








Biotechnology Bulletin ›› 2023, Vol. 39 ›› Issue (7): 1-12.doi: 10.13560/j.cnki.biotech.bull.1985.2023-0051
CHEN Xiao1( ), YU Ming-lan1, WU Long-kun2, ZHENG Xiao-ming3,4,5, PANG Hong-bo1(
), YU Ming-lan1, WU Long-kun2, ZHENG Xiao-ming3,4,5, PANG Hong-bo1( )
)
Received:2023-01-19
Online:2023-07-26
Published:2023-08-17
Contact:
PANG Hong-bo
E-mail:19861602091@163.com;panghb@synu.edu.cn
CHEN Xiao, YU Ming-lan, WU Long-kun, ZHENG Xiao-ming, PANG Hong-bo. Research Progress in lncRNA and Their Responses to Low Temperature Stress in Plant[J]. Biotechnology Bulletin, 2023, 39(7): 1-12.

Fig. 1 Structure and origin of lncRNA A: Gene structure of lncRNA. B: The mechanism of lncRNA formation.(1)The reading frame inserted into the intron of the protein-coding gene, and the inserted sequence is reintegrated with the previous sequence to form lncRNA.(2)Chromosome recombination, two long-distance(>10 Mb)non-coding regions of protein-coding genes connected together to form lncRNA containing multiple exons.(3)Non-coding gene retrotransposons use RNA as a medium to create a new copy at a new location in the genome through reverse transcription and paste it by copying and pasting, forming a lncRNA in conjunction with the previous non-coding sequences.(4)Tandem repeat events generate adjacent repeat sequences within non-coding RNA to produce new lncRNA.(5)Transposable elements inserted into non-coding genes, which forms a lncRNA in conjunction with the previous sequence
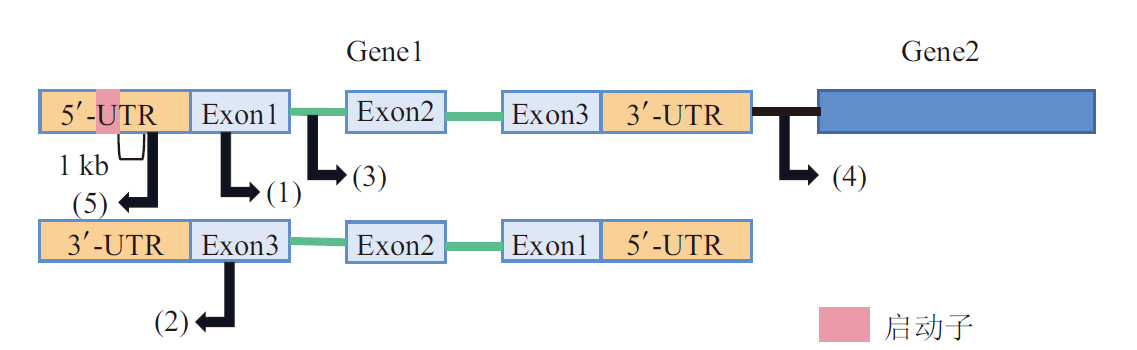
Fig. 2 Classification of lncRNAs based on genomic locations (1)Sense lncRNA is transcribed from the sense strand of protein-coding genes.(2)Antisense lncRNA is transcribed from the antisense strand of protein-coding genes in the opposite direction.(3)Intron lncRNA is transcribed from the intronic region of protein-coding genes.(4)Intergenic lncRNA is transcribed from the region between two genes.(5)Bidirectional lncRNA is transcribed in the opposite direction of protein-coding genes from the promoter region.
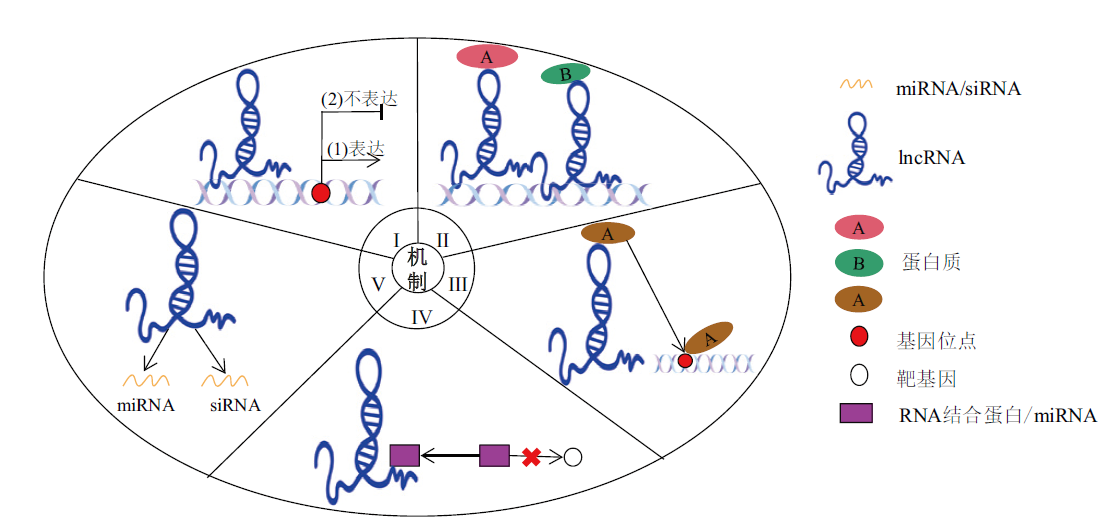
Fig. 3 Schematic diagram of lncRNA action mechanisms I: Signal molecule model, as signaling molecules, lncRNAs control neighboring genes to determine whether they are expressed; II: scaffold molecular model, the lncRNAs act as scaffolding molecules for a variety of proteins; III: guided molecular model, proteins are directed to specific sites for action by lncRNAs; IV: the bait molecule model, induced RNA-binding proteins or miRNAs bind to lncRNA, preventing them from binding to target genes; V: lncRNA acts as a precursor for the biosynthesis of miRNAs and siRNAs
| lncRNA的名称 Name of lncRNA | 物种 Species | 功能 Function | 机制 Mechanism | 参考文献 Reference |
|---|---|---|---|---|
| COLDAIR | 拟南芥 A. thaliana | 参与春化作用 Involved in vernalization | 信号、引导分子,抑制FLC的表达 Signal, guide molecules, and inhibit the expression of FLC | [ |
| COLDWRAP | 拟南芥 A. thaliana | 参与春化作用 Involved in vernalization | 抑制FLC的表达 Inhibit the expression of FLC | [ |
| COOLAIR | 拟南芥 A. thaliana | 参与春化作用 Involved in vernalization | 增加H3K27me3和降低H3K36me3转录来抑制FLC Inhibition of FLC by increasing H3K27me3 and decreasing H3K36me3 transcription | [ |
| Svalka | 拟南芥 A. thaliana | 参与冷胁迫 Involved in cold stress | Svalka-asCBF1级联控制CBF1表达 Svalka-asCBF1 cascade controls CBF1 expression | [ |
| CIL1 | 拟南芥 A. thaliana | 参与冷胁迫 Involved in cold stress | 影响活性氧或渗透调节物质来响应冷胁迫 Responding to cold stress by affecting reactive oxygen species or osmotic regulators | [ |
| APOLO | 拟南芥 A. thaliana | 寒冷条件调节根毛伸长 Regulates root hair elongation under cold conditions | 与转录因子WRKY42相互作用 Interacting with the TF WRKY42 | [ |
| TE-lincRNA11195 | 拟南芥 A. thaliana | 参与冷胁迫 Involved in cold stress | 水杨酸刺激反应基因可能是潜在靶标 Salicylic acid-induced response genes may be potential targets | [ |
| DPA lncRNAs | 水稻 O. sativa | 参与冷胁迫 Involved in cold stress | 多聚腺苷酸化来响应冷胁迫 Polyadenylation in response to cold stress | [ |
| lncRNA-Chr03G0008 | 水稻 O. sativa | 参与冷胁迫 Involved in cold stress | 在幼苗中表达响应冷胁迫 Expressing response to cold stress in seedlings | [ |
| lncRNA-SVR | 水稻 O. sativa | 低温下与种子活力相关 Associated with seed viability under low temperature | 顺式基因SAUR家族成员相互作用 Interacting with the cis-gene family member SAUR | [ |
| lncR9A, lncR117, lncR616 | 东农冬麦1号 T. aestivum(Dn1) | 参与冷胁迫 Involved in cold stress | 竞争性结合miR398间接调节CSD1的表达 Indirectly regulates the expression of CSD1 by competitively binding miR398 | [ |
| Traes_2BS_7A04BF5D5 | 小麦 Durum wheat | 参与冷胁迫 Involved in cold stress | 以WCOR413冷驯化基因为靶标响应冷胁迫 Responding to cold stress targeting the cold acclimation gene WCOR413 | [ |
| CRR5 | 木薯 M. esculenta | 参与冷胁迫 Involved in cold stress | 与蛋白激酶基因协同表达响应低温胁迫 Co-expression with protein kinase gene in response to low temperature stress | [ |
| CRIR1 | 木薯 M. esculenta | 参与冷胁迫 Involved in cold stress | 招募MeCSP5来提高mRNA的翻译效率 Recruit MeCSP5 to improve the translation efficiency of mRNA | [ |
| XH123 | 棉花 G. hirsutum | 参与冷胁迫 Involved in cold stress | 基因沉默引起冷调控基因PIF3等差异表达 Gene silencing induced differential expression of cold-regulated genes such as PIF3 | [ |
| 苜蓿 M. truncatula | 参与冷胁迫 Involved in cold stress | 构成lncRNA- MtCBFs调控网络 Construct lncRNA-MtCBFs regulatory network | [ | |
| 番茄 S. lycopersicum | 参与冷胁迫 Involved in cold stress | 竞争与共享miRNA来调控mRNA的表达 Competition and sharing of miRNAs to regulate mRNA expression | [ | |
| DE-lncRNAs | 甜椒 C. annuum | 参与冷胁迫 Involved in cold stress | 调控与冷损伤相关的靶基因 作为miRNAs前体响应冷胁迫 Regulating target genes related to cold damage Responding to cold stress as precursors of miRNAs | [ [ |
| 葡萄 V. vinifera | 参与冷胁迫 Involved in cold stress | 作为miRNAs的靶标 Being targets of miRNAs | [ | |
| 香蕉 M. balbisiana | 参与冷胁迫 Involved in cold stress | 调节类黄酮、蛋白激酶的生物合成以及TCA循环、硫传递系统等途径 Regulating the biosynthesis of flavonoids and protein kinases, as well as pathways such as the TCA cycle and sulfur transfer system | [ |
Table 1 Low temperature-related lncRNAs in plants
| lncRNA的名称 Name of lncRNA | 物种 Species | 功能 Function | 机制 Mechanism | 参考文献 Reference |
|---|---|---|---|---|
| COLDAIR | 拟南芥 A. thaliana | 参与春化作用 Involved in vernalization | 信号、引导分子,抑制FLC的表达 Signal, guide molecules, and inhibit the expression of FLC | [ |
| COLDWRAP | 拟南芥 A. thaliana | 参与春化作用 Involved in vernalization | 抑制FLC的表达 Inhibit the expression of FLC | [ |
| COOLAIR | 拟南芥 A. thaliana | 参与春化作用 Involved in vernalization | 增加H3K27me3和降低H3K36me3转录来抑制FLC Inhibition of FLC by increasing H3K27me3 and decreasing H3K36me3 transcription | [ |
| Svalka | 拟南芥 A. thaliana | 参与冷胁迫 Involved in cold stress | Svalka-asCBF1级联控制CBF1表达 Svalka-asCBF1 cascade controls CBF1 expression | [ |
| CIL1 | 拟南芥 A. thaliana | 参与冷胁迫 Involved in cold stress | 影响活性氧或渗透调节物质来响应冷胁迫 Responding to cold stress by affecting reactive oxygen species or osmotic regulators | [ |
| APOLO | 拟南芥 A. thaliana | 寒冷条件调节根毛伸长 Regulates root hair elongation under cold conditions | 与转录因子WRKY42相互作用 Interacting with the TF WRKY42 | [ |
| TE-lincRNA11195 | 拟南芥 A. thaliana | 参与冷胁迫 Involved in cold stress | 水杨酸刺激反应基因可能是潜在靶标 Salicylic acid-induced response genes may be potential targets | [ |
| DPA lncRNAs | 水稻 O. sativa | 参与冷胁迫 Involved in cold stress | 多聚腺苷酸化来响应冷胁迫 Polyadenylation in response to cold stress | [ |
| lncRNA-Chr03G0008 | 水稻 O. sativa | 参与冷胁迫 Involved in cold stress | 在幼苗中表达响应冷胁迫 Expressing response to cold stress in seedlings | [ |
| lncRNA-SVR | 水稻 O. sativa | 低温下与种子活力相关 Associated with seed viability under low temperature | 顺式基因SAUR家族成员相互作用 Interacting with the cis-gene family member SAUR | [ |
| lncR9A, lncR117, lncR616 | 东农冬麦1号 T. aestivum(Dn1) | 参与冷胁迫 Involved in cold stress | 竞争性结合miR398间接调节CSD1的表达 Indirectly regulates the expression of CSD1 by competitively binding miR398 | [ |
| Traes_2BS_7A04BF5D5 | 小麦 Durum wheat | 参与冷胁迫 Involved in cold stress | 以WCOR413冷驯化基因为靶标响应冷胁迫 Responding to cold stress targeting the cold acclimation gene WCOR413 | [ |
| CRR5 | 木薯 M. esculenta | 参与冷胁迫 Involved in cold stress | 与蛋白激酶基因协同表达响应低温胁迫 Co-expression with protein kinase gene in response to low temperature stress | [ |
| CRIR1 | 木薯 M. esculenta | 参与冷胁迫 Involved in cold stress | 招募MeCSP5来提高mRNA的翻译效率 Recruit MeCSP5 to improve the translation efficiency of mRNA | [ |
| XH123 | 棉花 G. hirsutum | 参与冷胁迫 Involved in cold stress | 基因沉默引起冷调控基因PIF3等差异表达 Gene silencing induced differential expression of cold-regulated genes such as PIF3 | [ |
| 苜蓿 M. truncatula | 参与冷胁迫 Involved in cold stress | 构成lncRNA- MtCBFs调控网络 Construct lncRNA-MtCBFs regulatory network | [ | |
| 番茄 S. lycopersicum | 参与冷胁迫 Involved in cold stress | 竞争与共享miRNA来调控mRNA的表达 Competition and sharing of miRNAs to regulate mRNA expression | [ | |
| DE-lncRNAs | 甜椒 C. annuum | 参与冷胁迫 Involved in cold stress | 调控与冷损伤相关的靶基因 作为miRNAs前体响应冷胁迫 Regulating target genes related to cold damage Responding to cold stress as precursors of miRNAs | [ [ |
| 葡萄 V. vinifera | 参与冷胁迫 Involved in cold stress | 作为miRNAs的靶标 Being targets of miRNAs | [ | |
| 香蕉 M. balbisiana | 参与冷胁迫 Involved in cold stress | 调节类黄酮、蛋白激酶的生物合成以及TCA循环、硫传递系统等途径 Regulating the biosynthesis of flavonoids and protein kinases, as well as pathways such as the TCA cycle and sulfur transfer system | [ |
| [1] | 谢虹, 杨兰, 李忠光. 脯氨酸在植物非生物胁迫耐性形成中的作用[J]. 生物技术通报, 2011(2): 23-27, 60. |
| Xie H, Yang L, Li ZG. The roles of proline in the formation of plant tolerance to abiotic stress[J]. Biotechnol Bull, 2011(2): 23-27, 60. | |
| [2] |
An D, Yang J, Zhang P. Transcriptome profiling of low temperature-treated cassava apical shoots showed dynamic responses of tropical plant to cold stress[J]. BMC Genomics, 2012, 13(1): 64.
doi: 10.1186/1471-2164-13-64 |
| [3] | 录亚丹, 赵战胜, 张庭军. 北疆棉花苗期低温冷害的预防及补救措施[J]. 中国棉花, 2019, 46(6): 41-42. |
| Lu YD, Zhao ZS, Zhang TJ. Prevention and remedial measures of low temperature chilling injury of cotton seedlings in north Xinjiang[J]. China Cotton, 2019, 46(6): 41-42. | |
| [4] |
项洪涛, 郑殿峰, 何宁, 等. 植物对低温胁迫的生理响应及外源脱落酸缓解胁迫效应的研究进展[J]. 草业学报, 2021, 30(1): 208-219.
doi: 10.11686/cyxb2020091 |
| Xiang HT, Zheng DF, He N, et al. Research progress on the physiological response of plants to low temperature and the amelioration effcectiveness of exogenous ABA[J]. Acta Prataculturae Sin, 2021, 30(1): 208-219. | |
| [5] |
Zhou TJ, Zhang WX, Zhang LX, et al. 2021: A year of unprecedented climate extremes in eastern Asia, North America, and Europe[J]. Adv Atmos Sci, 2022, 39(10): 1598-1607.
doi: 10.1007/s00376-022-2063-9 |
| [6] |
Miquel M, James D Jr, Dooner H, et al. Arabidopsis requires polyunsaturated lipids for low-temperature survival[J]. Proc Natl Acad Sci Usa, 1993, 90(13): 6208-6212.
doi: 10.1073/pnas.90.13.6208 pmid: 11607410 |
| [7] |
Los DA. Membrane fluidity and temperature perception[J]. Plant Physiol, 1997, 115(3): 875-879.
pmid: 12223851 |
| [8] |
Alonso A, Queiroz CS, Magalhães AC. Chilling stress leads to increased cell membrane rigidity in roots of coffee(Coffea arabica L.) seedlings[J]. BBA Biomembr, 1997, 1323(1): 75-84.
doi: 10.1016/S0005-2736(96)00177-0 URL |
| [9] | 王兆. 低温胁迫对彩叶草的生理效应及抗寒性研究[D]. 福州: 福建农林大学, 2014. |
| Wang Z. Study on physiological effects and cold resistance of colored leaf grass under low temperature stress[D]. Fuzhou: Fujian Agriculture and Forestry University, 2014. | |
| [10] |
Li XY, Yang Y, Zhang LF, et al. Regulation on contents of endogenous hormones and Asr1 gene expression of maize seedling by exogenous ABA under low-temperature stress[J]. Acta Agron Sin, 2017, 43(1): 141.
doi: 10.3724/SP.J.1006.2017.00141 URL |
| [11] |
Nievola CC, Carvalho CP, Carvalho V, et al. Rapid responses of plants to temperature changes[J]. Temperature, 2017, 4(4): 371-405.
doi: 10.1080/23328940.2017.1377812 pmid: 29435478 |
| [12] |
Ponting CP, Oliver PL, Reik W. Evolution and functions of long noncoding RNAs[J]. Cell, 2009, 136(4): 629-641.
doi: 10.1016/j.cell.2009.02.006 pmid: 19239885 |
| [13] | 原佳沛, 张浩文, 鲁志. 新型长链非编码RNA(lncRNA)的生物信息学研究进展[J]. 生物化学与生物物理进展, 2013, 40(7): 634-640. |
| Yuan JP, Zhang HW, Lu Z. Progress on bioinformatic research of lncRNA[J]. Prog Biochem Biophys, 2013, 40(7): 634-640. | |
| [14] |
Waititu JK, Zhang CY, Liu J, et al. Plant non-coding RNAs: origin, biogenesis, mode of action and their roles in abiotic stress[J]. Int J Mol Sci, 2020, 21(21): 8401.
doi: 10.3390/ijms21218401 URL |
| [15] |
Nejat N, Mantri N. Emerging roles of long non-coding RNAs in plant response to biotic and abiotic stresses[J]. Crit Rev Biotechnol, 2018, 38(1): 93-105.
doi: 10.1080/07388551.2017.1312270 pmid: 28423944 |
| [16] | 孙薏雯, 李金宝, 杨丹丹, 等. 长链非编码RNA在植物中的研究进展[J]. 山东农业大学学报: 自然科学版, 2020, 51(5): 968-974. |
| Sun YW, Li JB, Yang DD, et al. The research progress of long noncoding RNA in plants[J]. J Shandong Agric Univ Nat Sci Ed, 2020, 51(5): 968-974. | |
| [17] |
Wang JJ, Meng XW, Dobrovolskaya OB, et al. Non-coding RNAs and their roles in stress response in plants[J]. Genom Proteom Bioinform, 2017, 15(5): 301-312.
doi: 10.1016/j.gpb.2017.01.007 URL |
| [18] | 李睿, 罗云波. lncRNA及其生物学功能[J]. 农业生物技术学报, 2016, 24(4): 600-612. |
| Li R, Luo YB. LncRNA and its biological function[J]. Journal of Agricultural Biotechnology, 2016, 24(4): 600-612. | |
| [19] |
Hu HY, Wang MJ, Ding YH, et al. Transcriptomic repertoires depict the initiation of lint and fuzz fibres in cotton(Gossypium hirsutum L.)[J]. Plant Biotechnol J, 2018, 16(5): 1002-1012.
doi: 10.1111/pbi.2018.16.issue-5 URL |
| [20] | Salih IHI. 棉纤维发育起始和伸长阶段长链非编码RNA和mRNA的鉴定与表达[D]. 武汉: 华中农业大学, 2017. |
| Salih IHI. Identification and expression of long-chain non-coding RNA and mRNA in the initial and elongation stages of cotton fiber development[D]. Wuhan: Huazhong Agricultural University, 2017. | |
| [21] |
Wunderlich M, Groβ-Hardt R, Schöffl F. Heat shock factor HSFB2a involved in gametophyte development of Arabidopsis thaliana and its expression is controlled by a heat-inducible long non-coding antisense RNA[J]. Plant Mol Biol, 2014, 85(6): 541-550.
doi: 10.1007/s11103-014-0202-0 pmid: 24874772 |
| [22] | 刘琳营. 高温响应lncRNAs调控棉花花药育性的功能分析[D]. 武汉: 华中农业大学, 2021. |
| Liu LY. Functional analysis of high temperature response Lncrnas in regulating cotton anther fertility[D]. Wuhan: Huazhong Agricultural University, 2021. | |
| [23] |
Qin T, Zhao HY, Cui P, et al. A nucleus-localized long non-coding RNA enhances drought and salt stress tolerance[J]. Plant Physiol, 2017, 175(3): 1321-1336.
doi: 10.1104/pp.17.00574 pmid: 28887353 |
| [24] | 刘琳营, 苏晓俊, 闵玲. 植物中长链非编码RNA研究进展综述[J]. 江苏农业科学, 2021, 49(12): 12-19. |
| Liu LY, Su XJ, Min L. Review on research progress of long-chain non-coding RNA in plants[J]. Jiangsu Agric Sci, 2021, 49(12): 12-19. | |
| [25] | 李宁, 王柏柯, 王娟, 等. 植物长链非编码RNA的生物学功能研究进展[J]. 植物生理学报, 2019, 55(10): 1427-1435. |
| Li N, Wang BK, Wang J, et al. Advances in functional research of long non-coding RNAs in plants[J]. Plant Physiol J, 2019, 55(10): 1427-1435. | |
| [26] | 张爱晶, 郭兴玉, 何浩博, 等. 主要农作物lncRNA鉴定与分析研究进展[J]. 广东农业科学, 2020, 47(5): 1-10. |
| Zhang AJ, Guo XY, He HB, et al. Research progress on identification and analysis of lncRNA in major crops[J]. Guangdong Agric Sci, 2020, 47(5): 1-10. | |
| [27] |
Min L, Garbutt C, Tu CQ, et al. Potentials of long noncoding RNAs(LncRNAs)in sarcoma: From biomarkers to therapeutic targets[J]. Int J Mol Sci, 2017, 18(4): 731.
doi: 10.3390/ijms18040731 URL |
| [28] |
谭玉荣, 王丹, 高璇, 等. 植物长链非编码RNA研究进展[J]. 生物技术通报, 2018, 34(10): 1-10.
doi: 10.13560/j.cnki.biotech.bull.1985.2018-0167 |
| Tan YR, Wang D, Gao X, et al. Research advance on plant long noncoding RNAs[J]. Biotechnol Bull, 2018, 34(10): 1-10. | |
| [29] | 艾雯, 李娇卓, 闵德栋, 等. 果蔬作物中长链非编码RNA研究进展[J]. 北方园艺, 2020(17): 124-130. |
| Ai W, Li JZ, Min DD, et al. Research progress of long noncoding RNAs on fruits and vegetables crops[J]. North Hortic, 2020(17): 124-130. | |
| [30] |
Rai MI, Alam M, Lightfoot DA, et al. Classification and experimental identification of plant long non-coding RNAs[J]. Genomics, 2019, 111(5): 997-1005.
doi: S0888-7543(18)30245-3 pmid: 29679643 |
| [31] |
Gao CX, Zheng XW, Li HB, et al. Roles of lncRNAs in rice: advances and challenges[J]. Rice Sci, 2020, 27(5): 384-395.
doi: 10.1016/j.rsci.2020.03.003 |
| [32] | Li S, Nayar S, Jia H, et al. The Arabidopsis hypoxia inducible AtR8 long non-coding RNA also contributes to plant defense and root elongation coordinating with WRKY genes under low levels of salicylic acid[J]. Noncoding RNA, 2020, 6(1): E8. |
| [33] |
Ding JH, Lu Q, Ouyang YD, et al. A long noncoding RNA regulates photoperiod-sensitive male sterility, an essential component of hybrid rice[J]. Proc Natl Acad Sci USA, 2012, 109(7): 2654-2659.
doi: 10.1073/pnas.1121374109 pmid: 22308482 |
| [34] |
Ariel F, Jegu T, Latrasse D, et al. Noncoding transcription by alternative RNA polymerases dynamically regulates an auxin-driven chromatin loop[J]. Mol cell, 2014, 55(3): 383-396.
doi: 10.1016/j.molcel.2014.06.011 pmid: 25018019 |
| [35] |
Tian YK, Hou YK, Song Y. LncRNAs elevate plant adaptation under low temperature by maintaining local chromatin landscape[J]. Plant Signal Behav, 2022, 17(1): 2014677.
doi: 10.1080/15592324.2021.2014677 URL |
| [36] |
Campalans A, Kondorosi A, Crespi M. Enod40, a short open reading frame-containing mRNA, induces cytoplasmic localization of a nuclear RNA binding protein in Medicago truncatula[J]. Plant Cell, 2004, 16(4): 1047-1059.
pmid: 15037734 |
| [37] |
Wang Y, Luo XJ, Sun F, et al. Overexpressing lncRNA LAIR increases grain yield and regulates neighbouring gene cluster expression in rice[J]. Nat Commun, 2018, 9(1): 3516.
doi: 10.1038/s41467-018-05829-7 pmid: 30158538 |
| [38] |
Jha UC, Nayyar H, Jha R, et al. Long non-coding RNAs: Emerging players regulating plant abiotic stress response and adaptation[J]. BMC Plant Biol, 2020, 20(1): 466.
doi: 10.1186/s12870-020-02595-x pmid: 33046001 |
| [39] |
Franco-Zorrilla JM, Valli A, Todesco M, et al. Target mimicry provides a new mechanism for regulation of microRNA activity[J]. Nat Genet, 2007, 39(8): 1033-1037.
doi: 10.1038/ng2079 pmid: 17643101 |
| [40] | Fedak H, Palusinska M, Krzyczmonik K, et al. Control of seed dormancy in Arabidopsis by a cis-acting noncoding antisense transcript[J]. Proc Natl Acad Sci USA, 2016, 113(48): E7846-E7855. |
| [41] |
Brockdorff N, Ashworth A, Kay GF, et al. Conservation of position and exclusive expression of mouse Xist from the inactive X chromosome[J]. Nature, 1991, 351(6324): 329-331.
doi: 10.1038/351329a0 |
| [42] |
Brown CJ, Ballabio A, Rupert JL, et al. A gene from the region of the human X inactivation centre is expressed exclusively from the inactive X chromosome[J]. Nature, 1991, 349(6304): 38-44.
doi: 10.1038/349038a0 |
| [43] |
Lipshitz HD, Peattie DA, Hogness DS. Novel transcripts from the Ultrabithorax domain of the bithorax complex[J]. Genes Dev, 1987, 1(3): 307-322.
doi: 10.1101/gad.1.3.307 URL |
| [44] |
Moison M, Pacheco JM, Lucero L, et al. The lncRNA APOLO interacts with the transcription factor WRKY42 to trigger root hair cell expansion in response to cold[J]. Mol Plant, 2021, 14(6): 937-948.
doi: 10.1016/j.molp.2021.03.008 pmid: 33689931 |
| [45] | Liu WH, Cheng CZ, Lin YL, et al. Genome-wide identification and characterization of mRNAs and lncRNAs involved in cold stress in the wild banana(Musa itinerans)[J]. PLoS One, 2018, 13(7): e0200002. |
| [46] |
Jeon J, Kim J. Cold stress signaling networks in Arabidopsis[J]. J Plant Biol, 2013, 56(2): 69-76.
doi: 10.1007/s12374-013-0903-y URL |
| [47] |
Tang K, Zhao L, Ren Y, et al. The transcription factor ICE1 functions in cold stress response by binding to the promoters of CBF and COR genes[J]. J Integr Plant Biol, 2020, 62(3): 258-263.
doi: 10.1111/jipb.v62.3 URL |
| [48] |
Kindgren P, Ard R, Ivanov M, et al. Transcriptional read-through of the long non-coding RNA SVALKA governs plant cold acclimation[J]. Nat Commun, 2018, 9(1): 4561.
doi: 10.1038/s41467-018-07010-6 pmid: 30385760 |
| [49] |
Liu GC, Liu FX, Wang Y, et al. A novel long noncoding RNA CIL1 enhances cold stress tolerance in Arabidopsis[J]. Plant Sci, 2022, 323: 111370.
doi: 10.1016/j.plantsci.2022.111370 URL |
| [50] |
Jampala P, Garhewal A, Lodha M. Functions of long non-coding RNA in Arabidopsis thaliana[J]. Plant Signal Behav, 2021, 16(9): 1925440.
doi: 10.1080/15592324.2021.1925440 URL |
| [51] |
Swiezewski S, Liu FQ, Magusin A, et al. Cold-induced silencing by long antisense transcripts of an Arabidopsis polycomb target[J]. Nature, 2009, 462(7274): 799-802.
doi: 10.1038/nature08618 |
| [52] |
Ding YL, Shi YT, Yang SH. Molecular regulation of plant responses to environmental temperatures[J]. Mol plant, 2020, 13(4): 544-564.
doi: S1674-2052(20)30034-4 pmid: 32068158 |
| [53] |
Severing E, Faino L, Jamge S, et al. Arabidopsis thaliana ambient temperature responsive lncRNAs[J]. BMC Plant Biol, 2018, 18(1): 145.
doi: 10.1186/s12870-018-1362-x pmid: 30005624 |
| [54] |
Heo JB, Sung S. Vernalization-mediated epigenetic silencing by a long intronic noncoding RNA[J]. Science, 2011, 331(6013): 76-79.
doi: 10.1126/science.1197349 pmid: 21127216 |
| [55] |
Kim DH, Sung S. Vernalization-triggered intragenic chromatin loop formation by long noncoding RNAs[J]. Dev Cell, 2017, 40(3): 302-312.
doi: 10.1016/j.devcel.2016.12.021 URL |
| [56] | Tian Y, Zheng H, Zhang F, et al. PRC2 recruitment and H3K27me3 deposition at FLC require FCA binding of COOLAIR[J]. Sci Adv, 2019, 5(4): eaau7246. |
| [57] |
Csorba T, Questa JI, Sun QW, et al. Antisense COOLAIR mediates the coordinated switching of chromatin states at FLC during vernalization[J]. Proc Natl Acad Sci USA, 2014, 111(45): 16160-16165.
doi: 10.1073/pnas.1419030111 pmid: 25349421 |
| [58] |
Pan YH, Liang HF, Gao LJ, et al. Transcriptomic profiling of germinating seeds under cold stress and characterization of the cold-tolerant gene LTG5 in rice[J]. BMC Plant Biol, 2020, 20(1): 371.
doi: 10.1186/s12870-020-02569-z |
| [59] |
Yuan JP, Li JR, Yang Y, et al. Stress-responsive regulation of long non-coding RNA polyadenylation in Oryza sativa[J]. Plant J, 2018, 93(5): 814-827.
doi: 10.1111/tpj.2018.93.issue-5 URL |
| [60] |
Shin SY, Jeong JS, Lim JY, et al. Transcriptomic analyses of rice(Oryza sativa)genes and non-coding RNAs under nitrogen starvation using multiple omics technologies[J]. BMC genomics, 2018, 19(1): 532.
doi: 10.1186/s12864-018-4897-1 |
| [61] |
Gao QL, Liu JZ, Weng HB, et al. A long noncoding RNA derived from lncRNA-mRNA networks modulates seed vigor[J]. Int J Mol Sci, 2022, 23(16): 9472.
doi: 10.3390/ijms23169472 URL |
| [62] | Leng Y, Sun J, Wang JG, et al. Genome-wide lncRNAs identification and association analysis for cold-responsive genes at the booting stage in rice(Oryza sativa L.)[J]. Plant Genome, 2020, 13(2): e20020. |
| [63] | 卢秋巍. 冬小麦抗寒lncRNA筛选及与tae-miR398应答低温胁迫的互作研究[D]. 哈尔滨: 东北农业大学, 2018. |
| Lu QW. Screening of cold-resistant lncRNA in winter wheat and its interaction with tae-miR398 in response to low temperature stress[D]. Harbin:Northeast Agricultural University, 2018. | |
| [64] |
Díaz ML, Soresi DS, Basualdo J, et al. Transcriptomic response of durum wheat to cold stress at reproductive stage[J]. Mol Biol Rep, 2019, 46(2): 2427-2445.
doi: 10.1007/s11033-019-04704-y pmid: 30798485 |
| [65] |
Lu QW, Guo FY, Xu QH, et al. LncRNA improves cold resistance of winter wheat by interacting with miR398[J]. Funct Plant Biol, 2020, 47(6): 544-557.
doi: 10.1071/FP19267 pmid: 32345432 |
| [66] |
Suksamran R, Saithong T, Thammarongtham C, et al. Genomic and transcriptomic analysis identified novel putative cassava lncRNAs involved in cold and drought stress[J]. Genes, 2020, 11(4): 366.
doi: 10.3390/genes11040366 URL |
| [67] | 沈婕, 董世满, 李淑霞, 等. 木薯中lncRNA-CRR5在低温胁迫下的功能分析[J]. 热带作物学报, 2021, 42(1): 40-46. |
| Shen J, Dong SM, Li SX, et al. Identification of lncRNA-CRR5 in response to low temperature stress in cassava[J]. Chin J Trop Crops, 2021, 42(1): 40-46. | |
| [68] |
Li S, Cheng Z, Dong S, et al. Global identification of full-length cassava lncRNAs unveils the role of cold-responsive intergenic lncRNA 1 in cold stress response[J]. Plant Cell Environ, 2022, 45(2): 412-426.
doi: 10.1111/pce.v45.2 URL |
| [69] | 李玉强. 低温冷害对尉犁县棉花生长发育的影响分析[J]. 农业灾害研究, 2018, 8(5): 42-43. |
| Li YQ. Effect of chilling damage on growth and development of cotton in Yuli County[J]. J Agric Catastrophol, 2018, 8(5): 42-43. | |
| [70] | 曹泽毅. 棉花冷害相关长链非编码RNA的筛选与功能分析[D]. 杭州: 浙江大学, 2018. |
| Cao ZY. Screening and functional analysis of long-chain non-coding RNA related to cotton chilling injury[D]. Hangzhou: Zhejiang University, 2018. | |
| [71] |
Liu ZY, Li XL, Li F, et al. Mechanisms underlying the effects of fall dormancy on the cold acclimation and winter hard-iness of Medicago sativa[J]. Chin J Plant Ecol, 2015, 39(6): 635-648.
doi: 10.17521/cjpe.2015.0061 URL |
| [72] |
Zhao MG, Wang TZ, Sun TY, et al. Identification of tissue-specific and cold-responsive lncRNAs in Medicago truncatula by high-throughput RNA sequencing[J]. BMC Plant Biol, 2020, 20(1): 99.
doi: 10.1186/s12870-020-2301-1 |
| [73] | Zhao DY, Shen L, Fan B, et al. Physiological and genetic properties of tomato fruits from 2 cultivars differing in chilling tolerance at cold storage[J]. J Food Sci, 2009, 74(5): C348-C352. |
| [74] |
Wang YX, Gao LP, Zhu BZ, et al. Integrative analysis of long non-coding RNA acting as ceRNAs involved in chilling injury in tomato fruit[J]. Gene, 2018, 667: 25-33.
doi: S0378-1119(18)30513-4 pmid: 29753809 |
| [75] |
Zuo JH, Wang YX, Zhu BZ, et al. Analysis of the coding and non-coding RNA transcriptomes in response to bell pepper chilling[J]. Int J Mol Sci, 2018, 19(7): 2001.
doi: 10.3390/ijms19072001 URL |
| [76] |
Baruah PM, Krishnatreya DB, Bordoloi KS, et al. Genome wide identification and characterization of abiotic stress responsive lncRNAs in Capsicum annuum[J]. Plant Physiol Biochem, 2021, 162: 221-236.
doi: 10.1016/j.plaphy.2021.02.031 URL |
| [77] |
Wang P, Dai L, Ai J, et al. Identification and functional prediction of cold-related long non-coding RNA(lncRNA)in grapevine[J]. Sci Rep, 2019, 9(1): 1-15.
doi: 10.1038/s41598-018-37186-2 |
| [1] | FENG Shan-shan, WANG Lu, ZHOU Yi, WANG You-ping, FANG Yu-jie. Research Progresses on WOX Family Genes in Regulating Plant Development and Abiotic Stress Response [J]. Biotechnology Bulletin, 2023, 39(5): 1-13. |
| [2] | SHI Jian-lei, ZAI Wen-shan, SU Shi-wen, FU Cun-nian, XIONG Zi-li. Identification and Expression Analysis of miRNA Related to Bacterial Wilt Resistance in Tomato [J]. Biotechnology Bulletin, 2023, 39(5): 233-242. |
| [3] | ZHANG Xin-bo, CUI Hao-liang, SHI Pei-hua, GAO Jin-chun, ZHAO Shun-ran, TAO Chen-yu. Research Progress in Low-input Chromatin Immunoprecipitation Assay [J]. Biotechnology Bulletin, 2023, 39(4): 227-235. |
| [4] | WANG Hai-long, LI Yu-qian, WANG Bo, XING Guo-fang, ZHANG Jie-wei. Isolation and Expression Analysis of SiMAPK3 in Setaria italica L. [J]. Biotechnology Bulletin, 2023, 39(3): 123-132. |
| [5] | ZHAO Meng-liang, GUO Yi-ting, REN Yan-jing. Identification and Analysis of WRKY Transcription Factor Family Genes in Helianthus tuberosus [J]. Biotechnology Bulletin, 2023, 39(2): 116-125. |
| [6] | LV Yu-jing, WU Dan-dan, KONG Chun-yan, YANG Yu, GONG Ming. Genome-wide Identification of XTH Gene Family and Their Interacting miRNAs and Possible Roles in Low Temperature Adaptation in Jatropha curcas L. [J]. Biotechnology Bulletin, 2023, 39(2): 147-160. |
| [7] | ZHANG Xiao-yan, YANG Shu-hua, DING Yang-lin. Molecular Mechanism of Cold Signal Perception and Transduction in Plants [J]. Biotechnology Bulletin, 2023, 39(11): 28-35. |
| [8] | HAN Fang-ying, HU Xin, WANG Nan-nan, XIE Yu-hong, WANG Xiao-yan, ZHU Qiang. Research Progress in Response of DREBs to Abiotic Stress in Plant [J]. Biotechnology Bulletin, 2023, 39(11): 86-98. |
| [9] | FENG Ce-ting, JIANG Lyu, LIU Xing-ying, LUO Le, PAN Hui-tang, ZHANG Qi-xiang, YU Chao. Identification of the NAC Gene Family in Rosa persica and Response Analysis Under Drought Stress [J]. Biotechnology Bulletin, 2023, 39(11): 283-296. |
| [10] | CHEN Hao-ting, ZHANG Yu-jing, LIU Jie, DAI Ze-min, LIU Wei, SHI Yu, ZHANG Yi, LI Tian-lai. Functional Analysis of WRKY6 Gene in Tomato Under Low-phosphorus Stress [J]. Biotechnology Bulletin, 2023, 39(10): 136-147. |
| [11] | YIN Guo-ying, LIU Chang, CHANG Yong-chun, YU Wang-jie, WANG Bing, ZHANG Pan, GUO Yu-shuang. Identification of the Cysteine Protease Family and Corresponding miRNAs in Nicotiana tabacum L. and Their Responses to PVY [J]. Biotechnology Bulletin, 2023, 39(10): 184-196. |
| [12] | YU Xiao-ling, LI Wen-bin, LI Zhi-bo, RUAN Meng-bin. Cold Resistance Function Analysis of Cassava MeMYC2.2 [J]. Biotechnology Bulletin, 2023, 39(1): 224-231. |
| [13] | LI Jian-jian, HE Chen-jing, HUANG Xiao-ping, XIANG Tai-he. Research Progress in the Regulation of Development and Stress Response by Long Non-coding RNAs in Plants [J]. Biotechnology Bulletin, 2023, 39(1): 48-58. |
| [14] | WANG Nan-nan, WANG Wen-jia, ZHU Qiang. Research Progress of microRNAs in Plant Stress Responses [J]. Biotechnology Bulletin, 2022, 38(8): 1-11. |
| [15] | CHEN Gui-fang, YANG Jia-yi, GAO Yun-hua, REN Ge. Research Progress in Chromatin Immunoprecipitation Followed by Sequencing [J]. Biotechnology Bulletin, 2022, 38(7): 40-50. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||