








Biotechnology Bulletin ›› 2022, Vol. 38 ›› Issue (4): 253-260.doi: 10.13560/j.cnki.biotech.bull.1985.2021-0755
Previous Articles Next Articles
NIU Xin1( ), ZHANG Ying1, WANG Mao-jun1, LIU Wen-long2, LU Fu-ping1, LI Yu1(
), ZHANG Ying1, WANG Mao-jun1, LIU Wen-long2, LU Fu-ping1, LI Yu1( )
)
Received:2021-06-10
Online:2022-04-26
Published:2022-05-06
Contact:
LI Yu
E-mail:niuxin0968@outlook.com;liyu@tust.edu.cn
NIU Xin, ZHANG Ying, WANG Mao-jun, LIU Wen-long, LU Fu-ping, LI Yu. Effects of Different Integration Sites on the Expression of Exogenous Alkaline Protease in Bacillus amyloliquefaciens[J]. Biotechnology Bulletin, 2022, 38(4): 253-260.
| 菌株/质粒Strain/Plasmid | 特性/目的Characteristics/Purpose | 来源Source |
|---|---|---|
| 菌株Strain | ||
| E.coli EC135 pM.Bam | 对质粒DNA进行甲基化修饰 Plasmid DNA methylation modifcation | 中科院微生物研究所 Institute of Microbiology,Chinese Academy of Sciences |
| E.coli JM109 | 用于敲除质粒的构建Construction of knockout vectors | 本实验室Author’s lab |
| B.amyloliquefaciens111018 | 出发菌株Parent host | 本实验室Author’s lab |
| B.amyloliquefaciens18-ΔM | mpr基因缺失mpr gene deletion | 本研究This work |
| B.amyloliquefaciens18-ΔB | bpr基因缺失bpr gene deletion | 本研究This work |
| B.amyloliquefaciens18-ΔE | epr基因缺失epr gene deletion | 本研究This work |
| B.amyloliquefacien18-ΔV | vpr基因缺失vpr gene deletion | 本研究This work |
| B.amyloliquefaciens18-ΔN | nprE基因缺失nprE gene deletion | 本研究This work |
| B.amyloliquefaciens18-ΔA | apr基因缺失apr gene deletion | 本研究This work |
| B.amyloliquefaciens18-ΔN∷aprE | 在nprE位置整合aprE Integration of aprE in nprE site | 本研究This work |
| B.amyloliquefaciens18-Δα∷aprE | 在amyE位置整合aprE Integration of aprE in amyE site | 本研究This work |
| B.amyloliquefaciens18-ΔY∷aprE | 在yaah位置整合aprEIntegration of aprE in yaah site | 本研究This work |
| 质粒Plasmid | ||
| pWH-T2 | 穿梭表达载体Shuttle expression vector | 湖北大学Hubei University |
| pLY-2 | aprE表达载体aprE expression cassette | 本实验室Author’s lab |
Table 1 Bacterial strains and plasmids used in this study
| 菌株/质粒Strain/Plasmid | 特性/目的Characteristics/Purpose | 来源Source |
|---|---|---|
| 菌株Strain | ||
| E.coli EC135 pM.Bam | 对质粒DNA进行甲基化修饰 Plasmid DNA methylation modifcation | 中科院微生物研究所 Institute of Microbiology,Chinese Academy of Sciences |
| E.coli JM109 | 用于敲除质粒的构建Construction of knockout vectors | 本实验室Author’s lab |
| B.amyloliquefaciens111018 | 出发菌株Parent host | 本实验室Author’s lab |
| B.amyloliquefaciens18-ΔM | mpr基因缺失mpr gene deletion | 本研究This work |
| B.amyloliquefaciens18-ΔB | bpr基因缺失bpr gene deletion | 本研究This work |
| B.amyloliquefaciens18-ΔE | epr基因缺失epr gene deletion | 本研究This work |
| B.amyloliquefacien18-ΔV | vpr基因缺失vpr gene deletion | 本研究This work |
| B.amyloliquefaciens18-ΔN | nprE基因缺失nprE gene deletion | 本研究This work |
| B.amyloliquefaciens18-ΔA | apr基因缺失apr gene deletion | 本研究This work |
| B.amyloliquefaciens18-ΔN∷aprE | 在nprE位置整合aprE Integration of aprE in nprE site | 本研究This work |
| B.amyloliquefaciens18-Δα∷aprE | 在amyE位置整合aprE Integration of aprE in amyE site | 本研究This work |
| B.amyloliquefaciens18-ΔY∷aprE | 在yaah位置整合aprEIntegration of aprE in yaah site | 本研究This work |
| 质粒Plasmid | ||
| pWH-T2 | 穿梭表达载体Shuttle expression vector | 湖北大学Hubei University |
| pLY-2 | aprE表达载体aprE expression cassette | 本实验室Author’s lab |
| 引物Primer | 序列Sequence(5'-3') |
|---|---|
| Up-F | TTAACGAATTCCTGCAGCCCGGGCAGAAAGACCA- TCCACAC |
| Up-R | TCATCCGCCAAAGCAGATAACGGATGATTC |
| Down-F | CAGCCCTCAGCTGACTGAAAAACCATTTATCATTG |
| Down-R | ATCCTTTGATCTTTTCTACGAGCTCCCGCTTATTAA- G ACGGC |
| djh-up-F | TAGCACACGTTTCATTGCAAATC |
| djh-up-R | CCGACTGCGCAAAAGACATAATC |
| djh-down-F | CCGACTGCGCAAAAGACATAATC |
| djh-down-R | TCTGAGCGCAGAAACGG |
| sjh-F | TAGCACACGTTTCATTGCAAATC |
| sjh-R | TCTGAGCGCAGAAACGG |
Table 2 Primers for vpr gene deletion
| 引物Primer | 序列Sequence(5'-3') |
|---|---|
| Up-F | TTAACGAATTCCTGCAGCCCGGGCAGAAAGACCA- TCCACAC |
| Up-R | TCATCCGCCAAAGCAGATAACGGATGATTC |
| Down-F | CAGCCCTCAGCTGACTGAAAAACCATTTATCATTG |
| Down-R | ATCCTTTGATCTTTTCTACGAGCTCCCGCTTATTAA- G ACGGC |
| djh-up-F | TAGCACACGTTTCATTGCAAATC |
| djh-up-R | CCGACTGCGCAAAAGACATAATC |
| djh-down-F | CCGACTGCGCAAAAGACATAATC |
| djh-down-R | TCTGAGCGCAGAAACGG |
| sjh-F | TAGCACACGTTTCATTGCAAATC |
| sjh-R | TCTGAGCGCAGAAACGG |
| 引物Primer | 序列Sequence(5'-3') |
|---|---|
| 16S-F | GGGCTACACACGTGCTACAATGG |
| 16S-R | GTATTCACCGCGGCATGCTG |
| aprE-F | GAGCACATACCCAGGTTCAACG |
| aprE -R | GTTGCCGCTTCTGCATTGAC |
Table 3 Primers for real-time fluorescent quantitative PCR
| 引物Primer | 序列Sequence(5'-3') |
|---|---|
| 16S-F | GGGCTACACACGTGCTACAATGG |
| 16S-R | GTATTCACCGCGGCATGCTG |
| aprE-F | GAGCACATACCCAGGTTCAACG |
| aprE -R | GTTGCCGCTTCTGCATTGAC |
| RT | Mass | NCBI ID | Compound name |
|---|---|---|---|
| 1 | 58458.9 | P00692 | Alpha-amylase |
| 1.1 | 218451.1 | Q8TF72 | Protein Shroom3 |
| 1.2 | 77037.6 | O18740 | Keratin,type I cytoskeletal 9 |
| 1.3 | 56182.7 | B0TWF7 | Chromosomal replication initiator protein DnaA |
| 1.4 | 11230.8 | A5CR30 | 50S ribosomal protein L21 |
| 1.5 | 151424.3 | A4VHM3 | DNA-directed RNA polymerase subunit beta |
| 1.6 | 40412.9 | QQ4FN99 | Peptide chain release factor 1 |
| 1.7 | 18115.2 | A0NDK8 | Pro-corazonin |
| 1.8 | 100713.8 | Q65CL1 | Catenin alpha-3 |
| 1.9 | 35715.8 | G8GV69 | Dioxygenase easH |
| 2 | 22703.3 | Q6AGN3 | Nucleoside triphosphate pyrophosphatase |
| 2.1 | 33273.6 | A0R5C5 | UDP-glucose 4-epimerase |
| 2.2 | 35380.5 | Q7TVA4 | D-fructose 1,6-bisphosphatase class 2 |
| 2.3 | 107560.6 | A1A5S1 | Pre-mRNA-processing factor 6 |
| 2.4 | 75310.5 | A6TEG6 | Penicillin-binding protein activator LpoA |
| 2.5 | 229710 | P33144 | Separin |
| 2.6 | 43491.7 | A5GS98 | Ferrochelatase |
| 2.7 | 13835.7 | A9BNG3 | 30S ribosomal protein S6 |
| 2.8 | 52452.5 | A1UEN7 | Cobyric acid synthase |
| 2.9 | 48850.9 | Q28RY2 | Trigger factor |
| 3 | 40629.4 | Q5ZJ37 | Rab9 effector protein with kelch motifs |
| 3.1 | 86619.2 | P49051 | S-layer protein sap |
Table 4 Analysis of mass spectrometry results
| RT | Mass | NCBI ID | Compound name |
|---|---|---|---|
| 1 | 58458.9 | P00692 | Alpha-amylase |
| 1.1 | 218451.1 | Q8TF72 | Protein Shroom3 |
| 1.2 | 77037.6 | O18740 | Keratin,type I cytoskeletal 9 |
| 1.3 | 56182.7 | B0TWF7 | Chromosomal replication initiator protein DnaA |
| 1.4 | 11230.8 | A5CR30 | 50S ribosomal protein L21 |
| 1.5 | 151424.3 | A4VHM3 | DNA-directed RNA polymerase subunit beta |
| 1.6 | 40412.9 | QQ4FN99 | Peptide chain release factor 1 |
| 1.7 | 18115.2 | A0NDK8 | Pro-corazonin |
| 1.8 | 100713.8 | Q65CL1 | Catenin alpha-3 |
| 1.9 | 35715.8 | G8GV69 | Dioxygenase easH |
| 2 | 22703.3 | Q6AGN3 | Nucleoside triphosphate pyrophosphatase |
| 2.1 | 33273.6 | A0R5C5 | UDP-glucose 4-epimerase |
| 2.2 | 35380.5 | Q7TVA4 | D-fructose 1,6-bisphosphatase class 2 |
| 2.3 | 107560.6 | A1A5S1 | Pre-mRNA-processing factor 6 |
| 2.4 | 75310.5 | A6TEG6 | Penicillin-binding protein activator LpoA |
| 2.5 | 229710 | P33144 | Separin |
| 2.6 | 43491.7 | A5GS98 | Ferrochelatase |
| 2.7 | 13835.7 | A9BNG3 | 30S ribosomal protein S6 |
| 2.8 | 52452.5 | A1UEN7 | Cobyric acid synthase |
| 2.9 | 48850.9 | Q28RY2 | Trigger factor |
| 3 | 40629.4 | Q5ZJ37 | Rab9 effector protein with kelch motifs |
| 3.1 | 86619.2 | P49051 | S-layer protein sap |
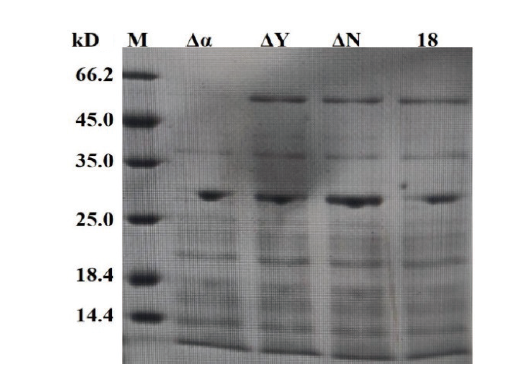
Fig.9 SDS-PAGE detection of AmyE in the integrated strains M:1 kb marker. Δα:Fermentation supernatant of strain 18-Δα∷aprE. ΔY:Fermentation supernatant of strain 18-ΔY∷aprE. ΔN:Fermentation supernatant of strain 18-ΔN∷aprE. 18:Fermentation supernatant of strain TCCC 111018
| [1] |
Ohmiya K, Tanimura S, Yashi TK, et al. Application of immobilized alkaline protease to cheese-making[J]. J Food Sci, 1979, 44(6):1584-1588.
doi: 10.1111/j.1365-2621.1979.tb09095.x URL |
| [2] |
Raval VH, Pillai S, Rawal CM, et al. Biochemical and structural characterization of a detergent-stable serine alkaline protease from seawater haloalkaliphilic bacteria[J]. Process Biochem, 2014, 49(6):955-962.
doi: 10.1016/j.procbio.2014.03.014 URL |
| [3] |
Mahmoudian M. Biocatalytic production of chiral pharmaceutical intermediates[J]. Biocatal Biotransformation, 2000, 18(2):105-118.
doi: 10.3109/10242420009015240 URL |
| [4] | Zhou C, Qin HL, Chen XJ, et al. A novel alkaline protease from alkaliphilic Idiomarina sp. C9-1 with potential application for eco-friendly enzymatic dehairing in the leather industry[J]. Sci Rep, 2018, 8(1):1-18. |
| [5] |
Omrane Benmrad M, Moujehed E, Ben Elhoul M, et al. A novel organic solvent- and detergent-stable serine alkaline protease from Trametes cingulata strain CTM10101[J]. Int J Biol Macromol, 2016, 91:961-972.
doi: 10.1016/j.ijbiomac.2016.06.025 pmid: 27296442 |
| [6] |
Baweja M, Tiwari R, Singh PK, et al. An alkaline protease from Bacillus pumilus MP 27:functional analysis of its binding model toward its applications as detergent additive[J]. Front Microbiol, 2016, 7:1195.
doi: 10.3389/fmicb.2016.01195 pmid: 27536284 |
| [7] |
Sauer C, Syvertsson S, Bohorquez LC, et al. Effect of genome position on heterologous gene expression in Bacillus subtilis:an unbiased analysis[J]. ACS Synth Biol, 2016, 5(9):942-947.
doi: 10.1021/acssynbio.6b00065 URL |
| [8] |
Bryant JA, Sellars LE, Busby SJ, et al. Chromosome position effects on gene expression in Escherichia coli K-12[J]. Nucleic Acids Res, 2014, 42(18):11383-11392.
doi: 10.1093/nar/gku828 pmid: 25209233 |
| [9] |
Kurat CF, Yeeles JTP, Patel H, et al. Chromatin controls DNA replication origin selection, lagging-strand synjournal, and replication fork rates[J]. Mol Cell, 2017, 65(1):117-130.
doi: 10.1016/j.molcel.2016.11.016 URL |
| [10] |
Couturier E, Rocha EPC. Replication-associated gene dosage effects shape the genomes of fast-growing bacteria but only for transcription and translation genes[J]. Mol Microbiol, 2006, 59(5):1506-1518.
pmid: 16468991 |
| [11] |
Zhou C, Zhou H, Li D, et al. Optimized expression and enhanced production of alkaline protease by genetically modified Bacillus licheniformis 2709[J]. Microb Cell Fact, 2020, 19(1):45.
doi: 10.1186/s12934-020-01307-2 URL |
| [12] |
Tsirigotaki A, De Geyter J, Šoštaric N, et al. Protein export through the bacterial Sec pathway[J]. Nat Rev Microbiol, 2017, 15(1):21-36.
doi: 10.1038/nrmicro.2016.161 pmid: 27890920 |
| [13] |
Mulder KCL, Bandola J, Schumann W. Construction of an artificial secYEG operon allowing high level secretion of α-amylase[J]. Protein Expr Purif, 2013, 89(1):92-96.
doi: 10.1016/j.pep.2013.02.008 URL |
| [14] |
Ren GH, Cao LC, Kong W, et al. Efficient secretion of the β-galactosidase Bgal1-3 via both tat-dependent and tat-independent pathways in Bacillus subtilis[J]. J Agric Food Chem, 2016, 64(28):5708-5716.
doi: 10.1021/acs.jafc.6b01735 URL |
| [15] |
Watzlawick H, Altenbuchner J. Multiple integration of the gene ganA into the Bacillus subtilis chromosome for enhanced β-galactosidase production using the CRISPR/Cas9 system[J]. AMB Express, 2019, 9(1):158.
doi: 10.1186/s13568-019-0884-4 pmid: 31571017 |
| [16] | Zhang G, Wang W, Deng A, et al. A mimicking-of-DNA-methylation-patterns pipeline for overcoming the restriction barrier of bacteria[J]. PLoS Genet, 2012, 8(9):e1002987. |
| [17] |
McKenney PT, Driks A, Eichenberger P. The Bacillus subtilis endospore:assembly and functions of the multilayered coat[J]. Nat Rev Microbiol, 2013, 11(1):33-44.
doi: 10.1038/nrmicro2921 pmid: 23202530 |
| [18] |
Zhang J, Xu X, Li X, et al. Reducing the cell Lysis to enhance yield of acid-stable alpha amylase by deletion of multiple peptidoglycan hydrolase-related genes in Bacillus amyloliquefaciens[J]. Int J Biol Macromol, 2021, 167:777-786.
doi: 10.1016/j.ijbiomac.2020.11.193 URL |
| [19] |
Ling Lin Fu, Zi Rong Xu, Wei Fen Li, et al. Protein secretion pathways in Bacillus subtilis:implication for optimization of heterologous protein secretion[J]. Biotechnol Adv, 2007, 25(1):1-12.
pmid: 16997527 |
| [20] |
Zhang X, Xu ZY, Liu S, et al. Improving the production of salt-tolerant glutaminase by integrating multiple copies of mglu into the protease and 16S rDNA genes of Bacillus subtilis 168[J]. Molecules, 2019, 24(3):592.
doi: 10.3390/molecules24030592 URL |
| [21] | Mo QS, Tian Y, Zhang HT, et al. Using 16S rDNA as target site for homologous recombination to improve the alkaline protease production of Bacillus alcalophilus[J]. Adv Mater Res, 2014, 886:349-354. |
| [22] |
Wang H, Zhang X, Qiu J, et al. Development of Bacillus amyloliquefaciens as a high-level recombinant protein expression system[J]. J Ind Microbiol Biotechnol, 2019, 46(1):113-123.
doi: 10.1007/s10295-018-2089-2 URL |
| [23] |
Kawamura F, Doi RH. Construction of a Bacillus subtilis double mutant deficient in extracellular alkaline and neutral proteases[J]. J Bacteriol, 1984, 160(1): 442-444.
doi: 10.1128/jb.160.1.442-444.1984 pmid: 6434524 |
| [24] |
Nakamura A, Toyama N, Kitamura A, et al. Use of a triple protease-deficient mutant of Bacillus subtilis as a host for secretion of a B. subtilis cellulase and TEM beta-lactamase[J]. Agric Biol Chem, 1991, 55(9):2367-2374.
pmid: 1368741 |
| [25] |
Abe S, Yasumura A, Tanaka T. Regulation of Bacillus subtilis aprE expression by glnA through inhibition of scoC and σ D -dependent degR expression[J]. J Bacteriol, 2009, 191(9):3050-3058.
doi: 10.1128/JB.00049-09 URL |
| [26] |
Kada S, Ishikawa A, Ohshima Y, et al. Alkaline serine protease AprE plays an essential role in poly-γ-glutamate production during natto fermentation[J]. Biosci Biotechnol Biochem, 2013, 77(4):802-809.
doi: 10.1271/bbb.120965 URL |
| [27] |
Gupta R, Gigras P, Mohapatra H, et al. Microbial α-amylases:a biotechnological perspective[J]. Process Biochem, 2003, 38(11):1599-1616.
doi: 10.1016/S0032-9592(03)00053-0 URL |
| [28] |
Wang P, Wang PL, Tian J, et al. A new strategy to express the extracellular α-amylase from Pyrococcus furiosus in Bacillus amyloliquefaciens[J]. Sci Rep, 2016, 6(1):1-10.
doi: 10.1038/s41598-016-0001-8 URL |
| [29] |
Reuβ DR, Altenbuchner J, Mäder U, et al. Large-scale reduction of the Bacillus subtilis genome:consequences for the transcriptional network, resource allocation, and metabolism[J]. Genome Res, 2017, 27(2):289-299.
doi: 10.1101/gr.215293.116 URL |
| [1] | WANG Shuai, FENG Yu-mei, BAI Miao, DU Wei-jun, YUE Ai-qin. Functional Analysis of Soybean Gene GmHMGR Responding to Exogenous Hormones and Abiotic Stresses [J]. Biotechnology Bulletin, 2023, 39(7): 131-142. |
| [2] | MA Yu-qian, SUN Dong-hui, YUE Hao-feng, XIN Jia-yu, LIU Ning, CAO Zhi-yan. Identification, Heterologous Expression and Functional Analysis of a GH61 Family Glycoside Hydrolase from Setosphaeria turcica with the Assisting Function in Degrading Cellulose [J]. Biotechnology Bulletin, 2023, 39(4): 124-135. |
| [3] | CHEN Nan-nan, WANG Chun-lai, JIANG Zhen-zhong, JIAO Peng, GUAN Shu-yan, MA Yi-yong. Genetic Transformation and Chilling Resistance Analysis of Maize ZmDHN15 Gene in Tobacco [J]. Biotechnology Bulletin, 2023, 39(4): 259-267. |
| [4] | WANG Bo-ya, JIANG Yong, HUANG Yan, CAO Ying, HU Shang-lian. Cloning and Functional Analysis of BeCesA4 in Bambusa emeiensis [J]. Biotechnology Bulletin, 2022, 38(11): 185-193. |
| [5] | WANG Xiao-tao, ZOU Hang, WU Yi, XIANG Shen-wei, LV Hua, LIU Chao-lan, LIN Jia-fu, WANG Xin-rong, CHU Yi-wen, SONG Tao. Heterologous Expression and Enzymatic Properties Analysis of Novel β-agarase Aga2 from Paraglaciecola hydrolytica [J]. Biotechnology Bulletin, 2022, 38(11): 258-268. |
| [6] | ZHANG Tong-tong, ZHENG Deng-yu, WU Zhong-yi, ZHANG Zhong-bao, YU Rong. Functional Analysis of ZmNF-YB13 Responding to Drought and Salt Stress [J]. Biotechnology Bulletin, 2022, 38(10): 115-123. |
| [7] | LIAO Zhao-min, CAI Jun, LIN Jian-guo, DU Xin, WANG Chang-gao. Expression of Glucose Oxidase Gene from Aspergillus niger in Pichia pastoris and Optimization of Enzyme Production Conditions [J]. Biotechnology Bulletin, 2021, 37(6): 97-107. |
| [8] | YUAN Yuan, WANG Lei, SHI Ya-wei. Research Advances in Strategies for Improving the Activity of Microbial-derived Alkaline Proteases [J]. Biotechnology Bulletin, 2021, 37(5): 231-236. |
| [9] | LI Xin-yue, ZHANG Jin-fang, XU Xiao-jian, LU Fu-ping, LI Yu. Effects of Spore Formation Related Gene Deletion on Biomass and Extracellular Enzyme Expression of Bacillus amyloliquefaciens [J]. Biotechnology Bulletin, 2021, 37(3): 35-43. |
| [10] | LIU Shan, YE Wei, ZHU Mu-zi, LI Sai-ni, DENG Zhang-shuang, ZHANG Wei-min. Cloning,Expression and Characterization of a Novel Acyltransferase GPAT [J]. Biotechnology Bulletin, 2021, 37(11): 257-266. |
| [11] | ZHAO Hai-yan, SONG Chen-bin, LIU Zheng-ya, MA Xing-rong, SHANG Hui-hui, LI An-hua, GUAN Xian-jun, WANG Jian-she. Cloning,Recombinant Expression and Enzymatic Properties of α-Amylase Gene from Laceyella sp. [J]. Biotechnology Bulletin, 2020, 36(8): 23-33. |
| [12] | LI Wei-na, SHEN Dong-ling, ZHANG Yu-xing, LIU Xue-tong, IRBIS Chagan. Cloning and Enzymatic Identification of Thermo-tolerant and Endotype Alginate Lyase Gene from Mangrovibacterium sp. SH-52 [J]. Biotechnology Bulletin, 2020, 36(12): 82-90. |
| [13] | ZHAO Jiang-hua, FANG Huan, ZHANG Da-wei. Research Progress in Biosynthesis of Secondary Metabolites of Microorganisms [J]. Biotechnology Bulletin, 2020, 36(11): 141-147. |
| [14] | ZHANG Yu-wen, YUAN Hang, YU Jiang-yue, MA Xiao-xiao, SHI Chao-shuo, LI Yu. Screening of a Bacterial Strain Efficiently Degrading Feather Waste and Optimization of Its Expression Condition [J]. Biotechnology Bulletin, 2019, 35(9): 93-98. |
| [15] | GU Yuan, XUN Zi-qi, ZHENG Fei, TU Tao, YAO Bin, LUO Hui-ying. Identification and Functional Exploration of the Expansin Gene from Thermophilic Talaromyces leycettanus JCM12802 [J]. Biotechnology Bulletin, 2018, 34(10): 172-181. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||