








Biotechnology Bulletin ›› 2026, Vol. 42 ›› Issue (2): 178-187.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0659
Previous Articles Next Articles
ZHANG Dong-ling1( ), ZHANG Yin-sheng2, WANG Jian-jun2, YE Fei-yu1, LU Zi-han1, MA Chen-chen1, LIU Hua-feng1, HU De-sheng1, DENG Ya-zhou1, CAO Li-ru1(
), ZHANG Yin-sheng2, WANG Jian-jun2, YE Fei-yu1, LU Zi-han1, MA Chen-chen1, LIU Hua-feng1, HU De-sheng1, DENG Ya-zhou1, CAO Li-ru1( )
)
Received:2025-06-07
Online:2026-02-26
Published:2026-03-17
Contact:
CAO Li-ru
E-mail:zhangdongling0626@163.com;caoliru008@126.com
ZHANG Dong-ling, ZHANG Yin-sheng, WANG Jian-jun, YE Fei-yu, LU Zi-han, MA Chen-chen, LIU Hua-feng, HU De-sheng, DENG Ya-zhou, CAO Li-ru. Expression Characteristics and Functions of HSFs Transcription Factors Family in Maize under Drought Stress[J]. Biotechnology Bulletin, 2026, 42(2): 178-187.
| 引物名称 Primer name | 正向引物 Forward primer (5′‒3′) | 反向引物 Reverse primer (5′‒3′) |
|---|---|---|
| HSF16 | ATGCAAGAGCTTGTCAGGCT | CAACGAAGCAGCAGCACTTT |
| HSF7 | GAGCTTGGGAAGGGAAGCAT | CCTCTCCACCACCAGTCAAC |
| HSF25 | TGAACGAAGGGATGAAGGGC | ACTACACGTCTGCACTCTGC |
| HSF26 | CACCCACCATGGAGAAGGAC | TGGTCTGGATCCCTCCTCAG |
Table 1 Quantitative primer sequences
| 引物名称 Primer name | 正向引物 Forward primer (5′‒3′) | 反向引物 Reverse primer (5′‒3′) |
|---|---|---|
| HSF16 | ATGCAAGAGCTTGTCAGGCT | CAACGAAGCAGCAGCACTTT |
| HSF7 | GAGCTTGGGAAGGGAAGCAT | CCTCTCCACCACCAGTCAAC |
| HSF25 | TGAACGAAGGGATGAAGGGC | ACTACACGTCTGCACTCTGC |
| HSF26 | CACCCACCATGGAGAAGGAC | TGGTCTGGATCCCTCCTCAG |
| 基因IDGene ID | 基因Gene | 蛋白长度 Protein length (aa) | 等电点pI | 分子量 Molecular weight (Da) | 不稳定指数 Instability index | 疏水指数 GRAVY | 带负电残基数 Asp+Glu | 带正电残基数 Arg+Lys |
|---|---|---|---|---|---|---|---|---|
| Zm00001eb353710 | ZmHSF1 | 449 | 5.15 | 49 978 | 58.13 | -0.586 | 62 | 42 |
| Zm00001eb293250 | ZmHSF2 | 469 | 5.41 | 51 624 | 60.51 | -0.533 | 62 | 48 |
| Zm00001eb017880 | ZmHSF3 | 331 | 9.11 | 37 173 | 61.83 | -0.783 | 40 | 45 |
| Zm00001eb100770 | ZmHSF4 | 298 | 9.13 | 32 270 | 55.75 | -0.556 | 36 | 41 |
| Zm00001eb056980 | ZmHSF5 | 371 | 5.12 | 41 282 | 48.30 | -0.604 | 59 | 47 |
| Zm00001eb160080 | ZmHSF6 | 398 | 5.84 | 43 463 | 42.50 | -0.461 | 52 | 47 |
| Zm00001eb047760 | ZmHSF7 | 357 | 4.99 | 40 502 | 59.15 | -0.722 | 61 | 46 |
| Zm00001eb328900 | ZmHSF8 | 323 | 6.57 | 35 561 | 59.80 | -0.623 | 36 | 31 |
| Zm00001eb399800 | ZmHSF91 | 407 | 4.97 | 45 318 | 53.67 | -0.590 | 55 | 36 |
| Zm00001eb242480 | ZmHSF10 | 508 | 4.96 | 56 062 | 55.94 | -0.536 | 77 | 49 |
| Zm00001eb358700 | ZmHSF11 | 348 | 8.09 | 37 410 | 53.05 | -0.302 | 35 | 37 |
| Zm00001eb100410 | ZmHSF12 | 394 | 7.81 | 41 743 | 66.21 | -0.372 | 36 | 37 |
| Zm00001eb159390 | ZmHSF13 | 331 | 5.94 | 35 884 | 60.93 | -0.345 | 40 | 35 |
| Zm00001eb198620 | ZmHSF14 | 383 | 6.23 | 41 003 | 58.83 | -0.515 | 47 | 43 |
| Zm00001eb239380 | ZmHSF15 | 257 | 5.85 | 27 837 | 46.60 | -0.402 | 33 | 30 |
| Zm00001eb064860 | ZmHSF16 | 500 | 5.01 | 54 420 | 59.29 | -0.507 | 59 | 44 |
| Zm00001eb384760 | ZmHSF17 | 267 | 6.99 | 28 329 | 42.41 | -0.228 | 30 | 30 |
| Zm00001eb371000 | ZmHSF18 | 433 | 5.25 | 48 648 | 51.50 | -0.842 | 69 | 51 |
| Zm00001eb009170 | ZmHSF19 | 417 | 5.09 | 46 816 | 55.59 | -0.693 | 57 | 39 |
| Zm00001eb301280 | ZmHSF20 | 375 | 4.7 | 42 044 | 53.81 | -0.716 | 63 | 37 |
| Zm00001eb060670 | ZmHSF21 | 359 | 5.57 | 40 587 | 53.80 | -0.835 | 53 | 41 |
| Zm00001eb314890 | ZmHSF22 | 298 | 9.53 | 32 258 | 53.74 | -0.567 | 35 | 43 |
| Zm00001eb037600 | ZmHSF23 | 379 | 6.27 | 40 646 | 57.77 | -0.434 | 45 | 42 |
| Zm00001eb319350 | ZmHSF24 | 394 | 5 | 41 468 | 54.72 | -0.433 | 50 | 35 |
| Zm00001eb004670 | ZmHSF25 | 384 | 5.3 | 43 268 | 71.81 | -0.759 | 62 | 51 |
| Zm00001eb241300 | ZmHSF26 | 528 | 5.57 | 58 139 | 51.23 | -0.714 | 74 | 56 |
| Zm00001eb428800 | ZmHSF27 | 318 | 5.7 | 33 948 | 45.96 | -0.707 | 50 | 45 |
| Zm00001eb210530 | ZmHSF28 | 497 | 5.06 | 54 105 | 63.64 | -0.484 | 58 | 44 |
| 平均值 Average | 384.75 | 6.08 | 42 206.51 | 55.59 | ‒‒ | ‒‒ | ‒‒ |
Table 2 Physicochemical properties of ZmHSFs protein
| 基因IDGene ID | 基因Gene | 蛋白长度 Protein length (aa) | 等电点pI | 分子量 Molecular weight (Da) | 不稳定指数 Instability index | 疏水指数 GRAVY | 带负电残基数 Asp+Glu | 带正电残基数 Arg+Lys |
|---|---|---|---|---|---|---|---|---|
| Zm00001eb353710 | ZmHSF1 | 449 | 5.15 | 49 978 | 58.13 | -0.586 | 62 | 42 |
| Zm00001eb293250 | ZmHSF2 | 469 | 5.41 | 51 624 | 60.51 | -0.533 | 62 | 48 |
| Zm00001eb017880 | ZmHSF3 | 331 | 9.11 | 37 173 | 61.83 | -0.783 | 40 | 45 |
| Zm00001eb100770 | ZmHSF4 | 298 | 9.13 | 32 270 | 55.75 | -0.556 | 36 | 41 |
| Zm00001eb056980 | ZmHSF5 | 371 | 5.12 | 41 282 | 48.30 | -0.604 | 59 | 47 |
| Zm00001eb160080 | ZmHSF6 | 398 | 5.84 | 43 463 | 42.50 | -0.461 | 52 | 47 |
| Zm00001eb047760 | ZmHSF7 | 357 | 4.99 | 40 502 | 59.15 | -0.722 | 61 | 46 |
| Zm00001eb328900 | ZmHSF8 | 323 | 6.57 | 35 561 | 59.80 | -0.623 | 36 | 31 |
| Zm00001eb399800 | ZmHSF91 | 407 | 4.97 | 45 318 | 53.67 | -0.590 | 55 | 36 |
| Zm00001eb242480 | ZmHSF10 | 508 | 4.96 | 56 062 | 55.94 | -0.536 | 77 | 49 |
| Zm00001eb358700 | ZmHSF11 | 348 | 8.09 | 37 410 | 53.05 | -0.302 | 35 | 37 |
| Zm00001eb100410 | ZmHSF12 | 394 | 7.81 | 41 743 | 66.21 | -0.372 | 36 | 37 |
| Zm00001eb159390 | ZmHSF13 | 331 | 5.94 | 35 884 | 60.93 | -0.345 | 40 | 35 |
| Zm00001eb198620 | ZmHSF14 | 383 | 6.23 | 41 003 | 58.83 | -0.515 | 47 | 43 |
| Zm00001eb239380 | ZmHSF15 | 257 | 5.85 | 27 837 | 46.60 | -0.402 | 33 | 30 |
| Zm00001eb064860 | ZmHSF16 | 500 | 5.01 | 54 420 | 59.29 | -0.507 | 59 | 44 |
| Zm00001eb384760 | ZmHSF17 | 267 | 6.99 | 28 329 | 42.41 | -0.228 | 30 | 30 |
| Zm00001eb371000 | ZmHSF18 | 433 | 5.25 | 48 648 | 51.50 | -0.842 | 69 | 51 |
| Zm00001eb009170 | ZmHSF19 | 417 | 5.09 | 46 816 | 55.59 | -0.693 | 57 | 39 |
| Zm00001eb301280 | ZmHSF20 | 375 | 4.7 | 42 044 | 53.81 | -0.716 | 63 | 37 |
| Zm00001eb060670 | ZmHSF21 | 359 | 5.57 | 40 587 | 53.80 | -0.835 | 53 | 41 |
| Zm00001eb314890 | ZmHSF22 | 298 | 9.53 | 32 258 | 53.74 | -0.567 | 35 | 43 |
| Zm00001eb037600 | ZmHSF23 | 379 | 6.27 | 40 646 | 57.77 | -0.434 | 45 | 42 |
| Zm00001eb319350 | ZmHSF24 | 394 | 5 | 41 468 | 54.72 | -0.433 | 50 | 35 |
| Zm00001eb004670 | ZmHSF25 | 384 | 5.3 | 43 268 | 71.81 | -0.759 | 62 | 51 |
| Zm00001eb241300 | ZmHSF26 | 528 | 5.57 | 58 139 | 51.23 | -0.714 | 74 | 56 |
| Zm00001eb428800 | ZmHSF27 | 318 | 5.7 | 33 948 | 45.96 | -0.707 | 50 | 45 |
| Zm00001eb210530 | ZmHSF28 | 497 | 5.06 | 54 105 | 63.64 | -0.484 | 58 | 44 |
| 平均值 Average | 384.75 | 6.08 | 42 206.51 | 55.59 | ‒‒ | ‒‒ | ‒‒ |
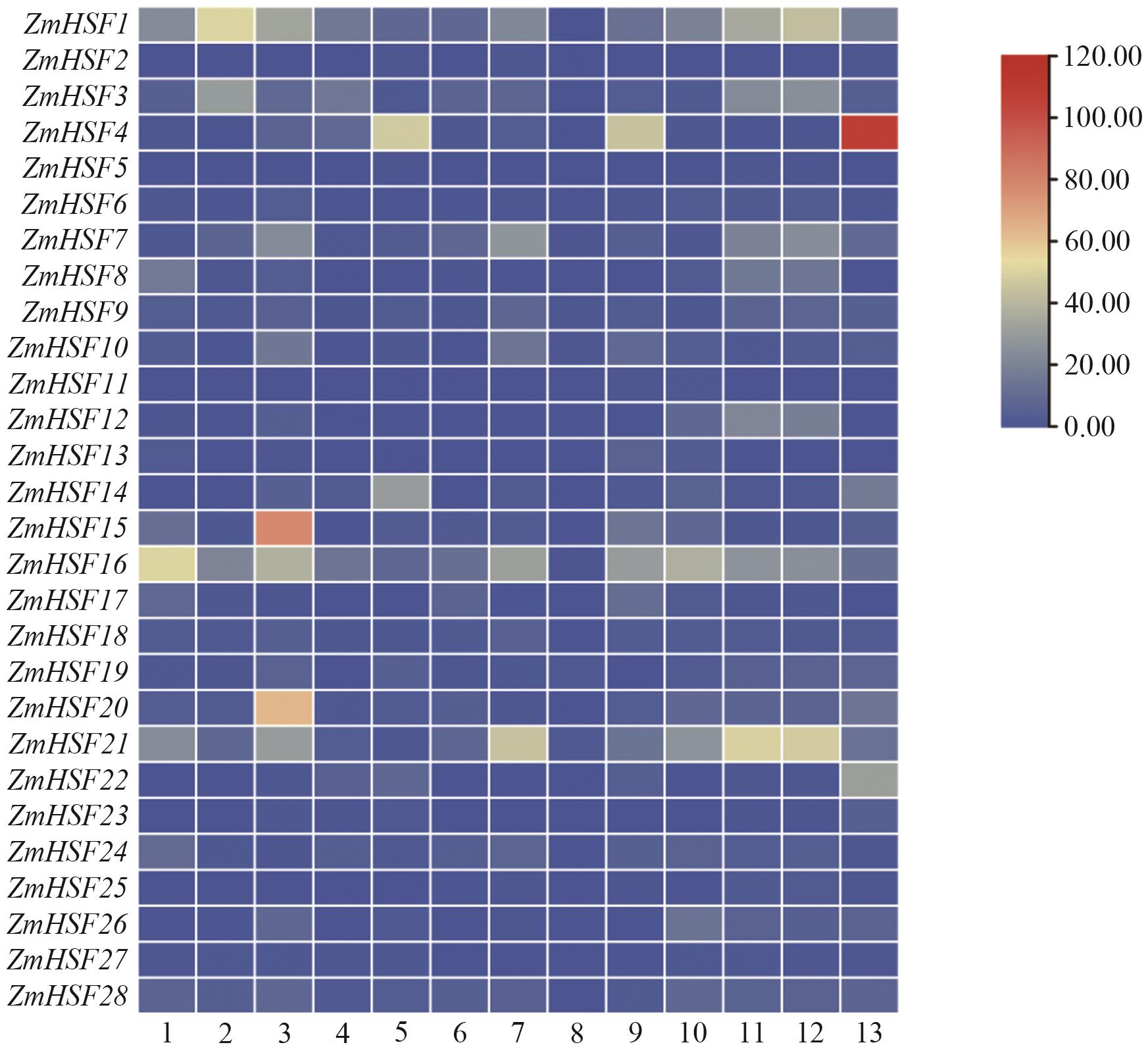
Fig. 7 Expressions of ZmHSFs gene family in different tissues1: Internode 7‒8. 2: Embryo 18 DAP. 3: Embryo 38 DAP. 4: Endosperm l6 DAP. 5: Endosperm 27 DAP. 6: Root. 7: Leaf. 8: Pollen. 9: Silk. 10: Female spike. l1: Ear primordium 2‒4 mm. 12: Ear primordium 6‒8 mm. 13: Pericarp/Aleurone 27 DAP
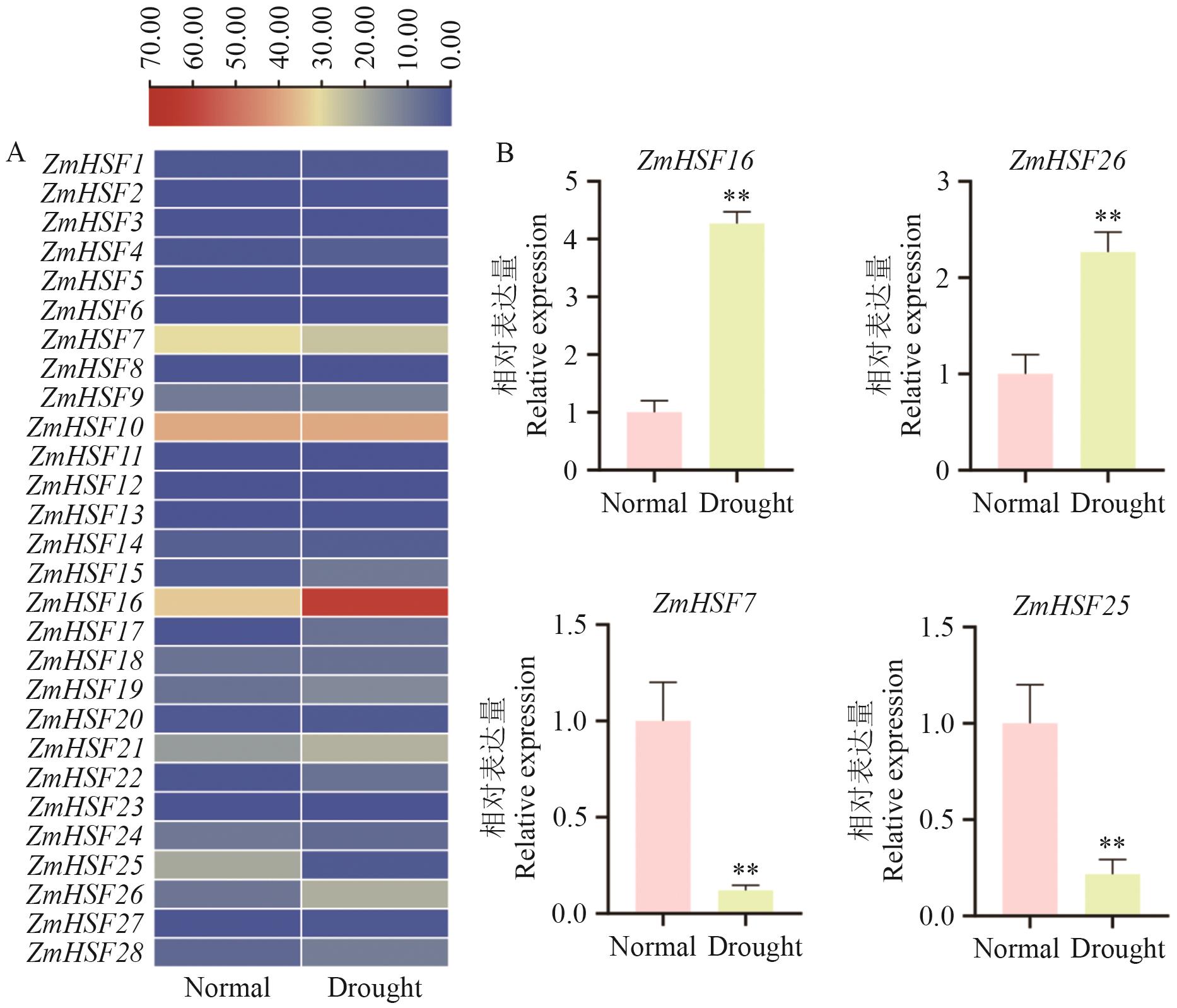
Fig. 8 Expressions of ZmHSFs gene family under drought stressA: Analysis of RNA-seq data of ZmHSFs gene family under drought stress. B: Detection of gene expressions. **: P<0.01. The same below
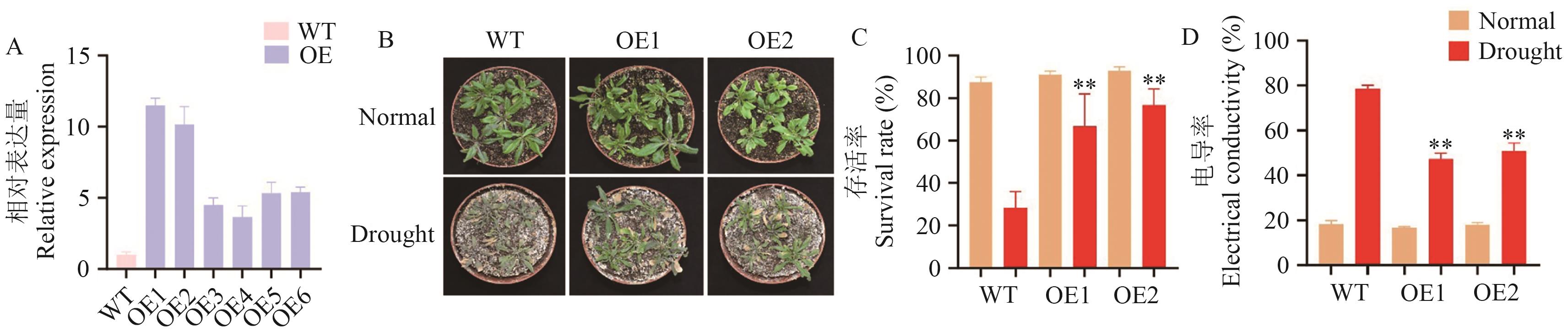
Fig. 9 ZmHSF16 enhances drought resistance in ArabidopsisA: The expressions of ZmHSF16 in WT and six independent ZmHSF16 overexpression lines. B: Phenotypic analysis of WT, ZmHSF16 overexpressing materials under normal conditions and drought stress conditions. C, D: Survival rate and conductivity of WT, ZmHSF16-overexpressing materials after 5 d of drought stress treatment
| [1] | Shi WJ, Wang ML, Liu YT. Climate change and crop drought stress management in China [J]. Sci Total Environ, 2021, 750: 141147. |
| [2] | Fahad S, Bajwa AA, Nazir U, et al. Crop production under drought and heat stress: Plant responses and management options [J]. Front Plant Sci, 2017, 8: 1147. |
| [3] | Wang X, Cai J, Zhou Q, et al. Physiological mechanisms of abiotic stress priming induced the crops stress tolerance: A review [J]. Sci Agric Sin, 2021, 54(11): 2287-2301. |
| [4] | Lesk C, Rowhani P, Ramankutty N. Influence of extreme weather disasters on global crop production [J]. Nature, 2016, 529(7584): 84-87. |
| [5] | Nover L, Bharti K, Döring P, et al. Arabidopsis and the heat stress transcription factor world: How many heat stress transcription factors do we need [J]. Cell Stress Chaperones, 2001, 6(3): 177-189. |
| [6] | Scharf KD, Berberich T, Ebersberger I, et al. The plant heat stress transcription factor (Hsf) family: Structure, function and evolution [J]. Biochim Biophys Acta Gene Regul Mech, 2012, 1819(2): 104-119. |
| [7] | Liu HC, Charng YY. Common and distinct functions of Arabidopsis class A1 and A2 heat shock factors in diverse abiotic stress responses and development [J]. Plant Physiol, 2013, 163(1): 276-290. |
| [8] | Mesihovic A, Ullrich S, Rosenkranz RRE, et al. HsfA7 coordinates the transition from mild to strong heat stress response by controlling the activity of the master regulator HsfA1a in tomato [J]. Cell Rep, 2022, 38(2): 110224. |
| [9] | Wang HR, Feng M, Jiang YJ, et al. Thermosensitive SUMOylation of TaHsfA1 defines a dynamic ON/OFF molecular switch for the heat stress response in wheat [J]. Plant Cell, 2023, 35(10): 3889-3910. |
| [10] | Wen JJ, Qin Z, Sun L, et al. Alternative splicing of TaHSFA6e modulates heat shock protein-mediated translational regulation in response to heat stress in wheat [J]. New Phytol, 2023, 239(6): 2235-2247. |
| [11] | Wang XY, Huang WL, Liu J, et al. Molecular regulation and physiological functions of a novel FaHsfA2c cloned from tall fescue conferring plant tolerance to heat stress [J]. Plant Biotechnol J, 2017, 15(2): 237-248. |
| [12] | Schmidt R, Schippers JHM, Welker A, et al. Transcription factor OsHsfC1b regulates salt tolerance and development in Oryza sativa ssp. Japonica [J]. AoB Plants, 2012, 2012: pls011. |
| [13] | Lin YX, Jiang HY, Chu ZX, et al. Genome-wide identification, classification and analysis of heat shock transcription factor family in maize [J]. BMC Genomics, 2011, 12: 76. |
| [14] | Jiang LY, Hu WJ, Qian YX, et al. Genome-wide identification, classification and expression analysis of the Hsf and Hsp70 gene families in maize [J]. Gene, 2021, 770: 145348. |
| [15] | Zhang HN, Li GL, Fu C, et al. Genome-wide identification, transcriptome analysis and alternative splicing events of Hsf family genes in maize [J]. Sci Rep, 2020, 10(1): 8073. |
| [16] | Song NN, Wang J, Qin QQ, et al. ZmHSFA2B self-regulatory loop is critical for heat tolerance in maize [J]. Plant Biotechnol J, 2025, 23(1): 284-301. |
| [17] | Ikeda M, Mitsuda N, Ohme-Takagi M. Arabidopsis HsfB1 and HsfB2b act as repressors of the expression of heat-inducible Hsfs but positively regulate the acquired thermotolerance [J]. Plant Physiol, 2011, 157(3): 1243-1254. |
| [18] | Li YF, Huang YM, Sun HY, et al. Heat shock protein 101 contributes to the thermotolerance of male meiosis in maize [J]. Plant Cell, 2022, 34(10): 3702-3717. |
| [19] | Li H, Wang J, Li ML, et al. The ZmHSF08-ZmUGT92A1 module regulates heat tolerance by altering reactive oxygen species levels in maize [J]. Crop J, 2024, 12(5): 1437-1446. |
| [20] | Huang Y, Li MY, Wang F, et al. Heat shock factors in carrot: genome-wide identification, classification, and expression profiles response to abiotic stress [J]. Mol Biol Rep, 2015, 42(5): 893-905. |
| [21] | Zhang HM, Zhu JH, Gong ZZ, et al. Abiotic stress responses in plants [J]. Nat Rev Genet, 2022, 23(2): 104-119. |
| [22] | Fujita Y, Yoshida T, Yamaguchi-Shinozaki K. Pivotal role of the AREB/ABF-SnRK2 pathway in ABRE-mediated transcription in response to osmotic stress in plants [J]. Physiol Plant, 2013, 147(1): 15-27. |
| [23] | Ambawat S, Sharma P, Yadav NR, et al. MYB transcription factor genes as regulators for plant responses: An overview [J]. Physiol Mol Biol Plants, 2013, 19(3): 307-321. |
| [24] | Ohama N, Sato H, Shinozaki K, et al. Transcriptional regulatory network of plant heat stress response [J]. Trends Plant Sci, 2017, 22(1): 53-65. |
| [25] | Li ZJ, Zhang LL, Wang AX, et al. Ectopic overexpression of SlHsfA3, a heat stress transcription factor from tomato, confers increased thermotolerance and salt hypersensitivity in germination in transgenic Arabidopsis [J]. PLoS One, 2013, 8(1): e54880. |
| [26] | Tardieu F, Parent B, Caldeira CF, et al. Genetic and physiological controls of growth under water deficit [J]. Plant Physiol, 2014, 164(4): 1628-1635. |
| [27] | Xue GP, Drenth J, McIntyre CL. TaHsfA6f is a transcriptional activator that regulates a suite of heat stress protection genes in wheat (Triticum aestivum L.) including previously unknown Hsf targets [J]. J Exp Bot, 2015, 66(3): 1025-1039. |
| [28] | Cheng Q, Zhou Y, Liu Z, et al. An alternatively spliced heat shock transcription factor, OsHSFA2dI, functions in the heat stress-induced unfolded protein response in rice [J]. Plant Biol, 2015, 17(2): 419-429. |
| [29] | Farooq MA, Niazi AK, Akhtar J, et al. Acquiring control: The evolution of ROS-Induced oxidative stress and redox signaling pathways in plant stress responses [J]. Plant Physiol Biochem, 2019, 141: 353-369. |
| [1] | PENG Shan-lin, LIAO Zhuo-cheng, WANG Tao, LIU Zhi-yu, LIU Hai-yi, WANG Ting-ting, YANG Qin, WANG Zhe, TAI Huan-huan. Diversity and Functional Differences of Rhizosphere Microbial Communities in Maize Varieties with Different Resistance to Stalk Rot [J]. Biotechnology Bulletin, 2026, 42(5): 1-13. |
| [2] | LIU Qing-yuan, WU Hong-qi, CHEN Xiu-e, CHEN Jian, JIANG Yuan-ze, HE Yan-zi, YU Qi-wei, LIU Ren-xiang. Function of Transcription Factor NtMYB96a in Regulating the Tolerance of Tobacco to Drought [J]. Biotechnology Bulletin, 2026, 42(4): 239-250. |
| [3] | CHEN Deng-ke, LAN Gang, XIA Zhi, HOU Bao-guo, YANG Liu-liu, CAO Cai-rong, LI Peng-bo, WU Cui-cui. Identification of ZF-HD Gene Family in Arachis hypogaea and Analysis in Response to Abiotic Stress [J]. Biotechnology Bulletin, 2026, 42(4): 114-128. |
| [4] | YIN Ya-long, ZHANG Ming-yang, WANG Jie-min, MIAO Xue-xue, CHEN Jin, WANG Wei-ping. Advances in Coordinated Tolerance Mechanisms to Abiotic Stresses in Rice [J]. Biotechnology Bulletin, 2026, 42(4): 26-37. |
| [5] | WANG Jing, LIU Zhuo-yan, ZHANG Xiao-lei, LIU Bao-hai, GUAN Hai-tao, WEN Hong-tao. Rapid Detection of Transgenic Maize DBN9501 Event-specificity Based on LAMP-CRISPR-Cas12a Technology [J]. Biotechnology Bulletin, 2026, 42(2): 149-157. |
| [6] | DONG Ya-ru, ZHU Hong, WANG Zhao-hong, ZHAO Dong-xiao, LIU Hui-fen. Cloning and Salt-drought Tolerance Analysis of MnDREB6E Gene in Mulberry [J]. Biotechnology Bulletin, 2026, 42(2): 306-316. |
| [7] | NONG Wei-you, ZHAO Chang-zu, QIAN Zhen-feng, DING Qian, WANG Yu-jie, CHEN Shu-ying, HE Li-lian, LI Fu-sheng. Identification of the EfBBX Gene Family in Erianthus fulvus and Analysis of Its Expression Patterns Under Cold Stress [J]. Biotechnology Bulletin, 2026, 42(2): 267-277. |
| [8] | REN Yun-er, WU Guo-qiang, CHENG Bin, WEI Ming. Genome-wide Identification of the BvATGs Genes Family in Sugar Beet (Beta vulgaris L.) and Analysis of Their Expression Pattern under Salt Stress [J]. Biotechnology Bulletin, 2026, 42(1): 184-197. |
| [9] | YANG Juan, FENG Hui, JI Nai-zhe, SUN Li-ping, WANG Yun, ZHANG Jia-nan, ZHAO Shi-wei. Cloning and Functional Analysis of AP2/ERF Transcription Factors RcERF4 and RcRAP2-12 in Rose [J]. Biotechnology Bulletin, 2026, 42(1): 150-160. |
| [10] | ZHANG Chi-hao, LIU Jin-nan, CHAO Yue-hui. Cloning and Functional Analysis of a bZIP Transcription Factor MtbZIP29 from Medicago truncatula [J]. Biotechnology Bulletin, 2026, 42(1): 241-250. |
| [11] | ZHANG Yue, DAI Yue-hua, ZHANG Ying-ying, LI Ao-hui, LI Chu-hui, XUE Jin-ai, QIN Hui-bin, CHEN Yan, NIE Meng-en, ZHANG Hai-ping. Cloning and Functional Analysis of the Soybean Enoyl-CoA Reductase ECR14 Gene [J]. Biotechnology Bulletin, 2026, 42(1): 95-104. |
| [12] | WU Cui-cui, CHEN Deng-ke, LAN Gang, XIA Zhi, LI Peng-bo. Bioinformatics Analysis of Peanut Transcription Factor AhHDZ70 and Its Tolerances to Salt and Drought [J]. Biotechnology Bulletin, 2026, 42(1): 198-207. |
| [13] | LYU Cheng-cong, HENG Meng, CHEN Si-qi, JIN Xue-hua. Cloning and Functional Analysis of ZhGSTF Related to Anthocyanin Transport Zantedeschia hybrida [J]. Biotechnology Bulletin, 2026, 42(1): 161-169. |
| [14] | YANG Dan, JIN Ya-rong, MAO Chun-li, WANG Bi-xian, ZHANG Ya-ning, YANG Zhi-yi, ZHOU Zhi-yao, YANG Rui-ming, FAN Heng-rui, HUANG Lin-kai, YAN Hai-dong. Identification and Expression Analysis of C2H2 Gene Family in Cenchrus purpureus [J]. Biotechnology Bulletin, 2026, 42(1): 251-261. |
| [15] | LIU Jia-li, SONG Jing-rong, ZHAO Wen-yu, ZHANG Xin-yuan, ZHAO Zi-yang, CAO Yi-bo, ZHANG Ling-yun. Identification of the R2R3-MYB Gene and Expression Analysis of Flavonoid Regulatory Genes in Blueberry [J]. Biotechnology Bulletin, 2025, 41(9): 124-138. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||