








生物技术通报 ›› 2024, Vol. 40 ›› Issue (10): 9-18.doi: 10.13560/j.cnki.biotech.bull.1985.2024-0669
贺文闯1( ), 许强1, 钱前1,2,3(
), 许强1, 钱前1,2,3( ), 商连光1,2(
), 商连光1,2( )
)
收稿日期:2024-07-14
出版日期:2024-10-26
发布日期:2024-11-20
通讯作者:
商连光,男,博士,研究员,研究方向:水稻种质资源挖掘与利用;E-mail: shanglianguang@caas.cn;作者简介:贺文闯,男,博士,副研究员,研究方向:水稻基因组学与优异基因挖掘;E-mail: hewenchuang@caas.cn
基金资助:
HE Wen-chuang1( ), XU Qiang1, QIAN Qian1,2,3(
), XU Qiang1, QIAN Qian1,2,3( ), SHANG Lian-guang1,2(
), SHANG Lian-guang1,2( )
)
Received:2024-07-14
Published:2024-10-26
Online:2024-11-20
摘要:
与单个基因组不同,泛基因组一般是指包含一个物种或群体中全部基因组信息的数据集。近十年来,泛基因组学在水稻中已逐步成为研究热点,相关泛基因组成果和工具已在群体遗传学、进化生物学和生物育种实践等多个下游研究领域中获得广泛应用。本文聚焦于水稻泛基因组学的发展历程与应用前景,回顾了水稻泛基因组学的内涵发展和研究成果的时间线,总结了现有的水稻泛基因组代表性重要成果工具及其在多个领域中的主要应用,展望了其面临的挑战和发展前景。
贺文闯, 许强, 钱前, 商连光. 水稻泛基因组学的发展与前景:重要工具与应用[J]. 生物技术通报, 2024, 40(10): 9-18.
HE Wen-chuang, XU Qiang, QIAN Qian, SHANG Lian-guang. Development and Prospects of Rice Pan-genomics: Important Tools and Applications[J]. Biotechnology Bulletin, 2024, 40(10): 9-18.
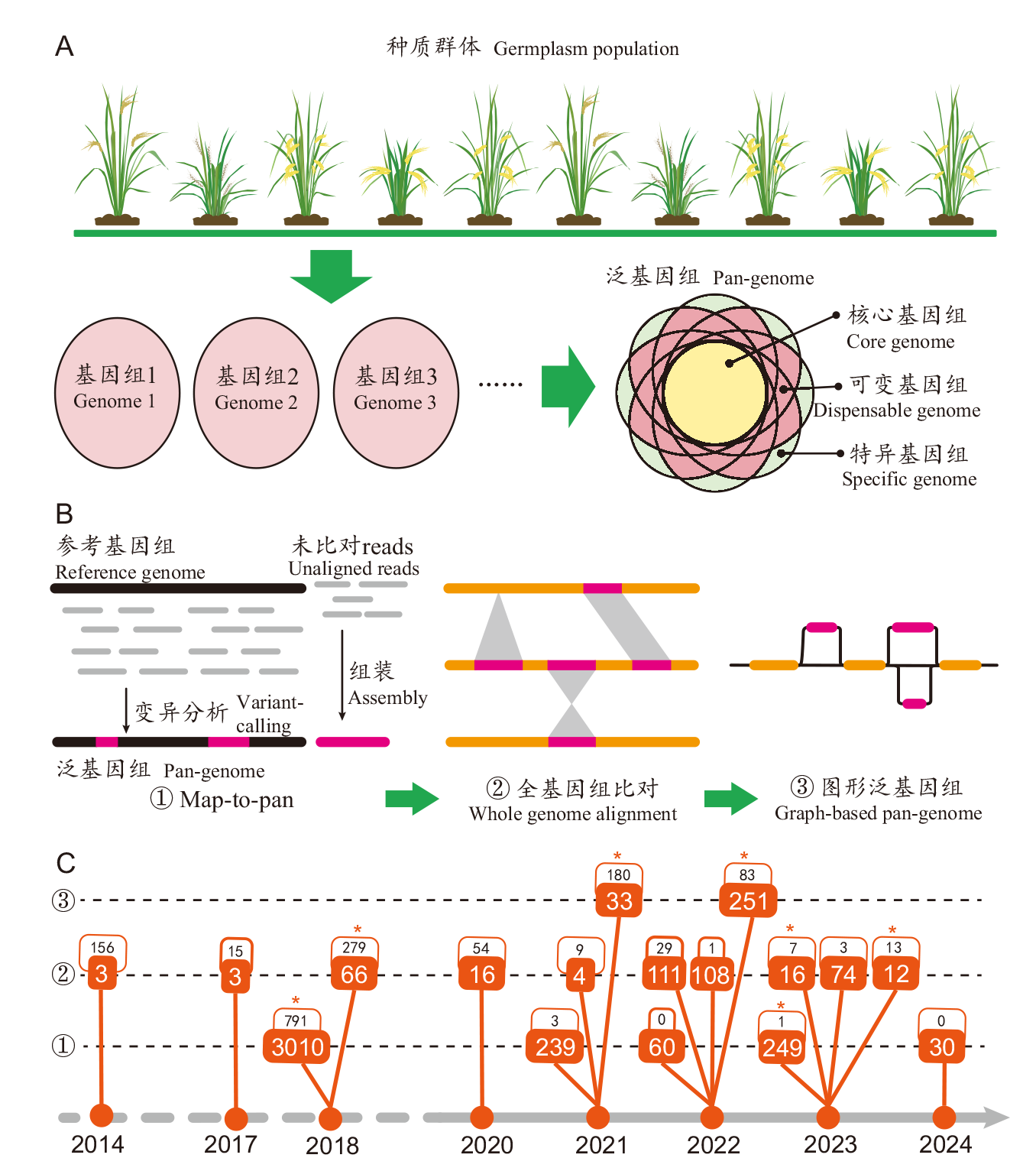
图1 水稻泛基因组学的发展历程 A:泛基因组定义的概念图示;B:泛基因组构建方法和组成形式的发展;C:水稻泛基因组学研究的发展时间线;时间线上的每个分支代表一个泛基因组数据集,两个数字展示了其对应的群体大小(下)和被非综述性论文引用的次数(上);*代表该泛基因组研究产出了可公开访问的数据库平台或友好分析工具;被引次数来源于Web of Science数据库,统计截至2024年6月17日
Fig. 1 Development of rice pan-genomics A: Illustration of the pan-genome definition. B: Development of pan-genome construction methods and constituent forms. C: Development timeline of rice pan-genomics studies. Each branch of the timeline iindicates a pan-genomic dataset, with two numbers showing its corresponding population size(bottom)and the number of citations to non-review papers(top). * indicates the production of a publicly accessible database or friendly analytical tool for the pan-genomic study. Number of citations are from the Web of Science database as of June 17, 2024
| 构建方法 Construction method | 发表时间 Published date | 群体大小 Population size | 群体组成 Species | 测序数据Sequencing data | 单基因组资源Genome resource | 泛基因组资源Pan-genome resource | 泛基因资源Pan-gene resource | 变异资源 Variant resource | 数据库 Database | 潜在应用 Potential application | 参考文献 Reference | |||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 单基因组参考 A single genomic reference | 泛基因组参考Pan-genomic reference | GWAS等研究GWAS etc. | 单倍型分析 Haplotype analysis | |||||||||||
| Map-to-pan | 2018 | 3 010 | Os | Illumina | × | √ | √ | SNP/InDel/ SV | √ | × | √ | √ | √ | [ |
| 2021 | 239 | Osj | Illumina | × | √ | √ | SNPs/PAV | × | × | √ | √ | √ | [ | |
| 2023 | 60 | Os | Illumina | × | √ | √ | SNP/PAV | × | × | √ | × | √ | [ | |
| 2024 | 249 | Ogla/Ob | Illumina | × | √ | √ | 无 | × | × | √ | × | × | [ | |
| 2024 | 30 | Oa/ Ogra/Ol | Illumina | × | √ | √ | SNP/PAV | × | × | √ | × | √ | [ | |
| 全基因组比对Whole genome alignment | 2014 | 3 | Os | Illumina | √ | × | 冗余 | 无 | × | √ | × | × | √ | [ |
| 2017 | 3 | Ogla | Illumina | √ | × | 冗余 | 无 | × | √ | × | × | √ | [ | |
| 2018 | 66 | Os/Or | Illumina | √ | × | √ | SNP/InDel | √ | √ | × | × | √ | [ | |
| 2020 | 16 | Os | Illumina/Pacbio/Bionano | √ | × | × | InDel/SV | × | √ | × | × | √ | [ | |
| 2021 | 4 | Os | Illumina/Pacbio | √ | × | √ | SNP/InDel/SV | × | √ | × | × | √ | [ | |
| 2022 | 111 | Or/Os | Illumina/Nanopore | √ | √ | √ | SV | × | √ | √ | × | √ | [ | |
| 2022 | 108 | Os | Illumina | √ | √ | √ | InDel/PAV | × | √ | √ | × | √ | [ | |
| 2023 | 74 | Or/Os/杂草稻 | PacBio(12)/Hi-C(4) | √ | × | √ | SNP/PAV | × | √ | × | × | √ | [ | |
| 2023 | 16 | Os | Iso-Seq/RNA-Seq | [16]* | × | √ | 可变剪切事件 | √ | √ | × | × | √ | [ | |
| 2023 | 12 | Os/Ogla | [21]* | [21]* | √ | √ | PAV | × | √ | √ | × | √ | [ | |
| 图形泛基因组Graph-based pan-genome | 2021 | 33 | Os/Ogla | Illumina/Pacbio/Bionano(3) | √ | √ | √ | SV/gCNV | √ | √ | √ | × | √ | [ |
| 2022 | 251 | Os/Or/ Ogla/Ob | Illumina/Nanopore/Hi-C(4) | √ | √ | √ | SV | √ | √ | √ | √ | √ | [ | |
表1 已发表的水稻泛基因组研究汇总
Table 1 Summary of published pan-genome studies in rice
| 构建方法 Construction method | 发表时间 Published date | 群体大小 Population size | 群体组成 Species | 测序数据Sequencing data | 单基因组资源Genome resource | 泛基因组资源Pan-genome resource | 泛基因资源Pan-gene resource | 变异资源 Variant resource | 数据库 Database | 潜在应用 Potential application | 参考文献 Reference | |||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 单基因组参考 A single genomic reference | 泛基因组参考Pan-genomic reference | GWAS等研究GWAS etc. | 单倍型分析 Haplotype analysis | |||||||||||
| Map-to-pan | 2018 | 3 010 | Os | Illumina | × | √ | √ | SNP/InDel/ SV | √ | × | √ | √ | √ | [ |
| 2021 | 239 | Osj | Illumina | × | √ | √ | SNPs/PAV | × | × | √ | √ | √ | [ | |
| 2023 | 60 | Os | Illumina | × | √ | √ | SNP/PAV | × | × | √ | × | √ | [ | |
| 2024 | 249 | Ogla/Ob | Illumina | × | √ | √ | 无 | × | × | √ | × | × | [ | |
| 2024 | 30 | Oa/ Ogra/Ol | Illumina | × | √ | √ | SNP/PAV | × | × | √ | × | √ | [ | |
| 全基因组比对Whole genome alignment | 2014 | 3 | Os | Illumina | √ | × | 冗余 | 无 | × | √ | × | × | √ | [ |
| 2017 | 3 | Ogla | Illumina | √ | × | 冗余 | 无 | × | √ | × | × | √ | [ | |
| 2018 | 66 | Os/Or | Illumina | √ | × | √ | SNP/InDel | √ | √ | × | × | √ | [ | |
| 2020 | 16 | Os | Illumina/Pacbio/Bionano | √ | × | × | InDel/SV | × | √ | × | × | √ | [ | |
| 2021 | 4 | Os | Illumina/Pacbio | √ | × | √ | SNP/InDel/SV | × | √ | × | × | √ | [ | |
| 2022 | 111 | Or/Os | Illumina/Nanopore | √ | √ | √ | SV | × | √ | √ | × | √ | [ | |
| 2022 | 108 | Os | Illumina | √ | √ | √ | InDel/PAV | × | √ | √ | × | √ | [ | |
| 2023 | 74 | Or/Os/杂草稻 | PacBio(12)/Hi-C(4) | √ | × | √ | SNP/PAV | × | √ | × | × | √ | [ | |
| 2023 | 16 | Os | Iso-Seq/RNA-Seq | [16]* | × | √ | 可变剪切事件 | √ | √ | × | × | √ | [ | |
| 2023 | 12 | Os/Ogla | [21]* | [21]* | √ | √ | PAV | × | √ | √ | × | √ | [ | |
| 图形泛基因组Graph-based pan-genome | 2021 | 33 | Os/Ogla | Illumina/Pacbio/Bionano(3) | √ | √ | √ | SV/gCNV | √ | √ | √ | × | √ | [ |
| 2022 | 251 | Os/Or/ Ogla/Ob | Illumina/Nanopore/Hi-C(4) | √ | √ | √ | SV | √ | √ | √ | √ | √ | [ | |

图2 水稻泛基因组研究的非综述文献引用的主要领域分布情况 * WOS数据库中MeSH Headings关键词的统计结果(参与统计的文献总数n=859,单个词条的引用量≥50)。此处仅展示水稻重要领域相关词条结果,描述相似或者领域相近的关键词仅展示其一
Fig. 2 Distribution of major fields of non-review citations for rice pan-genome studies * statistical results of MeSH Headings Keywords in the WOS database(the total number of literatures participating in the statistics n=859, the number of citations for a single entry ≥50). Only the results of terms related to important fields of rice are shown here, and only one keyword describing similar or similar fields is shown

图3 水稻泛基因组学工具的主要应用 进化和驯化图中元素根据Chen等[25]进行了修改;生物育种和智慧育种图中元素根据Ferrero-Serrano等[26]进行了修改
Fig. 3 Main applications of rice pan-genomics tools The elements in the evolution and domestication diagram are modified according to Chen et al[25]. The elements in the diagram for biological breeding and intelligent breeding are modified according to Ferrero-Serrano et al[26]
| [1] | Tettelin H, Masignani V, Cieslewicz MJ, et al. Genome analysis of multiple pathogenic isolates of Streptococcus agalactiae: implications for the microbial “pan-genome”[J]. Proc Natl Acad Sci U S A, 2005, 102(39): 13950-13955. |
| [2] |
Li RQ, Li YR, Zheng HC, et al. Building the sequence map of the human pan-genome[J]. Nat Biotechnol, 2010, 28(1): 57-63.
doi: 10.1038/nbt.1596 pmid: 19997067 |
| [3] |
Li YH, Zhou GY, Ma JX, et al. De novo assembly of soybean wild relatives for pan-genome analysis of diversity and agronomic traits[J]. Nat Biotechnol, 2014, 32(10): 1045-1052.
doi: 10.1038/nbt.2979 |
| [4] |
Bayer PE, Golicz AA, Scheben A, et al. Plant pan-genomes are the new reference[J]. Nat Plants, 2020, 6(8): 914-920.
doi: 10.1038/s41477-020-0733-0 pmid: 32690893 |
| [5] | Shi JP, Tian ZX, Lai JS, et al. Plant pan-genomics and its applications[J]. Mol Plant, 2023, 16(1): 168-186. |
| [6] |
Zhang F, Xue HZ, Dong XR, et al. Long-read sequencing of 111 rice genomes reveals significantly larger pan-genomes[J]. Genome Res, 2022, 32(5): 853-863.
doi: 10.1101/gr.276015.121 pmid: 35396275 |
| [7] |
Wang J, Yang W, Zhang SH, et al. A pangenome analysis pipeline provides insights into functional gene identification in rice[J]. Genome Biol, 2023, 24(1): 19.
doi: 10.1186/s13059-023-02861-9 pmid: 36703158 |
| [8] | Schatz MC, Maron LG, Stein JC, et al. Whole genome de novo assemblies of three divergent strains of rice, Oryza sativa, document novel gene space of aus and indica[J]. Genome Biol, 2014, 15(11): 506. |
| [9] | Monat C, Pera B, Ndjiondjop MN, et al. De novo assemblies of three Oryza glaberrima accessions provide first insights about pan-genome of African rices[J]. Genome Biol Evol, 2017, 9(1): 1-6. |
| [10] | Wang WS, Mauleon R, Hu ZQ, et al. Genomic variation in 3, 010 diverse accessions of Asian cultivated rice[J]. Nature, 2018, 557(7703): 43-49. |
| [11] |
Liu CX, Peng P, Li WG, et al. Deciphering variation of 239 elite Japonica rice genomes for whole genome sequences-enabled breeding[J]. Genomics, 2021, 113(5): 3083-3091.
doi: 10.1016/j.ygeno.2021.07.002 pmid: 34237377 |
| [12] | Woldegiorgis ST, Wu T, Gao LH, et al. Identification of heat-tolerant genes in non-reference sequences in rice by integrating pan-genome, transcriptomics, and QTLs[J]. Genes, 2022, 13(8): 1353. |
| [13] | Christine TD, Clothilde C, Mathieu B, et al. FrangiPANe, a tool for creating a panreference using left behind reads[J]. NAR Genom Bioinform, 2023, 5(1): lqad013. |
| [14] | Shivute FN, Zhong Y, Wu JW, et al. Genome-wide and pan-genomic analysis reveals rich variants of NBS-LRR genes in a newly developed wild rice line from Oryza alta Swallen[J]. Front Plant Sci, 2024, 15: 1345708. |
| [15] |
Zhao Q, Feng Q, Lu HY, et al. Pan-genome analysis highlights the extent of genomic variation in cultivated and wild rice[J]. Nat Genet, 2018, 50(2): 278-284.
doi: 10.1038/s41588-018-0041-z pmid: 29335547 |
| [16] |
Zhou Y, Chebotarov D, Kudrna D, et al. A platinum standard pan-genome resource that represents the population structure of Asian rice[J]. Sci Data, 2020, 7(1): 113.
doi: 10.1038/s41597-020-0438-2 pmid: 32265447 |
| [17] |
Panibe JP, Wang L, Li J, et al. Chromosomal-level genome assembly of the semi-dwarf rice Taichung Native 1, an initiator of green revolution[J]. Genomics, 2021, 113(4): 2656-2674.
doi: 10.1016/j.ygeno.2021.06.006 pmid: 34111524 |
| [18] | Singh PK, Rawal HC, Panda AK, et al. Pan-genomic, transcriptomic, and miRNA analyses to decipher genetic diversity and anthocyanin pathway genes among the traditional rice landraces[J]. Genomics, 2022, 114(5): 110436. |
| [19] |
Wu DY, Xie LJ, Sun YQ, et al. A syntelog-based pan-genome provides insights into rice domestication and de-domestication[J]. Genome Biol, 2023, 24(1): 179.
doi: 10.1186/s13059-023-03017-5 pmid: 37537691 |
| [20] | Yu ZC, Chen YM, Zhou Y, et al. Rice Gene Index: a comprehensive pan-genome database for comparative and functional genomics of Asian rice[J]. Mol Plant, 2023, 16(5): 798-801. |
| [21] |
Qin P, Lu HW, Du HL, et al. Pan-genome analysis of 33 genetically diverse rice accessions reveals hidden genomic variations[J]. Cell, 2021, 184(13): 3542-3558.e16.
doi: 10.1016/j.cell.2021.04.046 pmid: 34051138 |
| [22] |
Shang LG, Li XX, He HY, et al. A super pan-genomic landscape of rice[J]. Cell Res, 2022, 32(10): 878-896.
doi: 10.1038/s41422-022-00685-z pmid: 35821092 |
| [23] | Kawahara Y, de la Bastide M, Hamilton JP, et al. Improvement of the Oryza sativa Nipponbare reference genome using next generation sequence and optical map data[J]. Rice(N Y), 2013, 6(1): 4. |
| [24] | Sun C, Hu ZQ, Zheng TQ, et al. RPAN: rice pan-genome browser for -3000 rice genomes[J]. Nucleic Acids Res, 2017, 45(2): 597-605. |
| [25] | Chen WK, Chen L, Zhang X, et al. Convergent selection of a WD40 protein that enhances grain yield in maize and rice[J]. Science, 2022, 375(6587): eabg7985. |
| [26] | Ferrero-Serrano Á, Chakravorty D, Kirven KJ, et al. Oryza CLIMtools: a genome-environment association resource reveals adaptive roles for heterotrimeric G proteins in the regulation of rice agronomic traits[J]. Plant Commun, 2024, 5(4): 100813. |
| [27] | Wei H, Wang XM, Zhang ZP, et al. Uncovering key salt-tolerant regulators through a combined eQTL and GWAS analysis using the super pan-genome in rice[J]. Natl Sci Rev, 2024, 11(4): nwae043. |
| [28] |
Lv Y, Liu CC, Li XX, et al. A centromere map based on super pan-genome highlights the structure and function of rice centromeres[J]. J Integr Plant Biol, 2024, 66(2): 196-207.
doi: 10.1111/jipb.13607 |
| [29] | Cui YC, Lin YR, Wei H, et al. Identification of salt tolerance-associated presence-absence variations in the OsMADS56 gene through the integration of DEGs dataset and eQTL analysis[J]. New Phytol, 2024, 243(3): 833-838. |
| [30] |
Zhou Y, Yu ZC, Chebotarov D, et al. Pan-genome inversion index reveals evolutionary insights into the subpopulation structure of Asian rice[J]. Nat Commun, 2023, 14(1): 1567.
doi: 10.1038/s41467-023-37004-y pmid: 36944612 |
| [31] |
Wang TY, He WC, Li XX, et al. A rice variation map derived from 10 548 rice accessions reveals the importance of rare variants[J]. Nucleic Acids Res, 2023, 51(20): 10924-10933.
doi: 10.1093/nar/gkad840 pmid: 37843097 |
| [32] | Zhang H, Chen W, Zhu D, et al. Population-level exploration of alternative splicing and its unique role in controlling agronomic traits of rice[J]. Plant Cell, 2024: koae181. |
| [33] | Li XX, Dai XF, He HY, et al. A pan-TE map highlights transposable elements underlying domestication and agronomic traits in Asian rice[J]. Natl Sci Rev, 2024, 11(6): nwae188. |
| [34] |
He HY, Leng Y, Cao XL, et al. The pan-tandem repeat map highlights multiallelic variants underlying gene expression and agronomic traits in rice[J]. Nat Commun, 2024, 15: 7291.
doi: 10.1038/s41467-024-51854-0 pmid: 39181885 |
| [35] | Wang J, Hu HF, Jiang XY, et al. Pangenome-wide association study and transcriptome analysis reveal a novel QTL and candidate genes controlling both panicle and leaf blast resistance in rice[J]. Rice(N Y), 2024, 17(1): 27. |
| [36] | Singh G, Singh N, Ellur RK, et al. Genetic enhancement for biotic stress resistance in basmati rice through marker-assisted backcross breeding[J]. Int J Mol Sci, 2023, 24(22): 16081. |
| [37] | Sun Y, Zhang PT, Kou DR, et al. Terpene synthases in rice pan-genome and their responses to Chilo suppressalis larvae infesting[J]. Front Plant Sci, 2022, 13: 905982. |
| [38] |
Lin YR, Zhu YW, Cui YC, et al. Identification of natural allelic variation in TTL1 controlling thermotolerance and grain size by a rice super pan-genome[J]. J Integr Plant Biol, 2023, 65(12): 2541-2551.
doi: 10.1111/jipb.13568 |
| [39] |
Li WH, Wang DX, Hong XK, et al. Identification and validation of new MADS-box homologous genes in 3010 rice pan-genome[J]. Plant Cell Rep, 2023, 42(6): 975-988.
doi: 10.1007/s00299-023-03006-9 pmid: 37016094 |
| [40] | Daware A, Malik A, Srivastava R, et al. Rice Pangenome Genotyping Array: an efficient genotyping solution for pangenome-based accelerated genetic improvement in rice[J]. Plant J, 2023, 113(1): 26-46. |
| [41] | He WC, He HY, Yuan QL, et al. Widespread inversions shape the genetic and phenotypic diversity in rice[J]. Sci Bull, 2024, 69(5): 593-596. |
| [42] | Ramsbottom KA, Prakash A, Perez-Riverol Y, et al. Meta-analysis of rice phosphoproteomics data to understand variation in cell signaling across the rice pan-genome[J]. J Proteome Res, 2024, 23(7): 2518-2531. |
| [43] | Brotman Y, Llorente-Wiegand C, Oyong G, et al. The genetics underlying metabolic signatures in a brown rice diversity panel and their vital role in human nutrition[J]. Plant J, 2021, 106(2): 507-525. |
| [44] |
Wei X, Qiu J, Yong KC, et al. A quantitative genomics map of rice provides genetic insights and guides breeding[J]. Nat Genet, 2021, 53(2): 243-253.
doi: 10.1038/s41588-020-00769-9 pmid: 33526925 |
| [45] | Wang S, Qian YQ, Zhao RP, et al. Graph-based pan-genomes: increased opportunities in plant genomics[J]. J Exp Bot, 2023, 74(1): 24-39. |
| [46] | Chen RZ, Deng YW, Ding YL, et al. Rice functional genomics: decades’ efforts and roads ahead[J]. Sci China Life Sci, 2022, 65(1): 33-92. |
| [47] | Li HH, Li X, Zhang P, et al. Smart Breeding Platform: a web-based tool for high-throughput population genetics, phenomics, and genomic selection[J]. Mol Plant, 2024, 17(5): 677-681. |
| [1] | 冷燕, 马晓薇, 陈光, 任鹤, 李翔. 玉米高产竞赛助力中国玉米种业振兴[J]. 生物技术通报, 2023, 39(8): 4-10. |
| [2] | 解伟, 刘春明. 生物育种产业化面临的机遇与政策保障[J]. 生物技术通报, 2023, 39(1): 16-20. |
| [3] | 张雅涵, 朱丽霞, 胡静, 朱亚静, 张雪婧, 曹叶中. 草甘膦在我国生物育种产业化应用中的机遇与挑战[J]. 生物技术通报, 2022, 38(11): 1-9. |
| [4] | 张慧, 田方方, 吴毅. 合成型酵母基因组重排技术[J]. 生物技术通报, 2020, 36(4): 13-18. |
| [5] | 常丽, 邓欣, 唐慧娟, 李建军, 黄思齐, 陈安国, 张翠萍, 赵立宁, 李德芳. 基于专利分析的大麻研发态势及技术构成[J]. 生物技术通报, 2018, 34(12): 195-201. |
| [6] | 杨晶, 戴求仲, 侯振平, 王延周, 王满生. 基于专利分析的麻类作物饲料化利用技术态势及热点分析[J]. 生物技术通报, 2018, 34(12): 207-214. |
| [7] | 丁泽红, 付莉莉, 铁韦韦, 颜彦, 胡伟. 木薯MeTPS9基因克隆及表达特性分析[J]. 生物技术通报, 2017, 33(11): 84-91. |
| [8] | 宋娜娜, 柴志欣, 钟金城. 长链非编码RAN的研究进展[J]. 生物技术通报, 2016, 32(9): 23-31. |
| [9] | 丁泽红, 付莉莉, 铁韦韦, 颜彦, 胡伟. 木薯MeNCED3基因克隆、结构变异及其表达分析[J]. 生物技术通报, 2016, 32(10): 148-153. |
| [10] | 龚静, 柳纯洁, 缪小平, 郭安源. 人类长链非编码RNA相关SNP鉴定与功能预测的研究进展[J]. 生物技术通报, 2015, 31(11): 27-34. |
| [11] | 靳进朴, 郭安源, 何坤, 张禾, 朱其慧, 陈新, 高歌, 罗静初. 植物转录因子分类、预测和数据库构建[J]. 生物技术通报, 2015, 31(11): 68-77. |
| [12] | 王慧丽, 郭安源. 环境微生物宏基因组学数据库利用[J]. 生物技术通报, 2015, 31(11): 78-88. |
| [13] | 杨健, 蔡浩洋. 肿瘤生物信息学数据库[J]. 生物技术通报, 2015, 31(11): 89-101. |
| [14] | 罗静初. 实用生物信息技术课程教学实例[J]. 生物技术通报, 2015, 31(11): 102-111. |
| [15] | 郝心宁, 孙巍, 张学福. 生物育种领域知识结构演化分析[J]. 生物技术通报, 2013, 29(11): 186-192. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||