








Biotechnology Bulletin ›› 2026, Vol. 42 ›› Issue (4): 216-226.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0820
Previous Articles Next Articles
JIANG Xin-hua1,2,3( ), FANG Tian-yu1,2, ZHANG Jing-jing1,2, LI Xiang-yuan1,2, ZHANG Bang-yue1,2, LIAO Xiao-shan4, RONG Duo-yan1,2(
), FANG Tian-yu1,2, ZHANG Jing-jing1,2, LI Xiang-yuan1,2, ZHANG Bang-yue1,2, LIAO Xiao-shan4, RONG Duo-yan1,2( )
)
Received:2025-07-30
Online:2026-04-26
Published:2026-04-30
Contact:
RONG Duo-yan
E-mail:1978526350@qq.com;rongduoyan@163.com
JIANG Xin-hua, FANG Tian-yu, ZHANG Jing-jing, LI Xiang-yuan, ZHANG Bang-yue, LIAO Xiao-shan, RONG Duo-yan. Identification and Functional Analysis of the MpPP2A-C Gene in Marchantia polymorpha[J]. Biotechnology Bulletin, 2026, 42(4): 216-226.
基因名称 Gene name | 基因ID Locus ID | 分子量 Molecular weight (Da) | 等电点 Point isoelectric | 脂肪系数 Aliphatic index | 不稳定系数 Instability index | 亲水性 GRAVY | 亚细胞定位 Subcellular localization |
|---|---|---|---|---|---|---|---|
| MpPP2A-C1 | Mp6g15310.1 | 35 243.00 | 5.08 | 82.48 | 44.06 | -0.332 | 细胞质 Cytoplasm 线粒体 Mitochondrion 细胞核 Nucleus |
| MpPP2A-C2 | Mp2g13820.1 | 34 746.72 | 5.35 | 89.08 | 44.75 | -0.210 | 细胞质 Cytoplasm 线粒体 Mitochondrion 细胞核 Nucleus |
| MpPP2A-C3 | Mp5g22560.1 | 34 719.48 | 5.07 | 82.28 | 35.67 | -0.240 | 细胞质 Cytoplasm 细胞核 Nucleus |
Table 1 Basic information of PP2A-C in M. polymorpha
基因名称 Gene name | 基因ID Locus ID | 分子量 Molecular weight (Da) | 等电点 Point isoelectric | 脂肪系数 Aliphatic index | 不稳定系数 Instability index | 亲水性 GRAVY | 亚细胞定位 Subcellular localization |
|---|---|---|---|---|---|---|---|
| MpPP2A-C1 | Mp6g15310.1 | 35 243.00 | 5.08 | 82.48 | 44.06 | -0.332 | 细胞质 Cytoplasm 线粒体 Mitochondrion 细胞核 Nucleus |
| MpPP2A-C2 | Mp2g13820.1 | 34 746.72 | 5.35 | 89.08 | 44.75 | -0.210 | 细胞质 Cytoplasm 线粒体 Mitochondrion 细胞核 Nucleus |
| MpPP2A-C3 | Mp5g22560.1 | 34 719.48 | 5.07 | 82.28 | 35.67 | -0.240 | 细胞质 Cytoplasm 细胞核 Nucleus |
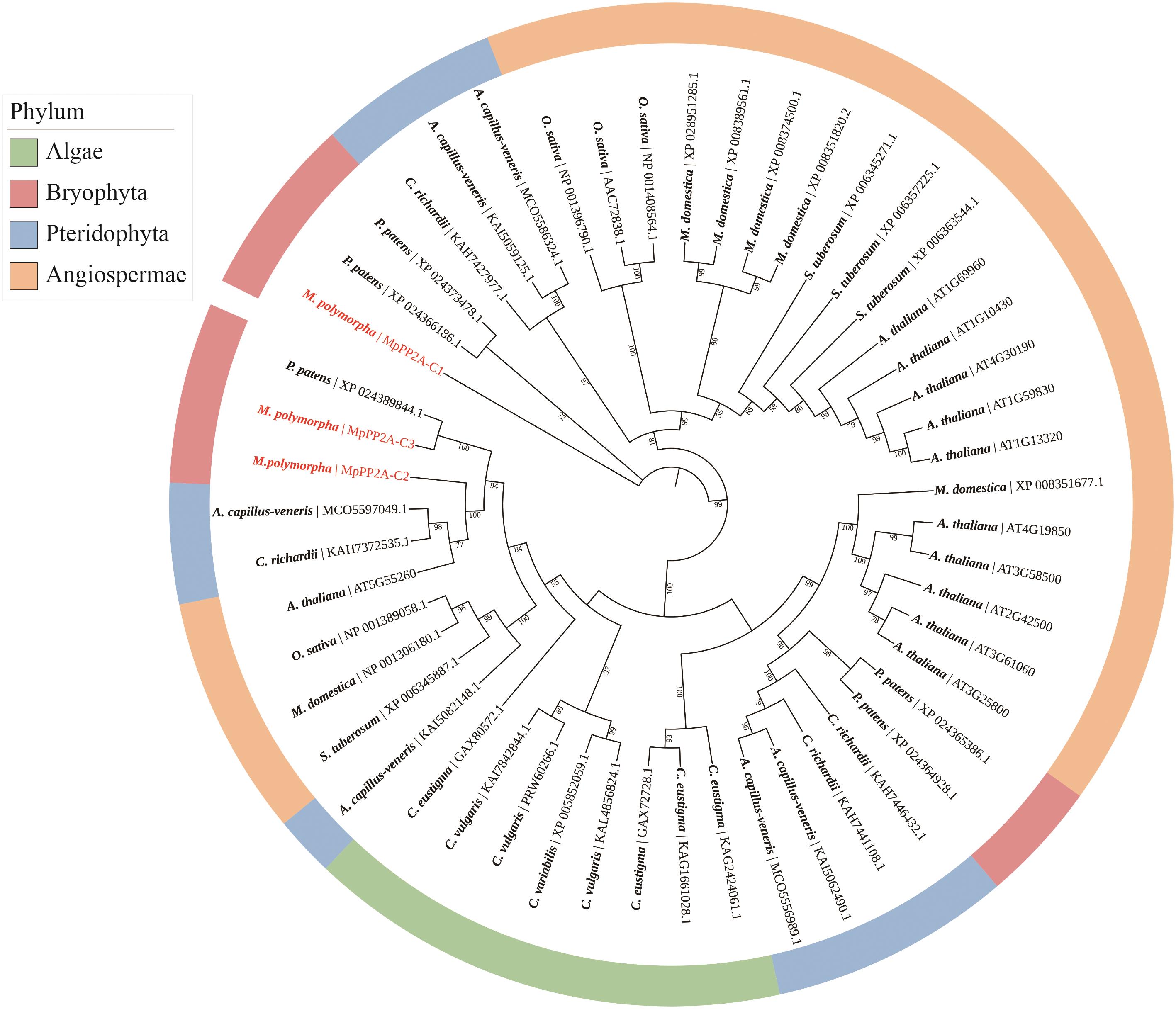
Fig. 1 Evolutionary analysis of PP2A-C amino acid sequences in M. polymorpha and other speciesC. Eustigma: Chlamydomonas eustigma; C. vulgaris: Chlorellavulgaris; M. Polymorpha: Marchantia polymorpha; P. patens: Physcomitrella patens; A. capillus-veneris: Adiantum capillus-veneris; C. richardii: Ceratopteris richardii; A. thaliana: Arabidopsis thaliana; O. sativa: Oryza sativa; M. domestica: Malus domestica; S. tuberosum: Solanum tuberosum
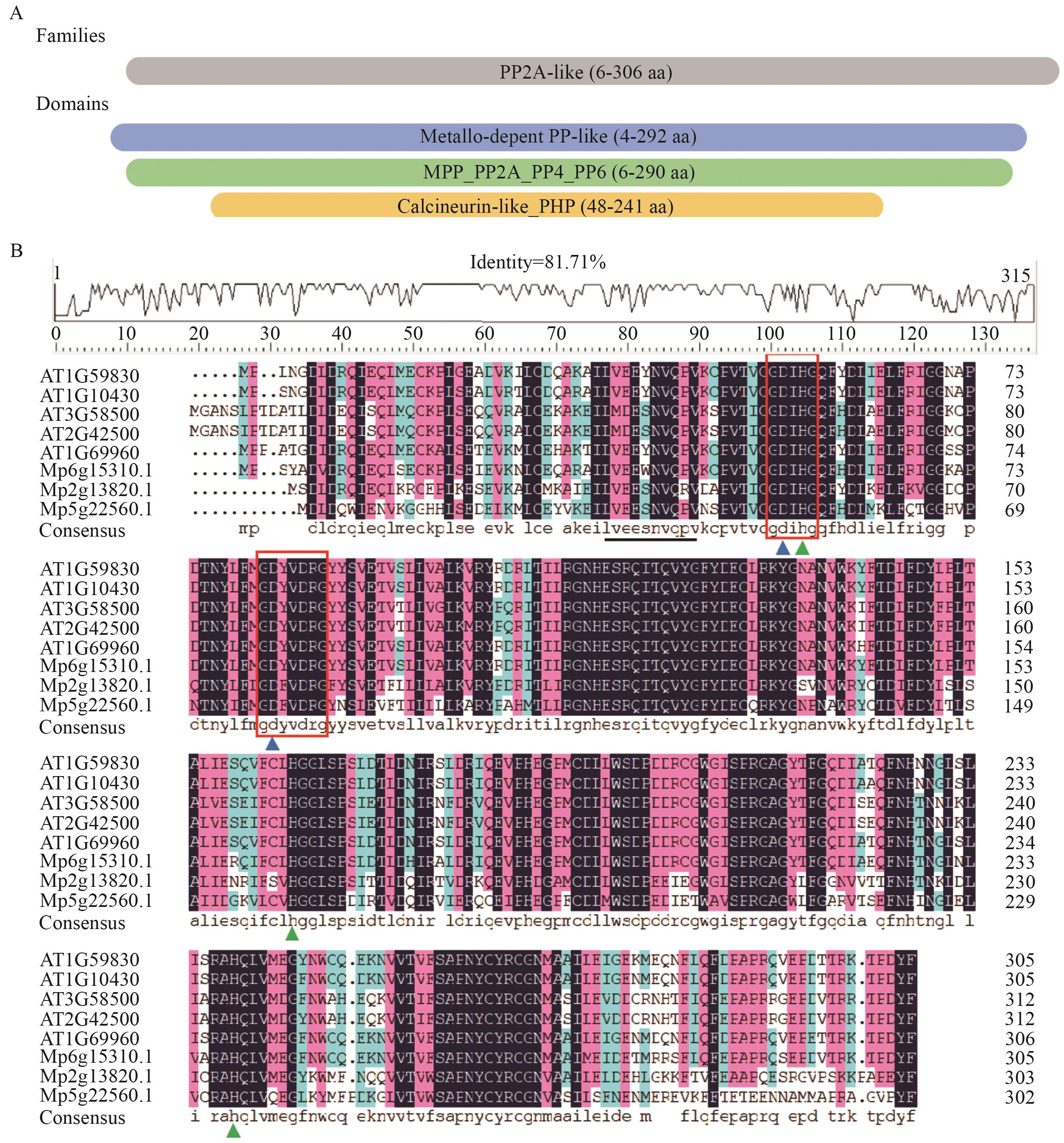
Fig. 2 Conserved domain and sequence alignment of PP2A-C proteins between A. thaliana and M. polymorphaA: Schematic diagram of structural domains in proteins. PP2A-like: Serine/Threonine protein phosphatase PP2A-like. Metallo-depent PP-like: Metallo-dependent phosphatase-like. MPP_PP2A_PP4_PP6: PP2A, PP4, and PP6 phosphoprotein phosphatases, metallophosphatase domain. Calcineurin-like_PHP: Calcineurin-like, phosphoesterase domain. B: Multiple sequence alignment of proteins. The conserved motifs of the catalytic core domain (GDXHG and GDXDRG) are indicated by red boxes; the conserved motif of the A-subunit binding domain (LVEEYNVQEV) is underlined in blue. The blue triangles indicate the metal ion binding site Asp, while the green triangles denote the metal ion binding site Glu
蛋白质名称 Protein name | 氨基酸数量 Number of amino acids (aa) | 蛋白质二级结构 Secondary structure of protein | 蛋白质三级结构 Tertiary structure of protein | |||||
|---|---|---|---|---|---|---|---|---|
α-螺旋 Alpha helix(%) | 延伸链 Extended strand(%) | 不规则卷曲 Random coil(%) | 相似性 Identity (%) | 全局模型质量评估 GMQE | 基于距离约束的定性模型能量分析全局分数 QMEANDisCo Global | |||
| MpPP2A-C1 | 306 | 37.25 | 15.03 | 47.71 | 82.89 | 0.91 | 0.89±0.05 | |
| MpPP2A-C2 | 304 | 39.47 | 14.80 | 45.72 | 65.35 | 0.88 | 0.83±0.05 | |
| MpPP2A-C3 | 303 | 38.28 | 16.18 | 45.54 | 58.14 | 0.88 | 0.83±0.05 | |
Table 2 Basic information on the secondary and tertiary structures of PP2A-C in M. polymorpha
蛋白质名称 Protein name | 氨基酸数量 Number of amino acids (aa) | 蛋白质二级结构 Secondary structure of protein | 蛋白质三级结构 Tertiary structure of protein | |||||
|---|---|---|---|---|---|---|---|---|
α-螺旋 Alpha helix(%) | 延伸链 Extended strand(%) | 不规则卷曲 Random coil(%) | 相似性 Identity (%) | 全局模型质量评估 GMQE | 基于距离约束的定性模型能量分析全局分数 QMEANDisCo Global | |||
| MpPP2A-C1 | 306 | 37.25 | 15.03 | 47.71 | 82.89 | 0.91 | 0.89±0.05 | |
| MpPP2A-C2 | 304 | 39.47 | 14.80 | 45.72 | 65.35 | 0.88 | 0.83±0.05 | |
| MpPP2A-C3 | 303 | 38.28 | 16.18 | 45.54 | 58.14 | 0.88 | 0.83±0.05 | |
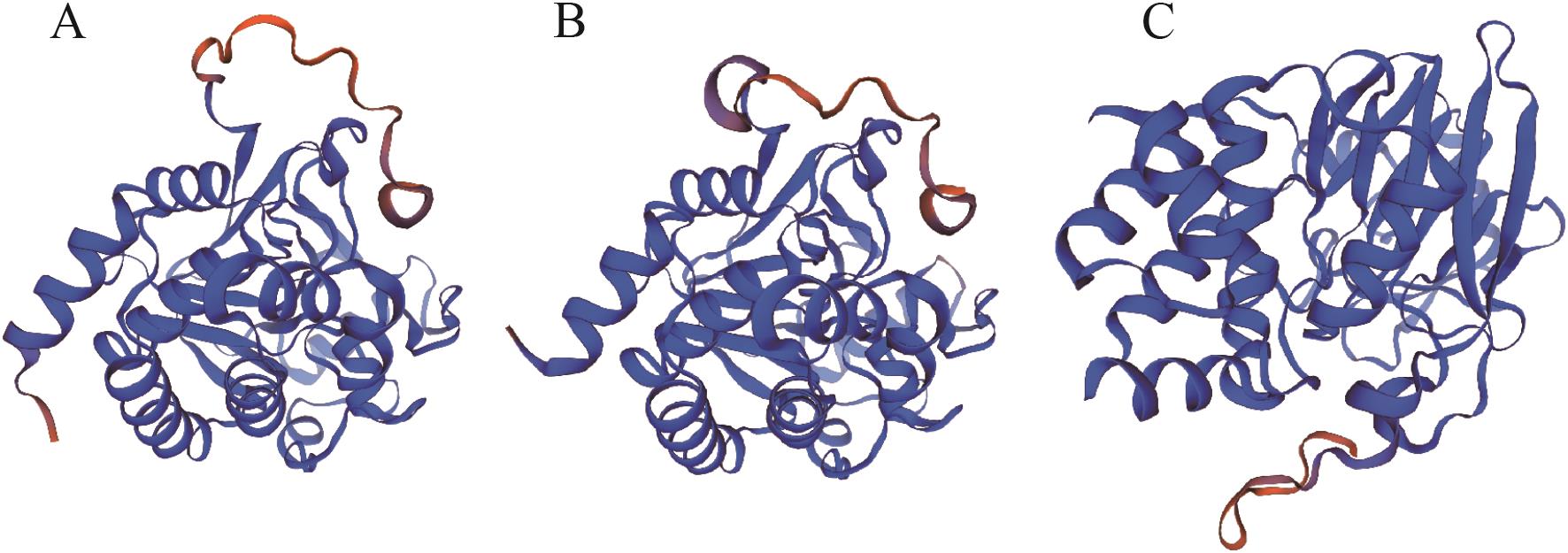
Fig. 3 Tertiary structure of MpPP2A-C proteinA: MpPP2A-C1. B: MpPP2A-C2. C: MpPP2A-C3. All structures are colored by the confidence gradient, with blue indicating high confidence (trustworthy) and red indicating low confidence (unreliable)
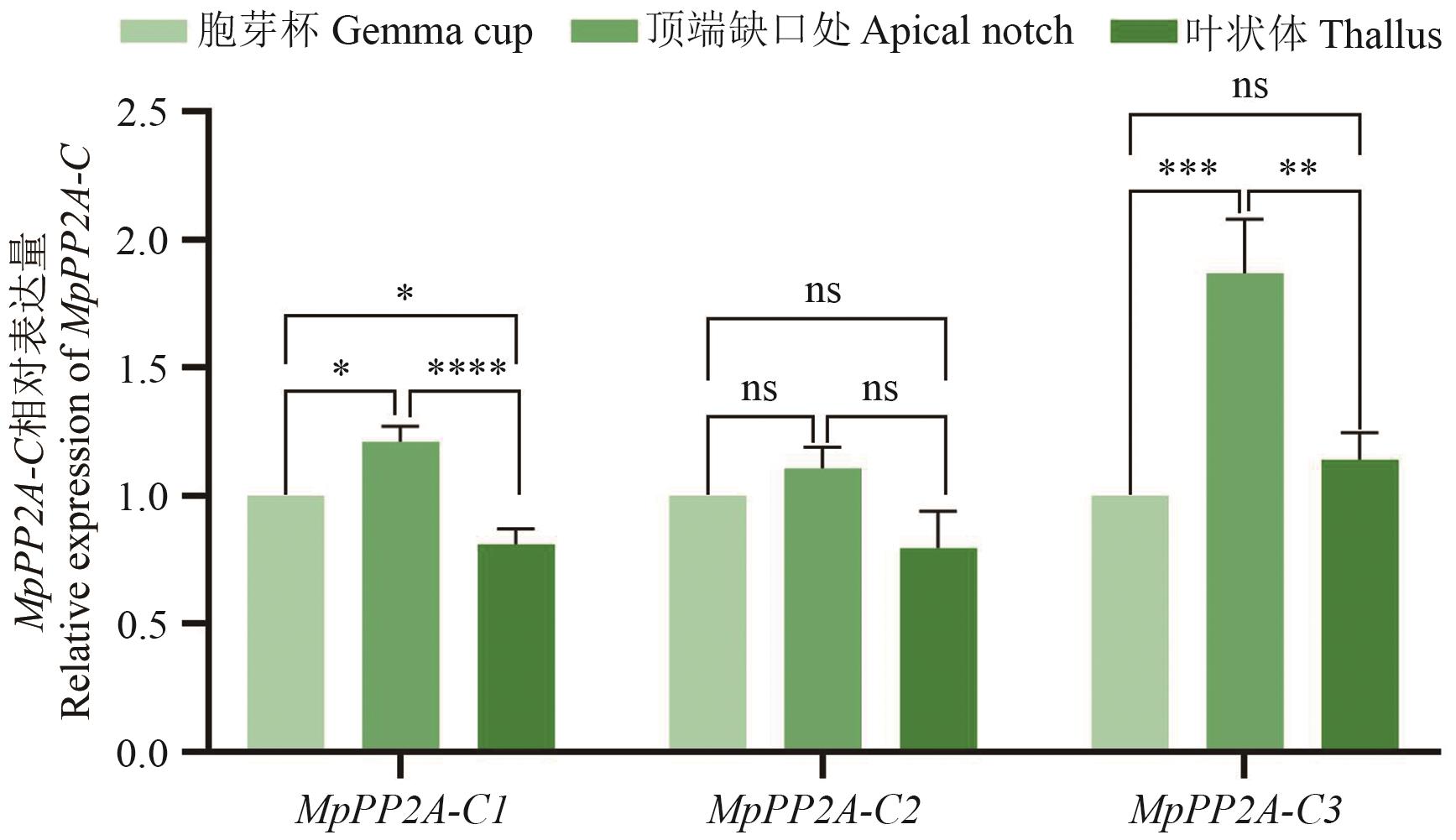
Fig. 4 Differences of expressions of MpPP2A-C in different tissue parts of M. polymorphaValues are the average±standard error, ****P<0.000 1 (Student's t-test)
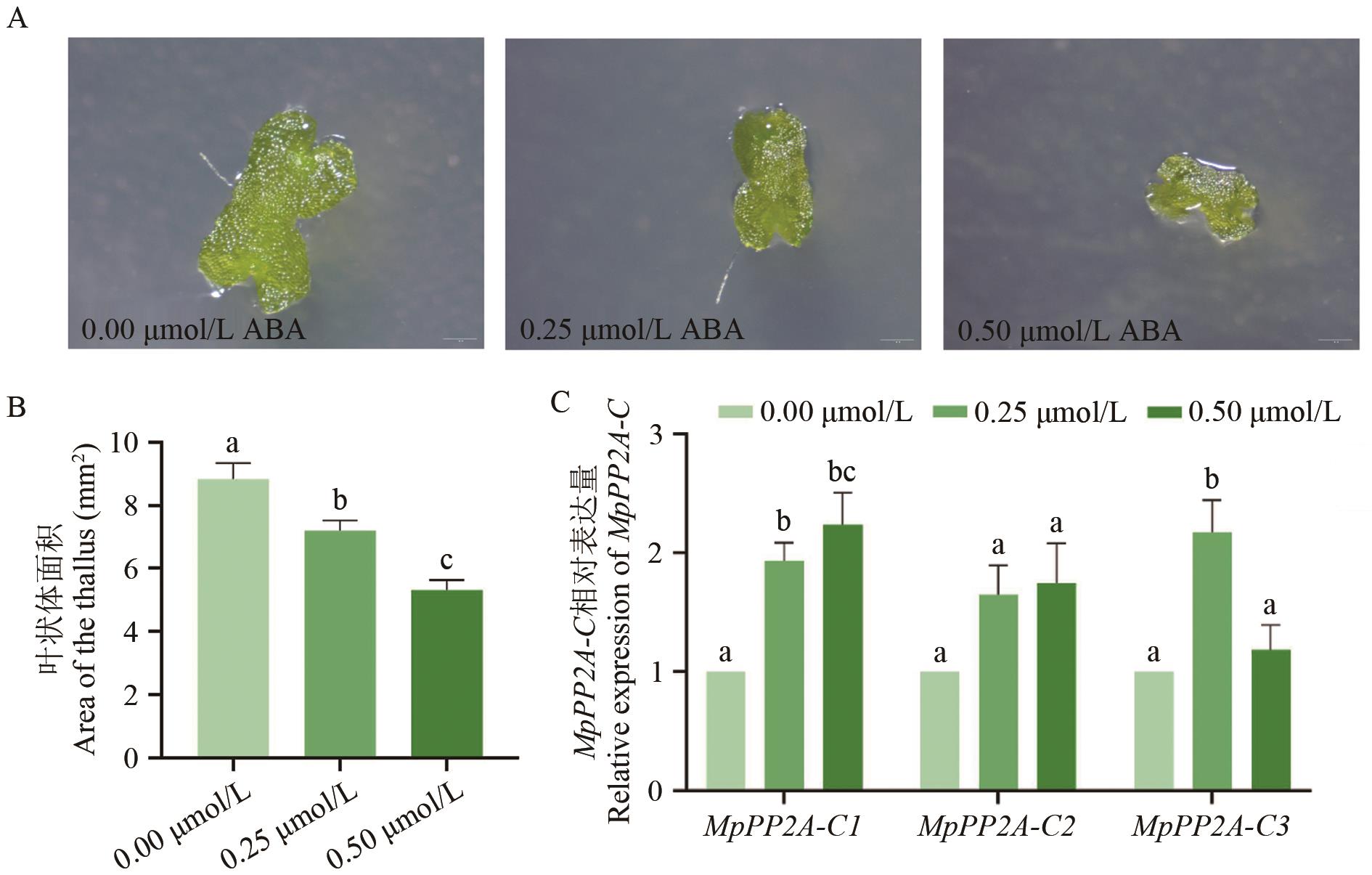
Fig. 5 Phenotype and expression analysis of MpPP2A-C gene treated with abscisic acid at different concentrationsA: The comparison of the surface type of the phyllodes of the plant under different concentrations of ABA, and the scale length is 1 mm. B: Statistics of the phyllosoma area under different concentrations of ABA treatment. C: Expressions of MpPP2A-C gene under different ABA concentrations. Values are the average ± standard error, the lowercase letters indicate significant differences between groups (P<0.05). Statistical analysis was performed using one-way analysis of variance (ANOVA) combined with Duncan’s multiple comparison test
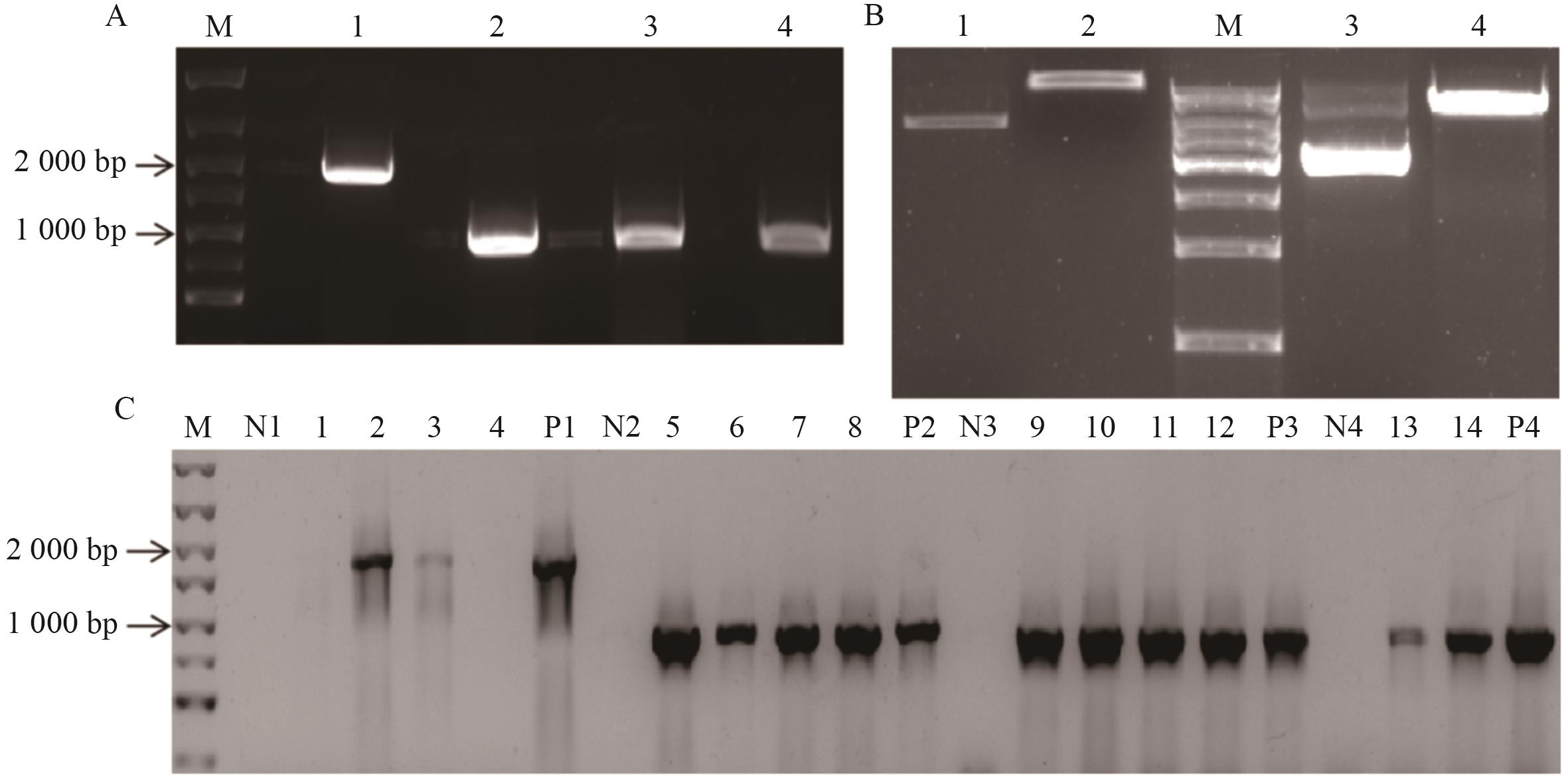
Fig. 6 Construction of yeast two-hybrid vectorsA: Amplification of the target fragment of the MpPP2A-A/C1/C2/C3 gene (M: DNA Marker; 1-4: PCR amplified products of the MpPP2A-A/C1/C2/C3 genes). B: Enzymatic digestion of pGADT7 and pGBKT7 plasmids (M: DNA Marker; 1: Undigested pGADT7 plasmid; 2: pGADT7 plasmid after enzymatic digestion; 3: Undigested pGBKT7 plasmid; 4: pGBKT7 plasmid after enzymatic digestion). C: Construction of the pGADT7-MpPP2A-A/C1 vector and pGBKT7-MpPP2A-C2/C3 vectors (M: DNA Marker; 1-4: DH5α-pGADT7-MpPP2A-A bacterial liquid, N1: Negative control, P1: Positive control (the target fragment of the MpPP2A-A); 5-8: DH5α-pGADT7-MpPP2A-C1 bacterial liquid, N2: Negative control, P2: Positive control (the target fragment of the MpPP2A-C1); 9-12: DH5α-pGBKT7-MpPP2A-C2 bacterial liquid. N3: Negative control. P3: Positive control (the target fragment of the MpPP2A-C2); 13-14: DH5α-pGBKT7-MpPP2A-C3 bacterial liquid. N4: Negative control. P4: Positive control (the target fragment of the MpPP2A-C3)
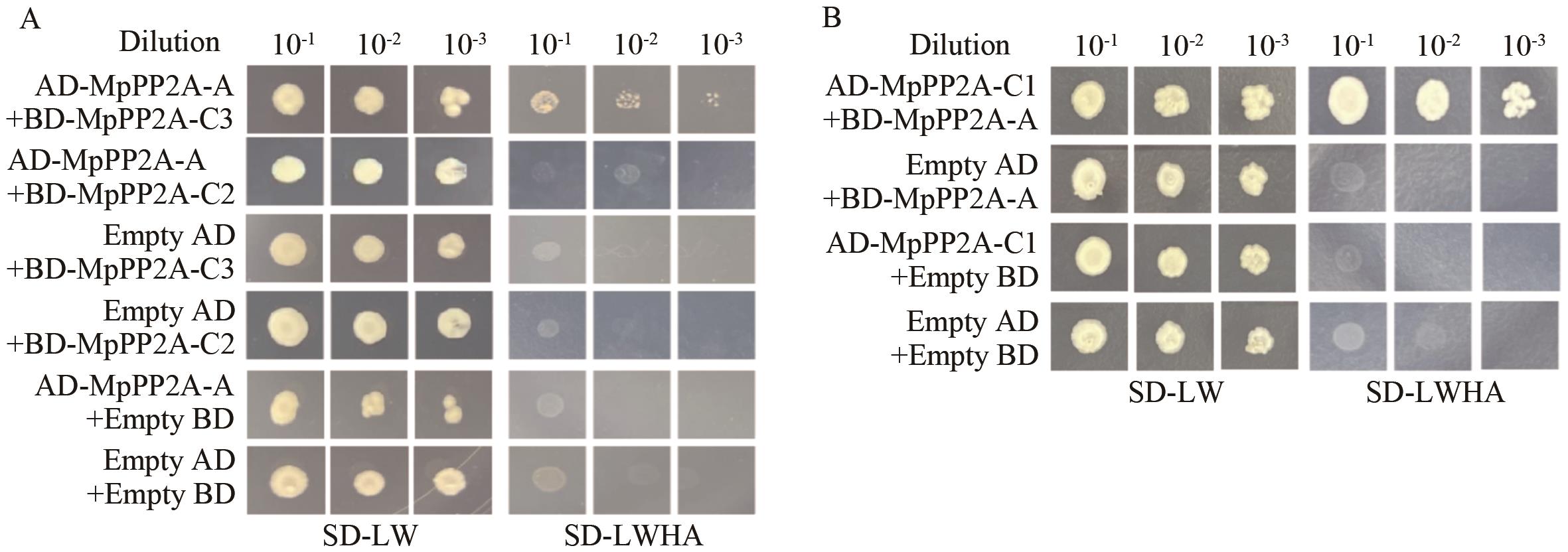
Fig. 7 Growth of colonies in the defective mediumA: Verification of the interaction between AD-MpPP2A-A and BD-MpPP2A-C2/C3. B: Verification of the interaction between AD-MpPP2A-C1 and BD-A. SD-LW: Dideficiency medium (lacking leucine and tryptophan). SD-LWHA: Four-Deficiency medium (lacking leucine, tryptophan, histidine and adenine)
| [1] | Qu L, Liu MY, Zheng LL, et al. Data-independent acquisition-based global phosphoproteomics reveal the diverse roles of casein kinase 1 in plant development [J]. Sci Bull, 2023, 68(18): 2077-2093. |
| [2] | Duan XX, Zhang YY, Huang XH, et al. GreenPhos, a universal method for in-depth measurement of plant phosphoproteomes with high quantitative reproducibility [J]. Mol Plant, 2024, 17(1): 199-213. |
| [3] | Zhang N, Liu HT. Switch on and off: Phospho-events in light signaling pathways [J]. J Integr Plant Biol, 2025, 67(7): 1756-1770. |
| [4] | Zhang WJ, Zhou YW, Zhang Y, et al. Protein phosphorylation: a molecular switch in plant signaling [J]. Cell Rep, 2023, 42(7): 112729. |
| [5] | Tan ST, Luschnig C, Friml J. Pho-view of auxin: reversible protein phosphorylation in auxin biosynthesis, transport and signaling [J]. Mol Plant, 2021, 14(1): 151-165. |
| [6] | Cohen PTW, Brewis ND, Hughes V, et al. Protein serine/threonine phosphatases; an expanding family [J]. FEBS Lett, 1990, 268(2): 355-359. |
| [7] | 刘卫军, 沈瑛, 丁健. 蛋白磷酸酶2A的结构、功能和活性调节 [J]. 生物化学与生物物理学报, 2003(2): 105-112. |
| Liu WJ, Shen Y, Ding J. Protein phosphatase 2A: its structure, function and activity regulation [J]. Acta Biochim Biophys Sin, 2003(2): 105-112. | |
| [8] | 韩琦, 潘超颖. 蛋白磷酸酶2A在真菌细胞中的研究进展 [J]. 微生物学通报, 2021, 48(6): 2214-2221. |
| Han Q, Pan CY. Research progress of fungal protein phosphatase 2A [J]. Microbiol China, 2021, 48(6): 2214-2221. | |
| [9] | Jiang TY, Cui XW, Zeng TM, et al. PTEN deficiency facilitates gemcitabine efficacy in cancer by modulating the phosphorylation of PP2Ac and DCK [J]. Sci Transl Med, 2023, 15(704): eadd7464. |
| [10] | Cai XB, Lee S, Gómez Jaime AP, et al. PHOSPHATASE 2A dephosphorylates PHYTOCHROME-INTERACTING FACTOR3 to modulate photomorphogenesis in Arabidopsis [J]. Plant Cell, 2024, 36(10): 4457-4471. |
| [11] | 丘韶鑫, 杨发恒, 何经纬, 等. 植物中PP2A功能研究进展 [J]. 植物生理学报, 2024, 60(3): 430-440. |
| Qiu SX, Yang FH, He JW, et al. Research advances on function of PP2A in plants [J]. Plant Physiol J, 2024, 60(3): 430-440. | |
| [12] | Durian G, Jeschke V, Rahikainen M, et al. PROTEIN PHOSPHATASE 2A-B'γ controls Botrytis cinerea resistance and developmental leaf senescence [J]. Plant Physiol, 2020, 182(2): 1161-1181. |
| [13] | Shimamura M. Marchantia polymorpha: taxonomy, phylogeny and morphology of a model system [J]. Plant Cell Physiol, 2016, 57(2): 230-256. |
| [14] | Naramoto S, Hata Y, Fujita T, et al. The bryophytes Physcomitrium patens and Marchantia polymorpha as model systems for studying evolutionary cell and developmental biology in plants [J]. Plant Cell, 2022, 34(1): 228-246. |
| [15] | Ishizaki K, Nishihama R, Yamato KT, et al. Molecular genetic tools and techniques for Marchantia polymorpha research [J]. Plant Cell Physiol, 2016, 57(2): 262-270. |
| [16] | Kubota A, Ishizaki K, Hosaka M, et al. Efficient Agrobacterium-mediated transformation of the liverwort Marchantia polymorpha using regenerating thalli [J]. Biosci Biotechnol Biochem, 2013, 77(1): 167-172. |
| [17] | Chang CR, Bowman JL, Meyerowitz EM. Field guide to plant model systems [J]. Cell, 2016, 167(2): 325-339. |
| [18] | Sugano SS, Shirakawa M, Takagi J, et al. CRISPR/Cas9-mediated targeted mutagenesis in the liverwort Marchantia polymorpha L [J]. Plant Cell Physiol, 2014, 55(3): 475-481. |
| [19] | 郝格格, 孙忠富, 张录强, 等. 脱落酸在植物逆境胁迫研究中的进展 [J]. 中国农学通报, 2009, 25(18): 212-215. |
| Hao GG, Sun ZF, Zhang LQ, et al. A research overview of the plant resistance to adverse environment by using abscisic acid [J]. Chin Agric Sci Bull, 2009, 25(18): 212-215. | |
| [20] | Kwak JM, Moon JH, Murata Y, et al. Disruption of a guard cell-expressed protein phosphatase 2A regulatory subunit, RCN1, confers abscisic acid insensitivity in Arabidopsis [J]. Plant Cell, 2002, 14(11): 2849-2861. |
| [21] | Pernas M, García-Casado G, Rojo E, et al. A protein phosphatase 2A catalytic subunit is a negative regulator of abscisic acid signalling [J]. Plant J, 2007, 51(5): 763-778. |
| [22] | Saint-Marcoux D, Proust H, Dolan L, et al. Identification of reference genes for real-time quantitative PCR experiments in the liverwort Marchantia polymorpha [J]. PLoS One, 2015, 10(3): e0118678. |
| [23] | Fu Y, Jia XF, Yuan JW, et al. Fam72a functions as a cell-cycle-controlled gene during proliferation and antagonizes apoptosis through reprogramming PP2A substrates [J]. Dev Cell, 2023, 58(5): 398-415.e7. |
| [24] | Shen C, Li H, Shu L, et al. Ancient large-scale gene duplications and diversification in bryophytes illuminate the plant terrestrialization [J]. New Phytol, 2025, 245(5): 2292-2308. |
| [25] | Rensing SA, Lang D, Zimmer AD, et al. The Physcomitrella Genome reveals evolutionary insights into the conquest of land by plants [J]. Science, 2008, 319(5859): 64-69. |
| [26] | Shao ZY, Zhao B, Kotla P, et al. Phosphorylation status of Bβ subunit acts as a switch to regulate the function of phosphatase PP2A in ethylene-mediated root growth inhibition [J]. New Phytol, 2022, 236(5): 1762-1778. |
| [27] | Yue K, Sandal P, Williams EL, et al. PP2A-3 interacts with ACR4 and regulates formative cell division in the Arabidopsis root [J]. Proc Natl Acad Sci U S A, 2016, 113(5): 1447-1452. |
| [28] | Zhu YF, Huang PC, Guo PC, et al. CDK8 is associated with RAP2.6 and SnRK2.6 and positively modulates abscisic acid signaling and drought response in Arabidopsis [J]. New Phytol, 2020, 228(5): 1573-1590. |
| [29] | Li FQ, Wang JY, Wang PC, et al. Dephosphorylation of bZIP59 by PP2A ensures appropriate shade avoidance response in Arabidopsis [J]. Dev Cell, 2025, 60(4): 551-566.e6. |
| [30] | Wang ZN, Song AX, Tao BL, et al. The phosphatase PP1 sustains global transcription by promoting RNA polymerase Ⅱ pause release [J]. Mol Cell, 2024, 84(24): 4824-4842.e7. |
| [31] | Umezawa T, Sugiyama N, Mizoguchi M, et al. Type 2C protein phosphatases directly regulate abscisic acid-activated protein kinases in Arabidopsis [J]. Proc Natl Acad Sci U S A, 2009, 106(41): 17588-17593. |
| [32] | Ahmad Z, Tian DY, Li Y, et al. Characterization, evolution, expression and functional divergence of the DMP gene family in plants [J]. Int J Mol Sci, 2024, 25(19): 10435. |
| [33] | Lyons SP, Greiner EC, Cressey LE, et al. Regulation of PP2A, PP4, and PP6 holoenzyme assembly by carboxyl-terminal methylation [J]. Sci Rep, 2021, 11: 23031. |
| [34] | Wu CG, Chen H, Guo F, et al. PP2A-B' holoenzyme substrate recognition, regulation and role in cytokinesis [J]. Cell Discov, 2017, 3: 17027. |
| [35] | Li H, Wang JL, Kuan TA, et al. Pathogen protein modularity enables elaborate mimicry of a host phosphatase [J]. Cell, 2023, 186(15): 3196-3207.e17. |
| [1] | LI Tian-yuan, QI Xin-liang, LIU Shan, ZHANG Jian-cheng, WANG Peng-fei, ZHANG Shuai, JIA Lu-ting, MU Xiao-peng. Identification of Cerasus humilisSPL Gene Family and Expression Analysis during Fruit Development [J]. Biotechnology Bulletin, 2026, 42(3): 362-373. |
| [2] | XU Ze, ZHOU Chen-ping, KUANG Rui-bin, WU Xia-ming, YANG Min, LIU Chuan-he, HE Han, WEI Yue-rong. Identification of PG Gene Family and Their Roles in Papaya Fruit Softening [J]. Biotechnology Bulletin, 2026, 42(3): 349-361. |
| [3] | CHEN Chang-lu, YANG Zhi-fang, CAO Song-xiao, LI Yang-qing, YE Jing-feng, LYU Hai-yan, CHEN Shan-shan, CHEN Hao. Mechanism of Abscisic Acid and Ethylene Collaboratively Regulating the Softening of Oriental Melon Fruits [J]. Biotechnology Bulletin, 2026, 42(3): 302-311. |
| [4] | LIU Bao-cai, HU Xue-bo, ZHANG Wu-jun, ZHAO Yun-qing, HUANG Ying-zhen, CHEN Jing-ying. Screening of Internal Reference Gene for Pholidota chinensis and Their Applications [J]. Biotechnology Bulletin, 2026, 42(2): 158-168. |
| [5] | NI Ying, LI Lei, WANG Jin-xuan, MA Bo, MENG Xin, LENG Ping-sheng, WU Jing, HU Zeng-hui. Cloning and Functional Analysis of So4CL Gene in Syringa oblata [J]. Biotechnology Bulletin, 2026, 42(1): 139-149. |
| [6] | CHEN Jing-huan, FANG Guo-nan, ZHU Wen-hao, YE Guang-ji, SU Wang, HE Miao-miao, YANG Sheng-long, ZHOU Yun. Starch Characterization and Related Gene Expression Analysis of Potato Germplasm Resources [J]. Biotechnology Bulletin, 2026, 42(1): 170-183. |
| [7] | ZENG Ting, ZHANG Lan, LUO Rui. Functional Analysis of the Transcription Factor MpR2R3-MYB17 in Regulating Gemma Development in Marchantia polymorpha L. [J]. Biotechnology Bulletin, 2026, 42(1): 208-217. |
| [8] | CHEN Qiang, YU Ying-fei, ZHANG Ying, ZHANG Chong. Regulatory Effect of Methyl Jasmonate on Postharvest Chilling Injury in Oriental Melon ‘Emerald’ [J]. Biotechnology Bulletin, 2025, 41(9): 105-114. |
| [9] | CHENG Xue, FU Ying, CHAI Xiao-jiao, WANG Hong-yan, DENG Xin. Identification of LHC Gene Family in Setaria italica and Expression Analysis under Abiotic Stresses [J]. Biotechnology Bulletin, 2025, 41(8): 102-114. |
| [10] | REN Rui-bin, SI Er-jing, WAN Guang-you, WANG Jun-cheng, YAO Li-rong, ZHANG Hong, MA Xiao-le, LI Bao-chun, WANG Hua-jun, MENG Ya-xiong. Identification and Expression Analysis of GH17 Gene Family of Pyrenophora graminea [J]. Biotechnology Bulletin, 2025, 41(8): 146-154. |
| [11] | ZENG Dan, HUANG Yuan, WANG Jian, ZHANG Yan, LIU Qing-xia, GU Rong-hui, SUN Qing-wen, CHEN Hong-yu. Genome-wide Identification and Expression Analysis of bZIP Transcription Factor Family in Dendrobium officinale [J]. Biotechnology Bulletin, 2025, 41(8): 197-210. |
| [12] | LI Xin-ni, LI Jun-yi, MA Xue-hua, HE Wei, LI Jia-li, YU Jia, CAO Xiao-ning, QIAO Zhi-jun, LIU Si-chen. Identification of the PMEI Gene Family of Pectin Methylesterase Inhibitor in Foxtail Millet and Analysis of Its Response to Abiotic Stress [J]. Biotechnology Bulletin, 2025, 41(7): 150-163. |
| [13] | HAN Yi, HOU Chang-lin, TANG Lu, SUN Lu, XIE Xiao-dong, LIANG Chen, CHEN Xiao-qiang. Cloning and Preliminary Functional Analysis of HvERECTA Gene in Hordeum vulgare [J]. Biotechnology Bulletin, 2025, 41(7): 106-116. |
| [14] | QU Shan, ZHAO Yue, LI Ya-hua, ZHENG Gui-ling, XIAN Hong-quan. A Study on the Interaction between Transcriptional Factor and Protein of Tachi2 Chitinase Gene in Trichoderma asperellum [J]. Biotechnology Bulletin, 2025, 41(5): 310-319. |
| [15] | LIU Zhuo-jun, CHAI Wen-ting, REN Yi-le, WANG Xin-yu, ZHU Li-xun, ZHAO Shan-shan, YANG Bo-hui, FAN Jia-li, LI Xin-feng, ZHAO Wei-jun, LYU Jin-hui, ZHANG Chun-lai. Analysis on Expression and DNA Variation of TGA Genes in Sorghum (Sorghum bicolor) in Response to Sporisorium reilianum Infection [J]. Biotechnology Bulletin, 2025, 41(5): 90-103. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||