








生物技术通报 ›› 2025, Vol. 41 ›› Issue (1): 143-156.doi: 10.13560/j.cnki.biotech.bull.1985.2024-0576
李禹欣1( ), 李苗1,2, 杜晓芬1,2, 韩康妮1, 连世超1, 王军1,2(
), 李苗1,2, 杜晓芬1,2, 韩康妮1, 连世超1, 王军1,2( )
)
收稿日期:2024-06-15
出版日期:2025-01-26
发布日期:2025-01-22
通讯作者:
王军,男,博士,研究员,研究方向:谷子分子育种;E-mail: 128wan@163.com作者简介:李禹欣,女,硕士,助理研究员,研究方向:谷子分子育种;E-mail: ecnulyx@163.com
基金资助:
LI Yu-xin1( ), LI Miao1,2, DU Xiao-fen1,2, HAN Kang-ni1, LIAN Shi-chao1, WANG Jun1,2(
), LI Miao1,2, DU Xiao-fen1,2, HAN Kang-ni1, LIAN Shi-chao1, WANG Jun1,2( )
)
Received:2024-06-15
Published:2025-01-26
Online:2025-01-22
摘要:
【目的】胁迫相关蛋白(stress associated protein, SAP)是一类具有A20/AN1锌指结构域的蛋白,在植物抵御非生物胁迫方面发挥重要作用,目前,谷子SiSAP基因家族的功能尚不明晰。对SiSAP基因家族进行鉴定和表达模式分析,为探究SiSAP基因的功能奠定基础。【方法】通过生物信息学方法对该基因家族进行鉴定及分析,经实时荧光定量PCR(RT-qPCR)验证SiSAP基因家族成员在非生物胁迫和激素处理下的表达模式。【结果】谷子SiSAP基因家族有17个成员,不均匀分布在6条染色体上,15个成员没有内含子,另外2个成员均含有1个内含子;氨基酸序列长度为150-291 aa,分子量介于15.05-32.12 kD,等电点为6.62-9.36;SiSAP基因家族成员与单子叶植物的亲缘关系更近;所有成员启动子序列均含有逆境胁迫和激素响应相关作用元件,以及ERF、Dof、C2H2等转录因子结合位点;转录组数据显示,SiSAP1、SiSAP2、SiSAP3、SiSAP4、SiSAP6、SiSAP7、SiSAP8、SiSAP11和SiSAP14等9个基因在各个组织器官中都有表达,其余8个基因几乎不表达;RT-qPCR结果证实,以上9个基因对低温、高盐和干旱3种非生物胁迫,以及脱落酸、茉莉酸甲酯、生长素和赤霉素4种激素处理的响应模式不同。【结论】SiSAP基因家族可能参与谷子抗旱、耐盐和耐低温胁迫的应答。
李禹欣, 李苗, 杜晓芬, 韩康妮, 连世超, 王军. 谷子SiSAP基因家族的鉴定与表达分析[J]. 生物技术通报, 2025, 41(1): 143-156.
LI Yu-xin, LI Miao, DU Xiao-fen, HAN Kang-ni, LIAN Shi-chao, WANG Jun. Identification and Expression Analysis of SiSAP Gene Family in Foxtail Millet(Setaria italica)[J]. Biotechnology Bulletin, 2025, 41(1): 143-156.
| 基因名称Gene name | 正向引物Forward primer(5'-3') | 反向引物Reverse primer(5'-3') |
|---|---|---|
| SiSAP1 | CTGCTGCTGTAATCCCCAAG | AGTGGCAGTCATGTTTGTCG |
| SiSAP2 | GTTCCTCGCAATTTTCTGGA | CCAAGTACGCGTTTGCATAG |
| SiSAP3 | ACGGTATATCTTGCGCTCGT | GCACTGTTGCCGAGATGTAA |
| SiSAP4 | GCAGATAGCCAAGCAGAACC | CCACCACCTGAAGGAACCTA |
| SiSAP6 | GTCTCCTCTCGCTCATCACG | CCTCACGCTTCTTCTTCTGC |
| SiSAP7 | AAGATTCCATCCAGCAGAGC | CATCACCGCCATCATTACAC |
| SiSAP8 | ATCCGGTGTTGCTAGTGGAG | ATGGGCAAATTTGAAGTCCA |
| SiSAP11 | ATACCGGACCCAAACAACTG | ACCGTCTCAAGACAGGTTCG |
| SiSAP14 | AACCCATTTCCTCTCCCTCA | GGCTCCTCCACCTTCTTGTC |
| SiACTIN | TGATCTCACTGACAGTCTGATG | GATGTCTCTTACAATTTCCCGC |
表1 RT-qPCR引物信息
Table 1 Primers for RT-qPCR in this study
| 基因名称Gene name | 正向引物Forward primer(5'-3') | 反向引物Reverse primer(5'-3') |
|---|---|---|
| SiSAP1 | CTGCTGCTGTAATCCCCAAG | AGTGGCAGTCATGTTTGTCG |
| SiSAP2 | GTTCCTCGCAATTTTCTGGA | CCAAGTACGCGTTTGCATAG |
| SiSAP3 | ACGGTATATCTTGCGCTCGT | GCACTGTTGCCGAGATGTAA |
| SiSAP4 | GCAGATAGCCAAGCAGAACC | CCACCACCTGAAGGAACCTA |
| SiSAP6 | GTCTCCTCTCGCTCATCACG | CCTCACGCTTCTTCTTCTGC |
| SiSAP7 | AAGATTCCATCCAGCAGAGC | CATCACCGCCATCATTACAC |
| SiSAP8 | ATCCGGTGTTGCTAGTGGAG | ATGGGCAAATTTGAAGTCCA |
| SiSAP11 | ATACCGGACCCAAACAACTG | ACCGTCTCAAGACAGGTTCG |
| SiSAP14 | AACCCATTTCCTCTCCCTCA | GGCTCCTCCACCTTCTTGTC |
| SiACTIN | TGATCTCACTGACAGTCTGATG | GATGTCTCTTACAATTTCCCGC |
| 基因Gene | 基因号Gene ID | 基因位置Gene location | 蛋白长度Protein length/aa | 分子量Molecular weight/kD | 等电点pI | 亲水性GRAVY |
|---|---|---|---|---|---|---|
| SiSAP1 | Seita.1G049700 | Chr.1:4 826 713-4828307 | 172 | 18.38 | 7.99 | -0.216 |
| SiSAP2 | Seita.1G050100 | Chr.1:4 844 574-4846624 | 172 | 18.21 | 8.28 | -0.198 |
| SiSAP3 | Seita.1G179600 | Chr.1:25 817 455-25 819 730 | 150 | 15.65 | 9.28 | -0.289 |
| SiSAP4 | Seita.2G045100 | Chr.2:3 644 125-3 646 735 | 165 | 18.09 | 8.02 | -0.479 |
| SiSAP5 | Seita.2G188600 | Chr.2:28 469 651-28 470 203 | 184 | 19.74 | 8.75 | -0.526 |
| SiSAP6 | Seita.2G195200 | Chr.2:29 175 966-29 176 521 | 185 | 19.76 | 9.04 | -0.546 |
| SiSAP7 | Seita.2G252300 | Chr.2:35 515 181-35 516 761 | 170 | 17.55 | 8.90 | -0.234 |
| SiSAP8 | Seita.2G358600 | Chr.2:43 671 478-43 674 297 | 291 | 32.12 | 8.66 | -0.627 |
| SiSAP9 | Seita.5G232000 | Chr.5:29 446 450-29 446 969 | 173 | 17.47 | 8.96 | -0.097 |
| SiSAP10 | Seita.5G232100 | Chr.5:29 462 246-29 462 780 | 178 | 17.86 | 9.06 | -0.063 |
| SiSAP11 | Seita.5G297700 | Chr.5:35 289 769-35 291 603 | 154 | 15.86 | 8.74 | -0.352 |
| SiSAP12 | Seita.5G330400 | Chr.5:37 812 781-37 813 282 | 167 | 17.11 | 8.76 | -0.089 |
| SiSAP13 | Seita.6G159700 | Chr.6:28 237 350-28 238 434 | 256 | 26.19 | 8.47 | -0.190 |
| SiSAP14 | Seita.6G203700 | Chr.6:32 355 017-32 361 096 | 179 | 18.82 | 9.14 | -0.357 |
| SiSAP15 | Seita.7G243600 | Chr.7:30 408 518-30 409 232 | 236 | 24.02 | 8.60 | -0.373 |
| SiSAP16 | Seita.9G061100 | Chr.9:3 504 533-3 504 971 | 146 | 15.05 | 9.36 | -0.312 |
| SiSAP17 | Seita.9G316800 | Chr.9:36 489 146-36 489 611 | 155 | 16.52 | 6.62 | -0.408 |
表2 SiSAP家族成员的鉴定
Table 2 Identification of SiSAP gene family members
| 基因Gene | 基因号Gene ID | 基因位置Gene location | 蛋白长度Protein length/aa | 分子量Molecular weight/kD | 等电点pI | 亲水性GRAVY |
|---|---|---|---|---|---|---|
| SiSAP1 | Seita.1G049700 | Chr.1:4 826 713-4828307 | 172 | 18.38 | 7.99 | -0.216 |
| SiSAP2 | Seita.1G050100 | Chr.1:4 844 574-4846624 | 172 | 18.21 | 8.28 | -0.198 |
| SiSAP3 | Seita.1G179600 | Chr.1:25 817 455-25 819 730 | 150 | 15.65 | 9.28 | -0.289 |
| SiSAP4 | Seita.2G045100 | Chr.2:3 644 125-3 646 735 | 165 | 18.09 | 8.02 | -0.479 |
| SiSAP5 | Seita.2G188600 | Chr.2:28 469 651-28 470 203 | 184 | 19.74 | 8.75 | -0.526 |
| SiSAP6 | Seita.2G195200 | Chr.2:29 175 966-29 176 521 | 185 | 19.76 | 9.04 | -0.546 |
| SiSAP7 | Seita.2G252300 | Chr.2:35 515 181-35 516 761 | 170 | 17.55 | 8.90 | -0.234 |
| SiSAP8 | Seita.2G358600 | Chr.2:43 671 478-43 674 297 | 291 | 32.12 | 8.66 | -0.627 |
| SiSAP9 | Seita.5G232000 | Chr.5:29 446 450-29 446 969 | 173 | 17.47 | 8.96 | -0.097 |
| SiSAP10 | Seita.5G232100 | Chr.5:29 462 246-29 462 780 | 178 | 17.86 | 9.06 | -0.063 |
| SiSAP11 | Seita.5G297700 | Chr.5:35 289 769-35 291 603 | 154 | 15.86 | 8.74 | -0.352 |
| SiSAP12 | Seita.5G330400 | Chr.5:37 812 781-37 813 282 | 167 | 17.11 | 8.76 | -0.089 |
| SiSAP13 | Seita.6G159700 | Chr.6:28 237 350-28 238 434 | 256 | 26.19 | 8.47 | -0.190 |
| SiSAP14 | Seita.6G203700 | Chr.6:32 355 017-32 361 096 | 179 | 18.82 | 9.14 | -0.357 |
| SiSAP15 | Seita.7G243600 | Chr.7:30 408 518-30 409 232 | 236 | 24.02 | 8.60 | -0.373 |
| SiSAP16 | Seita.9G061100 | Chr.9:3 504 533-3 504 971 | 146 | 15.05 | 9.36 | -0.312 |
| SiSAP17 | Seita.9G316800 | Chr.9:36 489 146-36 489 611 | 155 | 16.52 | 6.62 | -0.408 |
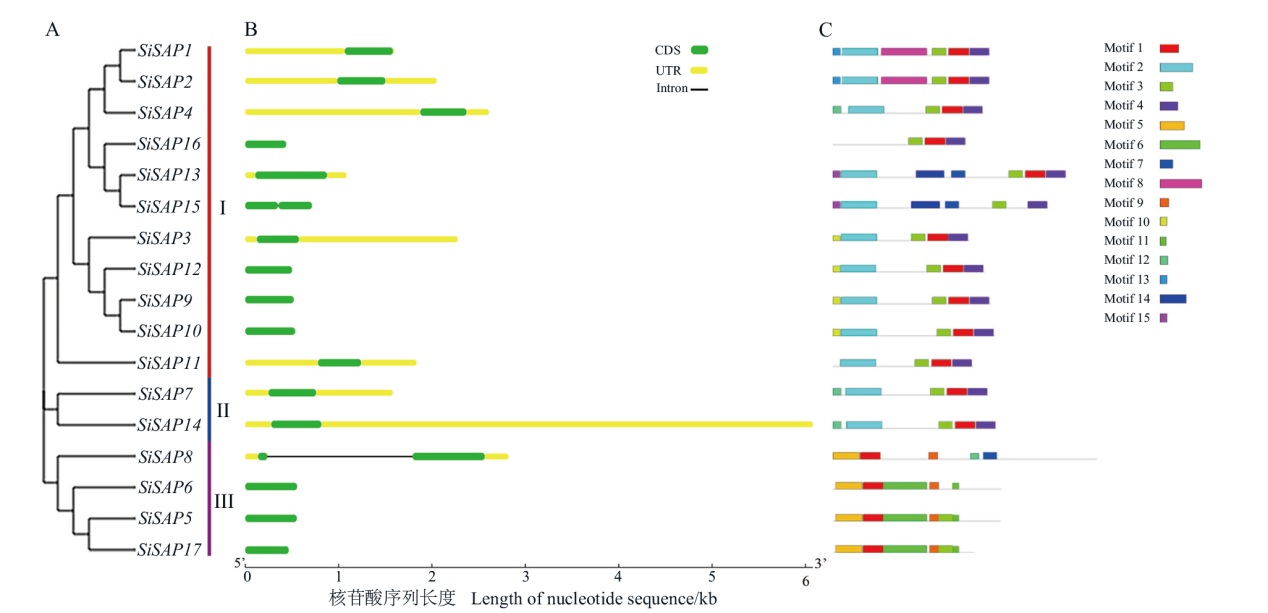
图2 SiSAP基因家族进化树、基因结构及motifs分析 A:进化关系;B:基因结构;C:蛋白保守基序;motif2、motif14-motif7为A20;motif3-motif1、motif5-motif1、motif9-motif3-motif11为AN1
Fig. 2 Analysis of SiSAP gene family evolutionary tree, gene structure and motifs A: Phylogenetic relationship. B: Gene structure. C: Conserved motifs. Motif2 and motif14-motif7 are A20; motif3-motif1, motif5-motif1 and motif9-motif3-motif11 are AN1

图3 SiSAP蛋白序列比对分析 蓝框代表A20保守结构域,红框代表AN1保守结构域
Fig. 3 Alignment and analysis of sequences of SiSAP protein A20 and AN1 are highlighted with blue and red squares, respectively
| 同源基因 Homologous gene | 非同义替换率(Ka) Non-synonymous substitution | 同义替换率(Ks) Synonymous substitution | 非同义替换率/同义替换率 Ka/Ks |
|---|---|---|---|
| SiSAP4-SiSAP16 | 0.41 | 1.21 | 0.34 |
| SiSAP7-SiSAP14 | 0.20 | 0.47 | 0.42 |
表3 SiSAP家族片段重复基因对的Ka/Ks值
Table 3 Ka/Ks ratios of segmental duplicated gene pairs in the SiSAP gene family
| 同源基因 Homologous gene | 非同义替换率(Ka) Non-synonymous substitution | 同义替换率(Ks) Synonymous substitution | 非同义替换率/同义替换率 Ka/Ks |
|---|---|---|---|
| SiSAP4-SiSAP16 | 0.41 | 1.21 | 0.34 |
| SiSAP7-SiSAP14 | 0.20 | 0.47 | 0.42 |

图6 植物SAP基因家族系统发育树 红色五角星:谷子;橙色对钩:水稻;蓝色三角形:小麦;绿色圆形:大麦;黄色正方形:拟南芥;紫色方框:刚毛柽柳
Fig. 6 Phylogenetic tree of SAP gene family in plants Red stars: Setaria italica. Orange checks: Oraza sativa. Blue triangles: Triticum aestivum. Green circles: Hordeum vulgare. Yellow rects: Arabidopsis thaliana. Purple squares: Tamarix hispida

图7 谷子SiSAP基因家族的顺式作用元件及转录因子结合位点预测分析 A:SiSAP家族基因启动子序列顺式作用元件分析。网格中不同强度的颜色和数量表示SiSAP基因中的顺式作用元件数量;B:不同颜色的直方图表示每个类别中的顺式作用元件的总和;C:显示前15个与SiSAP基因启动子结合的转录因子家族;网格中不同强度的颜色和数量表示转录因子结合位点的数量
Fig. 7 Prediction of cis-acting element and transcription factor binding sites in the SiSAP gene family of foxtail millet A: Analysis of cis-acting element in promoter sequence of SiSAP family genes. The different intensity colors and numbers of the grid indicate the cis-acting element numbers in the SiSAP genes. B: The different colored histogram represents the sum of the cis-acting elements in each category. C: Only the top 15 TFs that bind to the promoter of SiSAP genes are shown here. The different intensity colors and numbers of the grid indicate the numbers of putative TF sites
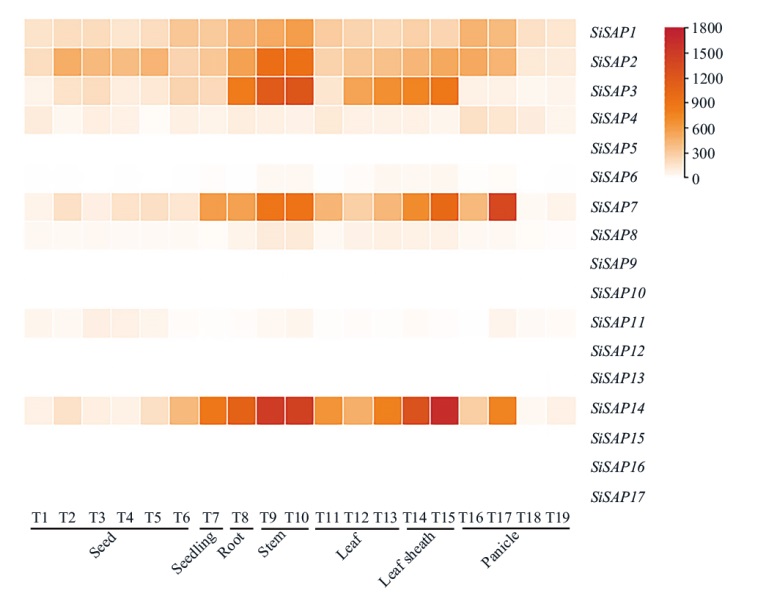
图8 谷子SiSAP基因家族的组织表达模式分析 T1:未成熟种子S1;T2:未成熟种子S2;T3:未成熟种子S3;T4:未成熟种子S4;T5:未成熟种子S5;T6:萌发3 d的种子;T7:两叶一心幼苗;T8:灌浆期根;T9:灌浆期穗下节;T10:灌浆期顶端第2茎;T11:抽穗2 d后的新叶(顶2和3叶);T12:灌浆期旗叶;T13:灌浆期顶端第4叶;T14:灌浆期旗叶叶鞘;T15:灌浆期顶端第4叶叶鞘;T16:分化初始阶段穗;T17:分化第3阶段穗;T18:未成熟小穗S2;T19:未成熟小穗S4
Fig. 8 Tissue expression pattern analysis of SiSAP gene family in foxtail millet T1: Immature seed at S1. T2: Immature seed at S2. T3: Immature seed at S3. T4: Immature seed at S4. T5: Immature seed at S5. T6: Seeds germinated for 3 d. T7: Plant of one-tip-two-leaf. T8: Root at filling stage. T9: Neck-panicle-internodes at filling stage. T10: Stem-top-second at filling stage. T11: Leaves after 2 d of heading(Leaf-top-2-3). T12: Flag leaf at filling stage. T13: Leaf-top-fourth at filling stage. T14: Flag leaf sheath at filling stage. T15: Leaf-sheath-top-fourth at filling stage. T16: Primary panicle at differentiation stage. T17: Third panicle at differentiation stage. T18: Immature spikelet at S2. T19: Immature spikelet at S4
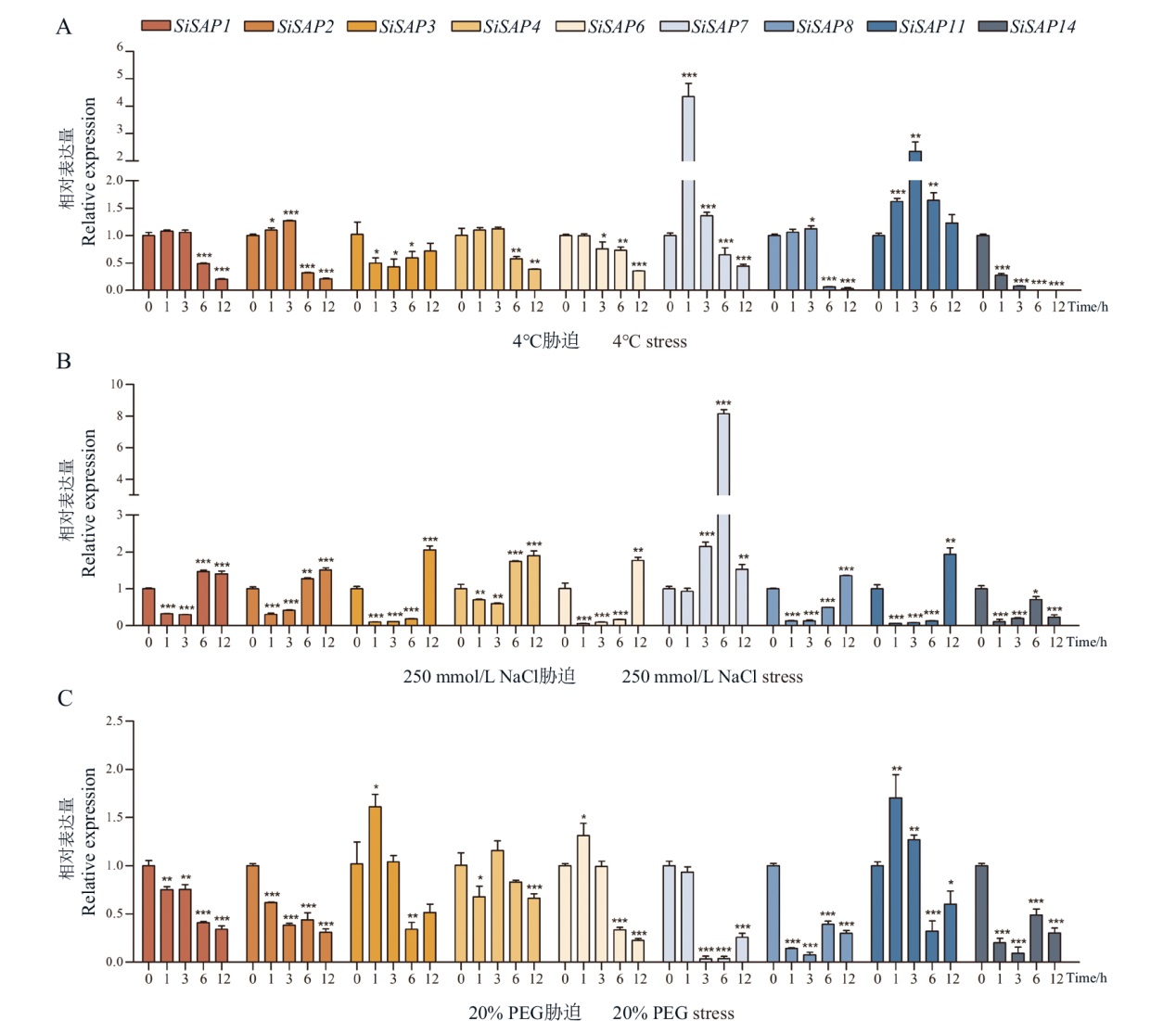
图9 非生物胁迫下SiSAP基因的表达模式分析 横坐标表示各胁迫处理后0、1、3、6和12 h。*,**,***分别表示P<0.05,P<0.01,P<0.001。下同
Fig. 9 Expression patterns of SiSAP gene in response to abiotic stress Abscissa indicates 0, 1, 3, 6 and 12 h after treatment. *, **, *** indicate P < 0.05, P < 0.01 and P < 0.001, respectively. The same below
| [1] |
Kanneganti V, Gupta AK. Overexpression of OsiSAP8, a member of stress associated protein(SAP)gene family of rice confers tolerance to salt, drought and cold stress in transgenic tobacco and rice[J]. Plant Mol Biol, 2008, 66(5): 445-462.
doi: 10.1007/s11103-007-9284-2 pmid: 18205020 |
| [2] | Yoon Y, Seo DH, Shin H, et al. The role of stress-responsive transcription factors in modulating abiotic stress tolerance in plants[J]. Agronomy, 2020, 10(6): 788. |
| [3] | Wan FX, Xu YH, Wang SL, et al. Identification and expression analysis of zinc finger A20/AN1 stress-associated genes SmSAP responding to abiotic stress in eggplant[J]. Horticulturae, 2022, 8(2): 108. |
| [4] | Zhao X, Wang R, Zhang Y, et al. Comprehensive analysis of the stress associated protein(SAP)gene family in Tamarix hispida and the function of ThSAP6 in salt tolerance[J]. Plant Physiol Biochem, 2021, 165: 1-9. |
| [5] |
丁兰, 顾宝, 李培楹, 等. 葡萄SAP家族的鉴定与表达分析[J]. 中国农业科学, 2019, 52(14): 2500-2514.
doi: 10.3864/j.issn.0578-1752.2019.14.009 |
| Ding L, Gu B, Li PY, et al. Genome-wide identification and expression analysis of SAP family in grape[J]. Sci Agric Sin, 2019, 52(14): 2500-2514. | |
| [6] |
杜志烨, 李明玉, 陈稷, 等. 植物胁迫相关蛋白功能研究进展[J]. 植物学报, 2024, 59(1): 110-121.
doi: 10.11983/CBB23029 |
| Du ZY, Li MY, Chen J, et al. Research advances in plant stress associated protein functions[J]. Chin Bull Bot, 2024, 59(1): 110-121. | |
| [7] |
Mukhopadhyay A, Vij S, Tyagi AK. Overexpression of a zinc-finger protein gene from rice confers tolerance to cold, dehydration, and salt stress in transgenic tobacco[J]. Proc Natl Acad Sci USA, 2004, 101(16): 6309-6314.
doi: 10.1073/pnas.0401572101 pmid: 15079051 |
| [8] | 苏晓帅, 张宝华, 刘佳静, 等. 小麦SAPs家族分析及TaSAP1;1耐盐和低磷胁迫功能研究[J]. 植物遗传资源学报, 2022, 23(3): 857-870. |
| Su XS, Zhang BH, Liu JJ, et al. Genomic analysis of stress associated proteins in wheat and functional study of TaSAP1;1 in salt and low-pi tolerance[J]. J Plant Genet Resour, 2022, 23(3): 857-870. | |
| [9] | Vij S, Tyagi AK. Genome-wide analysis of the stress associated protein(SAP)gene family containing A20/AN1 zinc-finger(s)in rice and their phylogenetic relationship with Arabidopsis[J]. Mol Genet Genom, 2006, 276(6): 565-575. |
| [10] |
Baidyussen A, Aldammas M, Kurishbayev A, et al. Identification, gene expression and genetic polymorphism of zinc finger A20/AN1 stress-associated genes, HvSAP, in salt stressed barley from Kazakhstan[J]. BMC Plant Biol, 2020, 20(Suppl 1): 156.
doi: 10.1186/s12870-020-02332-4 pmid: 33050881 |
| [11] | Zhu FJ, Wang K, Li DN, et al. OsSAP6 positively regulates soda saline-alkaline stress tolerance in rice[J]. Rice, 2022, 15(1): 69. |
| [12] | Wang X, Zou BH, Shao QL, et al. Natural variation reveals that OsSAP16 controls low-temperature germination in rice[J]. J Exp Bot, 2018, 69(3): 413-421. |
| [13] |
丁林云, 张微, 王晋成, 等. 过量表达棉花GhSAP1提高转基因烟草的耐盐性[J]. 中国农业科学, 2014, 47(8): 1458-1470.
doi: 10.3864/j.issn.0578-1752.2014.08.002 |
| Ding LY, Zhang W, Wang JC, et al. Overexpression of a Gossypium hirsutum stress-associated protein gene(GhSAP1)improves salt stress tolerance in transgenic tobacco[J]. Sci Agric Sin, 2014, 47(8): 1458-1470. | |
| [14] | 王亦学, 董艳辉, 张欢欢, 等. 陆地棉胁迫相关蛋白基因GhSAP8的克隆及其耐盐性分析[J]. 农业生物技术学报, 2020, 28(7): 1230-1239. |
| Wang YX, Dong YH, Zhang HH, et al. Cloning and salt-tolerance analysis of stress associated protein gene GhSAP8 in Gossypium hirsutum[J]. J Agric Biotechnol, 2020, 28(7): 1230-1239. | |
| [15] |
王亦学, 郝曜山, 张欢欢, 等. 小麦胁迫相关蛋白基因TaSAP12-D的耐盐性分析[J]. 核农学报, 2023, 37(1): 42-50.
doi: 10.11869/j.issn.1000-8551.2023.01.0042 |
| Wang YX, Hao YS, Zhang HH, et al. Identification and analysis of salt-tolerance of stress associated protein gene(TaSAP12-D)from wheat[J]. J Nucl Agric Sci, 2023, 37(1): 42-50. | |
| [16] | Xu QF, Mao XG, Wang YX, et al. A wheat gene TaSAP17-D encoding an AN1/AN1 zinc finger protein improves salt stress tolerance in transgenic Arabidopsis[J]. J Integr Agric, 2018, 17(3): 507-516. |
| [17] | Sharma G, Giri J, Tyagi AK. Rice OsiSAP7 negatively regulates ABA stress signalling and imparts sensitivity to water-deficit stress in Arabidopsis[J]. Plant Sci, 2015, 237: 80-92. |
| [18] | Ben Saad R, Ben Romdhane W, Mihoubi W, et al. A Lobularia maritima LmSAP protein modulates gibberellic acid homeostasis via its A20 domain under abiotic stress conditions[J]. PLoS One, 2020, 15(5): e0233420. |
| [19] |
李顺国, 刘斐, 刘猛, 等. 中国谷子产业和种业发展现状与未来展望[J]. 中国农业科学, 2021, 54(3): 459-470.
doi: 10.3864/j.issn.0578-1752.2021.03.001 |
|
Li SG, Liu F, Liu M, et al. Current status and future prospective of foxtail millet production and seed industry in China[J]. Sci Agric Sin, 2021, 54(3): 459-470.
doi: 10.3864/j.issn.0578-1752.2021.03.001 |
|
| [20] | Lai DL, Fan Y, Xue GX, et al. Genome-wide identification and characterization of the SPL gene family and its expression in the various developmental stages and stress conditions in foxtail millet(Setaria italica)[J]. BMC Genomics, 2022, 23(1): 389. |
| [21] |
Fan Y, Lai DL, Yang H, et al. Genome-wide identification and expression analysis of the bHLH transcription factor family and its response to abiotic stress in foxtail millet(Setaria italica L.)[J]. BMC Genomics, 2021, 22(1): 778.
doi: 10.1186/s12864-021-08095-y pmid: 34717536 |
| [22] |
Fan Y, Wei XB, Lai DL, et al. Genome-wide investigation of the GRAS transcription factor family in foxtail millet(Setaria italica L.)[J]. BMC Plant Biol, 2021, 21(1): 508.
doi: 10.1186/s12870-021-03277-y pmid: 34732123 |
| [23] | 雷彪, 曲瑞芳, 任超, 等. 谷子LBD基因家族的鉴定及其对逆境胁迫的响应[J]. 植物生理学报, 2023, 59(3): 527-542. |
| Lei B, Qu RF, Ren C, et al. Identification of SiLBDs gene family in foxtail millet and its participation in stress response[J]. Plant Physiol J, 2023, 59(3): 527-542. | |
| [24] | 桑璐曼, 汤沙, 张仁梁, 等. 谷子热激蛋白HSP90基因家族鉴定及分析[J]. 植物遗传资源学报, 2022, 23(4): 1085-1097. |
| Sang LM, Tang S, Zhang RL, et al. Identification and analysis of heat shock protein HSP90 family genes in foxtail millet[J]. J Plant Genet Resour, 2022, 23(4): 1085-1097. | |
| [25] | Chen CJ, Wu Y, Li JW, et al. TBtools-II: a “one for all, all for one” bioinformatics platform for biological big-data mining[J]. Mol Plant, 2023, 16(11): 1733-1742. |
| [26] |
Kumar S, Stecher G, Tamura K. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets[J]. Mol Biol Evol, 2016, 33(7): 1870-1874.
doi: 10.1093/molbev/msw054 pmid: 27004904 |
| [27] |
Jeffares DC, Penkett CJ, Bähler J. Rapidly regulated genes are intron poor[J]. Trends Genet, 2008, 24(8): 375-378.
doi: 10.1016/j.tig.2008.05.006 pmid: 18586348 |
| [28] |
Giri J, Dansana PK, Kothari KS, et al. SAPs as novel regulators of abiotic stress response in plants[J]. Bioessays, 2013, 35(7): 639-648.
doi: 10.1002/bies.201200181 pmid: 23640876 |
| [29] | Gao W, Long L, Tian XQ, et al. Genome-wide identification and expression analysis of stress-associated proteins(SAPs)containing A20/AN1 zinc finger in cotton[J]. Mol Genet Genomics, 2016, 291(6): 2199-2213. |
| [30] | Zhou Y, Zeng LM, Chen RR, et al. Genome-wide identification and characterization of stress-associated protein(SAP)gene family encoding A20/AN1 zinc-finger proteins in Medicago truncatula[J]. Arch Biol Sci, 2018, 70(1): 87-98. |
| [31] | Dong QL, Duan DY, Zhao S, et al. Genome-wide analysis and cloning of the apple stress-associated protein gene family reveals MdSAP15, which confers tolerance to drought and osmotic stresses in transgenic Arabidopsis[J]. Int J Mol Sci, 2018, 19(9): 2478. |
| [32] | Kong FL, Wang J, Cheng L, et al. Genome-wide analysis of the mitogen-activated protein kinase gene family in Solanum lycopersicum[J]. Gene, 2012, 499(1): 108-120. |
| [33] |
Matsukura S, Mizoi J, Yoshida T, et al. Comprehensive analysis of rice DREB2-type genes that encode transcription factors involved in the expression of abiotic stress-responsive genes[J]. Mol Genet Genomics, 2010, 283(2): 185-196.
doi: 10.1007/s00438-009-0506-y pmid: 20049613 |
| [34] |
Tyagi H, Jha S, Sharma M, et al. Rice SAPs are responsive to multiple biotic stresses and overexpression of OsSAP1, an A20/AN1 zinc-finger protein, enhances the basal resistance against pathogen infection in tobacco[J]. Plant Sci, 2014, 225: 68-76.
doi: 10.1016/j.plantsci.2014.05.016 pmid: 25017161 |
| [35] | Kothari KS, Dansana PK, Giri J, et al. Rice stress associated protein 1(OsSAP1)interacts with aminotransferase(OsAMTR1)and pathogenesis-related 1a protein(OsSCP)and regulates abiotic stress responses[J]. Front Plant Sci, 2016, 7: 1057. |
| [36] | Muthuramalingam P, Jeyasri R, Selvaraj A, et al. Global transcriptome analysis of novel stress associated protein(SAP)genes expression dynamism of combined abiotic stresses in Oryza sativa(L.)[J]. J Biomol Struct Dyn, 2021, 39(6): 2106-2117. |
| [1] | 殷缘, 程爽, 刘定豪, 邓晓霞, 李凯月, 王竞红, 蔺吉祥. 外源过氧化氢(H2O2)影响非生物胁迫下植物生长与生理代谢机制的研究进展[J]. 生物技术通报, 2025, 41(1): 1-13. |
| [2] | 武志健, 刘广洋, 林志豪, 盛彬, 陈鸽, 许晓敏, 王军伟, 徐东辉. 蔬菜种子萌发的纳米调控及其机制研究进展[J]. 生物技术通报, 2025, 41(1): 14-24. |
| [3] | 孔青洋, 张晓龙, 李娜, 张晨洁, 张雪云, 于超, 张启翔, 罗乐. 单叶蔷薇GRAS转录因子家族鉴定及表达分析[J]. 生物技术通报, 2025, 41(1): 210-220. |
| [4] | 申鹏, 高雅彬, 丁红. 马铃薯SAT基因家族的鉴定和表达分析[J]. 生物技术通报, 2024, 40(9): 64-73. |
| [5] | 宋兵芳, 柳宁, 程新艳, 徐晓斌, 田文茂, 高悦, 毕阳, 王毅. 马铃薯G6PDH基因家族鉴定及其在损伤块茎的表达分析[J]. 生物技术通报, 2024, 40(9): 104-112. |
| [6] | 吴慧琴, 王延宏, 刘涵, 司政, 刘雪晴, 王静, 阳宜, 成妍. 辣椒UGT基因家族的鉴定及表达分析[J]. 生物技术通报, 2024, 40(9): 198-211. |
| [7] | 满全财, 孟姿诺, 李伟, 蔡心汝, 苏润东, 付长青, 高顺娟, 崔江慧. 马铃薯AQP基因家族鉴定及表达分析[J]. 生物技术通报, 2024, 40(9): 51-63. |
| [8] | 周冉, 王兴平, 李彦霞, 罗仍卓么. 金黄色葡萄球菌型乳房炎奶牛乳腺组织的lncRNA差异表达分析[J]. 生物技术通报, 2024, 40(8): 320-328. |
| [9] | 崔原瑗, 王昭懿, 白双宇, 任毓昭, 豆飞飞, 刘彩霞, 刘凤楼, 王掌军, 李清峰. 大麦非特异性磷脂酶C基因家族全基因组鉴定及苗期胁迫表达分析[J]. 生物技术通报, 2024, 40(8): 74-82. |
| [10] | 刘丹丹, 王雷刚, 孙明慧, 焦小雨, 吴琼, 王文杰. 茶树海藻糖-6-磷酸合成酶(TPS)基因家族鉴定与表达分析[J]. 生物技术通报, 2024, 40(8): 152-163. |
| [11] | 杨巍, 赵丽芬, 唐兵, 周麟笔, 杨娟, 莫传园, 张宝会, 李飞, 阮松林, 邓英. 芥菜SRO基因家族全基因组鉴定与表达分析[J]. 生物技术通报, 2024, 40(8): 129-141. |
| [12] | 余纽, 柳帆, 杨锦昌. 油楠SgTPS7的克隆及其在萜类生物合成和非生物胁迫中的功能[J]. 生物技术通报, 2024, 40(8): 164-173. |
| [13] | 李亦君, 杨小贝, 夏琳, 罗朝鹏, 徐馨, 杨军, 宁黔冀, 武明珠. 烟草NtPRR37基因克隆及功能分析[J]. 生物技术通报, 2024, 40(8): 221-231. |
| [14] | 李勇慧, 鲍星星, 段一珂, 赵运霞, 于相丽, 陈尧, 张延召. 灵宝杜鹃bZIP家族全基因组鉴定及表达特征分析[J]. 生物技术通报, 2024, 40(8): 186-198. |
| [15] | 李雨晴, 吴楠, 罗建让. 卵叶牡丹花色苷合成相关基因bHLH的克隆与功能分析[J]. 生物技术通报, 2024, 40(8): 174-185. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||