








Biotechnology Bulletin ›› 2021, Vol. 37 ›› Issue (8): 111-120.doi: 10.13560/j.cnki.biotech.bull.1985.2020-1591
Previous Articles Next Articles
LIU Yuan-yuan( ), YANG Dong-jie, ZUO Dong-yun, CHENG Hai-liang, ZHANG You-ping, LV Li-min, WANG Qiao-lian, SONG Guo-li(
), YANG Dong-jie, ZUO Dong-yun, CHENG Hai-liang, ZHANG You-ping, LV Li-min, WANG Qiao-lian, SONG Guo-li( )
)
Received:2020-12-31
Online:2021-08-26
Published:2021-09-10
Contact:
SONG Guo-li
E-mail:yuanyl4956@163.com;sglzms@163.com
LIU Yuan-yuan, YANG Dong-jie, ZUO Dong-yun, CHENG Hai-liang, ZHANG You-ping, LV Li-min, WANG Qiao-lian, SONG Guo-li. Cloning and Functional Verification of GhD6PKL2 from Gossypium hirsutum[J]. Biotechnology Bulletin, 2021, 37(8): 111-120.
| 引物用途 Function of the primer | 引物名称 Name of the primer | 引物序列 Sequence of the primer(5'-3') |
|---|---|---|
| pRI101载体通用引物 The primers in pRI101 vector | 35ScexuPF | CCTTCGCAAGACCCTTCCTC |
| pRI101- AN- cexuRV | CAGGAAACAGCTATGAC | |
| cDNA扩增& 过表达载体构建引物 The primers for cDNA amplification and over-expression vector’s construction | D6PKPFCZ(Sal I) | TTGATACATATGCCCGTCGACATGGAGCCGTTTCTCGACGACT |
| D6PKPR1CZ(EcoR I) | GAATTCCTAATGATGATGATGATGATGATAATACTCTACTGGGGTCTCT | |
| D6PKPR2CZ(EcoR I) | AGAGTTGTTGATTCAGAATTCCTAATGATGATGATGATGATG | |
| 定量引物 The primers for quantitative real-time PCR | A07GD6PKPF1 | TGAGTTCCCTAAAGAACCCATTGT |
| A07GD6PKPR1 | GGTGGTGCTTGATGGCTGAT | |
| cDNA检测& 内参引物 The primers for cDNA quality’s verification and reference | AtactinePF | TGCTATTCTGCGTTTGGACCTTG |
| AtactinePR | ATCCCTTACGATTTCACGCTCTG | |
| Histone3PF1 | CCGTAAATCTGCCCCAACCA | |
| Histone3PR1 | GACCCACAAGGTATGCCTCTGC | |
| GFP& Linker片段扩增引物 The primers for GFP and Linker fragments amplification | GFPlinkPF1 | TGGCTCTGGCGGTGGCGGATCGATGGGTAAAGGAGAAGAACTTT |
| GFPPR1(EcoR I) | AGAGTTGTTGATTCAGAATTCTCATTTGTATAGTTCATCCATG | |
| LinkPF2(BamH I) | GGATCCGGTGGAGGCGGTTCAGGCGGAGGTGGCTCTGGCGGTGGCGGATCG | |
| 亚细胞定位中基因扩增引物 The primers for genes in subcellular location | AD6PKPF1(Sal I) | TTGATACATATGCCCGTCGACATGGAGCCGTTTCTCGACGACT |
| AD6PKPR1(BamH I) | CTGAACCGCCTCCACCGGATCCATAATACTCTACTGGGGTCTCT | |
| 诱饵载体引物 The primers for bait vector construction | A0D6PKPF1GB(Sal) | ATGCGGCCGCTGCAGGTCGACATGGAGCCGTTTCTCGACGACT |
| A0D6PKPR1GB(EcoR) | ATGGCCATGGAGGCCGAATTCATAATACTCTACTGGGGTCTCT |
Table1 Primers used in this study
| 引物用途 Function of the primer | 引物名称 Name of the primer | 引物序列 Sequence of the primer(5'-3') |
|---|---|---|
| pRI101载体通用引物 The primers in pRI101 vector | 35ScexuPF | CCTTCGCAAGACCCTTCCTC |
| pRI101- AN- cexuRV | CAGGAAACAGCTATGAC | |
| cDNA扩增& 过表达载体构建引物 The primers for cDNA amplification and over-expression vector’s construction | D6PKPFCZ(Sal I) | TTGATACATATGCCCGTCGACATGGAGCCGTTTCTCGACGACT |
| D6PKPR1CZ(EcoR I) | GAATTCCTAATGATGATGATGATGATGATAATACTCTACTGGGGTCTCT | |
| D6PKPR2CZ(EcoR I) | AGAGTTGTTGATTCAGAATTCCTAATGATGATGATGATGATG | |
| 定量引物 The primers for quantitative real-time PCR | A07GD6PKPF1 | TGAGTTCCCTAAAGAACCCATTGT |
| A07GD6PKPR1 | GGTGGTGCTTGATGGCTGAT | |
| cDNA检测& 内参引物 The primers for cDNA quality’s verification and reference | AtactinePF | TGCTATTCTGCGTTTGGACCTTG |
| AtactinePR | ATCCCTTACGATTTCACGCTCTG | |
| Histone3PF1 | CCGTAAATCTGCCCCAACCA | |
| Histone3PR1 | GACCCACAAGGTATGCCTCTGC | |
| GFP& Linker片段扩增引物 The primers for GFP and Linker fragments amplification | GFPlinkPF1 | TGGCTCTGGCGGTGGCGGATCGATGGGTAAAGGAGAAGAACTTT |
| GFPPR1(EcoR I) | AGAGTTGTTGATTCAGAATTCTCATTTGTATAGTTCATCCATG | |
| LinkPF2(BamH I) | GGATCCGGTGGAGGCGGTTCAGGCGGAGGTGGCTCTGGCGGTGGCGGATCG | |
| 亚细胞定位中基因扩增引物 The primers for genes in subcellular location | AD6PKPF1(Sal I) | TTGATACATATGCCCGTCGACATGGAGCCGTTTCTCGACGACT |
| AD6PKPR1(BamH I) | CTGAACCGCCTCCACCGGATCCATAATACTCTACTGGGGTCTCT | |
| 诱饵载体引物 The primers for bait vector construction | A0D6PKPF1GB(Sal) | ATGCGGCCGCTGCAGGTCGACATGGAGCCGTTTCTCGACGACT |
| A0D6PKPR1GB(EcoR) | ATGGCCATGGAGGCCGAATTCATAATACTCTACTGGGGTCTCT |

Fig.1 Analysis of the GhD6PKL2 sequence and protein sequence alignment A: The structure of the GhD6PKL2. B: Multiple alignment of amino acid sequences between GhD6PKL2 and other’s species(AtD6PKL2- 1: Arabidopsis; NtD6PKL2- 11: Nicotiana tabacum; OsD6PKL2-1: Oryza sativa; ZmHPK-1: Zea mays). The number Ⅰ to Ⅺ indicate 11 typical active sites of STPK family
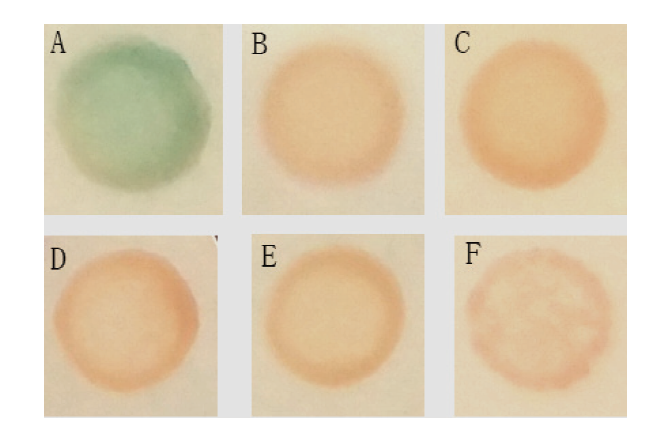
Fig.5 Detection of transcriptional activity of GhD6PKL2 A: Positive control: pGBKT7-53 and pGADT7-T, growth on SD-trp-leu/X-α-gal;B: nagetive control: pGBKT7-Lam and pGADT7-T growth on SD-trp-leu/X-α-gal; C: blank control: PGADT7-T growth on SD-trp; D: pGBKT7-GhD6PKL2 growth on SD-trp; E: pGBKT7-GhD6PKL2 growth on SD-trp/X-α-gal; F: pGBKT7-GhD6PKL2 growth on SD-trp/X-α-gal/AbA

Fig.7 Phynotype of Arabidopsis trichome between GhD6PKL2 trans-genic plants and wide type A: Arabidopsis trichome observation between transgenic plants and wide type;B: average arabidopsis trichome in transgenic plants and wide type. ***indicate that there is a significant difference at P= 0.001. The same below
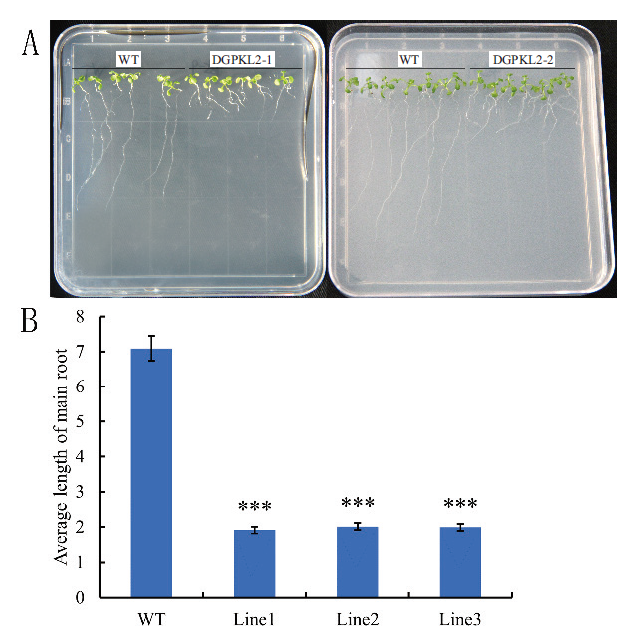
Fig.8 Phenotype of Arabidopsis main root between GhD6PKL2 trans-genic plants and wide type A: Arabidopsis root growths of GhD6PKL2 transgenic plants and WT type; B: Main root length in GhD6PKL2 transgenic plants and WT type.
| [1] |
Pitzschke A, Schikora A, Hirt H. MAPK cascade signalling networks in plant defence[J]. Curr Opin Plant Biol, 2009, 12(4):421-426.
doi: 10.1016/j.pbi.2009.06.008 pmid: 19608449 |
| [2] |
Goldberg J, Huang HB, Kwon YG, et al. Three- dimensional structure of the catalytic subunit of protein serine/threonine phosphatase- 1[J]. Nature, 1995, 376(6543):745-753.
doi: 10.1038/376745a0 URL |
| [3] |
Hanks SK, Quinn AM, Hunter T. The protein-kinase family:conserved features and deduced phylogeny of the catalytic domains[J]. Science, 1988, 241(4861):42-52.
pmid: 3291115 |
| [4] | Rademacher EH, Offringa R. Evolutionary adaptations of plant AGC kinases:from light signaling to cell polarity regulation[J]. Frontiers in Plant Science, 2012, 3(250). |
| [5] |
Huse M, Kuriyan J. The conformational plasticity of protein kinases[J]. Cell, 2002, 109(3):275-282.
doi: 10.1016/S0092-8674(02)00741-9 URL |
| [6] |
Hanks SK, Hunter T. The eukaryotic protein kinase superfamily:kinase(catalytic)domain structure and classification[J]. The FASEB Journal, 1995, 9(8):576-596.
doi: 10.1096/fsb2.v9.8 URL |
| [7] |
Bogre L, Okresz L, Henriques R, et al. Growth signalling pathways in Arabidopsis and the AGC protein kinases[J]. Trends in Plant Science, 2003, 8(9):424-431.
doi: 10.1016/S1360-1385(03)00188-2 URL |
| [8] | Zourelidou M, Muller I, Willige BC, et al. The polarly localized D6 PROTEIN KINASE is required for efficient auxin transport in Arabidopsis thaliana[J]. Develop, 2009, 136(4):627-636. |
| [9] |
Galvan-Ampudia CS, Offringa R. Plant evolution:AGC kinases tell the auxin tale[J]. Trends Plant Sci, 2007, 12(12):541-547.
doi: 10.1016/j.tplants.2007.10.004 URL |
| [10] |
Sakai T, Kagawa T, Kasahara M, et al. Arabidopsis nph1 and npl1:blue light receptors that mediate both phototropism and chloroplast relocation[J]. PNAS, 2001, 98(12):6969-6974.
pmid: 11371609 |
| [11] | Briggs WR, Huala E. Blue- light photoreceptors in higher plants[J]. Annu Rev Cell Devl Biol, 1999, 15:33-62. |
| [12] |
Santner AA, Watson JC. The WAG1 and WAG2 protein kinases negatively regulate root waving in Arabidopsis[J]. The Plant Journal, 2010, 45(5):752-764.
doi: 10.1111/tpj.2006.45.issue-5 URL |
| [13] |
Haga K, Hayashi KI, Sakai T. PINOID AGC kinases are necessary for phytochrome- mediated enhancement of hypocotyl phototropism in Arabidopsis[J]. Plant Physiol, 2014, 166(3):1535-1545.
doi: 10.1104/pp.114.244434 pmid: 25281709 |
| [14] |
Zhang Y, He JM, Mccormick S. Two Arabidopsis AGC kinases are critical for the polarized growth of pollen tubes[J]. The Plant Journal, 2010, 58(3):474-484.
doi: 10.1111/tpj.2009.58.issue-3 URL |
| [15] |
Enugutti B, Kirchhelle C, Oelschner M, et al. Regulation of planar growth by the Arabidopsis AGC protein kinase UNICORN[J]. Proc Natl Acad Sci USA, 2012, 109(37):15060-15065.
doi: 10.1073/pnas.1205089109 URL |
| [16] |
Haga K, Frank L, Kimura T, et al. Roles of AGC VIII kinases in the hypocotyl phototropism of Arabidopsis seedlings[J]. Plant Cell Physiology, 2018, 59(5):1060-1071.
doi: 10.1093/pcp/pcy048 URL |
| [17] |
Devarenne TP, Ekengren SK, Pedley KF, et al. Adi3 is a PDK1- interacting AGC kinase that negatively regulates plant cell death[J]. The EMBO Journal, 2006, 25(1):255-265.
doi: 10.1038/sj.emboj.7600910 URL |
| [18] |
Hammond RW, Zhao Y. Modification of tobacco plant development by sense and antisense expression of the tomato viroid- induced AGC VIIIa protein kinase PKV suggests involvement in gibberellin signaling[J]. BMC Plant Biology, 2009, 9:108.
doi: 10.1186/1471-2229-9-108 URL |
| [19] |
Zhang YY, et al. AGC protein kinase AGC1- 4 mediates seed size in Arabidopsis[J]. Plant Cell Rep, 2020, 39(6):825-837.
doi: 10.1007/s00299-020-02533-z URL |
| [20] |
Wan Q, Guan XY, Yang NN, et al. Small interfering RNAs from bidirectional transcripts of GhMML3_A12 regulate cotton fiber development[J]. New Phytologist, 2016, 210(4):1298-1310.
doi: 10.1111/nph.2016.210.issue-4 URL |
| [21] |
Serna L, Martin C. Trichomes:different regulatory networks lead to convergent structures[J]. Trends Plant Sci, 2006, 11(6):274-280.
doi: 10.1016/j.tplants.2006.04.008 URL |
| [22] |
Lee JJ, Woodward AW, Chen ZJ. Gene expression changes and early events in cotton fibre development[J]. Annals of Botany, 2008, 100(7):1391-1401.
doi: 10.1093/aob/mcm232 URL |
| [23] |
Guan XY, Pang MX, Nah G, et al. miR828 and miR858 regulate homoeologous MYB2 gene functions in Arabidopsis trichome and cotton fibre development[J]. Nat Commun, 2014, 5:3050.
doi: 10.1038/ncomms4050 URL |
| [24] |
Wang S, Wang JW, Yu N, et al. Control of plant trichome development by a cotton fiber MYB gene[J]. The Plant Cell, 2004, 16(9):2323-2334.
doi: 10.1105/tpc.104.024844 URL |
| [25] |
Guan XY, Lee JJ, Pang MX, et al. Activation of Arabidopsis seed hair development by cotton fiber- related genes[J]. PLoS One, 2011 6(7):e21301.
doi: 10.1371/journal.pone.0021301 URL |
| [26] |
Zourelidou M, Absmanner B, Weller B, et al. Auxin efflux by PIN-FORMED proteins is activated by two different protein kinases, D6 PROTEIN KINASE and PINOID[J]. eLife, 2014, 3:e02860.
doi: 10.7554/eLife.02860 URL |
| [27] |
Willige BC, Ahlers S, Zourelidou M, et al. D6PK AGCVIII kinases are required for auxin transport and phototropic hypocotyl bending in Arabidopsis[J]. The Plant Cell, 2013, 25(5):1674-1688.
doi: 10.1105/tpc.113.111484 pmid: 23709629 |
| [28] | Willige BC, Joanne C. A current perspective on the role of AGCVIII kinases in PIN- mediated apical hook development[J]. Frontiers in Plant science, 2015, 6(767):767-773. |
| [29] |
Stanislas T, Huser A, Barbosa IC, et al. Arabidopsis D6PK is a lipid domain- dependent mediator of root epidermal planar polarity[J]. Nature Plants, 2015, 1(11):15162.
doi: 10.1038/nplants.2015.162 URL |
| [30] |
Lee BH, Weber ZT, et al. Arabidopsis protein kinase D6PKL3 is involved in the formation of distinct plasma membrane aperture domains on the pollen surface[J]. Plant Cell, 2018, 30(9):2038-2056.
doi: 10.1105/tpc.18.00442 URL |
| [31] |
Daisuke M, Takashi N, Sato KI, et al. Activation of AtMEK1, an Arabidopsis mitogen- activated protein kinase kinase, in vitro and in vivo:analysis of active mutants expressed in E. coli and generation of the active form in stress response in seedlings[J]. The Plant Journal, 2002, 29(5):637-647.
doi: 10.1046/j.0960-7412.2001.01246.x URL |
| [32] |
Simon ML, Platre MP, Marques-Bueno MM, et al. A PtdIns(4)P- driven electrostatic field controls cell membrane identity and signalling in plants[J]. Nature Plants, 2016, 2:16089.
doi: 10.1038/nplants.2016.89 URL |
| [33] |
Ek-Ramos MJ, Julian A, Nelson-Dittrich AC, et al. The tomato cell death suppressor Adi3 is restricted to the endosomal system in response to the Pseudomonas syringae effector protein AvrPto[J]. PLoS One, 2014, 9(10):e110807.
doi: 10.1371/journal.pone.0110807 URL |
| [34] |
Xiao Y, Offringa R. PDK1 regulates auxin transport and Arabidopsis vascular development through AGC1 kinase PAX[J]. Nature Plants, 2020, 6(5):1-12.
doi: 10.1038/s41477-019-0586-6 URL |
| [35] |
Zeng JY, Zhang M, Hou L, et al. Cytokinin inhibits cotton fiber initiation by disrupting PIN3a- mediated asymmetric accumulation of auxin in the ovule epidermis[J]. Journal of Experimental Botany, 2019, 70(12):3139-3151.
doi: 10.1093/jxb/erz162 URL |
| [36] | Benjamins R, Quint A, et al. The PINOID protein kinase regulates organ development in Arabidopsis by enhancing polar auxin transport[J]. Develop, 2001, 128(20):4057-4067. |
| [37] | Weller B, Zourelidou M, Frank L, et al. Dynamic PIN- FORMED auxin efflux carrier phosphorylation at the plasma membrane controls auxin efflux- dependent growth[J]. Proc Natl Acad Sci. USA, 2017, 114(5):201614380. |
| [38] |
Tan ST, Zhang XX, Kong W, et al. A lipid code-dependent phosphoswitch directs PIN- mediated auxin efflux in Arabidopsis development[J]. Nature Plants, 2020, 6(5):556-569.
doi: 10.1038/s41477-020-0648-9 URL |
| [39] |
Zhu QK, Shao YM, Ge ST, et al. A MAPK cascade downstream of IDA-HAE/ HSL2 ligand- receptor pair in lateral root emergence[J]. Nature Plants, 2019, 5:414-423.
doi: 10.1038/s41477-019-0396-x URL |
| [40] |
Marhava P, Bassukas AE, Zourelidou M, et al. A molecular rheostat adjusts auxin flux to promote root protophloem differentiation[J]. Nature, 2018, 558(7709):297-300.
doi: 10.1038/s41586-018-0186-z URL |
| [41] |
Oyama T, Shimura Y, Okada K. The IRE gene encodes a protein kinase homologue and modulates root hair growth in Arabidopsis[J]. The Plant Journal, 2002, 30(3):289-299.
doi: 10.1046/j.1365-313X.2002.01290.x URL |
| [42] |
Anthony RG, Khan S, Costa J, et al. The Arabidopsis protein kinase PTI1-2 is activated by convergent phosphatidic acid and oxidative stress signaling pathways downstream of PDK1 and OXI1[J]. Journal of Biological Chemistry, 2006, 281(49):37536-37546.
doi: 10.1074/jbc.M607341200 URL |
| [1] | WANG Yi-fan, HOU Lin-hui, CHANG Yong-chun, YANG Ya-jie, CHEN Tian, ZHAO Zhu-yue, RONG Er-hua, WU Yu-xiang. Synthesis and Character Identification of Allohexaploid Between Gossypium hirsutum and G. gossypioides [J]. Biotechnology Bulletin, 2023, 39(5): 168-176. |
| [2] | LIU Meng-meng, HAN Li-jun, LIU Bao-ling, XUE Jin-ai, LI Run-zhi. Cloning and Expression Analysis of GhSDP1 and Its Promoter in Gossypium hirsutum [J]. Biotechnology Bulletin, 2022, 38(2): 34-43. |
| [3] | WANG Ya-li, WANG Na, CHENG Hong-mei. Comparison of Methods for Rapid Determination of Cotton Ploidy by Flow Cytometry [J]. Biotechnology Bulletin, 2022, 38(12): 144-148. |
| [4] | MA Qi, LI Ji-lian, XU Shou-zhen, CHEN Hong, LIU Wen-hao, NING Xinzhu, LIN Hai. Genetic Analysis of FBA Trait in Upland Cotton with Major Gene Plus Polygenes Mixed Genetic Model [J]. Biotechnology Bulletin, 2022, 38(10): 148-158. |
| [5] | FAN Ya-peng, RUI Cun, ZHANG Yue-xin, CHEN Xiu-gui, LU Xu-ke, WANG Shuai, ZHANG Hong, XU Nan, WANG Jing, CHEN Chao, YE Wu-wei. Cloning,Expression and Preliminary Bioinformatics Analysis of the Alkaline Tolerant Gene GhZAT12 in Gossypium hirsutum [J]. Biotechnology Bulletin, 2021, 37(8): 121-130. |
| [6] | YANG Xiao-min, WANG Jun-juan, WANG De-Long, LU Xu-ke, CHEN Xiu-gui, GUO Li-xue, WANG Shuai, CHEN Chao, WANG Xiao-ge, HAN Ming-ge, YE Wu-wei. Functional Verification and Bioinformatics Analysis of Cotton GhDMT3 Gene [J]. Biotechnology Bulletin, 2019, 35(1): 11-16. |
| [7] | LIU Yan, MENG Zhi-gang, SUN Guo-qing, WANG Yuan, ZHOU Tao, GUO San-dui, ZHANG Rui. Cloning and Function Analysis of Gene GhPYR1 in Gossypium hirsutum L. [J]. Biotechnology Bulletin, 2016, 32(2): 90-99. |
| [8] | Li Xin, Wang Zhengming, Xue Wei, Chu Mingguang. Identification and Characterization of a Novel Gene, GhWRI1, Encoding an AP2-type Transcription Factor in Gossypium hirsutum [J]. Biotechnology Bulletin, 2013, 0(6): 80-86. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||