








Biotechnology Bulletin ›› 2023, Vol. 39 ›› Issue (3): 101-115.doi: 10.13560/j.cnki.biotech.bull.1985.2022-0851
Previous Articles Next Articles
SONG Hai-na1( ), WU Xin-tong1, YANG Lu-yu1, GENG Xi-ning1, ZHANG Hua-min2(
), WU Xin-tong1, YANG Lu-yu1, GENG Xi-ning1, ZHANG Hua-min2( ), SONG Xiao-long3
), SONG Xiao-long3
Received:2022-07-10
Online:2023-03-26
Published:2023-04-10
SONG Hai-na, WU Xin-tong, YANG Lu-yu, GENG Xi-ning, ZHANG Hua-min, SONG Xiao-long. Selection and Validation of Reference Genes for RT-qPCR in Allium tuberosum Infected by Botrytis squamosa[J]. Biotechnology Bulletin, 2023, 39(3): 101-115.
| 基因名称Gene symbol | 基因ID Gene ID | 引物序列Primer sequence(5'-3') | 扩增长度Amplified length/bp |
|---|---|---|---|
| UBC1 | F02_cb4795_c6/f1p0/677 | F:CACTTCCCTCCTGACTATCCCT R:CAAATACTGCCATTGCTGTTGA | 92 |
| UBC2 | F02_cb10194_c5/f1p0/683 | F:AAGGTGCTCCTGTCTATCTGCT R:TGATTCATACTTGGCTCTGTCG | 108 |
| UBQ1 | F02_cb4331_c39/f4p2/2064 | F:GTCAACTCCGTCACCGTAAACA R:GCAACTTCGGACTTCTCCTCTA | 119 |
| UBQ2 | F02_cb3159_c105/f1p2/1236 | F:ACCTTGTGCTCCGTCTTCGTG R:AACATCTGGCTTACCGTCGTCT | 162 |
| GAPDH3 | F02_cb2521_c25/f1p0/1405 | F:GACAAGCCAGTTGCTGTATTCG R:CTTTGCACCACCCTTAATGTGA | 140 |
| GAPDH4 | F02_cb2521_c44/f7p1/1415 | F:AATCACTGCCACCCAGAAGACT R:ACGGAATGACATGCCAGTAAGC | 163 |
| TUB | F02_cb8195_c2075/f1p0/1651 | F:GGTGGTGGTACTGGATCTGG R:TTGTATGGCTCCACAACTGC | 134 |
| EF-1α | F02_cb8884_c10/f1p3/1571 | F:ACCGAGTCCATTCCTCCTGTTC R:GGGTTTAGGTGCCTCCTCTTCA | 147 |
| 40S RP | F02_cb13441_c1/f3p0/558 | F:TGACCTCTTGAATCCACCTGC R:AGCAACCTTGGCACTTGACA | 107 |
| PABP | F02_cb2343_c66/f1p0/2490 | F:TCCTGATATGGCTGGCTTGC R:CCTCGGGCGATTCAAGAAGA | 224 |
| DnaJ | F02_cb2629_c9/f1p0/682 | F:AGACTCCCTGCAATTCCAGC R:TCTTCATGAACTCCTCGGCG | 223 |
| DDX | F02_cb3613_c7/f1p2/2466 | F:CCATGTTCAAGAGCAGCACG R:CATGAAGAGCACGAGACCCA | 145 |
| eIF-1A | F02_cb13117_c3/f2p1/651 | F:GGCTTTGTCATATTCGGGGC R:ATCACCAGCACCATCGTCC | 227 |
| GST676 | F02_cb3237_c0/f37p2/676 | F:GCAAGTCGGTAAAGCTCGGA R:ACTGGCCAAAAGAGTAGCGA | 160 |
| PRP902 | F02_cb14527_c723/f1p0/902 | F:CGTCACAGACAAGCAAAGGC R:CTTACTGTGCCACGTGGGAT | 180 |
Table 1 Genes and primer sequences
| 基因名称Gene symbol | 基因ID Gene ID | 引物序列Primer sequence(5'-3') | 扩增长度Amplified length/bp |
|---|---|---|---|
| UBC1 | F02_cb4795_c6/f1p0/677 | F:CACTTCCCTCCTGACTATCCCT R:CAAATACTGCCATTGCTGTTGA | 92 |
| UBC2 | F02_cb10194_c5/f1p0/683 | F:AAGGTGCTCCTGTCTATCTGCT R:TGATTCATACTTGGCTCTGTCG | 108 |
| UBQ1 | F02_cb4331_c39/f4p2/2064 | F:GTCAACTCCGTCACCGTAAACA R:GCAACTTCGGACTTCTCCTCTA | 119 |
| UBQ2 | F02_cb3159_c105/f1p2/1236 | F:ACCTTGTGCTCCGTCTTCGTG R:AACATCTGGCTTACCGTCGTCT | 162 |
| GAPDH3 | F02_cb2521_c25/f1p0/1405 | F:GACAAGCCAGTTGCTGTATTCG R:CTTTGCACCACCCTTAATGTGA | 140 |
| GAPDH4 | F02_cb2521_c44/f7p1/1415 | F:AATCACTGCCACCCAGAAGACT R:ACGGAATGACATGCCAGTAAGC | 163 |
| TUB | F02_cb8195_c2075/f1p0/1651 | F:GGTGGTGGTACTGGATCTGG R:TTGTATGGCTCCACAACTGC | 134 |
| EF-1α | F02_cb8884_c10/f1p3/1571 | F:ACCGAGTCCATTCCTCCTGTTC R:GGGTTTAGGTGCCTCCTCTTCA | 147 |
| 40S RP | F02_cb13441_c1/f3p0/558 | F:TGACCTCTTGAATCCACCTGC R:AGCAACCTTGGCACTTGACA | 107 |
| PABP | F02_cb2343_c66/f1p0/2490 | F:TCCTGATATGGCTGGCTTGC R:CCTCGGGCGATTCAAGAAGA | 224 |
| DnaJ | F02_cb2629_c9/f1p0/682 | F:AGACTCCCTGCAATTCCAGC R:TCTTCATGAACTCCTCGGCG | 223 |
| DDX | F02_cb3613_c7/f1p2/2466 | F:CCATGTTCAAGAGCAGCACG R:CATGAAGAGCACGAGACCCA | 145 |
| eIF-1A | F02_cb13117_c3/f2p1/651 | F:GGCTTTGTCATATTCGGGGC R:ATCACCAGCACCATCGTCC | 227 |
| GST676 | F02_cb3237_c0/f37p2/676 | F:GCAAGTCGGTAAAGCTCGGA R:ACTGGCCAAAAGAGTAGCGA | 160 |
| PRP902 | F02_cb14527_c723/f1p0/902 | F:CGTCACAGACAAGCAAAGGC R:CTTACTGTGCCACGTGGGAT | 180 |
| 基因 名称 Gene symbol | 模拟接菌不同时间的样品 Mock-inoculated samples | 接种葱鳞葡萄孢菌不同时间的样品 Samples inoculated with B. squamosa | ||||||
|---|---|---|---|---|---|---|---|---|
| 24 h | 48 h | 72 h | 24 h | 48 h | 72 h | |||
| UBC1 | 14.63 | 14.47 | 15.48 | 21.21 | 14.66 | 21.60 | ||
| UBC2 | 45.80 | 53.35 | 59.64 | 72.14 | 51.73 | 83.00 | ||
| UBQ1 | 8.75 | 7.90 | 10.06 | 12.67 | 8.00 | 10.81 | ||
| UBQ2 | 187.15 | 189.18 | 197.98 | 223.79 | 139.12 | 245.41 | ||
| GAPDH3 | 17.86 | 28.28 | 23.98 | 25.81 | 27.96 | 15.07 | ||
| GAPDH4 | 111.50 | 168.16 | 154.12 | 214.92 | 132.19 | 155.86 | ||
| TUB | 49.37 | 58.77 | 50.06 | 58.57 | 52.65 | 47.37 | ||
| EF-1α | 28.34 | 18.20 | 26.84 | 48.63 | 13.83 | 22.83 | ||
| 40S RP | 183.95 | 252.63 | 198.60 | 206.80 | 235.64 | 163.86 | ||
| PABP | 87.77 | 97.44 | 88.98 | 97.07 | 97.91 | 72.27 | ||
| DnaJ | 516.66 | 480.78 | 478.24 | 429.37 | 332.07 | 522.24 | ||
| DDX | 48.00 | 43.42 | 35.19 | 51.43 | 38.59 | 33.68 | ||
| eIF-1A | 31.35 | 44.60 | 34.58 | 38.97 | 33.39 | 33.45 | ||
Table 2 FPKM values of candidate reference genes in the samples by RNA-Seq
| 基因 名称 Gene symbol | 模拟接菌不同时间的样品 Mock-inoculated samples | 接种葱鳞葡萄孢菌不同时间的样品 Samples inoculated with B. squamosa | ||||||
|---|---|---|---|---|---|---|---|---|
| 24 h | 48 h | 72 h | 24 h | 48 h | 72 h | |||
| UBC1 | 14.63 | 14.47 | 15.48 | 21.21 | 14.66 | 21.60 | ||
| UBC2 | 45.80 | 53.35 | 59.64 | 72.14 | 51.73 | 83.00 | ||
| UBQ1 | 8.75 | 7.90 | 10.06 | 12.67 | 8.00 | 10.81 | ||
| UBQ2 | 187.15 | 189.18 | 197.98 | 223.79 | 139.12 | 245.41 | ||
| GAPDH3 | 17.86 | 28.28 | 23.98 | 25.81 | 27.96 | 15.07 | ||
| GAPDH4 | 111.50 | 168.16 | 154.12 | 214.92 | 132.19 | 155.86 | ||
| TUB | 49.37 | 58.77 | 50.06 | 58.57 | 52.65 | 47.37 | ||
| EF-1α | 28.34 | 18.20 | 26.84 | 48.63 | 13.83 | 22.83 | ||
| 40S RP | 183.95 | 252.63 | 198.60 | 206.80 | 235.64 | 163.86 | ||
| PABP | 87.77 | 97.44 | 88.98 | 97.07 | 97.91 | 72.27 | ||
| DnaJ | 516.66 | 480.78 | 478.24 | 429.37 | 332.07 | 522.24 | ||
| DDX | 48.00 | 43.42 | 35.19 | 51.43 | 38.59 | 33.68 | ||
| eIF-1A | 31.35 | 44.60 | 34.58 | 38.97 | 33.39 | 33.45 | ||
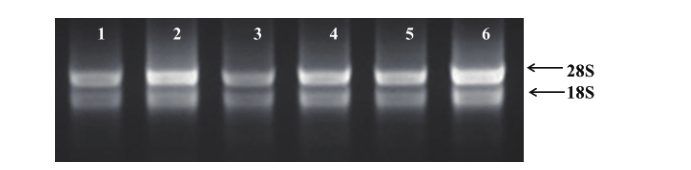
Fig. 1 Extracted total RNA in the leaves of Chinese chive A. tvberosum 1-3 are the mock-inoculated samples at 24, 48 and 72 h post inoculation respectively. 4-6 are the samples inoculated with B. squamosa at 24, 48 and 72 h post inoculation respectively

Fig. 2 Agarose electrophoresis analysis of PCR amplified products of candidate reference genes in Chinese chive 1: UBC1; 2: UBC2; 3: UBQ1; 4: UBQ2; 5: GAPDH3; 6: GAPDH4; 7: TUB; 8: EF-1a; 9: 40S RP; 10: DDX; 11: eIF-1A; 12: PABP; 13: DnaJ

Fig. 3 Melting curves of candidate reference genes in Chinese chive for RT-qPCR amplification -dF/dT is the first negative derivative of fluorescence(F)with respect to temperature(T)
| 基因名称 Gene symbol | 线性相关系数Linear correlation coefficient(R2) | 扩增效率Amplification efficiency/% |
|---|---|---|
| UBC1 | 0.993 9 | 104.83 |
| UBC2 | 0.963 0 | 101.12 |
| UBQ1 | 0.994 1 | 110.70 |
| UBQ2 | 0.990 7 | 106.64 |
| GAPDH3 | 0.995 5 | 109.88 |
| GAPDH4 | 0.996 6 | 110.65 |
| EF-1α | 0.991 2 | 110.36 |
| TUB | 0.991 7 | 107.54 |
| 40S RP | 0.995 3 | 100.80 |
| DDX | 0.990 3 | 99.19 |
| eIF-1A | 0.997 5 | 106.30 |
| PABP | 0.990 9 | 96.06 |
| DnaJ | 0.998 9 | 106.97 |
Table 3 Amplification efficiency of candidate reference genes
| 基因名称 Gene symbol | 线性相关系数Linear correlation coefficient(R2) | 扩增效率Amplification efficiency/% |
|---|---|---|
| UBC1 | 0.993 9 | 104.83 |
| UBC2 | 0.963 0 | 101.12 |
| UBQ1 | 0.994 1 | 110.70 |
| UBQ2 | 0.990 7 | 106.64 |
| GAPDH3 | 0.995 5 | 109.88 |
| GAPDH4 | 0.996 6 | 110.65 |
| EF-1α | 0.991 2 | 110.36 |
| TUB | 0.991 7 | 107.54 |
| 40S RP | 0.995 3 | 100.80 |
| DDX | 0.990 3 | 99.19 |
| eIF-1A | 0.997 5 | 106.30 |
| PABP | 0.990 9 | 96.06 |
| DnaJ | 0.998 9 | 106.97 |
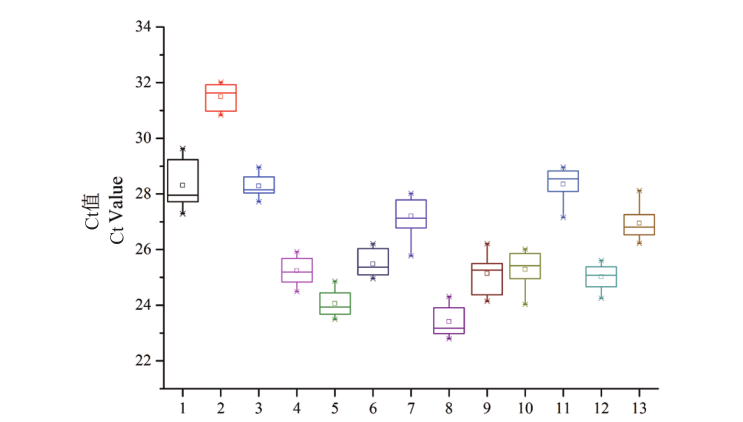
Fig. 4 Distribution of Ct values of candidate reference genes across all samples The horizontal line in the box indicates the average Ct values. 1: EF-1α; 2: UBQ1; 3: TUB; 4: UBQ2; 5: 40S RP; 6: eIF-1A; 7: DnaJ; 8: PABP; 9: DDX; 10: GAPDH4; 11: UBC1; 12: UBC2; 13: GAPDH3
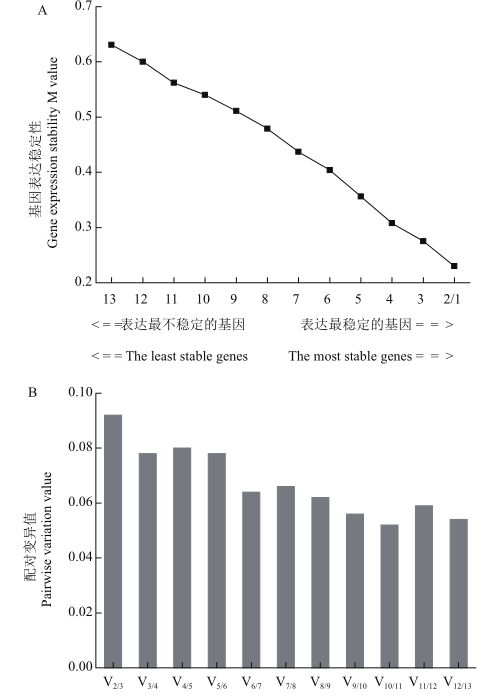
Fig. 5 Expression stabilities of candidate reference genes assayed by geNorm software A is the average expression stability M value of candidate reference genes. The least stable genes are on the left and the most stable genes on the right. B is the normalized number of reference genes. In figure A, 1: eIF-1A; 2: UBQ2; 3: UBC2; 4: 40S RP; 5: PABP; 6: UBC1; 7: UBQ1; 8: DnaJ; 9: GAPDH4; 10: TUB; 11: GAPDH3; 12: EF-1a; 13: DDX
| 基因名称Gene symbol | 稳定值 Stability | 排名Rank |
|---|---|---|
| UBQ1 | 0.195 | 1 |
| UBC2 | 0.261 | 2 |
| 40S RP | 0.28 | 3 |
| UBC1 | 0.374 | 4 |
| UBQ2 | 0.375 | 5 |
| eIF-1A | 0.443 | 6 |
| GAPDH4 | 0.456 | 7 |
| TUB | 0.478 | 8 |
| PABP | 0.48 | 9 |
| GAPDH3 | 0.51 | 10 |
| DnaJ | 0.581 | 11 |
| EF-1α | 0.642 | 12 |
| DDX | 0.654 | 13 |
Table 4 Expression stabilities of candidate reference genes assayed by NormFinder software
| 基因名称Gene symbol | 稳定值 Stability | 排名Rank |
|---|---|---|
| UBQ1 | 0.195 | 1 |
| UBC2 | 0.261 | 2 |
| 40S RP | 0.28 | 3 |
| UBC1 | 0.374 | 4 |
| UBQ2 | 0.375 | 5 |
| eIF-1A | 0.443 | 6 |
| GAPDH4 | 0.456 | 7 |
| TUB | 0.478 | 8 |
| PABP | 0.48 | 9 |
| GAPDH3 | 0.51 | 10 |
| DnaJ | 0.581 | 11 |
| EF-1α | 0.642 | 12 |
| DDX | 0.654 | 13 |
| 基因名称Gene symbol | 标准偏差Standard deviation(SD) | 变异系数Coefficient of variation(CV) | 调节系数标准差Std dev[± x-fold] | 排名Rank |
|---|---|---|---|---|
| TUB | 0.327 | 1.156 | 1.254 | 1 |
| UBC2 | 0.390 | 1.560 | 1.311 | 2 |
| 40S RP | 0.396 | 1.647 | 1.316 | 3 |
| eIF-1A | 0.411 | 1.615 | 1.330 | 4 |
| UBQ1 | 0.416 | 1.320 | 1.334 | 5 |
| PABP | 0.435 | 1.856 | 1.352 | 6 |
| UBQ2 | 0.436 | 1.726 | 1.353 | 7 |
| UBC1 | 0.472 | 1.666 | 1.387 | 8 |
| GAPDH3 | 0.477 | 1.768 | 1.391 | 9 |
| DnaJ | 0.494 | 1.811 | 1.408 | 10 |
| DDX | 0.560 | 2.225 | 1.474 | 11 |
| GAPDH4 | 0.596 | 2.359 | 1.512 | 12 |
| EF-1α | 0.727 | 2.567 | 1.655 | 13 |
Table 5 Expression stabilities of candidate reference genes assayed by BestKeeper software
| 基因名称Gene symbol | 标准偏差Standard deviation(SD) | 变异系数Coefficient of variation(CV) | 调节系数标准差Std dev[± x-fold] | 排名Rank |
|---|---|---|---|---|
| TUB | 0.327 | 1.156 | 1.254 | 1 |
| UBC2 | 0.390 | 1.560 | 1.311 | 2 |
| 40S RP | 0.396 | 1.647 | 1.316 | 3 |
| eIF-1A | 0.411 | 1.615 | 1.330 | 4 |
| UBQ1 | 0.416 | 1.320 | 1.334 | 5 |
| PABP | 0.435 | 1.856 | 1.352 | 6 |
| UBQ2 | 0.436 | 1.726 | 1.353 | 7 |
| UBC1 | 0.472 | 1.666 | 1.387 | 8 |
| GAPDH3 | 0.477 | 1.768 | 1.391 | 9 |
| DnaJ | 0.494 | 1.811 | 1.408 | 10 |
| DDX | 0.560 | 2.225 | 1.474 | 11 |
| GAPDH4 | 0.596 | 2.359 | 1.512 | 12 |
| EF-1α | 0.727 | 2.567 | 1.655 | 13 |

Fig. 6 Expression stabilities of candidate reference genes assayed by RefFinder 1: UBC2; 2: UBQ1; 3: 40S RP; 4: UBQ2; 5: eIF-1A; 6: TUB; 7: UBC1; 8: PABP;9: GAPDH4; 10: GAPDH3; 11: DnaJ; 12: EF-1α; 13: DDX
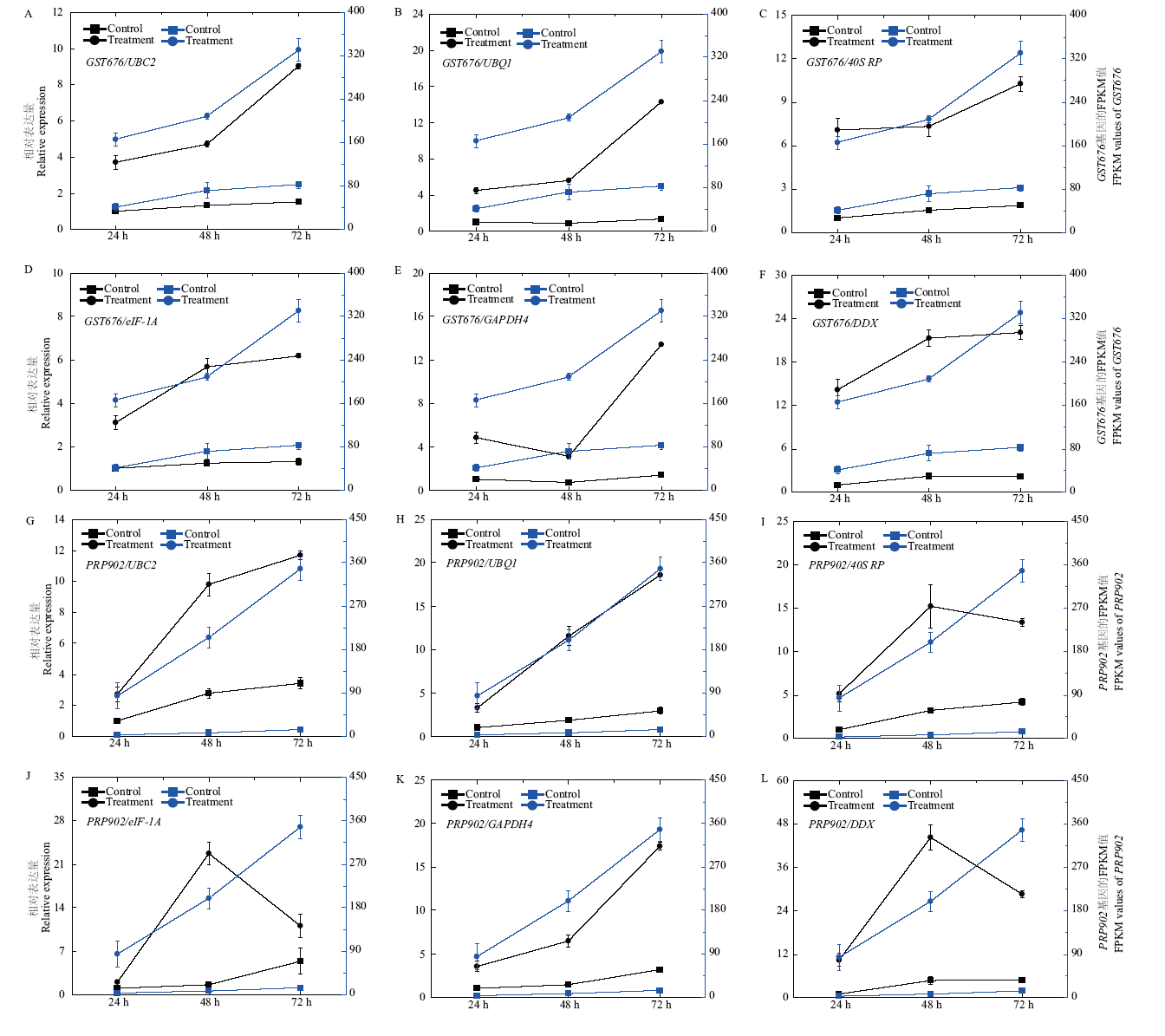
Fig. 7 Relative expressions of GST676 and PRP902 in Chinese chive leaves suffered by mock-inoculated and inoculated with B. squamosa when UBC2, UBQ1, 40S RP, eIF-1A, GAPDH4 or DDX used as the internal reference gene, and the FPKM values of GST676 and PRP902 in the samples by RNA-Seq The left side of Y axis is the relative expression, the right side of Y axis is the FPKM values of genes
| [1] | 黄征. 葱蒜类蔬菜真菌病害调查及其病原鉴定[J]. 中国农学通报, 2007, 23(5): 326-329. |
|
Huang Z. Diagnosis and identification on the fungal disease of bulb crops in Fujian[J]. Chin Agric Sci Bull, 2007, 23(5): 326-329.
doi: 10.11924/j.issn.1000-6850.0705326 |
|
| [2] | 崔蕴刚, 张华敏, 李延龙, 等. 韭菜灰霉病病原鉴定及其生物学特性[J]. 北方园艺, 2020(4): 14-19. |
| Cui YG, Zhang HM, Li YL, et al. Identification and biological characterization of the pathogen of Allium tuberosum grey mold[J]. North Hortic, 2020(4): 14-19. | |
| [3] |
Zhang J, Zhang L, Li GQ, et al. Botrytis sinoallii: a new species of the grey mould pathogen on Allium crops in China[J]. Mycoscience, 2010, 51(6): 421-431.
doi: 10.47371/mycosci.MYC51421 URL |
| [4] | 胡彬, 黄中乔, 刘西莉, 等. 9种杀菌剂对韭菜灰霉病的防治效果[J]. 中国农学通报, 2014, 30(4): 293-298. |
|
Hu B, Huang ZQ, Liu XL, et al. Control efficacy of 9 fungicides against Chinese chives gray mold[J]. Chin Agric Sci Bull, 2014, 30(4): 293-298.
doi: 10.11924/j.issn.1000-6850.2013-3021 |
|
| [5] | 胡彬, 李琳, 戚如诗, 等. 从韭菜腐霉利残留超标看农药登记及最大残留限量标准的科学制定[J]. 中国蔬菜, 2020(5): 9-11. |
| Hu B, Li L, Qi RS, et al. The scientific establishment of pesticide registration and maximum residue limit standard from the point of Chinese chives procymidone residue exceed the standard[J]. China Veg, 2020(5): 9-11. | |
| [6] | 高皓杰, 张兰云, 李桐桐, 等. 韭菜灰霉病防治烟剂的筛选与评价[J]. 农药学学报, 2022, 24(2): 315-325. |
| Gao HJ, Zhang LY, Li TT, et al. Screening and evaluation of smoke generators to control gray mold of Chinese chives[J]. Chin J Pestic Sci, 2022, 24(2): 315-325. | |
| [7] | 韩之琪, 贲海燕, 谢学文, 等. 灰葡萄孢对杀菌剂抗性研究进展[J]. 中国蔬菜, 2014(5): 6-10. |
| Han ZQ, Ben HY, Xie XW, et al. Research progress on Botrytis cinerea resistance to different fungicides[J]. China Veg, 2014(5): 6-10. | |
| [8] | Rupp S, Plesken C, Rumsey S, et al. Botrytis fragariae, a new species causing gray mold on strawberries, shows high frequencies of specific and efflux-based fungicide resistance[J]. Appl Environ Microbiol, 2017, 83(9): e00269-e00217. |
| [9] | 胡适宜. 被子植物胚胎学[M]. 北京: 人民教育出版社, 1982. |
| Hu SY. Angiosperm embryology[M]. Beijing: People’s Education Press, 1982. | |
| [10] | 张华敏, 陈建华, 尹守恒, 等. 韭菜无融合生殖种子形成机制研究进展[J]. 河南农业科学, 2020, 49(6): 1-7. |
| Zhang HM, Chen JH, Yin SH, et al. Research progress of the formation mechanism of apomictic seed in Chinese chive[J]. J Henan Agric Sci, 2020, 49(6): 1-7. | |
| [11] |
Gachon C, Mingam A, Charrier B. Real-time PCR: what relevance to plant studies?[J]. J Exp Bot, 2004, 55(402): 1445-1454.
doi: 10.1093/jxb/erh181 pmid: 15208338 |
| [12] |
Nolan T, Hands RE, Bustin SA. Quantification of mRNA using real-time RT-PCR[J]. Nat Protoc, 2006, 1(3): 1559-1582.
doi: 10.1038/nprot.2006.236 pmid: 17406449 |
| [13] |
Dheda K, Huggett JF, Chang JS, et al. The implications of using an inappropriate reference gene for real-time reverse transcription PCR data normalization[J]. Anal Biochem, 2005, 344(1): 141-143.
doi: 10.1016/j.ab.2005.05.022 pmid: 16054107 |
| [14] |
袁伟, 万红建, 杨悦俭. 植物实时荧光定量PCR内参基因的特点及选择[J]. 植物学报, 2012, 47(4): 427-436.
doi: 10.3724/SP.J.1259.2012.00427 |
| Yuan W, Wan HJ, Yang YJ. Characterization and selection of reference genes for real-time quantitative RT-PCR of plants[J]. Chin Bull Bot, 2012, 47(4): 427-436. | |
| [15] |
Brunner AM, Yakovlev IA, Strauss SH. Validating internal controls for quantitative plant gene expression studies[J]. BMC Plant Biol, 2004, 4: 14.
pmid: 15317655 |
| [16] |
Huggett J, Dheda K, Bustin S, et al. Real-time RT-PCR normalisation; strategies and considerations[J]. Genes Immun, 2005, 6(4): 279-284.
doi: 10.1038/sj.gene.6364190 pmid: 15815687 |
| [17] |
Jain M, Nijhawan A, Tyagi AK, et al. Validation of housekeeping genes as internal control for studying gene expression in rice by quantitative real-time PCR[J]. Biochem Biophys Res Commun, 2006, 345(2): 646-651.
doi: 10.1016/j.bbrc.2006.04.140 URL |
| [18] |
蒋婷婷, 高燕会, 童再康. 石蒜属植物实时荧光定量PCR内参基因的选择[J]. 园艺学报, 2015, 42(6): 1129-1138.
doi: 10.16420/j.issn.0513-353x.2014-0999 |
| Jiang TT, Gao YH, Tong ZK. Selection of reference genes for quantitative real-time PCR in Lycoris[J]. Acta Hortic Sin, 2015, 42(6): 1129-1138. | |
| [19] |
马璐琳, 段青, 崔光芬, 等. 钝裂银莲花花色素合成相关基因RT-qPCR内参基因的筛选[J]. 园艺学报, 2021, 48(2): 377-388.
doi: 10.16420/j.issn.0513-353x.2020-0306 |
| Ma LL, Duan Q, Cui GF, et al. Selection and validation of reference genes for RT-qPCR analysis of the correlated genes in flower pigments biosynthesis pathway of Anemone obtusiloba[J]. Acta Hortic Sin, 2021, 48(2): 377-388. | |
| [20] | 李梦倩, 刘敏, 张蒙, 等. 大蒜体细胞胚发生mRNA及miRNA qPCR内参基因的筛选和验证[J]. 农业生物技术学报, 2021, 29(12): 2449-2464. |
| Li MQ, Liu M, Zhang M, et al. Identification and verification of somatic embryogenesis mRNA and miRNA qPCR reference genes in garlic(Allium sativum)[J]. J Agric Biotechnol, 2021, 29(12): 2449-2464. | |
| [21] |
Gutierrez L, Mauriat M, Guénin S, et al. The lack of a systematic validation of reference genes: a serious pitfall undervalued in reverse transcription-polymerase chain reaction(RT-PCR)analysis in plants[J]. Plant Biotechnol J, 2008, 6(6): 609-618.
doi: 10.1111/j.1467-7652.2008.00346.x pmid: 18433420 |
| [22] |
Volkov RA, Panchuk II, Schöffl F. Heat-stress-dependency and developmental modulation of gene expression: the potential of house-keeping genes as internal standards in mRNA expression profiling using real-time RT-PCR[J]. J Exp Bot, 2003, 54(391): 2343-2349.
pmid: 14504302 |
| [23] |
Hu RB, Fan CM, Li HY, et al. Evaluation of putative reference genes for gene expression normalization in soybean by quantitative real-time RT-PCR[J]. BMC Mol Biol, 2009, 10: 93.
doi: 10.1186/1471-2199-10-93 pmid: 19785741 |
| [24] |
Die JV, Román B, Nadal S, et al. Evaluation of candidate reference genes for expression studies in Pisum sativum under different experimental conditions[J]. Planta, 2010, 232(1): 145-153.
doi: 10.1007/s00425-010-1158-1 URL |
| [25] |
Kozera B, Rapacz M. Reference genes in real-time PCR[J]. J Appl Genet, 2013, 54(4): 391-406.
pmid: 24078518 |
| [26] | 张玉芳, 赵丽娟, 曾幼玲. 基因表达研究中内参基因的选择与应用[J]. 植物生理学报, 2014, 50(8): 1119-1125. |
| Zhang YF, Zhao LJ, Zeng YL. Selection and application of reference genes for gene expression studies[J]. Plant Physiol J, 2014, 50(8): 1119-1125. | |
| [27] |
Deo RC, Bonanno JB, Sonenberg N, et al. Recognition of polyadenylate RNA by the poly(A)-binding protein[J]. Cell, 1999, 98(6): 835-845.
pmid: 10499800 |
| [28] |
Rocak S, Linder P. DEAD-box proteins: the driving forces behind RNA metabolism[J]. Nat Rev Mol Cell Biol, 2004, 5(3): 232-241.
doi: 10.1038/nrm1335 |
| [29] | 蔡敬, 孟小庆, 董婷婷, 等. DEAD-box解旋酶在植物非生物胁迫响应中的功能研究进展[J]. 生命科学, 2017, 29(5): 427-433. |
| Cai J, Meng XQ, Dong TT, et al. Progress of plant DEAD-box helicase in response to abiotic stress[J]. Chin Bull Life Sci, 2017, 29(5): 427-433. | |
| [30] | Vandesompele J, de Preter K, Pattyn F, et al. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes[J]. Genome Biol, 2002, 3(7): 467-470. |
| [31] |
Andersen CL, Jensen JL, Ørntoft TF. Normalization of real-time quantitative reverse transcription-PCR data: a model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets[J]. Cancer Res, 2004, 64(15): 5245-5250.
doi: 10.1158/0008-5472.CAN-04-0496 pmid: 15289330 |
| [32] |
Pfaffl MW, Tichopad A, Prgomet C, et al. Determination of stable housekeeping genes, differentially regulated target genes and sample integrity: BestKeeper-Excel-based tool using pair-wise correlations[J]. Biotechnol Lett, 2004, 26(6): 509-515.
doi: 10.1023/B:BILE.0000019559.84305.47 URL |
| [33] | 吴建阳, 何冰, 杜玉洁, 等. 利用geNorm、NormFinder和BestKeeper软件进行内参基因稳定性分析的方法[J]. 现代农业科技, 2017(5): 278-281. |
| Wu JY, He B, Du YJ, et al. Analysis method of systematically evaluating stability of reference genes using geNorm, NormFinder and BestKeeper[J]. Mod Agric Sci Technol, 2017(5): 278-281. | |
| [34] |
Xie F, Xiao P, Chen D, et al. miRDeepFinder: a miRNA analysis tool for deep sequencing of plant small RNAs[J]. Plant Mol Biol, 2012, 80(1):75-84.
doi: 10.1007/s11103-012-9885-2 URL |
| [35] | Ann M. Regulatory and functional interactions of plant growth regulators and plant glutathione S-transferases(GSTs)[J]. Vitam Horm, 2005, 72: 155-202. |
| [36] |
Agrawal GK, Rakwal R, Jwa NS, et al. Signalling molecules and blast pathogen attack activates rice OsPR1a and OsPR1b genes: a model illustrating components participating during defence/stress response[J]. Plant Physiol Biochem, 2001, 39(12): 1095-1103.
doi: 10.1016/S0981-9428(01)01333-X URL |
| [37] |
van Loon LC, Rep M, Pieterse CMJ. Significance of inducible defense-related proteins in infected plants[J]. Annu Rev Phytopathol, 2006, 44: 135-162.
pmid: 16602946 |
| [38] | 张亚琳, 张亚琨, 江春泉, 等. 人参锈腐病菌Ilyonectria robusta侵染人参根实时定量PCR内参基因的筛选与验证[J/OL]. 吉林农业大学学报, 2022. https://kns.cnki.net/kcms/detail/22.1100.S.20220331.1519.003.html. |
| Zhang YL, Zhang YK, Jiang CQ, et al. Screening and verification of internal reference genes in Panax ginseng roots infected by Ilyonectria robusta causing ginseng rusty rot by real-time quantitative PCR[J/OL]. J Jilin Agric Univ, 2022. https://kns.cnki.net/kcms/detail/22.1100.S.20220331.1519.003.html. | |
| [39] | 姚李祥, 潘春柳, 余丽莹, 等. 草果种子休眠解除过程中RT-qPCR内参基因筛选[J]. 中国中药杂志, 2021, 46(15): 3832-3837. |
| Yao LX, Pan CL, Yu LY, et al. Selection of RT-qPCR reference genes for Amomum tsaoko seeds during dormancy release[J]. China J Chin Mater Med, 2021, 46(15): 3832-3837. | |
| [40] | 赵艺蕊, 黄春颖, 王克涛, 等. 山核桃实时荧光定量PCR分析中内参基因的筛选与验证[J]. 果树学报, 2022, 39(1): 10-21. |
| Zhao YR, Huang CY, Wang KT, et al. Screening and verification of internal reference genes by real time quantitative PCR analysis in Carya cathayensis[J]. J Fruit Sci, 2022, 39(1): 10-21. | |
| [41] | 苏丹丹, 刘玉萍, 张雨, 等. 苦豆子实时荧光定量PCR内参基因筛选与验证[J]. 植物生理学报, 2022, 58(7): 1295-1306. |
| Su DD, Liu YP, Zhang Y, et al. Screening and validation of reference genes in Sophora alopecuroides for real-time quantitative PCR[J]. Plant Physiol J, 2022, 58(7): 1295-1306. | |
| [42] |
徐圆圆, 赵国春, 郝颖颖, 等. 无患子RT-qPCR内参基因的筛选与验证[J]. 生物技术通报, 2022, 38(10):80-89.
doi: 10.13560/j.cnki.biotech.bull.1985.2021-1616 |
| Xu YY, Zhao GC, Hao YY, et al. Reference genes selection and validation for RT-qPCR in Sapindus mukorossi[J]. Biotechnol Bull, 2022, 38(10):80-89. | |
| [43] |
庞强强, 李植良, 罗少波, 等. 高温胁迫下茄子RT-qPCR内参基因筛选及稳定性分析[J]. 园艺学报, 2017, 44(3): 475-486.
doi: 10.16420/j.issn.0513-353x.2016-0831 |
| Pang QQ, Li ZL, Luo SB, et al. Selection and stability analysis of reference gene for RT-qPCR in eggplant under high temperature stress[J]. Acta Hortic Sin, 2017, 44(3): 475-486. | |
| [44] |
Faccioli P, Ciceri GP, Provero P, et al. A combined strategy of “in silico” transcriptome analysis and web search engine optimization allows an agile identification of reference genes suitable for normalization in gene expression studies[J]. Plant Mol Biol, 2007, 63(5): 679-688.
doi: 10.1007/s11103-006-9116-9 URL |
| [45] |
Liu WG, Tang X, Qi XH, et al. The ubiquitin conjugating enzyme: an important ubiquitin transfer platform in ubiquitin-proteasome system[J]. Int J Mol Sci, 2020, 21(8): 2894.
doi: 10.3390/ijms21082894 URL |
| [46] |
Grumati P, Dikic I. Ubiquitin signaling and autophagy[J]. J Biol Chem, 2018, 293(15): 5404-5413.
doi: 10.1074/jbc.TM117.000117 pmid: 29187595 |
| [47] |
Nakamura N. Ubiquitin system[J]. International Journal of Molecular Sciences, 2018, 19(4):1080.
doi: 10.3390/ijms19041080 URL |
| [48] |
Hamey JJ, Wilkins MR. Methylation of elongation factor 1A: where, who, and why?[J]. Trends Biochem Sci, 2018, 43(3): 211-223.
doi: S0968-0004(18)30004-5 pmid: 29398204 |
| [49] |
Bevitori R, Oliveira MB, Grossi-de-Sá MF, et al. Selection of optimized candidate reference genes for RT-qPCR normalization in rice(Oryza sativa L.)during Magnaporthe oryzae infection and drought[J]. Genet Mol Res, 2014, 13(4): 9795-9805.
doi: 10.4238/2014.November.27.7 pmid: 25501189 |
| [50] | 代资举, 王艳, 王新涛, 等. 玉米粗缩病诱导下实时荧光定量PCR内参基因的选择[J]. 植物生理学报, 2019, 55(10): 1545-1553. |
| Dai ZJ, Wang Y, Wang XT, et al. Reference genes selection for real-time quantitative PCR under rough dwarf virus induced disease in maize[J]. Plant Physiol J, 2019, 55(10): 1545-1553. | |
| [51] | 唐枝娟, 刘秦, 肖晓蓉, 等. 白叶枯病菌侵染下的水稻内参基因稳定性[J]. 分子植物育种, 2017, 15(1): 300-306. |
| Tang ZJ, Liu Q, Xiao XR, et al. Selection of optimized candidate reference genes for RT-qPCR normalization in rice during Xanthomonas oryzae pv. oryzae infection[J]. Mol Plant Breed, 2017, 15(1): 300-306. | |
| [52] | 赵辉, 张春艳, 文艺, 等. 菜豆壳球孢侵染芝麻过程中内参基因的筛选[J]. 中国油料作物学报, 2017, 39(3): 393-398, 419. |
| Zhao H, Zhang CY, Wen Y, et al. Screening of reference genes in sesame during Macrophomina phaseolina infection[J]. Chin J Oil Crop Sci, 2017, 39(3): 393-398, 419. | |
| [53] | 姚全胜, 杨倩, 柳凤, 等. 芒果细菌性角斑病菌在侵染芒果叶片过程中内参基因筛选[J]. 分子植物育种, 2021, 19(18): 6088-6095. |
| Yao QS, Yang Q, Liu F, et al. Screening of reference genes in Xanthomonas citri pv. mangiferaeindicae during the infection of mango leaf[J]. Mol Plant Breed, 2021, 19(18): 6088-6095. | |
| [54] |
李恬静薇, 邹潇潇, 朱军, 等. 长茎葡萄蕨藻胁迫条件下RT-qPCR内参基因的筛选与验证[J]. 生物技术通报, 2021, 37(10): 266-276.
doi: 10.13560/j.cnki.biotech.bull.1985.2020-1585 |
| Li T, Zou XX, Zhu J, et al. Selection and validation of reference genes for quantitative real-time PCR in Caulerpa lentillifera under stress conditions[J]. Biotechnol Bull, 2021, 37(10): 266-276. | |
| [55] | 陈慧洁, 冯丽贞, 李慧敏, 等. 桉树焦枯病菌内参基因的筛选[J]. 森林与环境学报, 2018, 38(1): 98-103. |
| Chen HJ, Feng LZ, Li HM, et al. Screening and evaluation of reference genes of Calonectria pseudoreteaudii[J]. J For Environ, 2018, 38(1): 98-103. |
| [1] | YAO Zi-ting, CAO Xue-ying, XIAO Xue, LI Rui-fang, WEI Xiao-mei, ZOU Cheng-wu, ZHU Gui-ning. Screening of Reference Genes for RT-qPCR in Neoscytalidium dimidiatum [J]. Biotechnology Bulletin, 2023, 39(5): 92-102. |
| [2] | XU Yuan-yuan, ZHAO Guo-chun, HAO Ying-ying, WENG Xue-huang, CHEN Zhong, JIA Li-ming. Reference Genes Selection and Validation for RT-qPCR in Sapindus mukorossi [J]. Biotechnology Bulletin, 2022, 38(10): 80-89. |
| [3] | ZHANG Jing, XIONG Yan, HUA Yong-lin, GUO Yu, XIONG Xian-rong, ZI Xiang-dong, LI Jian. Screening of Reference Genes for Quantitative PCR of Skeletal Muscle Fiber Types in Mice [J]. Biotechnology Bulletin, 2021, 37(2): 71-79. |
| [4] | LI Tianjingwei, ZOU Xiao-xiao, ZHU Jun, BAO Shi-xiang. Selection and Validation of Reference Genes for Quantitative Real-time PCR in Caulerpa lentillifera Under Stress Conditions [J]. Biotechnology Bulletin, 2021, 37(10): 266-276. |
| [5] | PAN Mo-han, LU Tian-quan, TIAN Bo. Selection and Validation of Reference Genes in the Seeds of Paeonia delavayi in Quantitative Real-time PCR Analysis [J]. Biotechnology Bulletin, 2020, 36(9): 218-226. |
| [6] | ZHANG Rui-zhu, JIANG Yu-chen, HUANG Jun, YAN Jie. Cloning and Expression Analysis of SRPP2 Gene in Taraxacum kok-saghyz Rodin [J]. Biotechnology Bulletin, 2020, 36(1): 9-14. |
| [7] | KONG Chun-yan, CHEN Yong-kun, WANG Sha-sha, HAO Da-hai, YANG Yu, GONG Ming. Screening and Comparison of Reference Genes for microRNA Quantitative Real-time PCR in Jatropha curcas Under Chilling Stress [J]. Biotechnology Bulletin, 2019, 35(7): 25-32. |
| [8] | WANG Jie, ZHANG Yang, QIN Peng, HE Mao-lan, LI Jin, GU Yun-fu, ZENG Xian-fu, XIANG Quan-ju. Effects of pH on the Lentinan Content and the Transcriptional Levels of Genes Related to the Synthesis of Key Enzymes in Lentinula edodes [J]. Biotechnology Bulletin, 2019, 35(2): 39-45. |
| [9] | ZHANG Yan, XIA Geng-shou, LAI Zhi-bing. Recent Advances in Molecular Mechanisms of Plant Responses Against Botrytis cinerea [J]. Biotechnology Bulletin, 2018, 34(2): 10-24. |
| [10] | LI Yu ZHAO Lei CHEN Li ZHOU Yu-xun LI Kai XIAO Jun-hua. Accurate Detection Efficiency of SF2 Protein RIP Enriching RNA Using Exogenous RNA as Reference Gene with qPCR [J]. Biotechnology Bulletin, 2017, 33(5): 78-82. |
| [11] | LU Shou-liang LI Liang-yuan GAO Lei SHEN Min WAN Peng-cheng SHI Guo-qing DAI Rong. The Expression of LEPRb mRNA in the Estrus Cycle of Different Sheep Strains [J]. Biotechnology Bulletin, 2017, 33(5): 139-144. |
| [12] | REN Rui, DAI Peng-hui, CAO Fu-xiang, DONG Xu-jie, LI Meng. Cloning of a CAC Gene in Dove Tree(Davidia involucrata Baill.)and Evaluating its Potential as a Novel Reference Gene [J]. Biotechnology Bulletin, 2017, 33(4): 119-129. |
| [13] | ZHAO Xiao-bing, PAN Lei, LIU Wei, LIN Mao-yi, LI Hong-qing, LIU Zhong. Screening of Reference Genes for Quantitative Real-time PCR Analysis in Asarum sieboldii [J]. Biotechnology Bulletin, 2017, 33(11): 174-179. |
| [14] | HU Ya, LU Cheng, WEI Chuan-chuan, XIU Jiang-fan, WU Jian-wei. Cloning and Expression Pattern of Ribosomal Protein S18 Gene in Musca domestica [J]. Biotechnology Bulletin, 2016, 32(6): 135-142. |
| [15] | HE Ting-ting, SONG Ting, WANG Chao, ZHANG Chang-bin, WANG Hai-yan. Screening of Reference Genes in Bacillus pumilus by Real-time Fluorescence Quantitative PCR [J]. Biotechnology Bulletin, 2016, 32(11): 99-106. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||