








Biotechnology Bulletin ›› 2026, Vol. 42 ›› Issue (3): 48-59.doi: 10.13560/j.cnki.biotech.bull.1985.2025-1308
Previous Articles Next Articles
DU Dan1,2( ), GUO Xiang1, HU Xin1, PAN Yu1(
), GUO Xiang1, HU Xin1, PAN Yu1( )
)
Received:2025-12-01
Online:2026-03-26
Published:2026-04-23
Contact:
PAN Yu
E-mail:dudan199009@163.com;panyu1020@swu.edu.cn
DU Dan, GUO Xiang, HU Xin, PAN Yu. Advances in the Regulatory Mechanisms of Plastid Development on Fruit Ripening and Quality[J]. Biotechnology Bulletin, 2026, 42(3): 48-59.
基因名 Gene name | 物种 Species | 调控方式 Regulation way | 参考文献 Reference |
|---|---|---|---|
| GLK2 | 苹果、猕猴桃和番茄等 | 正调控果实质体发育 | [ |
| KNOXs | 番茄 | 正调控叶绿体发育 | [ |
| SlBEL2 | 番茄 | 负调控果实绿肩的形成 | [ |
| SlBLH7 | 番茄 | 负调控果实质体发育 | [ |
| APRR2s | 番茄、黄瓜和辣椒等 | 正调控果实质体发育 | [ |
| SlGRAS9 | 番茄 | 负调控果实质体叶绿素合成 | [ |
| SlZHD17 | 番茄 | 负调控果实质体叶绿素合成 | [ |
| SlGRAS38 | 番茄 | 正调控果实番茄红素合成 | [ |
| SlTGA2.2-SRDX | 番茄 | 果实质体数目与结构 | [ |
| PTOX | 番茄、拟南芥 | 正调控质体结构发育 | [ |
| TAGL1 | 番茄 | 正调控质体结构发育 | [ |
Table 1 Transcription factors involved in chloroplast development
基因名 Gene name | 物种 Species | 调控方式 Regulation way | 参考文献 Reference |
|---|---|---|---|
| GLK2 | 苹果、猕猴桃和番茄等 | 正调控果实质体发育 | [ |
| KNOXs | 番茄 | 正调控叶绿体发育 | [ |
| SlBEL2 | 番茄 | 负调控果实绿肩的形成 | [ |
| SlBLH7 | 番茄 | 负调控果实质体发育 | [ |
| APRR2s | 番茄、黄瓜和辣椒等 | 正调控果实质体发育 | [ |
| SlGRAS9 | 番茄 | 负调控果实质体叶绿素合成 | [ |
| SlZHD17 | 番茄 | 负调控果实质体叶绿素合成 | [ |
| SlGRAS38 | 番茄 | 正调控果实番茄红素合成 | [ |
| SlTGA2.2-SRDX | 番茄 | 果实质体数目与结构 | [ |
| PTOX | 番茄、拟南芥 | 正调控质体结构发育 | [ |
| TAGL1 | 番茄 | 正调控质体结构发育 | [ |
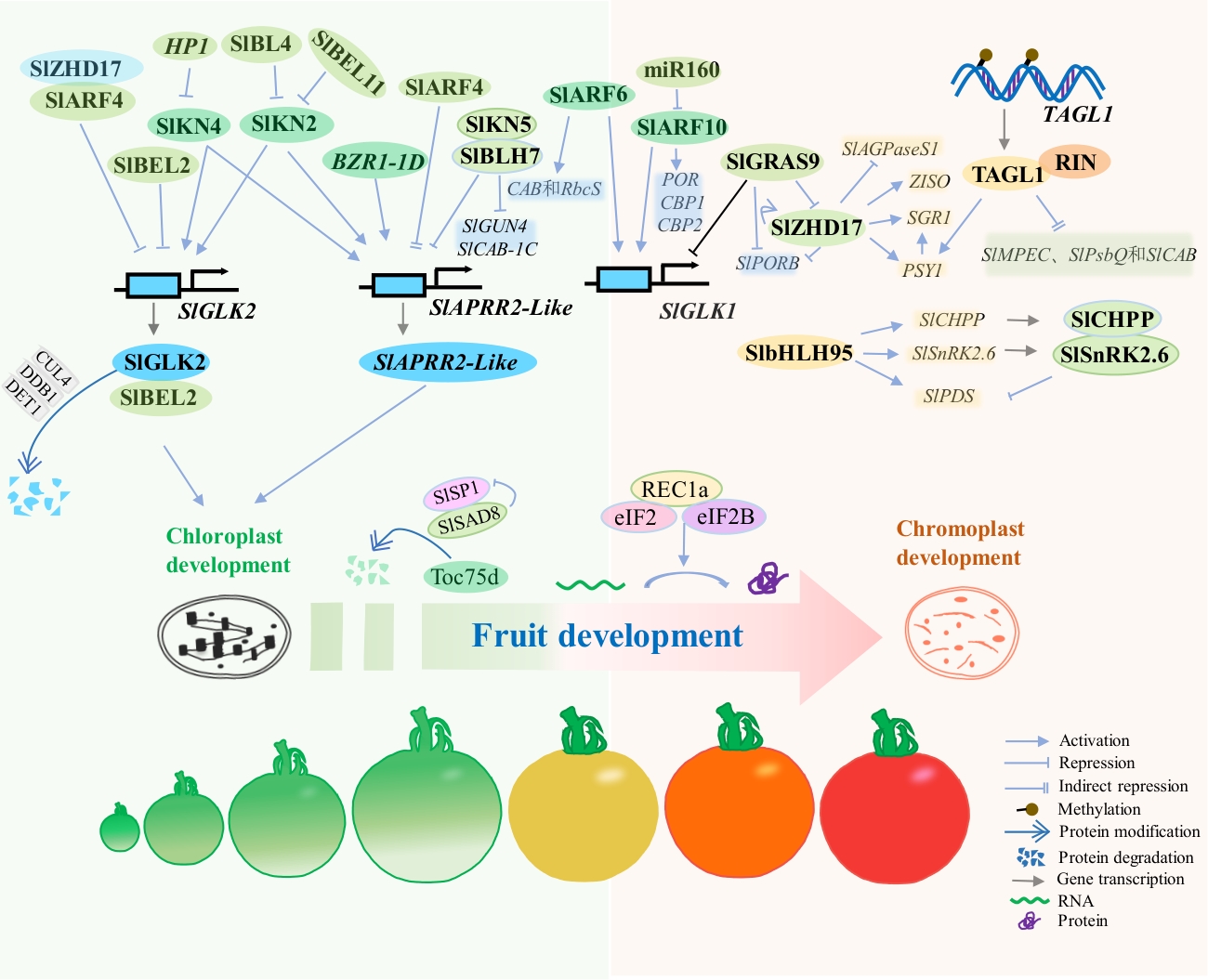
Fig. 2 Schematic diagram of transcriptional regulation during plastid development and differentiationThe development and maturation of fruit is also the process of chloroplasts developing into chromoplasts, mainly involving two aspects: structural transformation of chloroplasts and related metabolic processes. SlGLK2 and SlAPRR2 are primarily involved in regulating early chloroplast development in fruit. Both SlKN2 and SlKN4 can activate the expressions of SlGLK2 and SlAPRR2, while another transcriptional activator is the BR-responsive gene BZR1-1D. However, the expression of SlKN2 and SlKN4 is inhibited by HP1, SlBL4, and SlBEL11, respectively. Unlike the activation effects of SlKN2 and SlKN4, SlKN5 interacts with SlBLH7 and inhibits the transcription of SlAPRR2, as well as the chlorophyll synthesis-related gene SlGUN4 and the photosystem II-related gene SlCAB-1C. Furthermore, auxin also plays an important regulatory role in plastid development. The auxin response factor SlARF4 can inhibit the transcription of SlGLK2 and SlAPRR2, while SlARF6 and SlARF10 can activate the expression of the GLK1 gene and activate the transcriptional expression of CAB and RbcS, PORCBP1 and CBP2, respectively. Genes such as SlZHD17 and SlbHLH95 are mainly involved in chromoplast development, primarily by activating pigment synthesis genes (such as PSY1 and PDS) and inhibiting the expressions of sugar metabolism-related genes (such as SUS1). Despite this, SlZHD17 can interact with SlARF4 and inhibit the transcription of SlGLK2. As semi-autonomous organelles, SlSP1 and SlSAD8 mainly participate in the CHLORAD pathway of Toc proteins, involved in the transition from chloroplast to chromoplast structure, while REC1a participates in the chloroplast-chromoplast transition by participating in protein translation within chloroplasts. Fruit ripening-related transcription factors such as TAGL1 and RIN are mainly involved in the process of synergistic pigment accumulation in chromoplasts
| [1] | Ding WQ, Luo YS, Li WL, et al. Fine mapping and transcriptome profiling reveal CpAPRR2 to modulate immature fruit rind color formation in zucchini (Cucurbita pepo) [J]. Theor Appl Genet, 2024, 137(7): 167. |
| [2] | Wise RR. The diversity of plastid form and function [M]//Wise RR, Hoober JK. The Structure and Function of Plastids. Dordrecht: Springer Netherlands, 2006: 3-26. |
| [3] | Choi H, Yi T, Ha SH. Diversity of plastid types and their interconversions [J]. Front Plant Sci, 2021, 12: 692024. |
| [4] | Liebers M, Grübler B, Chevalier F, et al. Regulatory shifts in plastid transcription play a key role in morphological conversions of plastids during plant development [J]. Front Plant Sci, 2017, 8: 23. |
| [5] | Gajecka M, Marzec M, Chmielewska B, et al. Plastid differentiation during microgametogenesis determines green plant regeneration in barley microspore culture [J]. Plant Sci, 2020, 291: 110321. |
| [6] | Rottet S, Besagni C, Kessler F. The role of plastoglobules in thylakoid lipid remodeling during plant development [J]. Biochim Biophys Acta Bioenerg, 2015, 1847(9): 889-899. |
| [7] | Mortimer CL, Misawa N, Perez-Fons L, et al. The formation and sequestration of nonendogenous ketocarotenoids in transgenic Nicotiana glauca [J]. Plant Physiol, 2017, 173(3): 1617-1635. |
| [8] | Rottet S, Devillers J, Glauser G, et al. Identification of plastoglobules as a site of carotenoid cleavage [J]. Front Plant Sci, 2016, 7: 1855. |
| [9] | Garrido A, Conde A, Serôdio J, et al. Fruit photosynthesis: more to know about where, how and why [J]. Plants, 2023, 12(13): 2393. |
| [10] | Dall’Osto L, Cazzaniga S, North H, et al. The Arabidopsis Aba4-1 mutant reveals a specific function for neoxanthin in protection against photooxidative stress [J]. Plant Cell, 2007, 19(3): 1048-1064. |
| [11] | Egea I, Barsan C, Bian WP, et al. Chromoplast differentiation: current status and perspectives [J]. Plant Cell Physiol, 2010, 51(10): 1601-1611. |
| [12] | Kilcrease J, Collins AM, Richins RD, et al. Multiple microscopic approaches demonstrate linkage between chromoplast architecture and carotenoid composition in diverse Capsicum annuum fruit [J]. Plant J, 2013, 76(6): 1074-1083. |
| [13] | Nogueira M, Mora L, Enfissi EMA, et al. Subchromoplast sequestration of carotenoids affects regulatory mechanisms in tomato lines expressing different carotenoid gene combinations [J]. Plant Cell, 2013, 25(11): 4560-4579. |
| [14] | Yan F, Gao YS, Pang XQ, et al. BEL1-LIKE HOMEODOMAIN4 regulates chlorophyll accumulation, chloroplast development, and cell wall metabolism in tomato fruit [J]. J Exp Bot, 2020, 71(18): 5549-5561. |
| [15] | Sagar M, Chervin C, Mila I, et al. SlARF4, an auxin response factor involved in the control of sugar metabolism during tomato fruit development[J]. Plant Physiol, 2013, 161(3): 1362-1374. |
| [16] | Yuan YJ, Mei LH, Wu MB, et al. SlARF10, an auxin response factor, is involved in chlorophyll and sugar accumulation during tomato fruit development [J]. J Exp Bot, 2018, 69(22): 5507-5518. |
| [17] | Centeno DC, Osorio S, Nunes-Nesi A, et al. Malate plays a crucial role in starch metabolism, ripening, and soluble solid content of tomato fruit and affects postharvest softening [J]. Plant Cell, 2011, 23(1): 162-184. |
| [18] | Shi Y, Hu GJ, Wang Y, et al. The SlGRAS9-SlZHD17 transcriptional cascade regulates chlorophyll and carbohydrate metabolism contributing to fruit quality traits in tomato [J]. New Phytol, 2024, 241(6): 2540-2557. |
| [19] | Zita W, Bressoud S, Glauser G, et al. Chromoplast plastoglobules recruit the carotenoid biosynthetic pathway and contribute to carotenoid accumulation during tomato fruit maturation [J]. PLoS One, 2022, 17(12): e0277774. |
| [20] | Liu LH, Shao ZY, Zhang M, et al. Regulation of carotenoid metabolism in tomato [J]. Mol Plant, 2015, 8(1): 28-39. |
| [21] | Kilambi HV, Kumar R, Sharma R, et al. Chromoplast-specific carotenoid-associated protein appears to be important for enhanced accumulation of carotenoids in hp1 tomato fruits [J]. Plant Physiol, 2013, 161(4): 2085-2101. |
| [22] | Vishnevetsky M, Ovadis M, Zuker A, et al. Molecular mechanisms underlying carotenogenesis in the chromoplast: multilevel regulation of carotenoid-associated genes [J]. Plant J, 1999, 20(4): 423-431. |
| [23] | Wang Y, Tian C, Na QT, et al. The role of SlCHRC in carotenoid biosynthesis and plastid development in tomato fruit [J]. Int J Biol Macromol, 2024, 281: 136354. |
| [24] | Manning K, Tör M, Poole M, et al. A naturally occurring epigenetic mutation in a gene encoding an SBP-box transcription factor inhibits tomato fruit ripening [J]. Nat Genet, 2006, 38(8): 948-952. |
| [25] | Aprilyanto V, Wang XW, Wang RF, et al. A comprehensive model of tomato fruit ripening regulation by the transcription factors NOR-like1, NAC-NOR, and MADS-RIN [J]. Plant Physiol, 2025, 198(3): kiaf291. |
| [26] | Hu JH, Wang J, Muhammad T, et al. Integrative analysis of metabolome and transcriptome of carotenoid biosynthesis reveals the mechanism of fruit color change in tomato (Solanum lycopersicum) [J]. Int J Mol Sci, 2024, 25(12): 6493. |
| [27] | Obiadalla-Ali H, Fernie A, Lytovchenko A, et al. Inhibition of chloroplastic fructose 1, 6-bisphosphatase in tomato fruits leads to decreased fruit size, but only small changes in carbohydrate metabolism [J]. Planta, 2004, 219(3): 533-540. |
| [28] | Cackett L, Luginbuehl LH, Schreier TB, et al. Chloroplast development in green plant tissues: the interplay between light, hormone, and transcriptional regulation [J]. New Phytol, 2022, 233(5): 2000-2016. |
| [29] | Powell ALT, Nguyen CV, Hill T, et al. Uniform ripening encodes a Golden 2-like transcription factor regulating tomato fruit chloroplast development [J]. Science, 2012, 336(6089): 1711-1715. |
| [30] | Li GW, Chen DY, Tang XF, et al. Heterologous expression of kiwifruit (Actinidia chinensis) GOLDEN2-LIKE homolog elevates chloroplast level and nutritional quality in tomato (Solanum lycopersicum) [J]. Planta, 2018, 247(6): 1351-1362. |
| [31] | Brand A, Borovsky Y, Hill T, et al. CaGLK2 regulates natural variation of chlorophyll content and fruit color in pepper fruit [J]. Theor Appl Genet, 2014, 127(10): 2139-2148. |
| [32] | Tan YQ, Yang Y, Shen X, et al. Multiple cyclic nucleotide-gated channels function as ABA-activated Ca2+ channels required for ABA-induced stomatal closure in Arabidopsis [J]. Plant Cell, 2023, 35(1): 239-259. |
| [33] | Nakamura H, Muramatsu M, Hakata M, et al. Ectopic overexpression of the transcription factor OsGLK1 induces chloroplast development in non-green rice cells [J]. Plant Cell Physiol, 2009, 50(11): 1933-1949. |
| [34] | Waters MT, Wang P, Korkaric M, et al. GLK transcription factors coordinate expression of the photosynthetic apparatus in Arabidopsis [J]. Plant Cell, 2009, 21(4): 1109-1128. |
| [35] | Zheng MY, Wang XY, Luo J, et al. The pleiotropic functions of GOLDEN2-LIKE transcription factors in plants [J]. Front Plant Sci, 2024, 15: 1445875. |
| [36] | Nadakuduti SS, Holdsworth WL, Klein CL, et al. KNOX genes influence a gradient of fruit chloroplast development through regulation of GOLDEN2-LIKE expression in tomato [J]. Plant J, 2014, 78(6): 1022-1033. |
| [37] | Niu XL, Li HL, Li R, et al. Transcription factor SlBEL2 interferes with GOLDEN2-LIKE and influences green shoulder formation in tomato fruits [J]. Plant J, 2022, 112(4): 982-997. |
| [38] | Tang XF, Miao M, Niu XL, et al. Ubiquitin-conjugated degradation of golden 2-like transcription factor is mediated by CUL4-DDB1-based E3 ligase complex in tomato [J]. New Phytol, 2016, 209(3): 1028-1039. |
| [39] | Pan Y, Bradley G, Pyke K, et al. Network inference analysis identifies an APRR2-like gene linked to pigment accumulation in tomato and pepper fruits [J]. Plant Physiol, 2013, 161(3): 1476-1485. |
| [40] | Ezura K, Lu Y, Suzuki Y, et al. Class II knotted-like homeodomain protein SlKN5 with BEL1-like homeodomain proteins suppresses fruit greening in tomato fruit [J]. Plant J, 2024, 118(6): 2037-2054. |
| [41] | Chen LQ, Dong J, Qiu ZQ, et al. A 13-bp insertion in CmAPRR2 gene disrupts its function in regulating the green rind formation of immature melon fruit (Cucumis melo L.) [J]. Plant Sci, 2025, 359: 112590. |
| [42] | Shinozaki Y, Nicolas P, Fernandez-Pozo N, et al. High-resolution spatiotemporal transcriptome mapping of tomato fruit development and ripening [J]. Nat Commun, 2018, 9: 364. |
| [43] | Lemaire-Chamley M, Koutouan C, Jorly J, et al. A chimeric TGA repressor slows down fruit maturation and ripening in tomato [J]. Plant Cell Physiol, 2022, 63(1): 120-134. |
| [44] | Aluru MR, Stessman DJ, Spalding MH, et al. Alterations in photosynthesis in Arabidopsis lacking IMMUTANS, a chloroplast terminal oxidase [J]. Photosynth Res, 2007, 91(1): 11-23. |
| [45] | Shahbazi M, Gilbert M, Labouré AM, et al. Dual role of the plastid terminal oxidase in tomato [J]. Plant Physiol, 2007, 145(3): 691-702. |
| [46] | Liu GZ, Li CX, Yu HY, et al. GREEN STRIPE, encoding methylated TOMATO AGAMOUS-LIKE 1, regulates chloroplast development and Chl synthesis in fruit [J]. New Phytol, 2020, 228(1): 302-317. |
| [47] | Delannoy E, Le Ret M, Faivre-Nitschke E, et al. Arabidopsis tRNA adenosine deaminase arginine edits the wobble nucleotide of chloroplast tRNAArg (ACG) and is essential for efficient chloroplast translation [J]. Plant Cell, 2009, 21(7): 2058-2071. |
| [48] | Takenaka M, Zehrmann A, Verbitskiy D, et al. RNA editing in plants and its evolution [J]. Annu Rev Genet, 2013, 47: 335-352. |
| [49] | Kramer MC, Anderson SJ, Gregory BD. The nucleotides they are a-changin’: function of RNA binding proteins in post-transcriptional messenger RNA editing and modification in Arabidopsis [J]. Curr Opin Plant Biol, 2018, 45: 88-95. |
| [50] | Yang YF, Liu XY, Wang KR, et al. Molecular and functional diversity of organelle RNA editing mediated by RNA recognition motif-containing protein ORRM4 in tomato [J]. New Phytol, 2020, 228(2): 570-585. |
| [51] | Jia CP, Yang HT, Yang T, et al. Comprehensive metabolomic and transcriptomic analyses reveal SlSnRK2.6/SlCHPP-SlbHLH95-SlPDS module regulating tomato fruit carotene biosynthesis [J]. J Agric Food Chem, 2025, 73(44): 28493-28510. |
| [52] | Sadali NM, Sowden RG, Ling QH, et al. Differentiation of chromoplasts and other plastids in plants [J]. Plant Cell Rep, 2019, 38(7): 803-818. |
| [53] | Larkin RM, Stefano G, Ruckle ME, et al. REDUCED CHLOROPLAST COVERAGE genes from Arabidopsis thaliana help to establish the size of the chloroplast compartment [J]. Proc Natl Acad Sci U S A, 2016, 113(8): E1116-E1125. |
| [54] | Stanley LE, Ding BQ, Sun W, et al. A tetratricopeptide repeat protein regulates carotenoid biosynthesis and chromoplast development in monkeyflowers (Mimulus) [J]. Plant Cell, 2020, 32(5): 1536-1555. |
| [55] | Li L, Yuan H. Chromoplast biogenesis and carotenoid accumulation [J]. Arch Biochem Biophys, 2013, 539(2): 102-109. |
| [56] | D’Andrea L, Simon-Moya M, Llorente B, et al. Interference with Clp protease impairs carotenoid accumulation during tomato fruit ripening [J]. J Exp Bot, 2018, 69(7): 1557-1568. |
| [57] | Barsan C, Sanchez-Bel P, Rombaldi C, et al. Characteristics of the tomato chromoplast revealed by proteomic analysis [J]. J Exp Bot, 2010, 61(9): 2413-2431. |
| [58] | Barsan C, Zouine M, Maza E, et al. Proteomic analysis of chloroplast-to-chromoplast transition in tomato reveals metabolic shifts coupled with disrupted thylakoid biogenesis machinery and elevated energy-production components[J]. Plant Physiol, 2012, 160(2): 708-725. |
| [59] | Ling QH, Huang WH, Baldwin A, et al. Chloroplast biogenesis is regulated by direct action of the ubiquitin-proteasome system [J]. Science, 2012, 338(6107): 655-659. |
| [60] | Ling QH, Broad W, Trösch R, et al. Ubiquitin-dependent chloroplast-associated protein degradation in plants [J]. Science, 2019, 363(6429): eaav4467. |
| [61] | Schnell DJ. The TOC GTPase receptors: regulators of the fidelity, specificity and substrate profiles of the general protein import machinery of chloroplasts [J]. Protein J, 2019, 38(3): 343-350. |
| [62] | Ling QH, Sadali NM, Soufi Z, et al. The chloroplast-associated protein degradation pathway controls chromoplast development and fruit ripening in tomato [J]. Nat Plants, 2021, 7(5): 655-666. |
| [63] | Kessler F. Chloroplast delivery by UPS [J]. Science, 2012, 338(6107): 622-623. |
| [64] | Xu C, Li R, Chen XY, et al. Tomato ripening regulator SlSAD8 disturbs nuclear gene transcription and chloroplast-associated protein degradation [J]. Nat Plants, 2025, 11(11): 2230-2239. |
| [65] | Tan JJ, Zhou ZJ, Feng HQ, et al. Proteome and phosphoproteome of tomato fruit identify reduced chloroplast coverage 1a as a ripening regulator [J]. Genom Proteom Bioinform, 2025: qzaf050. |
| [66] | Guo Y, Bao ZR, Shi MY, et al. Autophagy plays a dual role in chromoplast transition and degradation and is essential for fruit coloration and ripening [J]. Autophagy, 2025, 21(9): 2058-2068. |
| [67] | Iglesias DJ, Cercós M, Colmenero-Flores JM, et al. Physiology of citrus fruiting [J]. Braz J Plant Physiol, 2007, 19(4): 333-362. |
| [68] | Zeng YL, Du JB, Wang L, et al. A comprehensive analysis of chromoplast differentiation reveals complex protein changes associated with plastoglobule biogenesis and remodeling of protein systems in sweet orange flesh [J]. Plant Physiol, 2015, 168(4): 1648-1665. |
| [69] | Li DX, Zhu F. Physicochemical, functional and nutritional properties of kiwifruit flour [J]. Food Hydrocoll, 2019, 92: 250-258. |
| [70] | Zhang MK, Zhang MP, Mazourek M, et al. Regulatory control of carotenoid accumulation in winter squash during storage [J]. Planta, 2014, 240(5): 1063-1074. |
| [71] | Li A, Lin JJ, Zeng ZB, et al. The kiwifruit amyloplast proteome (kfALP): a resource to better understand the mechanisms underlying amyloplast biogenesis and differentiation [J]. Plant J, 2024, 118(2): 565-583. |
| [72] | Song L, Xu W, Zha XX, et al. SlATG8f coordinates ethylene signaling and chloroplast turnover to drive tomato fruit ripening [J]. Plant Cell Rep, 2025, 44(9): 200. |
| [73] | Zheng H, Dong Y, Nong HL, et al. VvSUN may act in the auxin pathway to regulate fruit shape in grape [J]. Hortic Res, 2022, 9: uhac200. |
| [74] | De Jong M, Wolters-Arts M, Feron R, et al. The Solanum lycopersicum auxin response factor 7 (SlARF7) regulates auxin signaling during tomato fruit set and development [J]. Plant J, 2009, 57(1): 160-170. |
| [75] | Jones B, Frasse P, Olmos E, et al. Down-regulation of DR12, an auxin-response-factor homolog, in the tomato results in a pleiotropic phenotype including dark green and blotchy ripening fruit [J]. Plant J, 2002, 32(4): 603-613. |
| [76] | Yuan YJ, Xu X, Gong ZH, et al. Auxin response factor 6A regulates photosynthesis, sugar accumulation, and fruit development in tomato [J]. Hortic Res, 2019, 6: 85. |
| [77] | Hendelman A, Buxdorf K, Stav R, et al. Inhibition of Lamina outgrowth following Solanum lycopersicum auxin response factor 10 (SlARF10) derepression [J]. Plant Mol Biol, 2012, 78(6): 561-576. |
| [78] | Liu LH, Jia CG, Zhang M, et al. Ectopic expression of a BZR1-1D transcription factor in brassinosteroid signalling enhances carotenoid accumulation and fruit quality attributes in tomato [J]. Plant Biotechnol J, 2014, 12(1): 105-115. |
| [79] | Galpaz N, Wang Q, Menda N, et al. Abscisic acid deficiency in the tomato mutant high-pigment 3 leading to increased plastid number and higher fruit lycopene content [J]. Plant J, 2008, 53(5): 717-730. |
| [80] | Chuartzman SG, Nevo R, Shimoni E, et al. Thylakoid membrane remodeling during state transitions in Arabidopsis [J]. Plant Cell, 2008, 20(4): 1029-1039. |
| [81] | Kadirjan-Kalbach DK, Turmo A, Wang J, et al. Allelic variation in the chloroplast division gene FtsZ2-2 leads to natural variation in chloroplast size [J]. Plant Physiol, 2019, 181(3): 1059-1074. |
| [82] | Lira BS, Oliveira MJ, Shiose L, et al. Light and ripening-regulated BBX protein-encoding genes in Solanum lycopersicum [J]. Sci Rep, 2020, 10: 19235. |
| [83] | Gouws A, Steyn WJ. The effect of temperature, region and season on red colour development in apple peel under constant irradiance [J]. Sci Hortic, 2014, 173: 79-85. |
| [84] | Goldschmidt EE. Regulatory aspects of chloro-chromoplast interconversions in senescing citrus fruit peel [J]. Isr J Bot, 1988, 37(2/3/4): 123-130. |
| [85] | Gambi F, Pilkington SM, McAtee PA, et al. Fruit of three kiwifruit (Actinidia chinensis) cultivars differ in their degreening response to temperature after harvest [J]. Postharvest Biol Technol, 2018, 141: 16-23. |
| [86] | Lv JY, Ding SY, Zhang L, et al. Low temperature delays degreening of apple fruit by inhibiting pheophorbide a oxygenase (PAO) pathway and chlorophyll oxidation during ripening [J]. J Food Biochem, 2022, 46(8): e14173. |
| [87] | 葛笑笑, 邓淑芳, 曾凯芳, 等. 贮藏温度对采后蜜橘果皮转色过程中质体结构及相关基因表达的影响 [J]. 食品科学技术学报, 2022, 40(5): 51-61. |
| Ge XX, Deng SF, Zeng KF, et al. Effects of storage temperature on plastid structure and related gene expression during coloring of mandarin fruit peel after harvest [J]. J Food Sci Technol, 2022, 40(5): 51-61. | |
| [88] | Zeng XY, Ye LH, Zhang R, et al. GLK2 a GOLDEN2-LIKE transcription factor, directly regulates anthocyanin accumulation by binding with promoters of key anthocyanin biosynthetic genes in Arabidopsis [J]. Plant Cell Environ, 2025, 48(9): 7055-7071. |
| [1] | DU Lian-da, WEI Meng-meng, CHEN Ze, GUO Wei, ZHAO Ting-ting, HU Da-gang. Effects of Environmental Factors on Sugar-acid Quality in Apple Fruits: Multi-factor Interactions and Environmental Adaption Breeding [J]. Biotechnology Bulletin, 2026, 42(3): 79-95. |
| [2] | ZHAO Yan-xia, LI Qian, SUN Jia-bo, LIANG Hong-min, LI Bing-bing. Key Regulatory Genes and Molecular Networks Dissection Underlying Strawberry Fruit Quality Formation [J]. Biotechnology Bulletin, 2026, 42(3): 111-132. |
| [3] | LIU Yuan-yuan, CHEN Xi-feng, QIAN Qian, GAO Zhen-yu. Advances in Molecular Mechanisms Regulating Panicle Development in Rice [J]. Biotechnology Bulletin, 2025, 41(5): 1-13. |
| [4] | WANG Ya-ping, JIN Lan, HAO Jin-feng, CHANG Ming, WANG Yan-dan, GAO Feng. Identification and Expression Characteristics Analysis of CmRGLG Gene Family in Melon [J]. Biotechnology Bulletin, 2025, 41(12): 156-167. |
| [5] | JIANG Yun-bo, CHEN Xue-xue, ZHAO Yu-sheng. Research Progress in Temperature Regulation of Chloroplast Development in Crops [J]. Biotechnology Bulletin, 2025, 41(10): 20-31. |
| [6] | LEI Mei-ling, RAO Wen-hua, HU Jin-feng, YUE Qi, WU Zu-jian, FAN Guo-cheng. Bacterial Diversity and Structure in Rhizosphere Soil of Citrus Infested with Huanglongbing [J]. Biotechnology Bulletin, 2024, 40(2): 266-276. |
| [7] | LI Yuan-hong, GUO Yu-hao, CAO Yan, ZHU Zhen-zhou, WANG Fei-fei. Research Progress in the Microalgal Growth and Accumulation of Target Products Regulated by Exogenous Phytohormone [J]. Biotechnology Bulletin, 2023, 39(6): 61-72. |
| [8] | LIU Chuan-he, HE Han, HE Xiu-gu, CHEN Xin, LIU Kai, SHAO Xue-hua, LAI Duo, QIN Jian, ZHUANG Qing-li, KUANG Shi-zi, XIAO Wei-qiang. Physiological and Metabolitic Mechanisms of Different Pineapple Cultivars Responding to Low Temperature Stress [J]. Biotechnology Bulletin, 2023, 39(10): 219-230. |
| [9] | HAN Shao-jie, ZHENG Jing-wu. Research Advances on the Functional Study of Host Resistance Genes to Heterodera glycines [J]. Biotechnology Bulletin, 2021, 37(7): 14-24. |
| [10] | LI Ling, YANG Li-xia, GUO Mei. Function of Transcription Factor CNR in the Ripening Process of Tomato Fruit [J]. Biotechnology Bulletin, 2021, 37(2): 51-62. |
| [11] | KUANG Yong-jie, LIU Lang, YAN Fang, REN Bin, YAN Da-qi, ZHANG Da-wei, LIN Hong-hui, ZHOU Huan-bin. Functions of Phytohormones During the Interactions Between Rice, Pathogens [J]. Biotechnology Bulletin, 2018, 34(2): 74-86. |
| [12] | Bai Hua, Wang Peiyun, Tian Xiaowei, Yao Xinling. Review on Crosstalk Regulation Involving in Sugar Signal in Plant [J]. Biotechnology Bulletin, 2013, 29(10): 12-17. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||