








• 研究报告 • 下一篇
熊仕发1( ), 陈益存1,2, 吴立文1,2, 施翔1, 张盛剿3, 彭方有3, 陈涛梅3, 汪阳东1,2(
), 陈益存1,2, 吴立文1,2, 施翔1, 张盛剿3, 彭方有3, 陈涛梅3, 汪阳东1,2( )
)
收稿日期:2025-07-29
出版日期:2026-03-09
通讯作者:
汪阳东,男,博士,研究员,研究方向 :化工原料树种遗传育种;E-mail: wangyangdong@caf.ac.cn作者简介:熊仕发,男,博士,助理研究员,研究方向 :栎树遗传育种;E-mail: xiongshifa111@163.com
基金资助:
XIONG Shi-fa1( ), CHEN Yi-cun1,2, WU Li-wen1,2, SHI Xiang1, ZHANG Sheng-jiao3, PENG Fang-you3, CHEN Tao-mei3, WANG Yang-dong1,2(
), CHEN Yi-cun1,2, WU Li-wen1,2, SHI Xiang1, ZHANG Sheng-jiao3, PENG Fang-you3, CHEN Tao-mei3, WANG Yang-dong1,2( )
)
Received:2025-07-29
Published:2026-03-09
摘要:
目的 TCP(teosinte branched 1/cincinnata/proliferating cell factor)是植物特有的转录因子,在植物整个生长发育过程中都发挥着重要作用。通过鉴定白栎TCP基因家族,研究QfTCP基因在白栎分枝发育中的表达模式,验证关键基因QfTCP22的功能,为白栎株型改良提供分子基础。 方法 基于白栎基因组数据,通过生物信息学方法对白栎TCP基因家族进行鉴定,并分析该家族成员的理化性质、染色体分布、系统进化、基因结构、顺式作用元件和基因共线性。结合转录组数据和RT-qPCR对QfTCP基因家族成员在腋芽发育中的表达模式进行分析,并对QfTCP22进行异源过表达拟南芥验证基因功能。 结果 在白栎中共鉴定23个TCP基因,分为PCE、CIN和CYC/TB1 3个亚家族,不均匀地分布在10条染色体上,其中,第10染色体含有的成员数量最多。所有QfTCP成员都含有一个共同的保守基序Motif 1,基因结构较为简单。共线性分析发现10对QfTCP基因存在共线性关系。顺式作用元件分析发现QfTCP基因的功能较为复杂,在光信号响应、激素调控和逆境胁迫等方面均发挥作用。腋芽发育转录组数据和RT-qPCR结果表明,在腋芽发育中,CYC/TB1亚族的QfTCP1和QfTCP22均呈显著下调趋势,在拟南芥brc1突变体中过表达QfTCP22,能显著减少其分枝数量。 结论 鉴定白栎TCP家族成员,揭示QfTCP22是植物分枝发育中的抑制因子。
熊仕发, 陈益存, 吴立文, 施翔, 张盛剿, 彭方有, 陈涛梅, 汪阳东. 白栎TCP基因家族的鉴定及功能分析[J]. 生物技术通报, doi: 10.13560/j.cnki.biotech.bull.1985.2025-0818.
XIONG Shi-fa, CHEN Yi-cun, WU Li-wen, SHI Xiang, ZHANG Sheng-jiao, PENG Fang-you, CHEN Tao-mei, WANG Yang-dong. Identification and Functional Analysis of the TCP Gene Family in Quercus fabri[J]. Biotechnology Bulletin, doi: 10.13560/j.cnki.biotech.bull.1985.2025-0818.
引物名称 Primer name | 序列 Sequence (5′‒3′) | 用途 Usage |
|---|---|---|
| QfTCP1-F | TCACACTGGGCTCCTCTTTT | RT-qPCR |
| QfTCP1-R | TTTGGACCACCATTGCTTTT | RT-qPCR |
| QfTCP22-F | TTCCAACAAAAATCACATCA | RT-qPCR |
| QfTCP22-R | TTCAAGAGTAACCATTCAACG | RT-qPCR |
| QfTCP22-OE-F | TCAGCAGTCGAAGAGC | 过表达 |
| QfTCP22-OE-R | TTAGCGTGTGAAGAGC | 过表达 |
| QfActin-F | GATTCTGGTGATGGTGTGAGC | 内参 |
| QfActin-R | ATGAGAGATGGCTGGAAGAGT | 内参 |
表1 引物列表
Table 1 Primer list
引物名称 Primer name | 序列 Sequence (5′‒3′) | 用途 Usage |
|---|---|---|
| QfTCP1-F | TCACACTGGGCTCCTCTTTT | RT-qPCR |
| QfTCP1-R | TTTGGACCACCATTGCTTTT | RT-qPCR |
| QfTCP22-F | TTCCAACAAAAATCACATCA | RT-qPCR |
| QfTCP22-R | TTCAAGAGTAACCATTCAACG | RT-qPCR |
| QfTCP22-OE-F | TCAGCAGTCGAAGAGC | 过表达 |
| QfTCP22-OE-R | TTAGCGTGTGAAGAGC | 过表达 |
| QfActin-F | GATTCTGGTGATGGTGTGAGC | 内参 |
| QfActin-R | ATGAGAGATGGCTGGAAGAGT | 内参 |
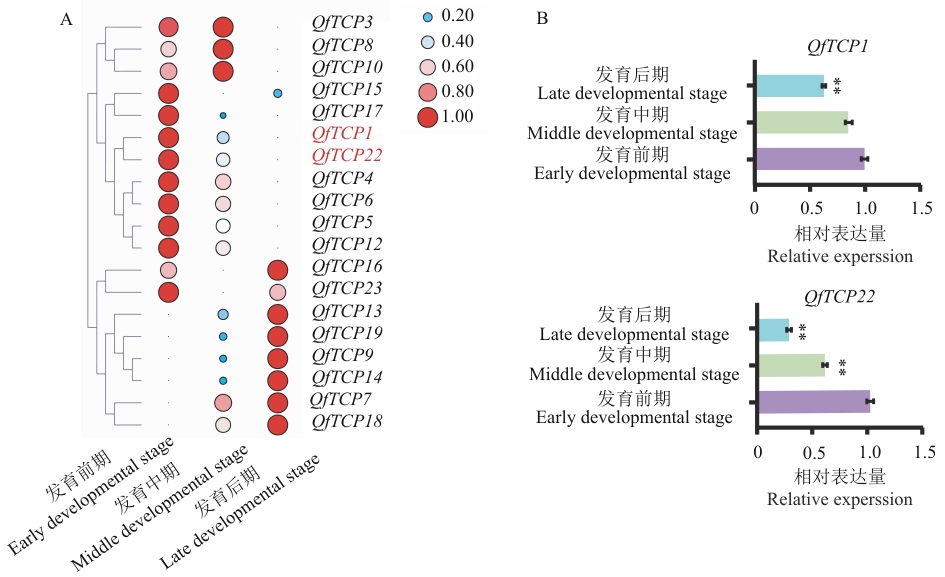
图6 白栎TCP基因家族在腋芽发育中的表达热图(A)及部分成员荧光定量PCR验证(B)* P<0.05, ** P<0.01. The same below
Fig. 6 Heatmap of TCP gene family expression in axillary bud development of Q. fabri (A) and RT-qPCR validation of some members (B)
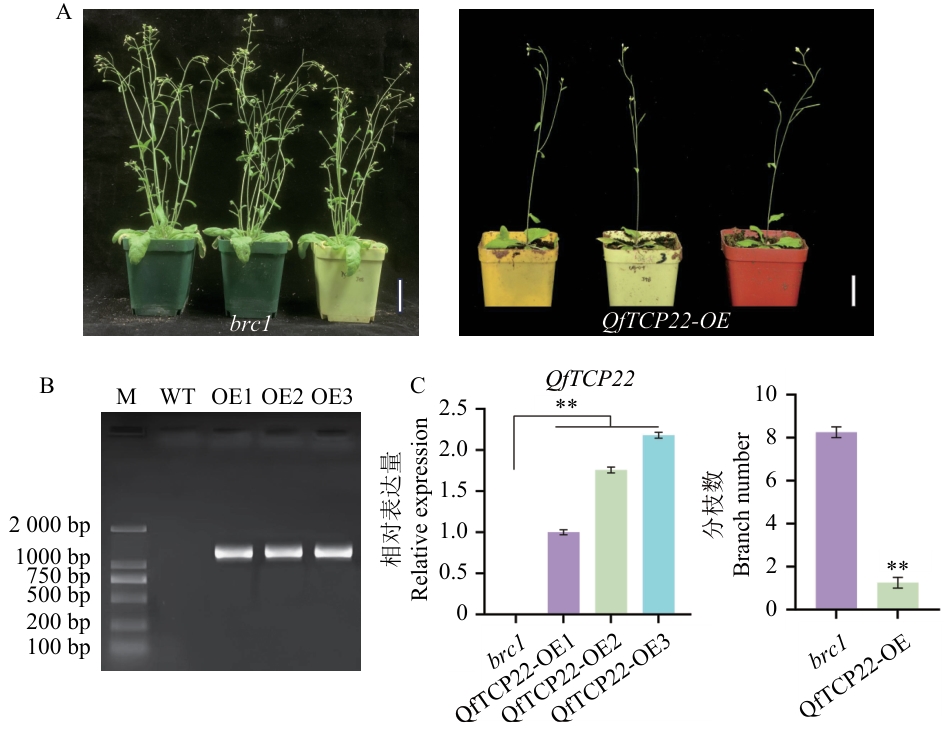
图7 过表达QfTCP22拟南芥植株的表型及RT-qPCRA:brc1突变体和QfTCP22回补brc1突变体的植株表型;B:PCR鉴定转基因阳性植株(M:DL2000 marker;WT:阴性对照;OE1‒OE3:转基因植株);C:brc1突变体和QfTCP22回补brc1突变体植株的分枝数及RT-qPCR
Fig. 7 Plant phenotype and RT-qPCR of Arabidopsis plants overexpressing QfTCP22A: Phenotypes of the brc1 mutant and QfTCP22-complemented brc1 mutant plants. B: PCR identification of transgenic positive plants (M: DL2000 marker; WT: negative control; OE1-OE3: transgenic lines). C: Branch number of the brc1 mutant and QfTCP22-complemented brc1 mutant plants and RT-qPCR
| [1] | Doebley J, Stec A, Hubbard L. The evolution of apical dominance in maize [J]. Nature, 1997, 386(6624): 485-488. |
| [2] | Luo D, Carpenter R, Vincent C, et al. Origin of floral asymmetry in Antirrhinum [J]. Nature, 1996, 383(6603): 794-799. |
| [3] | Kosugi S, Ohashi Y. PCF1 and PCF2 specifically bind to cis elements in the rice proliferating cell nuclear antigen gene [J]. Plant Cell, 1997, 9(9): 1607-1619. |
| [4] | Zhang Y, Xu YP, Nie JK, et al. DNA-TCP complex structures reveal a unique recognition mechanism for TCP transcription factor families [J]. Nucleic Acids Res, 2023, 51(1): 434-448. |
| [5] | Cubas P, Lauter N, Doebley J, et al. The TCP domain: a motif found in proteins regulating plant growth and development [J]. Plant J, 1999, 18(2): 215-222. |
| [6] | Howarth DG, Donoghue MJ. Phylogenetic analysis of the “ECE” (CYC/TB1) clade reveals duplications predating the core eudicots [J]. Proc Natl Acad Sci USA, 2006, 103(24): 9101-9106. |
| [7] | Martín-Trillo M, Cubas P. TCP genes: a family snapshot ten years later [J]. Trends Plant Sci, 2010, 15(1): 31-39. |
| [8] | Zhang W, Cochet F, Ponnaiah M, et al. The MPK8-TCP14 pathway promotes seed germination in Arabidopsis [J]. Plant J, 2019, 100(4): 677-692. |
| [9] | Giraud E, Ng S, Carrie C, et al. TCP transcription factors link the regulation of genes encoding mitochondrial proteins with the circadian clock in Arabidopsis thaliana [J]. Plant Cell, 2010, 22(12): 3921-3934. |
| [10] | Li X, Yang Q, Liao XQ, et al. A natural antisense RNA improves Chrysanthemum cold tolerance by regulating the transcription factor DgTCP1 [J]. Plant Physiol, 2022, 190(1): 605-620. |
| [11] | Wang YT, Cao Y, Qin GJ. Multifaceted roles of TCP transcription factors in fate determination [J]. New Phytol, 2025, 245(1): 95-101. |
| [12] | Yao X, Ma H, Wang J, et al. Genome-wide comparative analysis and expression pattern of TCP gene families in Arabidopsis thaliana and Oryza sativa [J]. J Integr Plant Biol, 2007, 49(6): 885-897. |
| [13] | Parapunova V, Busscher M, Busscher-Lange J, et al. Identification, cloning and characterization of the tomato TCP transcription factor family [J]. BMC Plant Biol, 2014, 14: 157. |
| [14] | 陈智华, 乔振升, 李嘉其, 等. 滇杨TCP基因家族的全基因组鉴定与分析 [J]. 生物技术通报, 2024, 40(11): 214-226. |
| Chen ZH, Qiao ZS, Li JQ, et al. Genome-wide identification and analysis of the TCP gene family in Populus yunnanensis [J]. Biotechnol Bull, 2024, 40(11): 214-226. | |
| [15] | 王苗苗, 赵相龙, 王召明, 等. 花苜蓿TCP基因家族的鉴定及其在干旱胁迫下的表达模式分析 [J]. 生物技术通报, 2025, 41(6): 179-190. |
| Wang MM, Zhao XL, Wang ZM, et al. Identification of TCP gene family in Medicago ruthenica and their expression pattern analysis under drought stress [J]. Biotechnol Bull, 2025, 41(6): 179-190. | |
| [16] | 宋姝熠, 蒋开秀, 刘欢艳, 等. ‘红阳’猕猴桃TCP基因家族鉴定及其在果实中的表达分析 [J]. 生物技术通报, 2025, 41(3): 190-201. |
| Song SY, Jiang KX, Liu HY, et al. Identification of the TCP gene family in Actinidia chinensis var. hongyang and their expression analysis in fruit [J]. Biotechnol Bull, 2025, 41(3): 190-201. | |
| [17] | 张雪莹, 尹一歌, 姜晶, 等. 参与番茄叶形发育的TCP转录因子的表达及生物信息分析 [J]. 中国蔬菜, 2021(10): 45-56. |
| Zhang XY, Yin YG, Jiang J, et al. Expression of SlTCPs transcription factors involved in tomato leaf shape development and bioinformatic analysis [J]. China Veg, 2021(10): 45-56. | |
| [18] | Pruneda-Paz JL, Breton G, Para A, et al. A functional genomics approach reveals CHE as a component of the Arabidopsis circadian clock [J]. Science, 2009, 323(5920): 1481-1485. |
| [19] | Aguilar-Martínez JA, Poza-Carrión C, Cubas P. Arabidopsis BRANCHED1 acts as an integrator of branching signals within axillary buds [J]. Plant Cell, 2007, 19(2): 458-472. |
| [20] | Finlayson SA. Arabidopsis Teosinte Branched1-like 1 regulates axillary bud outgrowth and is homologous to monocot Teosinte Branched1 [J]. Plant Cell Physiol, 2007, 48(5): 667-677. |
| [21] | Li GF, Tan M, Ma JJ, et al. Molecular mechanism of MdWUS2-MdTCP12 interaction in mediating cytokinin signaling to control axillary bud outgrowth [J]. J Exp Bot, 2021, 72(13): 4822-4838. |
| [22] | 李璇, 李垚, 方炎明. 基于优化的Maxent模型预测白栎在中国的潜在分布区 [J]. 林业科学, 2018, 54(8): 153-164. |
| Li X, Li Y, Fang YM. Prediction of potential suitable distribution areas of Quercus fabri in China based on an optimized maxent model [J]. Sci Silvae Sin, 2018, 54(8): 153-164. | |
| [23] | 熊仕发, 吴立文, 陈益存, 等. 不同种源白栎果实形态特征和营养成分含量变异分析 [J]. 林业科学研究, 2020, 33(2): 93-102. |
| Xiong SF, Wu LW, Chen YC, et al. Variation in morphological characters and nutrient contents of Quercus fabri fruits from different provenances [J]. For Res, 2020, 33(2): 93-102. | |
| [24] | Wu LW, Cai YQ, Jiang CG, et al. Uncovering the genetic basis for enhanced mushroom flavor in Quercus fabri through genome sequencing and metabolic profiling [J]. Hortic Res, 2025, 12(9): uhaf156. |
| [25] | Xiong SF, Zhao YX, Chen YC, et al. Transcriptome and proteome analyses reveal a critical role of the sucrose anabolic pathway in regulating Quercus fabri Hance axillary bud outgrowth [J]. Ind Crops Prod, 2023, 194: 116284. |
| [26] | Xiong SF, Wu LW, Chen YC, et al. Multi-omics analysis reveals the regulatory mechanism of branching development in Quercus fabri [J]. J Proteom, 2025, 313: 105373. |
| [27] | Chen CJ, Wu Y, Li JW, et al. TBtools-II: a “one for all, all for one” bioinformatics platform for biological big-data mining [J]. Mol Plant, 2023, 16(11): 1733-1742. |
| [28] | Kumar S, Stecher G, Tamura K. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets [J]. Mol Biol Evol, 2016, 33(7): 1870-1874. |
| [29] | 徐文鸾, 钟函江, 刘杏月, 等. 茶树树型相关CYC/TB1家族全基因组鉴定及表达分析 [J]. 植物生理学报, 2021, 57(11): 2179-2191. |
| Xu WL, Zhong HJ, Liu XY, et al. Genome-wide identification and expression analysis of tree architectures related CYC/TB1 family in tea plant(Camellia sinensis) [J]. Plant Physiol J, 2021, 57(11): 2179-2191. | |
| [30] | Wang S, Shen YR, Guo LY, et al. Innovation and emerging roles of Populus trichocarpa teosinte branched1/cycloidea/proliferating cell factor transcription factors in abiotic stresses by whole-genome duplication [J]. Front Plant Sci, 2022, 13: 850064. |
| [31] | 阮先乐, 张福丽, 王俊生. 番茄TCP基因家族全基因组鉴定和分析 [J]. 基因组学与应用生物学, 2017, 36(6): 2539-2547. |
| Ruan XL, Zhang FL, Wang JS. Genome-wide identification and analysis of TCP gene family in tomato [J]. Genom Appl Biol, 2017, 36(6): 2539-2547. | |
| [32] | 宋宇英, 江敏, 王世杰, 等. 蒙古栎TCP基因家族全基因组鉴定与分析 [J/OL]. 分子植物育种, 2023. . |
| Song YY, Jiang M, Wang SJ, et al. Genome-wide identification and analysis of TCP gene family in Quercus mongolica [J/OL]. Mol Plant Breed, 2023. . | |
| [33] | Wang YC, Li JJ, Chen YX, et al. Genome-wide identification of TCP transcription factors and their potential roles in hydrolyzable tannin production in Quercus variabilis cupule [J]. Front Plant Sci, 2024, 15: 1444081. |
| [34] | Chai WB, Jiang PF, Huang GY, et al. Identification and expression profiling analysis of TCP family genes involved in growth and development in maize [J]. Physiol Mol Biol Plants, 2017, 23(4): 779-791. |
| [35] | 刘俊, 陈玉龙, 刘燕, 等. 杜仲TIFY转录因子鉴定与表达分析 [J]. 中国实验方剂学杂志, 2021, 27(19): 165-174. |
| Liu J, Chen YL, Liu Y, et al. Identification and expression analysis of TIFY transcription factor in Eucommia ulmoides [J]. Chin J Exp Tradit Med Formulae, 2021, 27(19): 165-174. | |
| [36] | Yu L, Chen QW, Zheng JR, et al. Genome-wide identification and expression pattern analysis of the TCP transcription factor family in Ginkgo biloba [J]. Plant Signal Behav, 2022, 17(1): 1994248. |
| [37] | Lei N, Yu X, Li SX, et al. Phylogeny and expression pattern analysis of TCP transcription factors in cassava seedlings exposed to cold and/or drought stress [J]. Sci Rep, 2017, 7(1): 10016. |
| [38] | Poza-Carrión C, Aguilar-Martínez JA, Cubas P. Role of TCP gene BRANCHED1 in the control of shoot branching in Arabidopsis [J]. Plant Signal Behav, 2007, 2(6): 551-552. |
| [39] | Martín-Trillo M, Grandío EG, Serra F, et al. Role of tomato BRANCHED1-like genes in the control of shoot branching [J]. Plant J, 2011, 67(4): 701-714. |
| [40] | Amoo Olalekan, 胡利民, 翟云孤, 等. 利用基因编辑技术研究BRANCHED1参与油菜分枝过程的调控 [J]. 生物技术通报, 2022, 38(4): 97-105. |
| Amoo O, Hu LM, Zhai YG, et al. Regulation of shoot branching by BRANCHED1 in Brassica napus based on gene editing technology [J]. Biotechnol Bull, 2022, 38(4): 97-105. | |
| [41] | 冯爽爽, 罗嘉翼, 朱曦鉴, 等. 二倍体马铃薯StBRC1a功能缺失突变体的获得及其功能分析 [J]. 园艺学报, 2020, 47(1): 63-72. |
| Feng SS, Luo JY, Zhu XJ, et al. Homozygous mutant construction and function analysis of TCP transcription factor StBRC1a in diploid potato [J]. Acta Hortic Sin, 2020, 47(1): 63-72. |
| [1] | 陈登科, 兰刚, 夏芝, 侯保国, 杨六六, 曹彩荣, 李朋波, 吴翠翠. 花生ZF-HD基因家族的鉴定和非生物胁迫响应分析[J]. 生物技术通报, 2026, 42(4): 114-128. |
| [2] | 农韦优, 赵昌祖, 钱禛锋, 丁倩, 王誉洁, 陈疏影, 何丽莲, 李富生. 蔗茅EfBBX基因家族鉴定及冷胁迫表达模式分析[J]. 生物技术通报, 2026, 42(2): 1-11. |
| [3] | 吕呈聪, 衡蒙, 陈思琪, 金雪花. 彩色马蹄莲花青素苷转运相关ZhGSTF的克隆及功能分析[J]. 生物技术通报, 2026, 42(1): 161-169. |
| [4] | 杨丹, 靳雅荣, 毛春力, 王碧娴, 张雅宁, 杨智怡, 周芷瑶, 杨锐鸣, 范恒睿, 黄琳凯, 严海东. 象草C2H2基因家族鉴定及表达分析[J]. 生物技术通报, 2026, 42(1): 251-261. |
| [5] | 任云儿, 伍国强, 成斌, 魏明. 甜菜BvATGs基因家族全基因组鉴定及盐胁迫下表达模式分析[J]. 生物技术通报, 2026, 42(1): 184-197. |
| [6] | 杨娟, 冯慧, 吉乃喆, 孙丽萍, 王赟, 张佳楠, 赵世伟. 月季AP2/ERF转录因子RcERF4和RcRAP2-12的克隆及功能分析[J]. 生物技术通报, 2026, 42(1): 150-160. |
| [7] | 吴翠翠, 陈登科, 兰刚, 夏芝, 李朋波. 花生转录因子AhHDZ70的生物信息学分析及耐盐耐旱性研究[J]. 生物技术通报, 2026, 42(1): 198-207. |
| [8] | 刘佳丽, 宋经荣, 赵文宇, 张馨元, 赵子洋, 曹一博, 张凌云. 蓝莓R2R3-MYB基因鉴定及类黄酮调控基因表达分析[J]. 生物技术通报, 2025, 41(9): 124-138. |
| [9] | 程婷婷, 刘俊, 王利丽, 练从龙, 魏文君, 郭辉, 吴尧琳, 杨晶凡, 兰金旭, 陈随清. 杜仲查尔酮异构酶基因家族全基因组鉴定及其表达模式分析[J]. 生物技术通报, 2025, 41(9): 242-255. |
| [10] | 徐小萍, 杨成龙, 和兴, 郭文杰, 吴健, 方少忠. 百合LoAPS1克隆及其在休眠解除过程的功能分析[J]. 生物技术通报, 2025, 41(9): 195-206. |
| [11] | 董向向, 缪百灵, 许贺娟, 陈娟娟, 李亮杰, 龚守富, 朱庆松. 森林草莓FveBBX20基因的生物信息学分析及开花调控功能[J]. 生物技术通报, 2025, 41(9): 115-123. |
| [12] | 李珊, 马登辉, 马红义, 姚文孔, 尹晓. 葡萄SKP1基因家族鉴定与表达分析[J]. 生物技术通报, 2025, 41(9): 147-158. |
| [13] | 黄国栋, 邓宇星, 程宏伟, 但焱南, 周会汶, 吴兰花. 大豆ZIP基因家族鉴定及响应铝胁迫的表达分析[J]. 生物技术通报, 2025, 41(9): 71-81. |
| [14] | 巩慧玲, 邢玉洁, 马俊贤, 蔡霞, 冯再平. 马铃薯LAC基因家族的鉴定及盐胁迫下表达分析[J]. 生物技术通报, 2025, 41(9): 82-93. |
| [15] | 腊贵晓, 赵玉龙, 代丹丹, 余永亮, 郭红霞, 史贵霞, 贾慧, 杨铁钢. 红花质膜H+-ATPase基因家族成员鉴定及响应低氮低磷胁迫的表达分析[J]. 生物技术通报, 2025, 41(8): 220-233. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||