juan, ZHANG Kai, WEN Xiao-lei, CHU Jia-hao, SHI Feng-yu, WANG Yan-li. Mining the Core Genes Being Tolerant to Cadmium in Wild Soybean by WGCNA [J]. Biotechnology Bulletin, 2025, 41(8): 124-136. |
| [15] | LI Kai-jie, WU Yao, LI Dan-dan. Cloning of Gene CtbHLH128 in Safflower and Response Function Regulating Drought Stress [J]. Biotechnology Bulletin, 2025, 41(8): 234-241. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||