








生物技术通报 ›› 2024, Vol. 40 ›› Issue (9): 51-63.doi: 10.13560/j.cnki.biotech.bull.1985.2024-0439
• 薯类作物生物技术专题(专题主编:徐建飞,尚轶) • 上一篇 下一篇
满全财( ), 孟姿诺, 李伟, 蔡心汝, 苏润东, 付长青, 高顺娟, 崔江慧(
), 孟姿诺, 李伟, 蔡心汝, 苏润东, 付长青, 高顺娟, 崔江慧( )
)
收稿日期:2024-05-10
出版日期:2024-09-26
发布日期:2024-10-12
通讯作者:
崔江慧,女,博士,高级实验师,研究方向:马铃薯遗传育种;E-mail: cjianghui521@126.com作者简介:满全财,男,硕士,研究方向:马铃薯遗传育种;E-mail: 15715377671@163.com
基金资助:
MAN Quan-cai( ), MENG Zi-nuo, LI Wei, CAI Xin-ru, SU Run-dong, FU Chang-qing, GAO Shun-juan, CUI Jiang-hui(
), MENG Zi-nuo, LI Wei, CAI Xin-ru, SU Run-dong, FU Chang-qing, GAO Shun-juan, CUI Jiang-hui( )
)
Received:2024-05-10
Published:2024-09-26
Online:2024-10-12
摘要:
【目的】水通道蛋白(Aquaporins,AQPs)在植物非生物胁迫下的植物生长发育、水分运输和应激反应等生理过程中至关重要,本研究旨在揭示马铃薯StAQP家族的特性及功能。【方法】对马铃薯StAQP基因家族进行全基因组鉴定,利用生物信息学手段对基因结构、保守基序、顺式作用元件、染色体定位以及基因共线性等进行分析,并结合转录组数据以及RT-qPCR对基因表达模式进行验证。【结果】马铃薯StAQP基因家族的44个成员分布在PIP、TIP、NIP、SIP和XIP五个亚族中,相同亚族的家族成员的基因结构和保守基序基本一致,同时马铃薯AQP家族成员大多数分布在质膜上;马铃薯StAQP家族成员中含有大量和光反应以及胁迫响应有关的顺式作用元件;共线性结果表明,马铃薯AQP家族与拟南芥、烟草、番茄和辣椒4个物种的AQP家族之间存在大量同源基因对;转录组和RT-qPCR验证结果表明,马铃薯StAQP家族成员在干旱胁迫和盐胁迫下高度表达,不同家族成员表达存在差异,同时相同基因在不同胁迫下的表达情况也存在差异,其中StAQP7、StAQP28和StAQP31这3个基因是StAQP家族响应非生物胁迫的关键基因。【结论】StAQPs基因在马铃薯的生长发育和非生物胁迫响应中发挥重要作用。
满全财, 孟姿诺, 李伟, 蔡心汝, 苏润东, 付长青, 高顺娟, 崔江慧. 马铃薯AQP基因家族鉴定及表达分析[J]. 生物技术通报, 2024, 40(9): 51-63.
MAN Quan-cai, MENG Zi-nuo, LI Wei, CAI Xin-ru, SU Run-dong, FU Chang-qing, GAO Shun-juan, CUI Jiang-hui. Identification and Expression Analysis of AQP Gene Family in Potato[J]. Biotechnology Bulletin, 2024, 40(9): 51-63.
| 基因 Gene | 基因ID Gene ID | 染色体位置Chromosome localization | 氨基酸长度Amino acid length/aa | 相对分子量Molecular weight/Da | 理论等电点Theoretical isoelectric point | 亚细胞定位Subcellular localization |
|---|---|---|---|---|---|---|
| StAQP1 | PGSC0003DMG400034503 | Chr01 | 287 | 30698.7 | 6.50 | 质膜Plasma membrane |
| StAQP2 | PGSC0003DMG400000045 | Chr01 | 285 | 30573.2 | 8.84 | 质膜Plasma membrane |
| StAQP3 | PGSC0003DMG400024879 | Chr01 | 112 | 12379.6 | 9.83 | 细胞质Cytoplasm |
| StAQP4 | PGSC0003DMG400001640 | Chr01 | 248 | 26735.5 | 8.21 | 质膜Plasma membrane |
| StAQP5 | PGSC0003DMG400006183 | Chr01 | 283 | 30119.8 | 8.60 | 质膜Plasma membrane |
| StAQP6 | PGSC0003DMG400028456 | Chr02 | 181 | 19461.6 | 6.34 | 液泡Vacuole |
| StAQP7 | PGSC0003DMG401028457 | Chr02 | 290 | 30571 | 9.10 | 质膜Plasma membrane |
| StAQP8 | PGSC0003DMG400003587 | Chr02 | 251 | 26556.8 | 8.08 | 质膜Plasma membrane |
| StAQP9 | PGSC0003DMG400030648 | Chr02 | 267 | 28189.8 | 8.45 | 细胞骨架Cytoskeleton |
| StAQP10 | PGSC0003DMG400027819 | Chr03 | 277 | 29657.2 | 9.55 | 质膜Plasma membrane |
| StAQP11 | PGSC0003DMG400020742 | Chr03 | 286 | 30688.4 | 8.83 | 质膜Plasma membrane |
| StAQP12 | PGSC0003DMG400014818 | Chr03 | 251 | 25951 | 8.67 | 叶绿体Chloroplast |
| StAQP13 | PGSC0003DMG400015275 | Chr03 | 260 | 27777.9 | 9.22 | 细胞质Cytoplasm |
| StAQP14 | PGSC0003DMG400014123 | Chr03 | 306 | 31684.7 | 8.34 | 质膜Plasma membrane |
| StAQP15 | PGSC0003DMG400002552 | Chr03 | 249 | 25161.1 | 6.33 | 细胞质Cytoplasm |
| StAQP16 | PGSC0003DMG401030499 | Chr05 | 272 | 29260.3 | 8.20 | 细胞质Cytoplasm |
| StAQP17 | PGSC0003DMG400023466 | Chr05 | 275 | 29328.2 | 9.65 | 质膜Plasma membrane |
| StAQP18 | PGSC0003DMG400024197 | Chr06 | 286 | 30662.3 | 7.18 | 质膜Plasma membrane |
| StAQP19 | PGSC0003DMG400011875 | Chr06 | 250 | 25151 | 5.48 | 液泡Vacuole |
| StAQP20 | PGSC0003DMG400016528 | Chr06 | 249 | 25089 | 6.06 | 质膜Plasma membrane |
| StAQP21 | PGSC0003DMG400026969 | Chr06 | 259 | 27476.5 | 7.32 | 细胞质Cytoplasm |
| StAQP22 | PGSC0003DMG401005949 | Chr06 | 347 | 37712.9 | 8.32 | 质膜Plasma membrane |
| StAQP23 | PGSC0003DMG400007134 | Chr06 | 250 | 25749.5 | 6.13 | 细胞质Cytoplasm |
| StAQP24 | PGSC0003DMG400030308 | Chr06 | 253 | 26056.9 | 5.48 | 液泡Vacuole |
| StAQP25 | PGSC0003DMG400005780 | Chr08 | 287 | 30790.4 | 7.96 | 质膜Plasma membrane |
| StAQP26 | PGSC0003DMG400005803 | Chr08 | 296 | 30651.2 | 8.48 | 质膜Plasma membrane |
| StAQP27 | PGSC0003DMG400009604 | Chr08 | 247 | 26042.5 | 6.22 | 液泡Vacuole |
| StAQP28 | PGSC0003DMG400012337 | Chr08 | 286 | 30686.6 | 9.34 | 质膜Plasma membrane |
| StAQP29 | PGSC0003DMG402020908 | Chr09 | 196 | 20811.2 | 9.74 | 细胞质Cytoplasm |
| StAQP30 | PGSC0003DMG401020908 | Chr09 | 307 | 32802.8 | 8.11 | 质膜Plasma membrane |
| StAQP31 | PGSC0003DMG400020906 | Chr09 | 279 | 29848.6 | 8.54 | 质膜Plasma membrane |
| StAQP32 | PGSC0003DMG400034609 | Chr10 | 183 | 19587.9 | 8.91 | 液泡Vacuole |
| StAQP33 | PGSC0003DMG400026706 | Chr10 | 262 | 27681.7 | 7.92 | 液泡Vacuole |
| StAQP34 | PGSC0003DMG400026705 | Chr10 | 328 | 35105.8 | 7.47 | 质膜Plasma membrane |
| StAQP35 | PGSC0003DMG400004438 | Chr10 | 326 | 34807.5 | 7.80 | 质膜Plasma membrane |
| StAQP36 | PGSC0003DMG400026718 | Chr10 | 283 | 30409.3 | 9.23 | 质膜Plasma membrane |
| StAQP37 | PGSC0003DMG400010475 | Chr10 | 281 | 30179.7 | 8.42 | 质膜Plasma membrane |
| StAQP38 | PGSC0003DMG400028182 | Chr10 | 248 | 25341.3 | 6.34 | 细胞质Cytoplasm |
| StAQP39 | PGSC0003DMG400008358 | Chr10 | 243 | 25717.4 | 10.24 | 液泡Vacuole |
| StAQP40 | PGSC0003DMG400008078 | Chr11 | 287 | 30628.4 | 7.99 | 质膜Plasma membrane |
| StAQP41 | PGSC0003DMG400018823 | Chr12 | 239 | 25734.9 | 6.67 | 液泡Vacuole |
| StAQP42 | PGSC0003DMG400026463 | Chr12 | 248 | 24998.9 | 5.95 | 液泡Vacuole |
| StAQP43 | PGSC0003DMG400004980 | Chr12 | 288 | 30942.6 | 8.30 | 质膜Plasma membrane |
| StAQP44 | PGSC0003DMG400008556 | Chr12 | 260 | 27844.6 | 8.68 | 液泡Vacuole |
表1 马铃薯AQP基因家族成员及理化性质分析
Table 1 Analysis of potato AQP gene family members and physicochemical properties
| 基因 Gene | 基因ID Gene ID | 染色体位置Chromosome localization | 氨基酸长度Amino acid length/aa | 相对分子量Molecular weight/Da | 理论等电点Theoretical isoelectric point | 亚细胞定位Subcellular localization |
|---|---|---|---|---|---|---|
| StAQP1 | PGSC0003DMG400034503 | Chr01 | 287 | 30698.7 | 6.50 | 质膜Plasma membrane |
| StAQP2 | PGSC0003DMG400000045 | Chr01 | 285 | 30573.2 | 8.84 | 质膜Plasma membrane |
| StAQP3 | PGSC0003DMG400024879 | Chr01 | 112 | 12379.6 | 9.83 | 细胞质Cytoplasm |
| StAQP4 | PGSC0003DMG400001640 | Chr01 | 248 | 26735.5 | 8.21 | 质膜Plasma membrane |
| StAQP5 | PGSC0003DMG400006183 | Chr01 | 283 | 30119.8 | 8.60 | 质膜Plasma membrane |
| StAQP6 | PGSC0003DMG400028456 | Chr02 | 181 | 19461.6 | 6.34 | 液泡Vacuole |
| StAQP7 | PGSC0003DMG401028457 | Chr02 | 290 | 30571 | 9.10 | 质膜Plasma membrane |
| StAQP8 | PGSC0003DMG400003587 | Chr02 | 251 | 26556.8 | 8.08 | 质膜Plasma membrane |
| StAQP9 | PGSC0003DMG400030648 | Chr02 | 267 | 28189.8 | 8.45 | 细胞骨架Cytoskeleton |
| StAQP10 | PGSC0003DMG400027819 | Chr03 | 277 | 29657.2 | 9.55 | 质膜Plasma membrane |
| StAQP11 | PGSC0003DMG400020742 | Chr03 | 286 | 30688.4 | 8.83 | 质膜Plasma membrane |
| StAQP12 | PGSC0003DMG400014818 | Chr03 | 251 | 25951 | 8.67 | 叶绿体Chloroplast |
| StAQP13 | PGSC0003DMG400015275 | Chr03 | 260 | 27777.9 | 9.22 | 细胞质Cytoplasm |
| StAQP14 | PGSC0003DMG400014123 | Chr03 | 306 | 31684.7 | 8.34 | 质膜Plasma membrane |
| StAQP15 | PGSC0003DMG400002552 | Chr03 | 249 | 25161.1 | 6.33 | 细胞质Cytoplasm |
| StAQP16 | PGSC0003DMG401030499 | Chr05 | 272 | 29260.3 | 8.20 | 细胞质Cytoplasm |
| StAQP17 | PGSC0003DMG400023466 | Chr05 | 275 | 29328.2 | 9.65 | 质膜Plasma membrane |
| StAQP18 | PGSC0003DMG400024197 | Chr06 | 286 | 30662.3 | 7.18 | 质膜Plasma membrane |
| StAQP19 | PGSC0003DMG400011875 | Chr06 | 250 | 25151 | 5.48 | 液泡Vacuole |
| StAQP20 | PGSC0003DMG400016528 | Chr06 | 249 | 25089 | 6.06 | 质膜Plasma membrane |
| StAQP21 | PGSC0003DMG400026969 | Chr06 | 259 | 27476.5 | 7.32 | 细胞质Cytoplasm |
| StAQP22 | PGSC0003DMG401005949 | Chr06 | 347 | 37712.9 | 8.32 | 质膜Plasma membrane |
| StAQP23 | PGSC0003DMG400007134 | Chr06 | 250 | 25749.5 | 6.13 | 细胞质Cytoplasm |
| StAQP24 | PGSC0003DMG400030308 | Chr06 | 253 | 26056.9 | 5.48 | 液泡Vacuole |
| StAQP25 | PGSC0003DMG400005780 | Chr08 | 287 | 30790.4 | 7.96 | 质膜Plasma membrane |
| StAQP26 | PGSC0003DMG400005803 | Chr08 | 296 | 30651.2 | 8.48 | 质膜Plasma membrane |
| StAQP27 | PGSC0003DMG400009604 | Chr08 | 247 | 26042.5 | 6.22 | 液泡Vacuole |
| StAQP28 | PGSC0003DMG400012337 | Chr08 | 286 | 30686.6 | 9.34 | 质膜Plasma membrane |
| StAQP29 | PGSC0003DMG402020908 | Chr09 | 196 | 20811.2 | 9.74 | 细胞质Cytoplasm |
| StAQP30 | PGSC0003DMG401020908 | Chr09 | 307 | 32802.8 | 8.11 | 质膜Plasma membrane |
| StAQP31 | PGSC0003DMG400020906 | Chr09 | 279 | 29848.6 | 8.54 | 质膜Plasma membrane |
| StAQP32 | PGSC0003DMG400034609 | Chr10 | 183 | 19587.9 | 8.91 | 液泡Vacuole |
| StAQP33 | PGSC0003DMG400026706 | Chr10 | 262 | 27681.7 | 7.92 | 液泡Vacuole |
| StAQP34 | PGSC0003DMG400026705 | Chr10 | 328 | 35105.8 | 7.47 | 质膜Plasma membrane |
| StAQP35 | PGSC0003DMG400004438 | Chr10 | 326 | 34807.5 | 7.80 | 质膜Plasma membrane |
| StAQP36 | PGSC0003DMG400026718 | Chr10 | 283 | 30409.3 | 9.23 | 质膜Plasma membrane |
| StAQP37 | PGSC0003DMG400010475 | Chr10 | 281 | 30179.7 | 8.42 | 质膜Plasma membrane |
| StAQP38 | PGSC0003DMG400028182 | Chr10 | 248 | 25341.3 | 6.34 | 细胞质Cytoplasm |
| StAQP39 | PGSC0003DMG400008358 | Chr10 | 243 | 25717.4 | 10.24 | 液泡Vacuole |
| StAQP40 | PGSC0003DMG400008078 | Chr11 | 287 | 30628.4 | 7.99 | 质膜Plasma membrane |
| StAQP41 | PGSC0003DMG400018823 | Chr12 | 239 | 25734.9 | 6.67 | 液泡Vacuole |
| StAQP42 | PGSC0003DMG400026463 | Chr12 | 248 | 24998.9 | 5.95 | 液泡Vacuole |
| StAQP43 | PGSC0003DMG400004980 | Chr12 | 288 | 30942.6 | 8.30 | 质膜Plasma membrane |
| StAQP44 | PGSC0003DMG400008556 | Chr12 | 260 | 27844.6 | 8.68 | 液泡Vacuole |
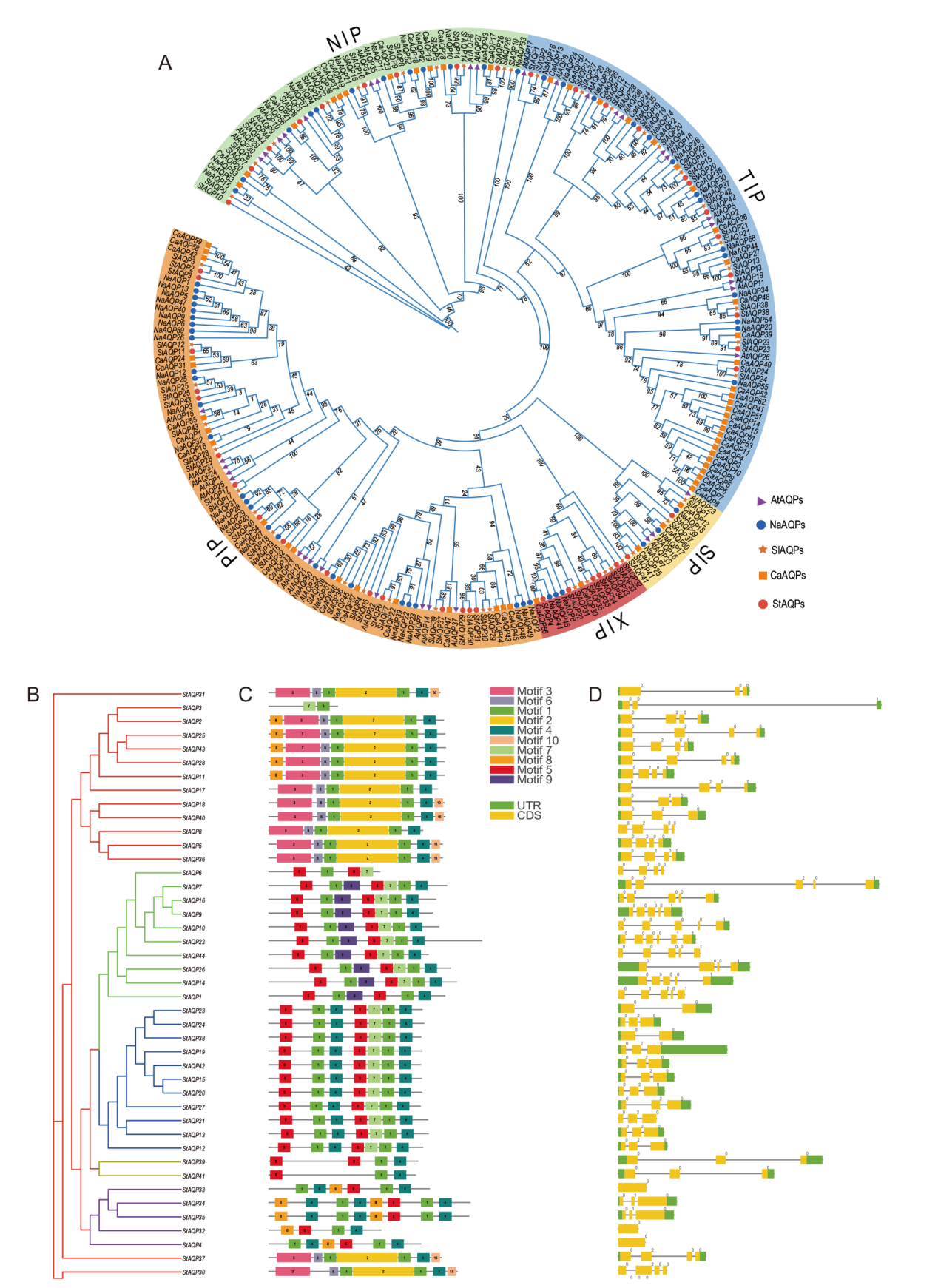
图1 AQP基因家族系统发育、基因结构与保守基序分析 A:拟南芥、烟草、马铃薯、番茄、辣椒AQP家族系统发育树;B:StAQP基因家族系统发育树;C:StAQP保守基序分析;D:StAQP基因结构分析
Fig. 1 Phylogenetic, gene structure and conserved motif analysis of the AQP gene family A: Phylogenetic trees of AQP families in the A. thaliana, N. attenuata, S. tuberosum, S. lycopersicum, and Capsicum annuum. B: Phylogenetic tree of the StAQP gene family. C: Conserved motif analysis of the StAQP gene. D: Analysis of StAQP gene structure

图3 马铃薯AQP家族基因复制事件 A:StAQP基因染色体定位图;B:StAQP基因共线性分析;C:多物种间AQP基因的共线性分析(At:拟南芥;St:马铃薯;Na:烟草;Ca:辣椒;Sl:番茄)
Fig. 3 Gene duplication events in the AQP family of S. tuberosum A: Chromosomal localization map of StAQP gene. B: Collinearity analysis of StAQP gene. C: Collinearity analysis of AQP genes in multiple species(At: A. thaliana; St: S. tuberosum; Na: N. attenuata; Ca: C. annuum; Sl: S. lycopersicum)
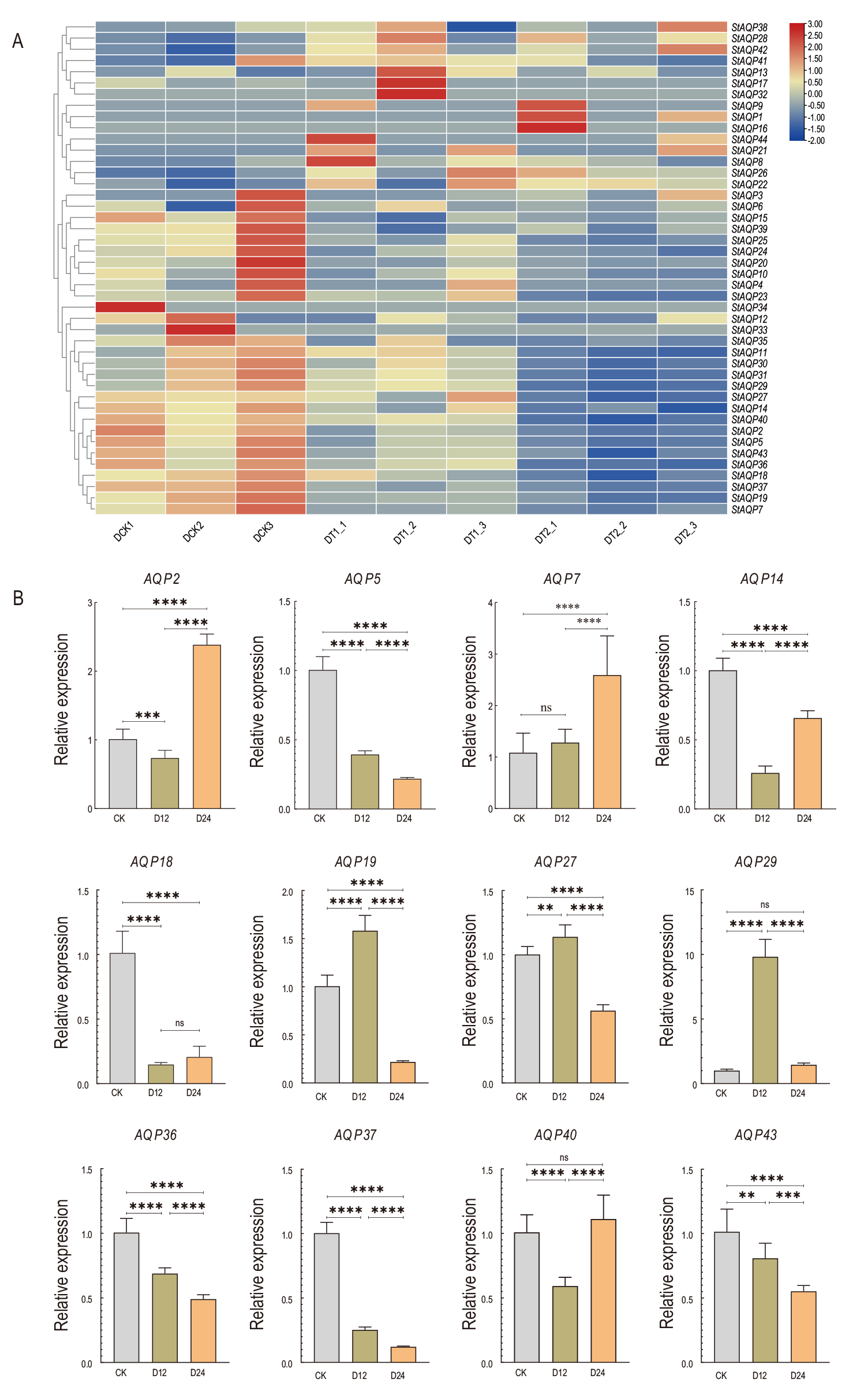
图5 StAQP基因 在干旱胁迫下的表达 A: StAQP在干旱胁迫下的表达水平热图;B: StAQP在干旱胁迫下的相对表达量,*P<0.05,**P<0.01,***P<0.001,****P<0.0001。下同
Fig. 5 Expression of StAQP gene under drought stress A: Expression of StAQP under drought stress. B: Relative expression of StAQP under drought stress, *P<0.05, **P<0.01, ***P<0.001, ****P<0.0001. The same below
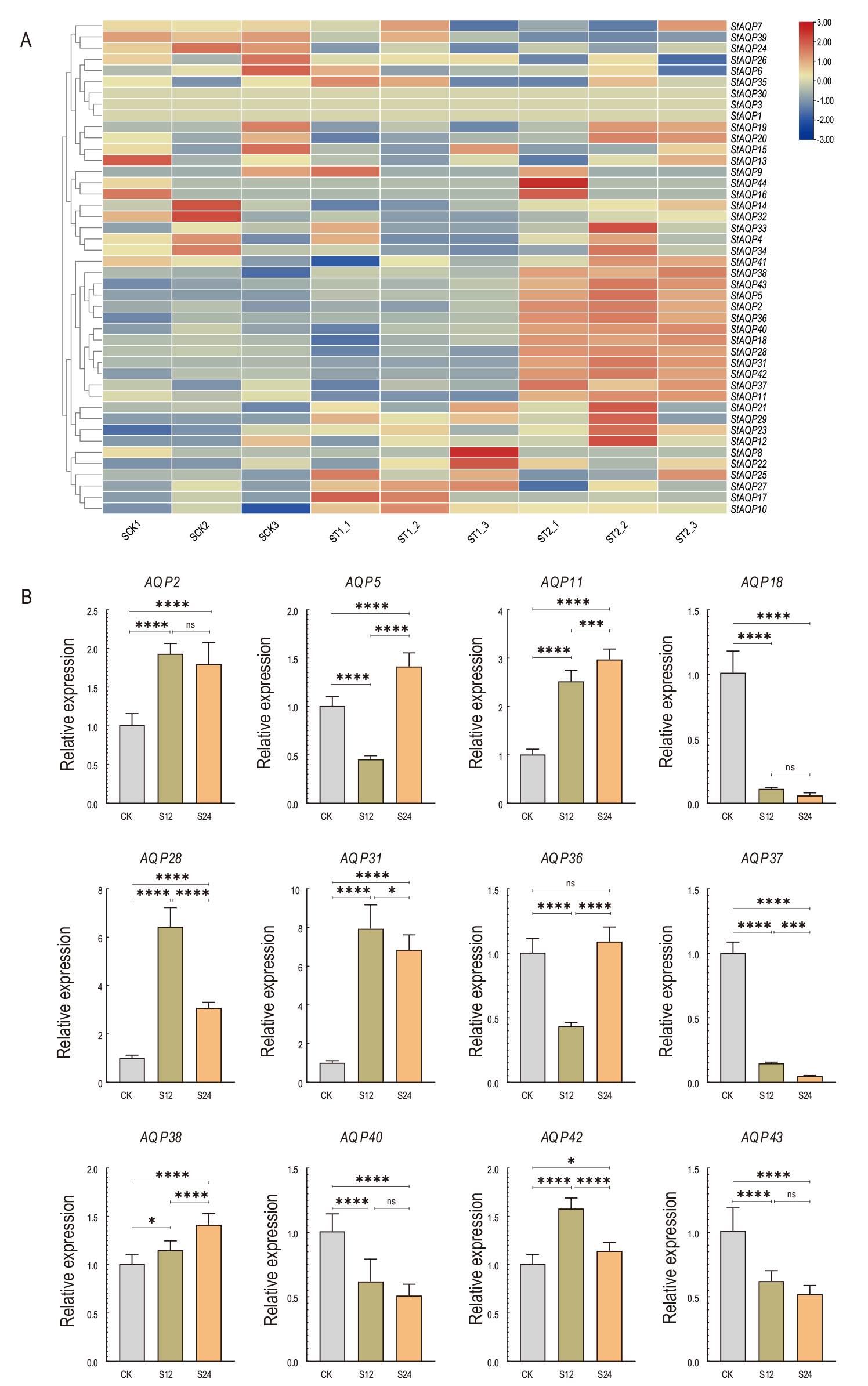
图6 StAQP基因在盐胁迫下的表达 A: StAQP在盐胁迫下的表达水平热图;B: StAQP在盐胁迫下的相对表达量
Fig. 6 Expression of StAQP gene under salt stress A: Expression of StAQP under salt stress. B: Relative expression of StAQP under salt stress
| [1] | Kopecká R, Kameniarová M, Černý M, et al. Abiotic stress in crop production[J]. Int J Mol Sci, 2023, 24(7): 6603. |
| [2] |
Chen XX, Ding YL, Yang YQ, et al. Protein kinases in plant responses to drought, salt, and cold stress[J]. J Integr Plant Biol, 2021, 63(1): 53-78.
doi: 10.1111/jipb.13061 |
| [3] | Dietz KJ, Zörb C, Geilfus CM. Drought and crop yield[J]. Plant Biol, 2021, 23(6): 881-893. |
| [4] | Riyazuddin R, Nisha NS, Singh K, et al. Involvement of dehydrin proteins in mitigating the negative effects of drought stress in plants[J]. Plant Cell Rep, 2022, 41(3): 519-533. |
| [5] | Sánchez-Bermúdez M, Del Pozo JC, Pernas M. Effects of combined abiotic stresses related to climate change on root growth in crops[J]. Front Plant Sci, 2022, 13: 918537. |
| [6] | Zhao SS, Zhang QK, Liu MY, et al. Regulation of plant responses to salt stress[J]. Int J Mol Sci, 2021, 22(9): 4609. |
| [7] | Kausar R, Komatsu S. Proteomic approaches to uncover salt stress response mechanisms in crops[J]. Int J Mol Sci, 2022, 24(1): 518. |
| [8] | Colin L, Ruhnow F, Zhu JK, et al. The cell biology of primary cell walls during salt stress[J]. Plant Cell, 2023, 35(1): 201-217. |
| [9] | Xu J, Li Y, Kaur L, et al. Functional food based on potato[J]. Foods, 2023, 12(11): 2145. |
| [10] | Hill D, Nelson D, Hammond J, et al. Morphophysiology of potato(Solanum tuberosum)in response to drought stress: paving the way forward[J]. Front Plant Sci, 2021, 11: 597554. |
| [11] | Li Q, Qin YZ, Hu XX, et al. Physiology and gene expression analysis of potato(Solanum tuberosum L.) in salt stress[J]. Plants, 2022, 11(12): 1565. |
| [12] | Wang Y, Zhao ZJ, Liu F, et al. Versatile roles of aquaporins in plant growth and development[J]. Int J Mol Sci, 2020, 21(24): 9485. |
| [13] |
Li SC, Li CL, Wang WD. Molecular aspects of aquaporins[J]. Vitam Horm, 2020, 113: 129-181.
doi: S0083-6729(19)30079-2 pmid: 32138947 |
| [14] | Vats S, Sudhakaran S, Bhardwaj A, et al. Targeting aquaporins to alleviate hazardous metal(loid)s imposed stress in plants[J]. J Hazard Mater, 2021, 408: 124910. |
| [15] | Byrt CS, Zhang RY, Magrath I, et al. Exploring aquaporin functions during changes in leaf water potential[J]. Front Plant Sci, 2023, 14: 1213454. |
| [16] |
Patel J, Mishra A. Plant aquaporins alleviate drought tolerance in plants by modulating cellular biochemistry, root-architecture, and photosynthesis[J]. Physiol Plant, 2021, 172(2): 1030-1044.
doi: 10.1111/ppl.13324 pmid: 33421148 |
| [17] |
Ahmed S, Kouser S, Asgher M, et al. Plant aquaporins: a frontward to make crop plants drought resistant[J]. Physiol Plant, 2021, 172(2): 1089-1105.
doi: 10.1111/ppl.13416 pmid: 33826759 |
| [18] | Kumari A, Bhatla SC. Regulation of salt-stressed sunflower(Helianthus annuus)seedling's water status by the coordinated action of Na+/K+ accumulation, nitric oxide, and aquaporin expression[J]. Funct Plant Biol, 2021, 48(6): 573-587. |
| [19] | Feng ZJ, Xu SC, Liu N, et al. Identification of the AQP members involved in abiotic stress responses from Arabidopsis[J]. Gene, 2018, 646: 64-73. |
| [20] | De Rosa A, Watson-Lazowski A, Evans JR, et al. Genome-wide identification and characterisation of aquaporins in Nicotiana tabacum and their relationships with other Solanaceae species[J]. BMC Plant Biol, 2020, 20(1): 266. |
| [21] | Yi XF, Sun XC, Tian R, et al. Genome-wide characterization of the aquaporin gene family in radish and functional analysis of RsPIP2-6 involved in salt stress[J]. Front Plant Sci, 2022, 13: 860742. |
| [22] |
Stajich JE. An introduction to BioPerl[J]. Methods Mol Biol, 2007, 406: 535-548.
pmid: 18287711 |
| [23] | Horton P, Park KJ, Obayashi T, et al. WoLF PSORT: protein localization predictor[J]. Nucleic Acids Res, 2007, 35(Web Server issue): W585-W587. |
| [24] | Nguyen LT, Schmidt HA, von Haeseler A, et al. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies[J]. Mol Biol Evol, 2015, 32(1): 268-274. |
| [25] | Letunic I, Bork P. Interactive Tree Of Life(iTOL)v5: an online tool for phylogenetic tree display and annotation[J]. Nucleic Acids Res, 2021, 49(W1): W293-W296. |
| [26] |
Hu B, Jin JP, Guo AY, et al. GSDS 2.0: an upgraded gene feature visualization server[J]. Bioinformatics, 2015, 31(8): 1296-1297.
doi: 10.1093/bioinformatics/btu817 pmid: 25504850 |
| [27] | Bailey TL, Boden M, Buske FA, et al. MEME SUITE: tools for motif discovery and searching[J]. Nucleic Acids Res, 2009, 37(Web Server issue): W202-W208. |
| [28] |
Lescot M, Déhais P, Thijs G, et al. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences[J]. Nucleic Acids Res, 2002, 30(1): 325-327.
doi: 10.1093/nar/30.1.325 pmid: 11752327 |
| [29] | Wang YP, Tang HB, Debarry JD, et al. MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity[J]. Nucleic Acids Res, 2012, 40(7): e49. |
| [30] |
Krzywinski M, Schein J, Birol I, et al. Circos: an information aesthetic for comparative genomics[J]. Genome Res, 2009, 19(9): 1639-1645.
doi: 10.1101/gr.092759.109 pmid: 19541911 |
| [31] | Wang DP, Zhang YB, Zhang Z, et al. KaKs_Calculator 2.0: a toolkit incorporating gamma-series methods and sliding window strategies[J]. Genomics Proteomics Bioinformatics, 2010, 8(1): 77-80. |
| [32] | Szklarczyk D, Kirsch R, Koutrouli M, et al. The STRING database in 2023: protein-protein association networks and functional enrichment analyses for any sequenced genome of interest[J]. Nucleic Acids Res, 2023, 51(D1): D638-D646. |
| [33] | Franz M, Lopes CT, Fong D, et al. Cytoscape.js 2023 update: a graph theory library for visualization and analysis[J]. Bioinformatics, 2023, 39(1): btad031. |
| [34] | Chen CJ, Wu Y, Li JW, et al. TBtools-II: a “one for all, all for one” bioinformatics platform for biological big-data mining[J]. Mol Plant, 2023, 16(11): 1733-1742. |
| [35] |
Tang X, Zhang N, Si HJ, et al. Selection and validation of reference genes for RT-qPCR analysis in potato under abiotic stress[J]. Plant Methods, 2017, 13: 85.
doi: 10.1186/s13007-017-0238-7 pmid: 29075311 |
| [36] |
Maren NA, Duduit JR, Huang DB, et al. Stepwise optimization of real-time RT-PCR analysis[J]. Methods Mol Biol, 2023, 2653: 317-332.
doi: 10.1007/978-1-0716-3131-7_20 pmid: 36995635 |
| [37] | Ishibashi K, Kondo S, Hara S, et al. The evolutionary aspects of aquaporin family[J]. Am J Physiol Regul Integr Comp Physiol, 2011, 300(3): R566-R576. |
| [38] |
Maurel C, Boursiac Y, Luu DT, et al. Aquaporins in plants[J]. Physiol Rev, 2015, 95(4): 1321-1358.
doi: 10.1152/physrev.00008.2015 pmid: 26336033 |
| [39] | Yaghobi M, Heidari P. Genome-wide analysis of aquaporin gene family in Triticum turgidum and its expression profile in response to salt stress[J]. Genes, 2023, 14(1): 202. |
| [40] | He W, Liu MY, Qin XY, et al. Genome-wide identification and expression analysis of the aquaporin gene family in Lycium barbarum during fruit ripening and seedling response to heat stress[J]. Curr Issues Mol Biol, 2022, 44(12): 5933-5948. |
| [41] |
Wang R, Li RZ, Cheng LN, et al. SlERF52 regulates SlTIP1;1 expression to accelerate tomato pedicel abscission[J]. Plant Physiol, 2021, 185(4): 1829-1846.
doi: 10.1093/plphys/kiab026 pmid: 33638643 |
| [1] | 申鹏, 高雅彬, 丁红. 马铃薯SAT基因家族的鉴定和表达分析[J]. 生物技术通报, 2024, 40(9): 64-73. |
| [2] | 王超, 白如仟, 管俊梅, 罗稷林, 何雪姣, 迟绍轶, 马玲. 马铃薯块茎变绿中StHY5对龙葵素合成的促进作用[J]. 生物技术通报, 2024, 40(9): 113-122. |
| [3] | 夏士轩, 耿泽栋, 祝光涛, 张春芝, 李大伟. 基于深度学习的马铃薯花粉活力快速检测[J]. 生物技术通报, 2024, 40(9): 123-130. |
| [4] | 毛向红, 卢瑶, 范向斌, 杜培兵, 白小东. 基于SSR荧光标记毛细管电泳的马铃薯品种遗传多样性分析及分子身份证构建[J]. 生物技术通报, 2024, 40(9): 131-140. |
| [5] | 袁兰, 黄娅楠, 张贝妮, 熊雨萌, 王洪洋. 基于流式细胞仪鉴定马铃薯倍性的高通量样品制备方法[J]. 生物技术通报, 2024, 40(9): 141-147. |
| [6] | 吴慧琴, 王延宏, 刘涵, 司政, 刘雪晴, 王静, 阳宜, 成妍. 辣椒UGT基因家族的鉴定及表达分析[J]. 生物技术通报, 2024, 40(9): 198-211. |
| [7] | 谭博文, 张懿, 张鹏, 王振宇, 马秋香. 木薯镁离子转运蛋白家族基因的鉴定及生物信息学分析[J]. 生物技术通报, 2024, 40(9): 20-32. |
| [8] | 宋倩娜, 段永红, 冯瑞云. CRISPR/Cas9介导的高效四倍体马铃薯试管薯基因编辑体系的建立[J]. 生物技术通报, 2024, 40(9): 33-41. |
| [9] | 王柯然, 闫俊杰, 刘建凤, 高玉林. RNAi技术在马铃薯害虫防控中的应用和风险[J]. 生物技术通报, 2024, 40(9): 4-10. |
| [10] | 张小妹, 周南伶, 张赛行, 王超, 沈玉龙, 管俊梅, 马玲. 马铃薯StDREBs基因的克隆及其表达分析[J]. 生物技术通报, 2024, 40(9): 42-50. |
| [11] | 吴娟, 武小娟, 王沛捷, 谢锐, 聂虎帅, 李楠, 马艳红. 彩色马铃薯花青素合成相关ERF基因筛选及表达分析[J]. 生物技术通报, 2024, 40(9): 82-91. |
| [12] | 乔岩, 杨芳, 任盼荣, 祁伟亮, 安沛沛, 李茜, 李丹, 肖俊飞. 马铃薯野生种烯酰水合酶超家族基因ScDHNS的克隆与功能分析[J]. 生物技术通报, 2024, 40(9): 92-103. |
| [13] | 宋兵芳, 柳宁, 程新艳, 徐晓斌, 田文茂, 高悦, 毕阳, 王毅. 马铃薯G6PDH基因家族鉴定及其在损伤块茎的表达分析[J]. 生物技术通报, 2024, 40(9): 104-112. |
| [14] | 周冉, 王兴平, 李彦霞, 罗仍卓么. 金黄色葡萄球菌型乳房炎奶牛乳腺组织的lncRNA差异表达分析[J]. 生物技术通报, 2024, 40(8): 320-328. |
| [15] | 武帅, 辛燕妮, 买春海, 穆晓娅, 王敏, 岳爱琴, 赵晋忠, 吴慎杰, 杜维俊, 王利祥. 大豆GS基因家族全基因组鉴定及胁迫响应分析[J]. 生物技术通报, 2024, 40(8): 63-73. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||