








Biotechnology Bulletin ›› 2026, Vol. 42 ›› Issue (2): 278-292.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0585
Previous Articles Next Articles
YANG Zi-han1,2,3( ), LI Kui-xiu1,2,3, LI Jun-liang1,2,3, LYU Wen-hui1,2,3, HE Wen-ting4, LIU Xu-yan5, LIU Guan-ze1,2(
), LI Kui-xiu1,2,3, LI Jun-liang1,2,3, LYU Wen-hui1,2,3, HE Wen-ting4, LIU Xu-yan5, LIU Guan-ze1,2( )
)
Received:2025-06-06
Online:2026-02-26
Published:2026-03-17
Contact:
LIU Guan-ze
E-mail:1439104670@qq.com;guanzeliu@ynau.edu.cn
YANG Zi-han, LI Kui-xiu, LI Jun-liang, LYU Wen-hui, HE Wen-ting, LIU Xu-yan, LIU Guan-ze. Identification and Expression Analysis of the RALF Gene Family in Panax notoginseng[J]. Biotechnology Bulletin, 2026, 42(2): 278-292.
| 基因 Gene | 上游引物Forward primer (5′-3′) | 下游引物序列Reverse primer (5′-3′) |
|---|---|---|
| PnACT2 | GATTGATCTTGGCACCGGGA | TGACCAACCCACGACCTTTC |
| PnoRALF1 | GCTACATGCAACGGTCTGATC | TGCAGTTGTAGTAGGAGGCTC |
| PnoRALF2 | CCACCTGAACTCTATTGGGT | CGCACCGTAGCTGATGTA |
| PnoRALF3 | AGTGATGCTGATGGACTCGG | CGTCTGTAGGGATTGGCTCT |
| PnoRALF4 | CCACCTGAACTCTATTGGGT | GGGAGTACGGATTTGCCTG |
| PnoRALF5 | CGACATGGATTCAGAGAGC | ACGGTTGTAAGGGTTAGCTT |
| PnoRALF6 | CGCCTCCTACTACAACTGTA | ACCAGCATCCATCACAAATC |
| PnoRALF7 | GCAAACAGTCGGTACATCAG | GGGAGTACGGATTTGCCTG |
| PnoRALF8 | GATTCATGGTCGTTTGTCAC | CTGGCCTCGTTTGTTACATG |
| PnoRALF9 | TGATCTTGCTCCCATTTCCA | GGCTGATCTCCGAATCCATA |
| PnoRALF10 | TGTTTCGACGGTGGTGTTTC | CACACACCTGCTCGTTTCTC |
| PnoRALF11 | ATGATGTTGGATTCGGAGGC | CGTCTGTAGGGATTGGCTCT |
| PnoRALF12 | AGTCCTAATCCTGCTGTGTTTC | CGGTTTATCTCCGAATCCATCA |
| PnoRALF13 | CGGAGGTTTCTTGCAGCAAA | CGGAGGTTTCTTGCAGCAAA |
| pEAQ-HT-PnoRALF9 | cccaaattcgcgaccggtATGGGAAAG TCTACTGGTTTTCTCG | atgaaaccagagttaaaggcctcgagACGCC GGCACCGAGTGAT |
Table 1 Primers for the RT-qPCR analysis of PnoRALFs
| 基因 Gene | 上游引物Forward primer (5′-3′) | 下游引物序列Reverse primer (5′-3′) |
|---|---|---|
| PnACT2 | GATTGATCTTGGCACCGGGA | TGACCAACCCACGACCTTTC |
| PnoRALF1 | GCTACATGCAACGGTCTGATC | TGCAGTTGTAGTAGGAGGCTC |
| PnoRALF2 | CCACCTGAACTCTATTGGGT | CGCACCGTAGCTGATGTA |
| PnoRALF3 | AGTGATGCTGATGGACTCGG | CGTCTGTAGGGATTGGCTCT |
| PnoRALF4 | CCACCTGAACTCTATTGGGT | GGGAGTACGGATTTGCCTG |
| PnoRALF5 | CGACATGGATTCAGAGAGC | ACGGTTGTAAGGGTTAGCTT |
| PnoRALF6 | CGCCTCCTACTACAACTGTA | ACCAGCATCCATCACAAATC |
| PnoRALF7 | GCAAACAGTCGGTACATCAG | GGGAGTACGGATTTGCCTG |
| PnoRALF8 | GATTCATGGTCGTTTGTCAC | CTGGCCTCGTTTGTTACATG |
| PnoRALF9 | TGATCTTGCTCCCATTTCCA | GGCTGATCTCCGAATCCATA |
| PnoRALF10 | TGTTTCGACGGTGGTGTTTC | CACACACCTGCTCGTTTCTC |
| PnoRALF11 | ATGATGTTGGATTCGGAGGC | CGTCTGTAGGGATTGGCTCT |
| PnoRALF12 | AGTCCTAATCCTGCTGTGTTTC | CGGTTTATCTCCGAATCCATCA |
| PnoRALF13 | CGGAGGTTTCTTGCAGCAAA | CGGAGGTTTCTTGCAGCAAA |
| pEAQ-HT-PnoRALF9 | cccaaattcgcgaccggtATGGGAAAG TCTACTGGTTTTCTCG | atgaaaccagagttaaaggcctcgagACGCC GGCACCGAGTGAT |
| 基因 | 基因ID | 氨基酸数 | 分子量 | 等电点 | 亲水性 | 亚细胞定位 | 信号肽 |
|---|---|---|---|---|---|---|---|
| Gene name | Gene ID | Number of amino acids | Molecular weight (kD) | pl | GRAVY | Subcellular localization | Signal peptide |
| PnoRALF1 | Pno01G005926.t1 | 128 | 14.06 | 9.03 | -0.30 | 细胞外Extracellular | Yes |
| PnoRALF2 | Pno03G001289.t1 | 126 | 13.74 | 8.91 | -0.11 | 细胞外Extracellular | Yes |
| PnoRALF3 | Pno03G005478.t1 | 131 | 14.39 | 0.24 | -0.22 | 细胞外Extracellular | Yes |
| PnoRALF4 | Pno03G005479.t1 | 125 | 13.75 | 9.54 | -0.44 | 细胞外Extracellular | Yes |
| PnoRALF5 | Pno06G000518.t1 | 114 | 12.33 | 8.60 | -0.20 | 细胞外Extracellular | Yes |
| PnoRALF6 | Pno07G004032.t1 | 158 | 17.28 | 9.15 | 0.24 | 质膜 Plasma membrane | Yes |
| PnoRALF7 | Pno09G001048.t1 | 109 | 12.07 | 7.50 | -0.35 | 细胞外Extracellular | Yes |
| PnoRALF8 | Pno10G000839.t1 | 123 | 13.77 | 9.83 | -0.19 | 细胞外Extracellular | No |
| PnoRALF9 | Pno11G003888.t1 | 133 | 14.33 | 8.99 | -0.08 | 细胞外Extracellular | Yes |
| PnoRALF10 | Pno12G001525.t1 | 110 | 12.31 | 7.54 | -0.07 | 细胞外Extracellular | Yes |
| PnoRALF11 | Pno284G000001.t1 | 125 | 12.75 | 9.54 | -0.44 | 细胞外Extracellular | Yes |
| PnoRALF12 | Pno606G000002.t1 | 110 | 12.30 | 7.54 | -0.07 | 细胞外Extracellular | Yes |
| PnoRALF13 | Pno606G000003.t1 | 113 | 12.46 | 8.69 | -0.29 | 细胞外Extracellular | Yes |
Table 2 Basic physicochemical properties of PnoRALFs gene family
| 基因 | 基因ID | 氨基酸数 | 分子量 | 等电点 | 亲水性 | 亚细胞定位 | 信号肽 |
|---|---|---|---|---|---|---|---|
| Gene name | Gene ID | Number of amino acids | Molecular weight (kD) | pl | GRAVY | Subcellular localization | Signal peptide |
| PnoRALF1 | Pno01G005926.t1 | 128 | 14.06 | 9.03 | -0.30 | 细胞外Extracellular | Yes |
| PnoRALF2 | Pno03G001289.t1 | 126 | 13.74 | 8.91 | -0.11 | 细胞外Extracellular | Yes |
| PnoRALF3 | Pno03G005478.t1 | 131 | 14.39 | 0.24 | -0.22 | 细胞外Extracellular | Yes |
| PnoRALF4 | Pno03G005479.t1 | 125 | 13.75 | 9.54 | -0.44 | 细胞外Extracellular | Yes |
| PnoRALF5 | Pno06G000518.t1 | 114 | 12.33 | 8.60 | -0.20 | 细胞外Extracellular | Yes |
| PnoRALF6 | Pno07G004032.t1 | 158 | 17.28 | 9.15 | 0.24 | 质膜 Plasma membrane | Yes |
| PnoRALF7 | Pno09G001048.t1 | 109 | 12.07 | 7.50 | -0.35 | 细胞外Extracellular | Yes |
| PnoRALF8 | Pno10G000839.t1 | 123 | 13.77 | 9.83 | -0.19 | 细胞外Extracellular | No |
| PnoRALF9 | Pno11G003888.t1 | 133 | 14.33 | 8.99 | -0.08 | 细胞外Extracellular | Yes |
| PnoRALF10 | Pno12G001525.t1 | 110 | 12.31 | 7.54 | -0.07 | 细胞外Extracellular | Yes |
| PnoRALF11 | Pno284G000001.t1 | 125 | 12.75 | 9.54 | -0.44 | 细胞外Extracellular | Yes |
| PnoRALF12 | Pno606G000002.t1 | 110 | 12.30 | 7.54 | -0.07 | 细胞外Extracellular | Yes |
| PnoRALF13 | Pno606G000003.t1 | 113 | 12.46 | 8.69 | -0.29 | 细胞外Extracellular | Yes |
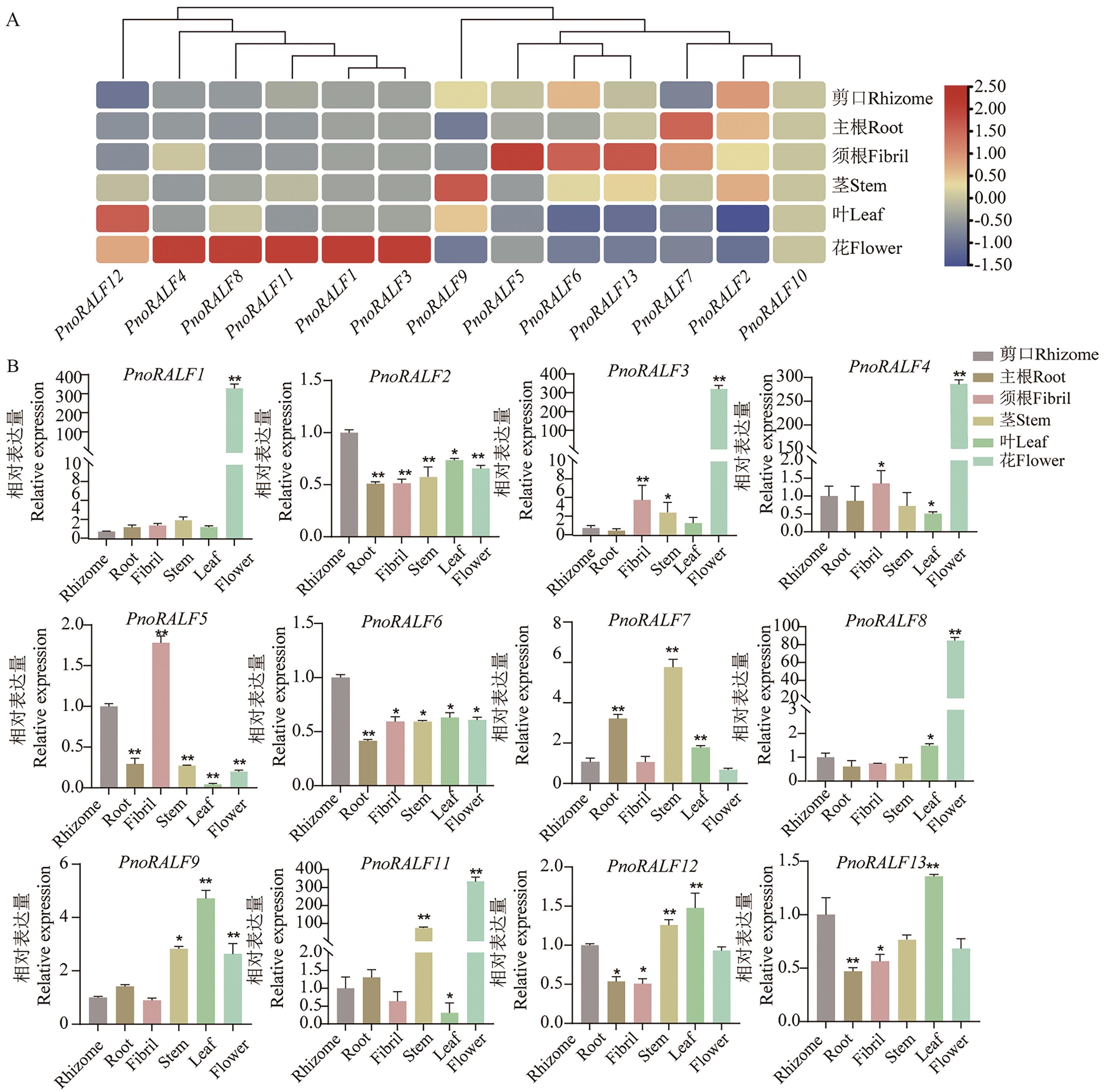
Fig. 6 Expression profiling of PnoRALFs gene in different tissuesA: Transcriptome expression data of PnoRALFs gene in different tissues. B: Relative expressions of PnoRALFs gene in different tissues. *P<0.05, **P<0.01. The same below
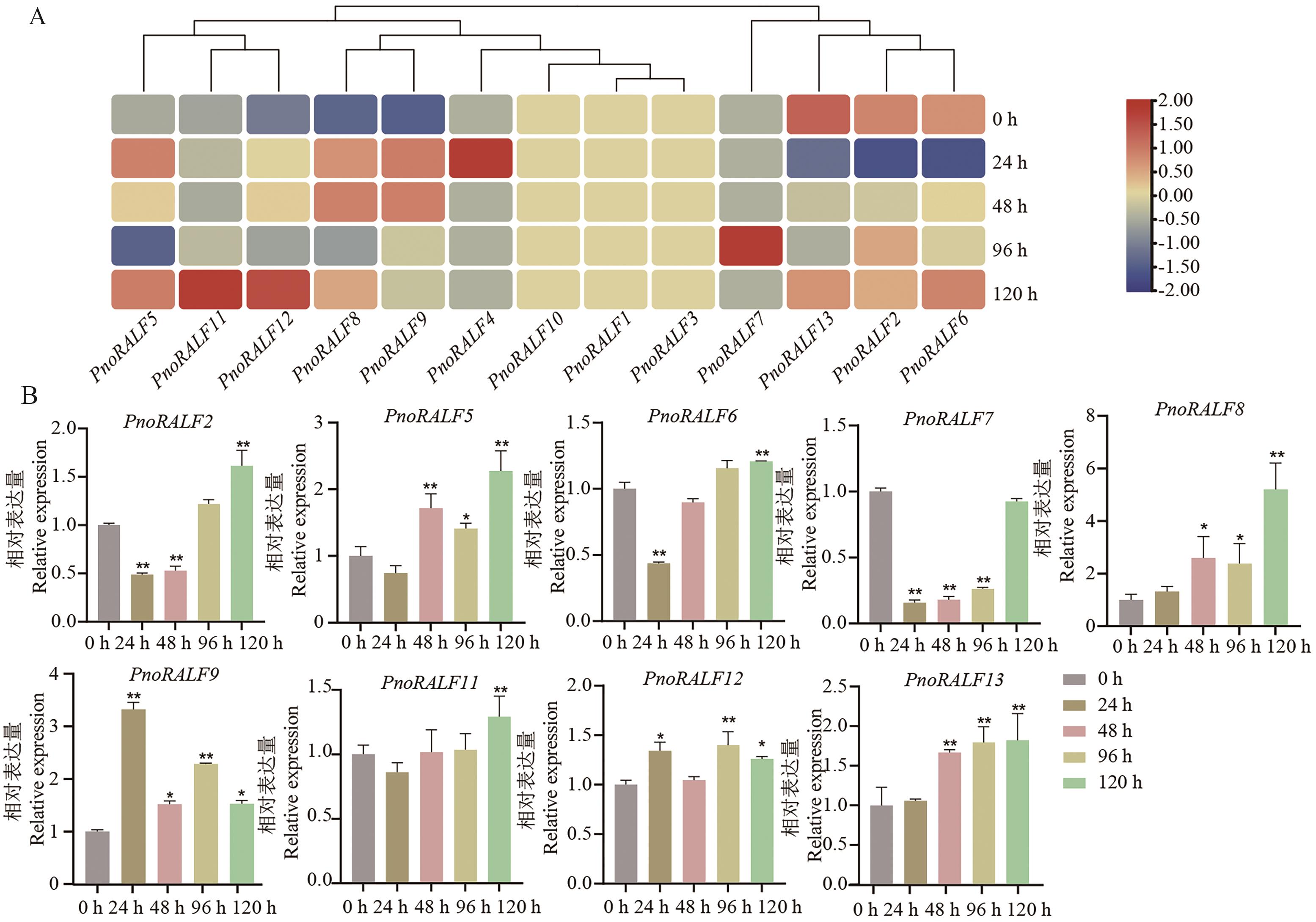
Fig. 11 Expression profiling of PnoRALFs gene in response to C. destructans infectionA: Transcriptomic data of PnoRALFs gene in response to C. destructans infection.B: Relative expression of PnoRALFs gene reponding to C. destructans infection
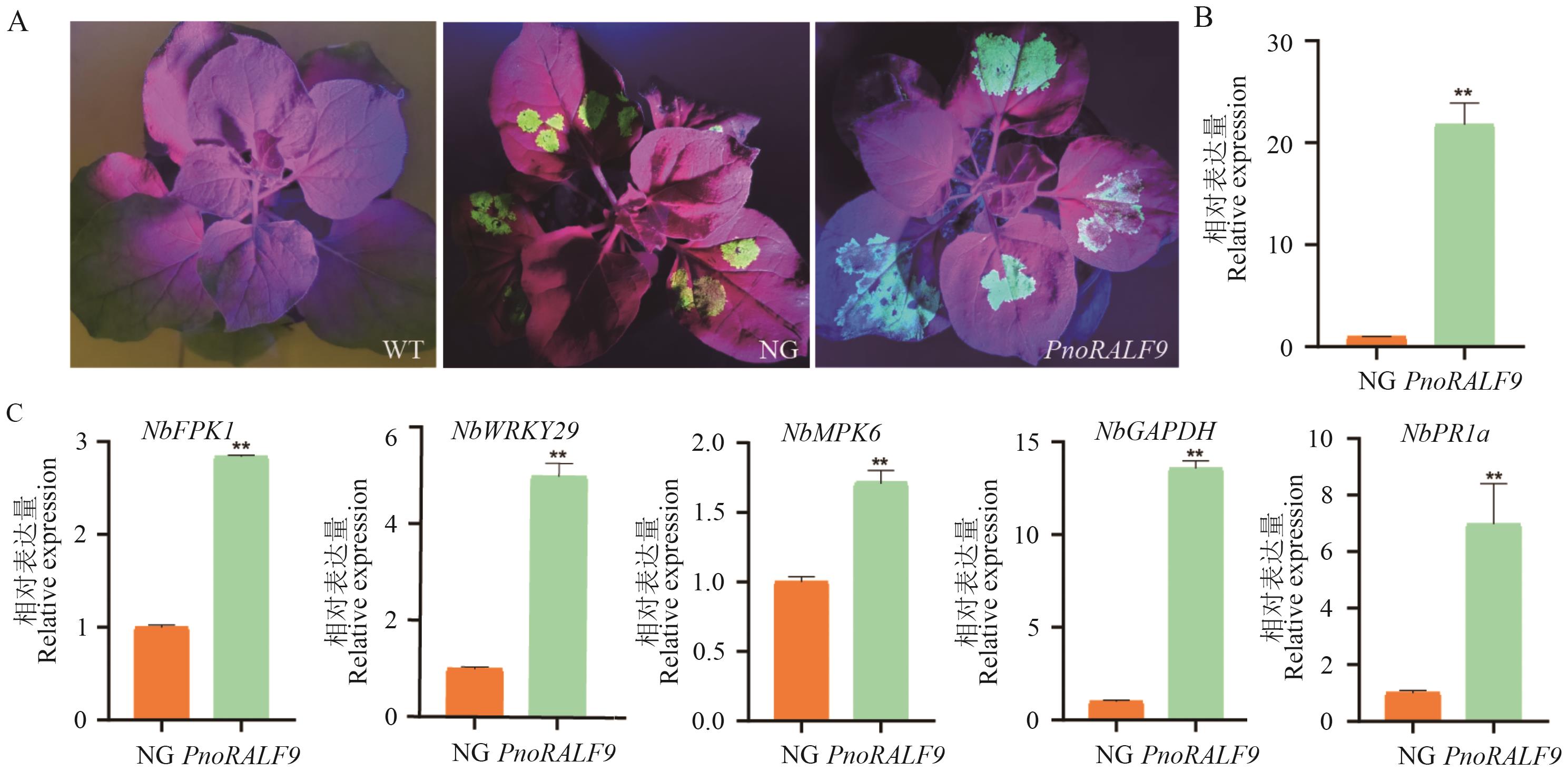
Fig. 12 Analysis of phenotypic and gene expression in N. benthamiana following transient transformation of PnoRALF9A: Phenotype of GFP expression in N. benthamiana following transient expression of PnoRALF9. B: Analysis of PnoRALF9 expression after transient transformation. C: Expression analysis of immune-related genes in N. benthamiana
| [1] | 李满桥, 梁绮文, 闫静, 等. 三七遗传改良的研究进展 [J]. 中草药, 2022, 53(10): 3241-3250. |
| Li MQ, Liang QW, Yan J, et al. Research progress on genetic improvement of Panax notoginseng [J]. Chin Tradit Herb Drugs, 2022, 53(10): 3241-3250. | |
| [2] | 谭勇, 崔尹赡, 季秀玲, 等. 三七连作的根际、根内微生物变化与生态学研究进展 [J]. 中草药, 2017, 48(2): 391-399. |
| Tan Y, Cui YS, Ji XL, et al. Research progress in microorganism changes of rhizospheric soil and root endogenous and ecology during continuous cropping of Panax notoginseng [J]. Chin Tradit Herb Drugs, 2017, 48(2): 391-399. | |
| [3] | Li JB, Ai MT, Hou JE, et al. Plant-pathogen interaction with root rot of Panax notoginseng as a model: Insight into pathogen pathogenesis, plant defence response and biological control [J]. Mol Plant Pathol, 2024, 25(2): e13427. |
| [4] | 张晋豪, 杨俊, 姬广海, 等. 云南三七根腐病病原细菌的鉴定 [J]. 南方农业学报, 2020, 51(3): 586-592. |
| Zhang JH, Yang J, Ji GH, et al. Bacterial pathogen identification of Panax notoginseng root rot in Yunnan Province [J]. J South Agric, 2020, 51(3): 586-592. | |
| [5] | Pearce G, Moura DS, Stratmann J, et al. RALF, a 5-kDa ubiquitous polypeptide in plants, arrests root growth and development [J]. Proc Natl Acad Sci U S A, 2001, 98(22): 12843-12847. |
| [6] | Blackburn MR, Haruta M, Moura DS. Twenty years of progress in physiological and biochemical investigation of RALF peptides [J]. Plant Physiol, 2020, 182(4): 1657-1666. |
| [7] | Abarca A, Franck CM, Zipfel C. Family-wide evaluation of rapid alkalinization factor peptides [J]. Plant Physiol, 2021, 187(2): 996-1010. |
| [8] | Campbell L, Turner SR. A comprehensive analysis of RALF proteins in green plants suggests there are two distinct functional groups [J]. Front Plant Sci, 2017, 8: 37. |
| [9] | Xiao Y, Stegmann M, Han ZF, et al. Mechanisms of RALF peptide perception by a heterotypic receptor complex [J]. Nature, 2019, 572(7768): 270-274. |
| [10] | 蔺欢, 王俊娟, 孙振婷, 等. 植物小分子肽的研究进展 [J]. 西北植物学报, 2021, 41(1): 168-180. |
| Lin H, Wang JJ, Sun ZT, et al. The research progress of plant small molecular peptides [J]. Acta Bot Boreali Occidentalia Sin, 2021, 41(1): 168-180. | |
| [11] | 何语涵. RALF22的抗核盘菌免疫功能及机制分析 [D]. 杭州: 浙江大学, 2022. |
| He YH. Functions and molecular mechanisms of RALF22 in plant immunity against Sclerotinia sclerotiorum [D]. Hangzhou: Zhejiang University, 2022. | |
| [12] | Zhang R, Shi PT, Zhou M, et al. Rapid alkalinization factor: function, regulation, and potential applications in agriculture [J]. Stress Biol, 2023, 3(1): 16. |
| [13] | Xue BP, Liang ZC, Liu Y, et al. Genome-wide identification of the RALF gene family and expression pattern analysis in Zea mays L. under abiotic stresses [J]. Plants, 2024, 13(20): 2883. |
| [14] | Moussu S, Lee HK, Haas KT, et al. Plant cell wall patterning and expansion mediated by protein-peptide-polysaccharide interaction [J]. Science, 2023, 382(6671): 719-725. |
| [15] | 强晓楠, 李鑫, 陈佳, 等. 拟南芥RALF多肽家族的功能多样性初步分析 [J]. 生物技术通报, 2019, 35(1): 2-10. |
| Qiang XN, Li X, Chen J, et al. Preliminary analysis of functional diversity of RALF peptide family in Arabidopsis thaliana [J]. Biotechnol Bull, 2019, 35(1): 2-10. | |
| [16] | Liu LN, Liu X, Bai ZK, et al. Small but powerful: RALF peptides in plant adaptive and developmental responses [J]. Plant Sci, 2024, 343: 112085. |
| [17] | Galindo-Trigo S, Gray JE, Smith LM. Conserved roles of CrRLK1L receptor-like kinases in cell expansion and reproduction from algae to angiosperms [J]. Front Plant Sci, 2016, 7: 1269. |
| [18] | Gonneau M, Desprez T, Martin M, et al. Receptor kinase THESEUS1 is a rapid alkalinization factor 34 receptor in Arabidopsis [J]. Curr Biol, 2018, 28(15): 2452-2458.e4. |
| [19] | Stegmann M, Monaghan J, Smakowska-Luzan E, et al. The receptor kinase FER is a RALF-regulated scaffold controlling plant immune signaling [J]. Science, 2017, 355(6322): 287-289. |
| [20] | Rui PH, Jia ZX, Fang XX, et al. A plant viral effector subverts FER-RALF1 module-mediated intracellular immunity [J]. Plant Biotechnol J, 2025, 23(7): 2734-2751. |
| [21] | Yang ZJ, Liu GZ, Zhang GH, et al. The chromosome-scale high-quality genome assembly of Panax notoginseng provides insight into dencichine biosynthesis [J]. Plant Biotechnol J, 2021, 19(5): 869-871. |
| [22] | Lu SN, Wang JY, Chitsaz F, et al. CDD/SPARCLE: the conserved domain database in 2020 [J]. Nucleic Acids Res, 2020, 48(D1): D265-D268. |
| [23] | Wilkins MR, Gasteiger E, Bairoch A, et al. Protein identification and analysis tools in the ExPASy server [M]//2-D Proteome Analysis Protocols. New Jersey: Humana Press, 2003: 531-552. |
| [24] | Tamura K, Stecher G, Kumar S. MEGA11: molecular evolutionary genetics analysis version 11 [J]. Mol Biol Evol, 2021, 38(7): 3022-3027. |
| [25] | Lescot M, Déhais P, Thijs G, et al. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences [J]. Nucleic Acids Res, 2002, 30(1): 325-327. |
| [26] | 梁童瑶, 麻鹏达, 夏鹏国, 等. pEAQ载体介导的烟草叶片瞬时遗传转化体系[C]. 陕西安康: 陕西省植物学会, 2016. |
| Liang CY, Ma PD, Xia PG, et al. Transient genetic transformation system of tobacco leaves mediated by pEAQ vector [C]. Ankang, Shanxi: Shanxi Prov Bot Soc, 2016. | |
| [27] | Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT Method [J]. Methods (San Diego, Calif.), 2001, 25(4): 402-408. |
| [28] | Chen XY, Chen J, Xu F, et al. RALF-FER, a master ligand-receptor pair in plant health [J]. Crop Health, 2025, 3(1): 4. |
| [29] | Cao J, Shi F. Evolution of the RALF gene family in plants: gene duplication and selection patterns [J]. Evol Bioinform Online, 2012, 8: EBO.S9652. |
| [30] | Waadt R, Seller CA, Hsu PK, et al. Plant hormone regulation of abiotic stress responses [J]. Nat Rev Mol Cell Biol, 2022, 23(10): 680-694. |
| [31] | Xia JZ, Wang Y, Zhang TT, et al. Genome-wide identification, expression profiling, and functional analysis of ammonium transporter 2 (AMT2) gene family in cassava (Manihot esculenta Crantz) [J]. Front Genet, 2023, 14: 1145735. |
| [32] | Zhao CZ, Jiang W, Zayed O, et al. The LRXs-RALFs-FER module controls plant growth and salt stress responses by modulating multiple plant hormones [J]. Natl Sci Rev, 2021, 8(1): nwaa149. |
| [33] | Iqbal N, Khan NA, Ferrante A, et al. Ethylene role in plant growth, development and senescence: interaction with other phytohormones [J]. Front Plant Sci, 2017, 8: 475. |
| [34] | Hou SG, Liu ZY, Shen HX, et al. Damage-associated molecular pattern-triggered immunity in plants [J]. Front Plant Sci, 2019, 10: 646. |
| [35] | 胡海琳, 徐黎, 李晓旭, 等. 小肽激素调控植物生长发育及逆境生理研究进展 [J]. 生物技术通报, 2023, 39(7): 13-25. |
| Hu HL, Xu L, Li XX, et al. Advances in the regulation of plant growth, development and stress physiology by small peptide hormones [J]. Biotechnol Bull, 2023, 39(7): 13-25. | |
| [36] | Zhao CZ, Zayed O, Yu ZP, et al. Leucine-rich repeat extensin proteins regulate plant salt tolerance in Arabidopsis [J]. Proc Natl Acad Sci U S A, 2018, 115(51): 13123-13128. |
| [37] | He YH, Zhang ZR, Xu YP, et al. Genome-wide identification of rapid alkalinization factor family in Brassica napus and functional analysis of BnRALF10 in immunity to Sclerotinia sclerotiorum [J]. Front Plant Sci, 2022, 13: 877404. |
| [38] | Guo HQ, Nolan TM, Song GY, et al. FERONIA receptor kinase contributes to plant immunity by suppressing jasmonic acid signaling in Arabidopsis thaliana [J]. Curr Biol, 2018, 28(20): 3316-3324.e6. |
| [39] | Li MQ, Che XL, Liang QW, et al. Genome-wide identification and characterization of WRKYs family involved in responses to Cylindrocarpon destructans in Panax notoginseng [J]. BMC Genomics, 2025, 26(1): 104. |
| [40] | Sun H, Li MQ, Liu XY, et al. Isolation and functional characterization of a pathogenesis-related protein 4 gene from Panax notoginseng [J]. Australas Plant Pathol, 2024, 53(6): 539-550. |
| [41] | Tang XY, Frederick RD, Zhou JM, et al. Initiation of plant disease resistance by physical interaction of AvrPto and pto kinase [J]. Science, 1996, 274(5295): 2060-2063. |
| [42] | Islam MM, El-Sappah AH, Ali HM, et al. Pathogenesis-related proteins (PRs) countering environmental stress in plants: a review [J]. S Afr N J Bot, 2023, 160: 414-427. |
| [1] | SU Yan-zhu, LI Da, ZHANG Ai-ai, LIU Yong-guang, ZHANG Xiu-rong, XUE Qi-qin. Identification and Expression Analysis of CAD Gene Family in Soybean(Glycine max (L.) Merr.) [J]. Biotechnology Bulletin, 2026, 42(4): 101-113. |
| [2] | LIU Qing-yuan, WU Hong-qi, CHEN Xiu-e, CHEN Jian, JIANG Yuan-ze, HE Yan-zi, YU Qi-wei, LIU Ren-xiang. Function of Transcription Factor NtMYB96a in Regulating the Tolerance of Tobacco to Drought [J]. Biotechnology Bulletin, 2026, 42(4): 239-250. |
| [3] | CHEN Deng-ke, LAN Gang, XIA Zhi, HOU Bao-guo, YANG Liu-liu, CAO Cai-rong, LI Peng-bo, WU Cui-cui. Identification of ZF-HD Gene Family in Arachis hypogaea and Analysis in Response to Abiotic Stress [J]. Biotechnology Bulletin, 2026, 42(4): 114-128. |
| [4] | REN Yun-er, WU Guo-qiang, CHENG Bin, WEI Ming. Genome-wide Identification of the BvATGs Genes Family in Sugar Beet (Beta vulgaris L.) and Analysis of Their Expression Pattern under Salt Stress [J]. Biotechnology Bulletin, 2026, 42(1): 184-197. |
| [5] | YANG Yue-qin, XING Ying, ZHONG Zi-he, TIAN Wei-jun, YANG Xue-qing, WANG Jian-xu. Expression and Functional Analysis of OsMATE34 in Rice under Mercury Stress [J]. Biotechnology Bulletin, 2026, 42(1): 86-94. |
| [6] | LI Jian-bin, HOU Jia-e, LI Lei-lin, AI Ming-tao, LIU Tian-tai, CUI Xiu-ming, YANG Qian. Genome-wide Identification of Panax notoginseng Lipoxygenases Coupled in Response to Methyl-jasmonate and Wounding [J]. Biotechnology Bulletin, 2026, 42(1): 218-229. |
| [7] | CHEN Jing-huan, FANG Guo-nan, ZHU Wen-hao, YE Guang-ji, SU Wang, HE Miao-miao, YANG Sheng-long, ZHOU Yun. Starch Characterization and Related Gene Expression Analysis of Potato Germplasm Resources [J]. Biotechnology Bulletin, 2026, 42(1): 170-183. |
| [8] | LI Shan, MA Deng-hui, MA Hong-yi, YAO Wen-kong, YIN Xiao. Identification and Expression Analysis of SKP1 Gene Family in Grapevine (Vitis vinifera L.) [J]. Biotechnology Bulletin, 2025, 41(9): 147-158. |
| [9] | GONG Hui-ling, XING Yu-jie, MA Jun-xian, CAI Xia, FENG Zai-ping. Identification of Laccase (LAC) Gene Family in Potato (Solanum tuberosum L.) and Its Expression Analysis under Salt Stresses [J]. Biotechnology Bulletin, 2025, 41(9): 82-93. |
| [10] | CHENG Ting-ting, LIU Jun, WANG Li-li, LIAN Cong-long, WEI Wen-jun, GUO Hui, WU Yao-lin, YANG Jing-fan, LAN Jin-xu, CHEN Sui-qing. Genome-wide Identification of the Chalcone Isomerase Gene Family in Eucommia ulmoides and Analysis of Their Expression Patterns [J]. Biotechnology Bulletin, 2025, 41(9): 242-255. |
| [11] | BAI Yu-guo, LI Wan-di, LIANG Jian-ping, SHI Zhi-yong, LU Geng-long, LIU Hong-jun, NIU Jing-ping. Growth-promoting Mechanism of Trichoderma harzianum T9131 on Astragalus membranaceus Seedlings [J]. Biotechnology Bulletin, 2025, 41(8): 175-185. |
| [12] | REN Rui-bin, SI Er-jing, WAN Guang-you, WANG Jun-cheng, YAO Li-rong, ZHANG Hong, MA Xiao-le, LI Bao-chun, WANG Hua-jun, MENG Ya-xiong. Identification and Expression Analysis of GH17 Gene Family of Pyrenophora graminea [J]. Biotechnology Bulletin, 2025, 41(8): 146-154. |
| [13] | ZENG Dan, HUANG Yuan, WANG Jian, ZHANG Yan, LIU Qing-xia, GU Rong-hui, SUN Qing-wen, CHEN Hong-yu. Genome-wide Identification and Expression Analysis of bZIP Transcription Factor Family in Dendrobium officinale [J]. Biotechnology Bulletin, 2025, 41(8): 197-210. |
| [14] | HUA Wen-ping, LIU Fei, HAO Jia-xin, CHEN Chen. Identification and Expression Patterns Analysis of ADH Gene Family in Salvia miltiorrhiza [J]. Biotechnology Bulletin, 2025, 41(8): 211-219. |
| [15] | LI Kai-yue, DENG Xiao-xia, YIN Yuan, DU Ya-tong, XU Yuan-jing, WANG Jing-hong, YU Song, LIN Ji-xiang. Identification of LEA Gene Family and Analysis on Its Response to Aluminum Stress in Ricinus communis L. [J]. Biotechnology Bulletin, 2025, 41(7): 128-138. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||