








生物技术通报 ›› 2026, Vol. 42 ›› Issue (4): 153-160.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0837
孙婷1( ), 张艳1, 刘玉珊1, 冯媛媛2, 秦恒山1, 张军1, 何小岗1, 张景荣1(
), 张艳1, 刘玉珊1, 冯媛媛2, 秦恒山1, 张军1, 何小岗1, 张景荣1( )
)
收稿日期:2025-08-02
出版日期:2026-04-26
发布日期:2026-04-30
通讯作者:
张景荣,男,博士,副研究员,研究方向 :植物生物技术与遗传改良;E-mail: zjr_sc@163.com作者简介:孙婷,女,硕士,研究员,研究方向 :蔬菜育种与栽培;E-mail: 5796946@qq.com
基金资助:
SUN Ting1( ), ZHANG Yan1, LIU Yu-shan1, FENG Yuan-yuan2, QIN Heng-shan1, ZHANG Jun1, HE Xiao-gang1, ZHANG Jing-rong1(
), ZHANG Yan1, LIU Yu-shan1, FENG Yuan-yuan2, QIN Heng-shan1, ZHANG Jun1, HE Xiao-gang1, ZHANG Jing-rong1( )
)
Received:2025-08-02
Published:2026-04-26
Online:2026-04-30
摘要:
目的 从黄秋葵(Abelmoschus esculentus L.)中克隆黄烷酮3-羟化酶基因(AeF3H),通过拟南芥遗传转化鉴定其生物学功能,为阐明黄秋葵类黄酮累积的调控机理提供理论基础。 方法 根据黄秋葵转录组数据设计引物,以秋葵籽cDNA为模板,通过PCR扩增获得AeF3H基因;通过生物信息学分析AeF3H的基本特征;采用实时荧光定量PCR(RT-qPCR)分析其组织表达特性;构建35::AeF3H::YFP过表达载体,通过花序浸染法转化拟南芥,采用硝酸铝比色法测定T3代拟南芥转基因株系中总黄酮含量变化;利用拟南芥原生质体瞬时表达技术分析AeF3H亚细胞定位。 结果 AeF3H的ORF长1 101 bp,编码366个氨基酸;基因组序列长1 304 bp,含3个外显子和2个内含子。推测其蛋白的相对分子量为41.08 kD,理论等电点为5.32。系统进化树分析发现,AeF3H与木槿(Hibiscus trionum)的F3H亲缘关系最近。RT-qPCR结果表明,AeF3H具有一定的组织表达特异性,在嫩籽和花蕾中的表达量最高,在茎和果荚中的表达量最低。亚细胞定位显示其蛋白同时定位于细胞核和细胞质中。过表达AeF3H的拟南芥植株,其总黄酮含量为野生型的4.75倍。 结论 AeF3H具有一定的组织表达特异性,其蛋白定位于细胞核和细胞质中。AeF3H在黄秋葵类黄酮生物合成代谢途径中发挥着重要作用,过量表达AeF3H能够显著增加转基因拟南芥的总黄酮含量。
孙婷, 张艳, 刘玉珊, 冯媛媛, 秦恒山, 张军, 何小岗, 张景荣. 黄秋葵AeF3H基因的克隆与功能分析[J]. 生物技术通报, 2026, 42(4): 153-160.
SUN Ting, ZHANG Yan, LIU Yu-shan, FENG Yuan-yuan, QIN Heng-shan, ZHANG Jun, HE Xiao-gang, ZHANG Jing-rong. Cloning and Functional Analysis of AeF3H Gene in Okra[J]. Biotechnology Bulletin, 2026, 42(4): 153-160.
引物名称 Primer name | 引物序列 Primer sequence (5′-3′) | 用途 Purpose |
|---|---|---|
| F3H-F | CCCAACTAAACATAGAAACACTCAC | ORF序列扩增 |
| F3H-R | ACTAAGACAATGGCACCAAGCAC | Amplification of ORF sequence |
| F3HYFP-F | CCGCTCGAGATGGCTCCTTCCACTCTCAC (Xoh I) | 载体构建 |
| F3HYFP-R | CGCGGATCCTGCAAGGATTTGCTCTAGCG (BamH I) | Vector construction |
| YFP-F | GCGACGTAAACGGCCACAAGT | 转基因拟南芥的PCR鉴定 |
| YFP-R | CAGCTCGTCCATGCCGAGAGT | PCR identification of transgenic Arabidopsis |
| qeIF4a-F | ATGCATATGGTTTTGAGAAGCC | 秋葵内参基因 |
| qeIF4a-R | AAAGTTGCAGTCTTCCCAGTTC | Reference gene in A. esculentus |
| qF3H-F | AGTTTTTCGCTTTGCCTGCT | 实时荧光定量PCR |
| qF3H-R | TCTCGCTGTATTCCTTTGTCAC | Real-time quantitative PCR |
| qAtEF1α-F | ATTGACAGGCGTTCTGGTAAG | 拟南芥内参基因 |
| qAtEF1α-R | CAGCAACGGTCTGCCTCAT | Reference gene in Arabidopsis |
表 1 本实验所用的引物序列
Table 1 Primers' sequences used in this study
引物名称 Primer name | 引物序列 Primer sequence (5′-3′) | 用途 Purpose |
|---|---|---|
| F3H-F | CCCAACTAAACATAGAAACACTCAC | ORF序列扩增 |
| F3H-R | ACTAAGACAATGGCACCAAGCAC | Amplification of ORF sequence |
| F3HYFP-F | CCGCTCGAGATGGCTCCTTCCACTCTCAC (Xoh I) | 载体构建 |
| F3HYFP-R | CGCGGATCCTGCAAGGATTTGCTCTAGCG (BamH I) | Vector construction |
| YFP-F | GCGACGTAAACGGCCACAAGT | 转基因拟南芥的PCR鉴定 |
| YFP-R | CAGCTCGTCCATGCCGAGAGT | PCR identification of transgenic Arabidopsis |
| qeIF4a-F | ATGCATATGGTTTTGAGAAGCC | 秋葵内参基因 |
| qeIF4a-R | AAAGTTGCAGTCTTCCCAGTTC | Reference gene in A. esculentus |
| qF3H-F | AGTTTTTCGCTTTGCCTGCT | 实时荧光定量PCR |
| qF3H-R | TCTCGCTGTATTCCTTTGTCAC | Real-time quantitative PCR |
| qAtEF1α-F | ATTGACAGGCGTTCTGGTAAG | 拟南芥内参基因 |
| qAtEF1α-R | CAGCAACGGTCTGCCTCAT | Reference gene in Arabidopsis |
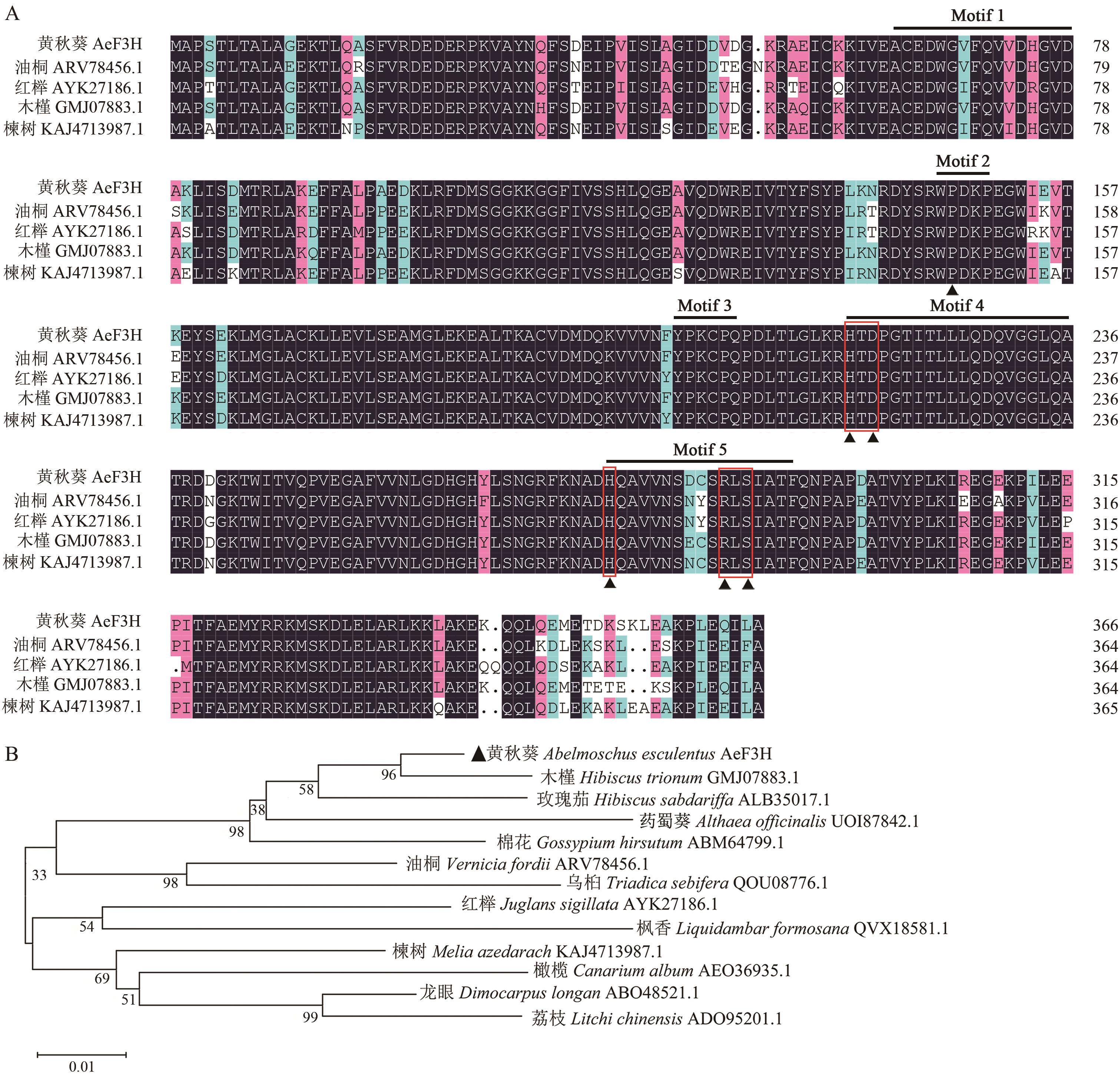
图2 AeF3H的序列比对(A)与系统进化树分析(B)图A中,红色方框标示出亚铁离子配位基序HXDX55H及2-氧代戊二酸(2-ODD)结合基序RLS,保守氨基酸残基用黑色三角形(▲)标记;同时标记出植物2-氧戊二酸依赖型双加氧酶中的5个保守基序
Fig. 2 Sequence alignment (A) and phylogenetic tree analysis (B) of AeF3HIn Fig. A, red boxes indicate the ferrous iron ligation motif HXDX55H and a 2oxoglutarate (2-ODD) binding motif RLS. Black triangles (▲) indicate the conserved amino acid residues in F3Hs. Five conserved motifs in plant 2-oxoglutarate-dependent dioxygenase are also marked
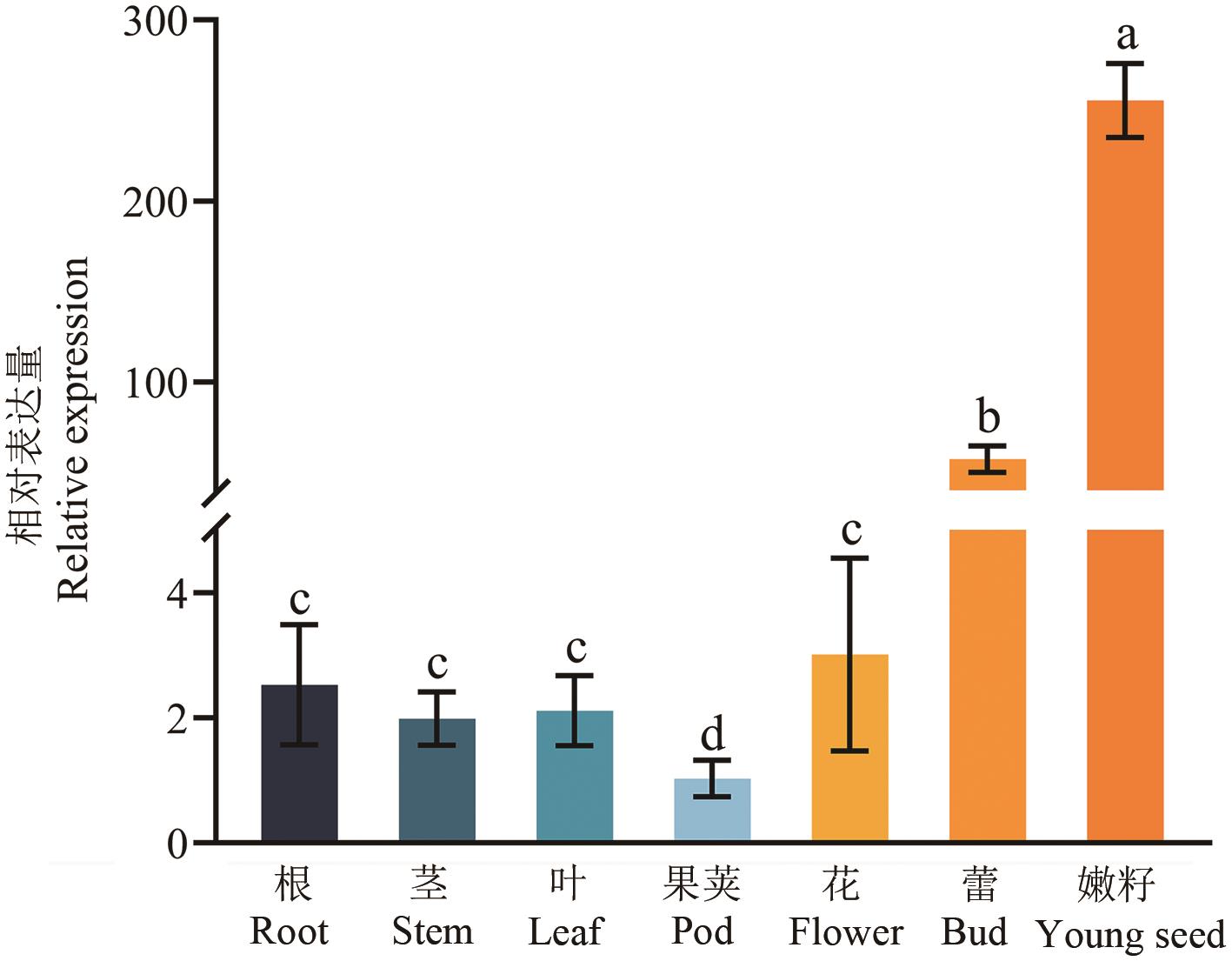
图3 AeF3H基因的表达模式分析不同字母表示差异显著(P<0.05)。下同
Fig. 3 Expression pattern analysis of the AeF3H geneDifferent lowercase letters indicate significant differences (P<0.05). The same below
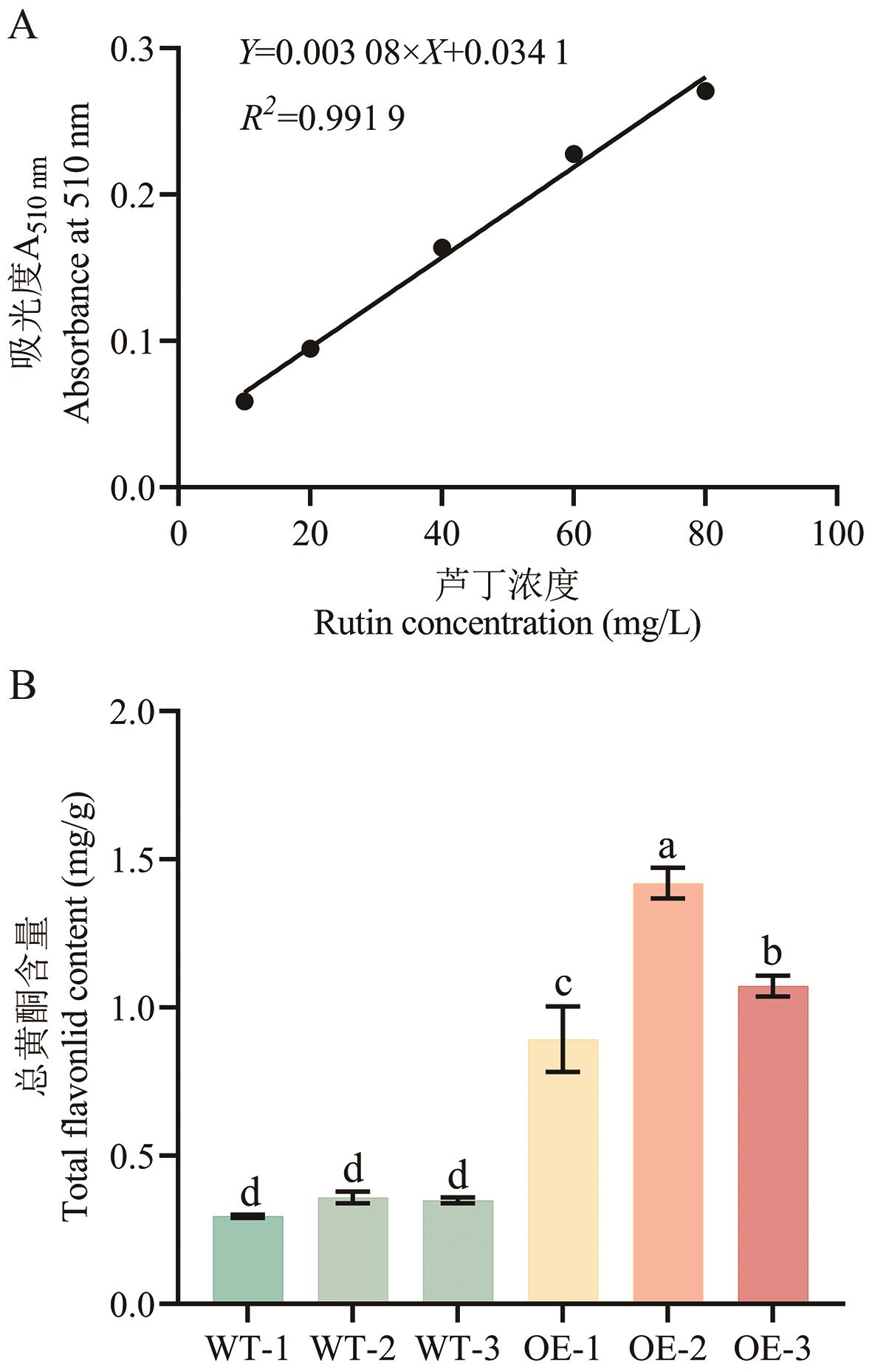
图6 过量表达AeF3H对拟南芥总黄酮含量的影响A:芦丁标准曲线;B:野生型植株和转基因植株总黄酮含量的测定
Fig. 6 Effects of overexpressing AeF3H on the total flavonoids contents in ArabidopsisA: Standard curve of rutin. B: Determination of total flavonoids in wild-type and transgenic plants
| [1] | Romdhane MH, Chahdoura H, Barros L, et al. Chemical composition, nutritional value, and biological evaluation of Tunisian okra pods (Abelmoschus esculentus L. moench) [J]. Molecules, 2020, 25(20): 4739. |
| [2] | Gakhar M, Singh L, Routh S, et al. Evaluation of anti-cancer potential of Abelmoschus esculentus (Okra) [J]. Protein Pept Lett, 2025: 32(8) : 575-583. |
| [3] | Panighel G, Ferrarese I, Lupo MG, et al. Investigating the in vitro mode of action of okra (Abelmoschus esculentus) as hypocholesterolemic, anti-inflammatory, and antioxidant food [J]. Food Chem Mol Sci, 2022, 5: 100126. |
| [4] | Saatchi A, Aghamohammadzadeh N, Beheshtirouy S, et al. Anti-hyperglycemic effect of Abelmoschus culentesus (Okra) on patients with diabetes type 2: a randomized clinical trial [J]. Phytother Res, 2022, 36(4): 1644-1651. |
| [5] | 郑晟, 毛玉珊, 张腾国, 等. 柠条锦鸡儿F3H基因克隆及功能分析 [J]. 广西植物, 2017, 37(6): 723-733. |
| Zheng S, Mao YS, Zhang TG, et al. Cloning and expression analysis of F3H gene in Caragana korshinskii [J]. Guihaia, 2017, 37(6): 723-733. | |
| [6] | 朱三明, 郑敏敏, 田恬, 等. 植物次生代谢途径与调控研究进展 [J]. 植物生理学报, 2023, 59(12): 2188-2216. |
| Zhu SM, Zheng MM, Tian T, et al. Research progress on plant secondary metabolism and regulation [J]. Plant Physiol J, 2023, 59(12): 2188-2216. | |
| [7] | Liu WX, Feng Y, Yu SH, et al. The flavonoid biosynthesis network in plants [J]. Int J Mol Sci, 2021, 22(23): 12824. |
| [8] | 段玥彤, 王鹏年, 张春宝, 等. 植物黄烷酮-3-羟化酶基因研究进展 [J]. 生物技术通报, 2022, 38(6): 27-33. |
| Duan YT, Wang PN, Zhang CB, et al. Research progress in plant flavanone-3-hydroxylase gene [J]. Biotechnol Bull, 2022, 38(6): 27-33. | |
| [9] | Zhu SM, Cui MY, Zhao Q. Characterization of the 2ODD genes of DOXC subfamily and its members involved in flavonoids biosynthesis in Scutellaria baicalensis [J]. BMC Plant Biol, 2024, 24(1): 804. |
| [10] | Li XH, Park NI, Kim YB, et al. Accumulation of flavonoids and expression of flavonoid biosynthetic genes in Tartary and rice-Tartary buckwheat [J]. Process Biochem, 2012, 47(12): 2306-2310. |
| [11] | Sunil L, Shetty NP. Biosynthesis and regulation of anthocyanin pathway genes [J]. Appl Microbiol Biotechnol, 2022, 106(5/6): 1783-1798. |
| [12] | Dai MJ, Kang XR, Wang YQ, et al. Functional characterization of flavanone 3-hydroxylase (F3H) and its role in anthocyanin and flavonoid biosynthesis in mulberry [J]. Molecules, 2022, 27(10): 3341. |
| [13] | Xiong S, Tian N, Long JH, et al. Molecular cloning and characterization of a flavanone 3-hydroxylase gene from Artemisia annua L [J]. Plant Physiol Biochem, 2016, 105: 29-36. |
| [14] | 曹鑫娴, 石佳琪, 王霜, 等. 苦荞黄烷酮3-羟化酶基因FtF3H及其启动子的克隆与功能分析 [J]. 四川农业大学学报, 2023, 41(4): 582-591, 639. |
| Cao XX, Shi JQ, Wang S, et al. Cloning and functional analysis of flavanone 3-hydroxylase gene FtF3H and its promoter in Fagopyrum tataricum [J]. J Sichuan Agric Univ, 2023, 41(4): 582-591, 639. | |
| [15] | Song XY, Diao JJ, Ji J, et al. Molecular cloning and identification of a flavanone 3-hydroxylase gene from Lycium chinense, and its overexpression enhances drought stress in tobacco [J]. Plant Physiol Biochem, 2016, 98: 89-100. |
| [16] | 赵莹, 杨欣宇, 赵晓丹, 等. 植物类黄酮化合物生物合成调控研究进展 [J]. 食品工业科技, 2021, 42(21): 454-463. |
| Zhao Y, Yang XY, Zhao XD, et al. Research progress on regulation of plant flavonoids biosynthesis [J]. Sci Technol Food Ind, 2021, 42(21): 454-463. | |
| [17] | Yang J, Chen XQ, Rao SQ, et al. Identification and quantification of flavonoids in okra (Abelmoschus esculentus L. moench) and antiproliferative activity in vitro of four main components identified [J]. Metabolites, 2022, 12(6): 483. |
| [18] | Cui YT, Wu JG, Chen YY, et al. Optimization of near-infrared reflectance models in determining flavonoid composition of okra (Abelmoschus esculentus L.) pods [J]. Food Chem, 2023, 418: 135953. |
| [19] | Guebebia S, Espinosa-Ruiz C, Zourgui L, et al. Effects of okra (Abelmoschus esculentus L.) leaves, fruits and seeds extracts on European sea bass (Dicentrarchus labrax) leukocytes, and their cytotoxic, bactericidal and antioxidant properties [J]. Fish Shellfish Immunol, 2023, 138: 108799. |
| [20] | Wang RY, Li W, He Q, et al. The genome of okra (Abelmoschus esculentus) provides insights into its genome evolution and high nutrient content [J]. Hortic Res, 2023, 10(8): uhad120. |
| [21] | Clough SJ, Bent AF. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana [J]. Plant J, 1998, 16(6): 735-743. |
| [22] | Yoo SD, Cho YH, Sheen J. Arabidopsis mesophyll protoplasts: a versatile cell system for transient gene expression analysis [J]. Nat Protoc, 2007, 2(7): 1565-1572. |
| [23] | Zhao FF, Zhao T, Deng LB, et al. Visualizing the essential role of complete virion assembly machinery in efficient hepatitis C virus cell-to-cell transmission by a viral infection-activated split-intein-mediated reporter system [J]. J Virol, 2017, 91(2): e01720-16. |
| [24] | 杨飞芸, 武燕燕, 崔爽, 等. 异源表达CiRS基因通过生成白藜芦醇增强拟南芥的抗氧化能力 [J]. 中国生物工程杂志, 2017, 37(12): 27-33. |
| Yang FY, Wu YY, Cui S, et al. Heterologous expression of CiRS gene enhances the antioxidant capacity of Arabidopsis by increasing the content of resveratrol [J]. China Biotechnol, 2017, 37(12): 27-33. | |
| [25] | Zhang JR, Feng YY, Yang MJ, et al. Systematic screening and validation of reliable reference genes for qRT-PCR analysis in okra (Abelmoschus esculentus L.) [J]. Sci Rep, 2022, 12(1): 12913. |
| [26] | Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method [J]. Methods, 2001, 25(4): 402-408. |
| [27] | 李萍, 王若曦, 王欢. 黄酮3-羟化酶(F3H)的生物信息学分析 [J]. 生物学杂志, 2015, 32(6): 25-29. |
| Li P, Wang RX, Wang H. Bioinformatics analysis of flavanone-3-hydroxylase (F3H) [J]. J Biol, 2015, 32(6): 25-29. | |
| [28] | 张欣宇, 董潞娜, 曹浩, 等. 不同植物源的黄烷酮-3-羟化酶的催化特性 [J]. 微生物学通报, 2022, 49(3): 875-887. |
| Zhang XY, Dong LN, Cao H, et al. Catalytic properties of flavanone-3-hydroxylase from different plants [J]. Microbiol China, 2022, 49(3): 875-887. | |
| [29] | Li CC, Liu SH, Yao XH, et al. PnF3H, a flavanone 3-hydroxylase from the Antarctic moss Pohlia nutans, confers tolerance to salt stress and ABA treatment in transgenic Arabidopsis [J]. Plant Growth Regul, 2017, 83(3): 489-500. |
| [30] | Mao YY, Luo JJ, Cai ZP. Biosynthesis and regulatory mechanisms of plant flavonoids: a review [J]. Plants, 2025, 14(12): 1847. |
| [31] | Quattrocchio F, Wing JF, Va K, et al. Analysis of bHLH and MYB domain proteins: species-specific regulatory differences are caused by divergent evolution of target anthocyanin genes [J]. Plant J, 1998, 13(4): 475-488. |
| [1] | 刘梦茹, 夏琳, 刘瑞霞, 闫新可, 杨军, 张菁华, 武明珠. 烟草多酚氧化酶NtPPO1-2基因克隆及功能分析[J]. 生物技术通报, 2026, 42(2): 218-227. |
| [2] | 倪莹, 李雷, 汪进萱, 马波, 孟昕, 冷平生, 吴静, 胡增辉. 紫丁香So4CL的克隆及功能分析[J]. 生物技术通报, 2026, 42(1): 139-149. |
| [3] | 徐小萍, 杨成龙, 和兴, 郭文杰, 吴健, 方少忠. 百合LoAPS1克隆及其在休眠解除过程的功能分析[J]. 生物技术通报, 2025, 41(9): 195-206. |
| [4] | 董向向, 缪百灵, 许贺娟, 陈娟娟, 李亮杰, 龚守富, 朱庆松. 森林草莓FveBBX20基因的生物信息学分析及开花调控功能[J]. 生物技术通报, 2025, 41(9): 115-123. |
| [5] | 刘佳丽, 宋经荣, 赵文宇, 张馨元, 赵子洋, 曹一博, 张凌云. 蓝莓R2R3-MYB基因鉴定及类黄酮调控基因表达分析[J]. 生物技术通报, 2025, 41(9): 124-138. |
| [6] | 张永艳, 郭思健, 李晶, 郝思怡, 李瑞得, 刘嘉鹏, 程春振. 蓝莓花青素相关VcGSTF19基因的克隆及功能研究[J]. 生物技术通报, 2025, 41(9): 139-146. |
| [7] | 李玉珍, 李梦丹, 张蔚, 彭婷. 基于月季扩展蛋白基因家族鉴定的野蔷薇RmEXPB2基因功能研究[J]. 生物技术通报, 2025, 41(9): 182-194. |
| [8] | 李开杰, 吴瑶, 李丹丹. 红花CtbHLH128基因克隆及调控干旱胁迫应答功能研究[J]. 生物技术通报, 2025, 41(8): 234-241. |
| [9] | 康琴, 汪霞, 谌明洋, 徐静天, 陈诗兰, 廖平杨, 许文志, 吴卫, 徐东北. 薄荷UV-B受体基因McUVR8的克隆与表达分析[J]. 生物技术通报, 2025, 41(8): 255-266. |
| [10] | 王芳, 乔帅, 宋伟, 崔鹏娟, 廖安忠, 谭文芳, 杨松涛. 甘薯IbNRT2基因家族全基因组鉴定和表达分析[J]. 生物技术通报, 2025, 41(7): 193-204. |
| [11] | 魏雨佳, 李岩, 康语涵, 弓晓楠, 杜敏, 涂岚, 石鹏, 于子涵, 孙彦, 张昆. 白颖苔草CrMYB4基因的克隆和表达分析[J]. 生物技术通报, 2025, 41(7): 248-260. |
| [12] | 许慧珍, SHANTWANA Ghimire, RAJU Kharel, 岳云, 司怀军, 唐勋. 马铃薯SUMO E3连接酶基因家族分析及StSIZ1基因的克隆与表达模式分析[J]. 生物技术通报, 2025, 41(6): 119-129. |
| [13] | 裴景琪, 赵梦然, 黄晨阳, 邬向丽. 一个影响糙皮侧耳生长发育的功能基因的发现与验证[J]. 生物技术通报, 2025, 41(6): 327-334. |
| [14] | 刘鑫, 王嘉雯, 李进伟, 牟策, 杨盼盼, 明军, 徐雷锋. 兰州百合三个LdBBXs基因的克隆与表达分析[J]. 生物技术通报, 2025, 41(5): 186-196. |
| [15] | 彭绍智, 王登科, 张祥, 戴雄泽, 徐昊, 邹学校. 辣椒CaFD1基因克隆、表达特征及功能验证[J]. 生物技术通报, 2025, 41(5): 153-164. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||