








生物技术通报 ›› 2022, Vol. 38 ›› Issue (2): 205-217.doi: 10.13560/j.cnki.biotech.bull.1985.2021-0447
邱益彬1,3( ), 马艳琴2, 沙媛媛2, 朱逸凡2, 苏二正1, 雷鹏2, 李莎2, 徐虹2
), 马艳琴2, 沙媛媛2, 朱逸凡2, 苏二正1, 雷鹏2, 李莎2, 徐虹2
收稿日期:2021-04-07
出版日期:2022-02-26
发布日期:2022-03-09
作者简介:邱益彬,男,博士,讲师,研究方向:微生物合成生物学与代谢工程;E-mail: 基金资助:
QIU Yi-bin1,3( ), MA Yan-qin2, SHA Yuan-yuan2, ZHU Yi-fan2, SU Er-zheng1, LEI Peng2, LI Sha2, XU Hong2
), MA Yan-qin2, SHA Yuan-yuan2, ZHU Yi-fan2, SU Er-zheng1, LEI Peng2, LI Sha2, XU Hong2
Received:2021-04-07
Published:2022-02-26
Online:2022-03-09
摘要:
解淀粉芽孢杆菌是FDA认定的安全级(generally recognized as safe,GRAS)菌株,在工业酶制剂、高分子聚合物、大宗化学品、绿色生物农药等方面的生产具有突出的优势。近年来,随着解淀粉芽孢杆菌的分子遗传操作技术越来越成熟,对利用该菌开发成微生物发酵平台化菌株用于合成生物学制造领域提出了更迫切的需求。文中围绕解淀粉芽孢杆菌的遗传操作工具、代谢改造应用和未来发展前景等方面进行了详细综述,为进一步推动解淀粉芽孢杆菌合成生物学技术的创新与发展提供借鉴和 参考。
邱益彬, 马艳琴, 沙媛媛, 朱逸凡, 苏二正, 雷鹏, 李莎, 徐虹. 解淀粉芽孢杆菌分子遗传操作及其应用研究进展[J]. 生物技术通报, 2022, 38(2): 205-217.
QIU Yi-bin, MA Yan-qin, SHA Yuan-yuan, ZHU Yi-fan, SU Er-zheng, LEI Peng, LI Sha, XU Hong. Research Progress in Molecular Genetic Manipulation Technology of Bacillus amyloliquefaciens and Its Application[J]. Biotechnology Bulletin, 2022, 38(2): 205-217.
| Plasmid | Promoter | Terminator | Replicon | Reference |
|---|---|---|---|---|
| pUBXC | PxylA | t0 | repB | [ |
| pDR | PHpaII | Tfd | repF | [ |
| pMA5 | PHpaII | Tfd | repB | |
| pNX01 | P43 | Tamy | Ori-rep(p2Sip) | [ |
| pWH1520 | PxylA | pBC16 ori | [ | |
| pKSV7 | P43 | Temperature-sensitive replication from pE194ts | ||
| pNW33N | ctaB、qcr、qox、resD promoter | repB | [ | |
| pLY-3 | Promoter from amyE of B. subtilis 168 | [ | ||
| pLakr | PQ | [ | ||
| pHT01 | Pgrac | Theta replicon | [ |
表1 B. amyloliquefaciens中所使用的合成生物学基因元件
Table 1 Synthetic biological genetic elements used in B. amyloliquefaciens
| Plasmid | Promoter | Terminator | Replicon | Reference |
|---|---|---|---|---|
| pUBXC | PxylA | t0 | repB | [ |
| pDR | PHpaII | Tfd | repF | [ |
| pMA5 | PHpaII | Tfd | repB | |
| pNX01 | P43 | Tamy | Ori-rep(p2Sip) | [ |
| pWH1520 | PxylA | pBC16 ori | [ | |
| pKSV7 | P43 | Temperature-sensitive replication from pE194ts | ||
| pNW33N | ctaB、qcr、qox、resD promoter | repB | [ | |
| pLY-3 | Promoter from amyE of B. subtilis 168 | [ | ||
| pLakr | PQ | [ | ||
| pHT01 | Pgrac | Theta replicon | [ |
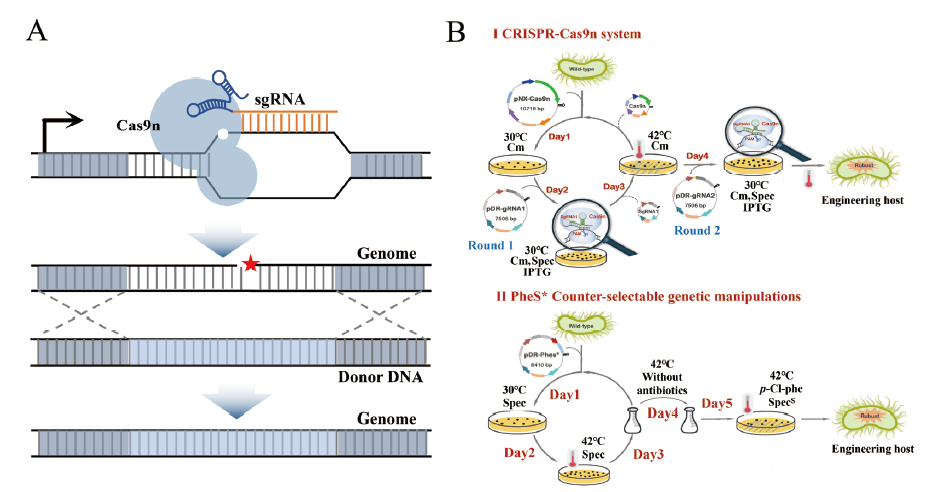
图2 CRISPR-Cas9n基因编辑技术在B. amyloliquefaciens中的操作流程示意图 A:CRISPR-Cas9n系统的工作原理;B:CRISPR-Cas9n与传统同源重组技术操作流程的对比
Fig. 2 A schematic diagram of operation while CRISPR-Cas9n gene editing technology in B. amyloliquefaciens A:The gene editing mechanism of CRISPR-Cas9n. B:Comparison of CRISPR-Cas9n and conventional homologous recombination techniques

图3 CRISPR-dCas9n基因沉默技术在B. amyloliquefaciens中的构建 A:CRISPR-dCas9系统的工作原理;B:CRISPR-dCas9双质粒系统组成;C:靶向egfp表达框的sgRNA设计;不同sgRNA介导的egfp基因的转录水平;不同sgRNA调控下荧光共聚焦显微镜观察
Fig. 3 Construction of CRISPR-Cas9n gene silencing technology in B. amyloliquefaciens A:The catalytic mechanism of CRISPR-dCas9. B:Schematic representation of the CRISPR-dCas9 double plasmid system. C:Design of sgRNA targeting to egfp expression region. Transcription level of different sgRNA-mediated egfp genes,observed by fluorescence confocal microscopy under different sgRNA regulation
| Strain | Transformation method | Concrete operation | Transformation plasmid | Transformation efficiency /(CFU·μg-1) |
|---|---|---|---|---|
| B. amyloliquefaciens TA208[ | Electroproration | Adding DL-threonine or glycine to weaken the cell wall in hypertonic medium | pUB110 | (1.13 ± 0.34)×107 |
| Adding DL-threonine or glycine to weaken the cell wall in hypertonic medium;heat shock treatment | pHCMC02 | (8.94 ± 0.77)×105 | ||
| B. amyloliquefaciens NB[ | Electroproration | Demethylation | pDR | 4.94 ± 0.42)×104 |
| Demethylation | pMA5 | (6.15 ± 0.19)×103 | ||
| B. amyloliquefaciens LL3[ | Electroproration | Demethylation | pWH1520 | 7.6×102 |
| B. amyloliquefaciens[ | Chemical transformation | Overexpressing ComK regulator to induce the formation of competent | pUBXC | [(129 ± 20.6)- (1.7 ± 0.1)]×105 |
| PCR fragment-mediated knockout | [(3.2 ± 0.76)- (3.5 ± 0.42)]×104 | |||
| B. amyloliquefaciens[ | Transformation of protoplasts | Mediating by polyethylene glycol | pUB110 | (2-4)×105 |
表2 不同方法转化B. amyloliquefaciens转化效率的研究
Table 2 Study on the transformation efficiency of B. amyloliquefaciens by different methods
| Strain | Transformation method | Concrete operation | Transformation plasmid | Transformation efficiency /(CFU·μg-1) |
|---|---|---|---|---|
| B. amyloliquefaciens TA208[ | Electroproration | Adding DL-threonine or glycine to weaken the cell wall in hypertonic medium | pUB110 | (1.13 ± 0.34)×107 |
| Adding DL-threonine or glycine to weaken the cell wall in hypertonic medium;heat shock treatment | pHCMC02 | (8.94 ± 0.77)×105 | ||
| B. amyloliquefaciens NB[ | Electroproration | Demethylation | pDR | 4.94 ± 0.42)×104 |
| Demethylation | pMA5 | (6.15 ± 0.19)×103 | ||
| B. amyloliquefaciens LL3[ | Electroproration | Demethylation | pWH1520 | 7.6×102 |
| B. amyloliquefaciens[ | Chemical transformation | Overexpressing ComK regulator to induce the formation of competent | pUBXC | [(129 ± 20.6)- (1.7 ± 0.1)]×105 |
| PCR fragment-mediated knockout | [(3.2 ± 0.76)- (3.5 ± 0.42)]×104 | |||
| B. amyloliquefaciens[ | Transformation of protoplasts | Mediating by polyethylene glycol | pUB110 | (2-4)×105 |
| Products | Strategies | Yield | Reference | |
|---|---|---|---|---|
| γ-PGA | Knock out genes involved in by-products,degrading enzymes and optimize the endogenous glutamate synthesis | 20.3 g/L | [ | |
| Modification of substrate inulin utilization,sugar metabolism and by-product pathways | 32.14 g/L | [ | ||
| Using the CRISPRi system’s multiple sgRNA combination strategy to control the expression of degrading enzymes,that achieve multiple molecular weights of γ-PGA | Different molecular weights of γ-PGA High(>800 kD)、middle(400-600 kD)、low(50-100 kD);25-27 g/L | [ | ||
| To control the stereochemical configuration | Low-molecular-weight(<10 kD)γ-PGA;28.35 g/L | [ | ||
| EPS | Optimization of fermentation conditions | 4.46 g/L | [ | |
| Levan | Removing six protease-related genes,the biofilm matrix protein TasA and γ-PGA synthase genes | 31.1 g/L | [ | |
| Optimization of levansucrase expression based on promoters and signal peptides | 102 g/L | [ | ||
| Enzymes production | α-amylase | Optimization of the promoters and host | 2714 U/mL | [ |
| Pullulanase | Cloning and expression optimization | 2.8 ASPU/mL | [ | |
| Keratinase | Cloning and expression optimization | 1 361.54 U/mL | [ | |
| lytic polysaccharide monooxygenase | Establishing a high-throughput screening system for expression and secretion | 12.17 U/g | [ | |
| Nucleotides | Guanosine | Relieving the purine operon of the purine biosynthetic pathway and optimizing the energy of the respiratory chain | 19 g/L | [ |
| Inosine | Protoplast fusion | 6 g/L | [ | |
| Antimicrobial lipopeptides | Surfactin | Using nano iron particles to improve the permeability of cell membranes | 7.15 g/L | [ |
| Iturin A | Overexpression of the iturin A biosynthesis genes | 37.35 mg/L | [ | |
| Fengycin | Genome shuffling | 450.51 mg/L | [ | |
| Bacillomycin D | Knocking out of the gene rapC | (360.8±30.7)mg/L | [ | |
| Bulk chemicals | Acetoin | Compound mutagenesis | 85.2 g/L | [ |
| 2, 3-butanediol | Overexpression of glyceraldehyde-3-phosphate dehydrogenase and 2, 3-butanediol dehydrogenase | 132.9 g/L | [ | |
表3 基于B. amyloliquefaciens合成的生物基产品汇总
Table 3 Summary of the synthesis of biobased products based on B. amyloliquefaciens species
| Products | Strategies | Yield | Reference | |
|---|---|---|---|---|
| γ-PGA | Knock out genes involved in by-products,degrading enzymes and optimize the endogenous glutamate synthesis | 20.3 g/L | [ | |
| Modification of substrate inulin utilization,sugar metabolism and by-product pathways | 32.14 g/L | [ | ||
| Using the CRISPRi system’s multiple sgRNA combination strategy to control the expression of degrading enzymes,that achieve multiple molecular weights of γ-PGA | Different molecular weights of γ-PGA High(>800 kD)、middle(400-600 kD)、low(50-100 kD);25-27 g/L | [ | ||
| To control the stereochemical configuration | Low-molecular-weight(<10 kD)γ-PGA;28.35 g/L | [ | ||
| EPS | Optimization of fermentation conditions | 4.46 g/L | [ | |
| Levan | Removing six protease-related genes,the biofilm matrix protein TasA and γ-PGA synthase genes | 31.1 g/L | [ | |
| Optimization of levansucrase expression based on promoters and signal peptides | 102 g/L | [ | ||
| Enzymes production | α-amylase | Optimization of the promoters and host | 2714 U/mL | [ |
| Pullulanase | Cloning and expression optimization | 2.8 ASPU/mL | [ | |
| Keratinase | Cloning and expression optimization | 1 361.54 U/mL | [ | |
| lytic polysaccharide monooxygenase | Establishing a high-throughput screening system for expression and secretion | 12.17 U/g | [ | |
| Nucleotides | Guanosine | Relieving the purine operon of the purine biosynthetic pathway and optimizing the energy of the respiratory chain | 19 g/L | [ |
| Inosine | Protoplast fusion | 6 g/L | [ | |
| Antimicrobial lipopeptides | Surfactin | Using nano iron particles to improve the permeability of cell membranes | 7.15 g/L | [ |
| Iturin A | Overexpression of the iturin A biosynthesis genes | 37.35 mg/L | [ | |
| Fengycin | Genome shuffling | 450.51 mg/L | [ | |
| Bacillomycin D | Knocking out of the gene rapC | (360.8±30.7)mg/L | [ | |
| Bulk chemicals | Acetoin | Compound mutagenesis | 85.2 g/L | [ |
| 2, 3-butanediol | Overexpression of glyceraldehyde-3-phosphate dehydrogenase and 2, 3-butanediol dehydrogenase | 132.9 g/L | [ | |
| [1] | 张娟, 杨彩梅, 曹广添, 等. 解淀粉芽孢杆菌及其作为益生菌的应用[J]. 动物营养学报, 2014, 26(4):863-867. |
| Zhang J, Yang CM, Cao GT, et al. Bacillus amyloliquefaciens and its application as a probiotic[J]. Chin J Animal Nutr, 2014, 26(4):863-867. | |
| [2] |
Chen XT, Ji JB, Liu YC, et al. Artificial induction of genetic competence in Bacillus amyloliquefaciens isolates[J]. Biotechnol Lett, 2016, 38(12):2109-2117.
doi: 10.1007/s10529-016-2194-0 URL |
| [3] |
Qiu Y, Zhu Y, Zhang Y, et al. Characterization of a Regulator pgsR on Endogenous Plasmid p2Sip and Its Complementation for poly(γ-glutamic acid)accumulation in Bacillus amyloliquefaciens[J]. J Agric Food Chem, 2019, 67(13):3711-3722.
doi: 10.1021/acs.jafc.9b00332 URL |
| [4] |
Sha Y, Zhang Y, Qiu Y, et al. Efficient biosynjournal of low-molecular-weight poly-γ-glutamic acid by stable overexpression of PgdS hydrolase in Bacillus amyloliquefaciens NB[J]. J Agric Food Chem, 2019, 67(1):282-290.
doi: 10.1021/acs.jafc.8b05485 URL |
| [5] |
Zhang W, Xie H, He Y, et al. Chromosome integration of the Vitreoscilla hemoglobin gene(vgb)mediated by temperature-sensitive plasmid enhances γ-PGA production in Bacillus amyloliquefaciens[J]. FEMS Microbiol Lett, 2013, 343(2):127-134.
doi: 10.1111/1574-6968.12139 pmid: 23521121 |
| [6] | Zhou X, Zhang N, Xia LM, et al. ResDE two-component regulatory system mediates oxygen limitation-induced biofilm formation by Bacillus amyloliquefaciens SQR9[J]. Appl Environ Microbiol, 2018, 84(8):e02744-17. |
| [7] |
Guo X, Chai CC, An YJ, et al. Rational design of signal peptides for improved MtC1LPMO production in Bacillus amyloliquefaciens[J]. Int J Biol Macromol, 2021, 175:262-269.
doi: 10.1016/j.ijbiomac.2021.02.034 URL |
| [8] | 陈玉娟, 沈微, 陈献忠, 等. 解淀粉芽孢杆菌β-1, 3-1, 4-葡聚糖酶的高效表达[J]. 生物技术, 2011, 21(2):22-26. |
| Chen YJ, Shen W, Chen XZ, et al. Overexpression of Bacillus amyloliquefaciens β-1, 3-1, 4-glucanase[J]. Biotechnology, 2011, 21(2):22-26. | |
| [9] |
Gu Y, Zheng J, Feng J, et al. Improvement of levan production in Bacillus amyloliquefaciens through metabolic optimization of regulatory elements[J]. Appl Microbiol Biotechnol, 2017, 101(10):4163-4174.
doi: 10.1007/s00253-017-8171-2 URL |
| [10] |
Qiao JQ, Tian DW, Huo R, et al. Functional analysis and application of the cryptic plasmid pBSG3 harboring the RapQ-PhrQ system in Bacillus amyloliquefaciens B3[J]. Plasmid, 2011, 65(2):141-149.
doi: 10.1016/j.plasmid.2010.11.008 URL |
| [11] |
Liao Y, Huang L, Wang B, et al. The global transcriptional landscape of Bacillus amyloliquefaciens XH7 and high-throughput screening of strong promoters based on RNA-seq data[J]. Gene, 2015, 571(2):252-262.
doi: 10.1016/j.gene.2015.06.066 URL |
| [12] |
Liao Y, Wang B, Ye Y, et al. Determination and optimization of a strong promoter element from Bacillus amyloliquefaciens by using a promoter probe vector[J]. Biotechnol Lett, 2018, 40(1):119-126.
doi: 10.1007/s10529-017-2449-4 URL |
| [13] |
Qiu YB, Zhu YF, Sha YY, et al. Development of a robust Bacillus amyloliquefaciens cell factory for efficient poly(γ-glutamic acid)production from Jerusalem artichoke[J]. ACS Sustainable Chem Eng, 2020, 8(26):9763-9774.
doi: 10.1021/acssuschemeng.0c02107 URL |
| [14] |
Novikov AA, Borukhov SI, Strongin AY. Bacillusamyloliquefaciens α -amylase signal sequence fused in frame with human proinsulin is properly processed by Bacillus subtilis cells[J]. Biochem Biophys Res Commun, 1990, 169(1):297-301.
doi: 10.1016/0006-291X(90)91467-7 URL |
| [15] |
Zakataeva NP, Nikitina OV, Gronskiy SV, et al. A simple method to introduce marker-free genetic modifications into the chromosome of naturally nontransformable Bacillus amyloliquefaciens strains[J]. Appl Microbiol Biotechnol, 2010, 85(4):1201-1209.
doi: 10.1007/s00253-009-2276-1 pmid: 19820923 |
| [16] | 杨慧林, 王坤, 廖瑜玲, 等. 解淀粉芽孢杆菌ptsGHI基因的敲除及缺陷株生长特性[J]. 华南理工大学学报:自然科学版, 2012, 40(8):95-100. |
| Yang HL, Wang K, Liao YL, et al. Knockout of ptsGHI gene of Bacillus amyloliquefaciens and growth characteristics of corresponding deficient strain[J]. J South China Univ Technol:Nat Sci Ed, 2012, 40(8):95-100. | |
| [17] |
Wu L, Wu H, Chen L, et al. Bacilysin overproduction in Bacillus amyloliquefaciens FZB42 markerless derivative strains FZBREP and FZBSPA enhances antibacterial activity[J]. Appl Microbiol Biotechnol, 2015, 99(10):4255-4263.
doi: 10.1007/s00253-014-6251-0 URL |
| [18] |
Zhang W, Gao WX, Feng J, et al. A markerless gene replacement method for B. amyloliquefaciens LL3 and its use in genome reduction and improvement of poly-γ-glutamic acid production[J]. Appl Microbiol Biotechnol, 2014, 98(21):8963-8973.
doi: 10.1007/s00253-014-5824-2 pmid: 24859524 |
| [19] |
Zhou C, Shi L, Ye B, et al. pheS *, an effective host-genotype-independent counter-selectable marker for marker-free chromosome deletion in Bacillus amyloliquefaciens[J]. Appl Microbiol Biotechnol, 2017, 101(1):217-227.
doi: 10.1007/s00253-016-7906-9 URL |
| [20] |
Hsu PD, Lander ES, Zhang F. Development and applications of CRISPR-Cas9 for genome engineering[J]. Cell, 2014, 157(6):1262-1278.
doi: 10.1016/j.cell.2014.05.010 URL |
| [21] |
Sha YY, Qiu YB, Zhu YF, et al. CRISPRi-based dynamic regulation of hydrolase for the synjournal of poly-γ-glutamic acid with variable molecular weights[J]. ACS Synth Biol, 2020, 9(9):2450-2459.
doi: 10.1021/acssynbio.0c00207 URL |
| [22] |
Zhang GQ, Bao P, Zhang Y, et al. Enhancing electro-transformation competency of recalcitrant Bacillus amyloliquefaciens by combining cell-wall weakening and cell-membrane fluidity disturbing[J]. Anal Biochem, 2011, 409(1):130-137.
doi: 10.1016/j.ab.2010.10.013 URL |
| [23] |
Vehmaanperä J. Transformation of Bacillus amyloliquefaciens protoplasts with plasmid DNA[J]. FEMS Microbiol Lett, 1988, 49(1):101-105.
doi: 10.1111/fml.1988.49.issue-1 URL |
| [24] |
Xue GP, Johnson JS, Dalrymple BP. High osmolarity improves the electro-transformation efficiency of the gram-positive bacteria Bacillus subtilis and Bacillus licheniformis[J]. J Microbiol Methods, 1999, 34(3):183-191.
doi: 10.1016/S0167-7012(98)00087-6 URL |
| [25] | Akamatsu T, Sekiguchi J. Transformation of Bacillus protoplasts by plasmid pTP4 DNA[J]. Agricultural and Biological Chemistry, 1982, 46(6):1617-1621. |
| [26] |
Wang P, Wang P, Tian J, et al. A new strategy to express the extracellular α-amylase from Pyrococcus furiosus in Bacillus amyloliquefaciens[J]. Sci Rep, 2016, 6:22229.
doi: 10.1038/srep22229 URL |
| [27] | 孙娟娟, 沈微, 石贵阳, 等. 普鲁兰酶基因在解淀粉芽孢杆菌BF7658菌株中的分泌表达[J]. 工业微生物, 2012, 42(2):18-22. |
| Sun JJ, Shen W, Shi GY, et al. Secreted expression of the pullulanase in Bacillus amyloliquefaciens BF7658[J]. Ind Microbiol, 2012, 42(2):18-22. | |
| [28] | 辛青龙, 刘业学, 何光明, 等. 地衣芽孢杆菌角蛋白酶kerA基因的异源表达及性能研究[J]. 中国皮革, 2018, 47(12):28-34. |
| Xin QL, Liu YX, He GM, et al. Overexpression of Bacillus licheniformis keratinase gene(kerA)and its performance[J]. China Leather, 2018, 47(12):28-34. | |
| [29] | 王慧. 解淀粉芽孢杆菌Bacillus amyloliquefaciens K11高效表达体系的建立及其高效表达元件的优化[D]. 北京:中国农业科学院, 2018. |
| Wang H. Establishment of the highly effective expression system in Bacillus amyloliquefaciens K11 and optimization of its efficient expression elements[D]. Beijing:Chinese Academy of Agricultural Sciences, 2018. | |
| [30] | 刘刚, 王清明, 陈惠鹏. 非经典的蛋白质分泌途径[J]. 生物技术通讯, 2005, 16(1):53-55. |
| Liu G, Wang QM, Chen HP. Non-classical secretory pathway[J]. Lett Biotechnol, 2005, 16(1):53-55. | |
| [31] | 徐虹, 冯小海, 徐得磊, 等. 聚氨基酸功能高分子的发展状况与应用前景[J]. 生物产业技术, 2017(6):92-99. |
| Xu H, Feng XH, Xu DL, et al. Recent developments and potential applications of poly(amino acid)-based functional polymers[J]. Biotechnol Bus, 2017(6):92-99. | |
| [32] |
Qiu Y, Sha Y, Zhang Y, et al. Development of Jerusalem artichoke resource for efficient one-step fermentation of poly-(γ-glutamic acid)using a novel strain Bacillus amyloliquefaciens NX-2S[J]. Bioresour Technol, 2017, 239:197-203.
doi: 10.1016/j.biortech.2017.05.005 URL |
| [33] |
Qiu YB, Zhang YT, Zhu YF, et al. Improving poly-(γ-glutamic acid)production from a glutamic acid-independent strain from inulin substrate by consolidated bioprocessing[J]. Bioprocess Biosyst Eng, 2019, 42(10):1711-1720.
doi: 10.1007/s00449-019-02167-w URL |
| [34] |
Sha Y, Huang Y, Zhu Y, et al. Efficient biosynjournal of low-molecular-weight poly-γ-glutamic acid based on stereochemistry regulation in Bacillus amyloliquefaciens[J]. ACS Synth Biol, 2020, 9(6):1395-1405.
doi: 10.1021/acssynbio.0c00080 URL |
| [35] |
Zhao W, Zhang J, Jiang YY, et al. Characterization and antioxidant activity of the exopolysaccharide produced by Bacillus amyloliquefaciens GSBa-1[J]. J Microbiol Biotechnol, 2018, 28(8):1282-1292.
doi: 10.4014/jmb.1801.01012 URL |
| [36] |
Han YZ, Liu EQ, Liu LS, et al. Rheological, emulsifying and thermostability properties of two exopolysaccharides produced by Bacillus amyloliquefaciens LPL061[J]. Carbohydr Polym, 2015, 115:230-237.
doi: 10.1016/j.carbpol.2014.08.044 URL |
| [37] |
Chen YT, Yuan Q, Shan LT, et al. Antitumor activity of bacterial exopolysaccharides from the endophyte Bacillus amyloliquefaciens sp. isolated from Ophiopogon japonicus.[J]. Oncol Lett, 2013, 5(6):1787-1792.
doi: 10.3892/ol.2013.1284 URL |
| [38] |
Yang H, Deng J, Yuan Y, et al. Two novel exopolysaccharides from Bacillus amyloliquefaciens C-1:antioxidation and effect on oxidative stress[J]. Curr Microbiol, 2015, 70(2):298-306.
doi: 10.1007/s00284-014-0717-2 URL |
| [39] |
Cai G, Liu Y, Li X, et al. New levan-type exopolysaccharide from Bacillus amyloliquefaciens as an antiadhesive agent against enterotoxigenic Escherichia coli[J]. J Agric Food Chem, 2019, 67(28):8029-8034.
doi: 10.1021/acs.jafc.9b03234 URL |
| [40] |
Feng J, Gu Y, Quan Y, et al. Recruiting a new strategy to improve levan production in Bacillus amyloliquefaciens[J]. Sci Rep, 2015, 5:13814.
doi: 10.1038/srep13814 pmid: 26347185 |
| [41] |
Phengnoi P, Charoenwongpaiboon T, Wangpaiboon K, et al. Levansucrase from Bacillus amyloliquefaciens KK9 and its Y237S variant producing the high bioactive levan-type fructooligosaccharides[J]. Biomolecules, 2020, 10(5):692.
doi: 10.3390/biom10050692 URL |
| [42] | 吴飞, 史建明, 谢希贤, 等. 鸟苷产生菌解淀粉芽孢杆菌的选育[J]. 天津科技大学学报, 2010, 25(2):1-4. |
| Wu F, Shi JM, Xie XX, et al. Breeding of strain Bacillus amyloliquefaciens for guanosine-producing[J]. J Tianjin Univ Sci Technol, 2010, 25(2):1-4. | |
| [43] | 何逵夫, 马跃超, 杜姗姗, 等. 解淀粉芽胞杆菌关键酶基因过表达对鸟苷积累的影响[J]. 微生物学报, 2012, 52(6):718-725. |
| He KF, Ma YC, Du SS, et al. Effects of overexpression of key enzyme genes on guanosine accumulation in Bacillus amyloliquefaciens[J]. Acta Microbiol Sin, 2012, 52(6):718-725. | |
| [44] |
Liao YL, Ye YR, Wang B, et al. Optimization of the purine operon and energy generation in Bacillus amyloliquefaciens for guanosine production[J]. Biotechnol Lett, 2017, 39(11):1675-1682.
doi: 10.1007/s10529-017-2412-4 URL |
| [45] |
Wong JH, Hao J, Cao Z, et al. An antifungal protein from Bacillus amyloliquefaciens[J]. J Appl Microbiol, 2008, 105(6):1888-1898.
doi: 10.1111/j.1365-2672.2008.03917.x pmid: 19120637 |
| [46] | 吴江, 吴梧桐, 黄为一, 等. 通过原生质体融合选育直接利用淀粉的肌苷产生菌[J]. 药物生物技术, 1995, 2(2):7-10 |
| Wu J, Wu WT, Huang WY, et al. Selection of inosine producing fusants by protoplast fusion[J]. Pharm Biotechnol, 1995, 2(2):7-10 | |
| [47] |
Wibisana A, Sumaryono W, Mirawati Sudiro T, et al. Optimization of surfactin production by Bacillus amyloliquefaciens MD4-12 using response surface methodology[J]. Microbiol Indones, 2015, 9(3):120-128.
doi: 10.5454/mi.9.3.4 URL |
| [48] | 周泽宇, 张婉茹, 张柔萱, 等. 代谢工程改造Bacillus amyloliquefaciens提高Surfactin产量[J]. 南开大学学报:自然科学版, 2018, 51(5):18-26. |
| Zhou ZY, Zhang WR, Zhang RX, et al. Metabolic engineering of Bacillus amyloliquefaciens to improve surfactin production[J]. Acta Sci Nat Univ Nankaiensis, 2018, 51(5):18-26. | |
| [49] |
Yang N, Wu Q, Xu Y. Fe nanoparticles enhanced surfactin production in Bacillus amyloliquefaciens[J]. ACS Omega, 2020, 5(12):6321-6329.
doi: 10.1021/acsomega.9b03648 pmid: 32258866 |
| [50] |
Dang Y, Zhao F, Liu X, et al. Enhanced production of antifungal lipopeptide iturin A by Bacillus amyloliquefaciens LL3 through metabolic engineering and culture conditions optimization[J]. Microb Cell Fact, 2019, 18(1):68.
doi: 10.1186/s12934-019-1121-1 URL |
| [51] |
Sang-Cheol L, Kim SH, Park IH, et al. Isolation, purification, and characterization of novel fengycin S from Bacillus amyloliquefaciens LSC04 degrading-crude oil[J]. Biotechnol Bioprocess Eng, 2010, 15(2):246-253.
doi: 10.1007/s12257-009-0037-8 URL |
| [52] |
Zhao JF, Zhang C, Lu J, et al. Enhancement of fengycin production in Bacillus amyloliquefaciens by genome shuffling and relative gene expression analysis using RT-PCR[J]. Can J Microbiol, 2016, 62(5):431-436.
doi: 10.1139/cjm-2015-0734 URL |
| [53] | Sun HG, Lu FX, Zhang C, et al. Improvement of fengycin production by Bacillus amyloliquefaciens via promoter replacement at the fengycin operon with the P59 and PrepU promoters[J]. Journal of Pure and Applied Microbiology, 2014, 8(2):1071-1077. |
| [54] | Sun J, Liu Y, Lin F, et al. CodY, ComA, DegU and Spo0A controlling lipopeptides biosynjournal in Bacillus amyloliquefaciens fmb[J][J]. J Appl Microbiol, 2021:1364-5072. |
| [55] |
Zhang YJ, Li SB, Liu LM, et al. Acetoin production enhanced by manipulating carbon flux in a newly isolated Bacillus amyloliquefaciens[J]. Bioresour Technol, 2013, 130:256-260.
doi: 10.1016/j.biortech.2012.10.036 URL |
| [56] |
Luo QL, Wu J, Wu MC. Enhanced acetoin production by Bacillus amyloliquefaciens through improved acetoin tolerance[J]. Process Biochem, 2014, 49(8):1223-1230.
doi: 10.1016/j.procbio.2014.05.005 URL |
| [57] | 王诗卉, 罗秋玲, 刘佳, 等. 高产3-羟基丁酮解淀粉芽孢杆菌的选育及发酵优化[J]. 生物工程学报, 2018, 34(5):803-811. |
| Wang SH, Luo QL, Liu J, et al. Mutation and fermentation optimization of Bacillus amyloliquefaciens for acetoin production[J]. Chin J Biotechnol, 2018, 34(5):803-811. | |
| [58] |
Yang TW, Rao ZM, Zhang X, et al. Production of 2, 3-butanediol from glucose by GRAS microorganism Bacillus amyloliquefaciens[J]. J Basic Microbiol, 2011, 51(6):650-658.
doi: 10.1002/jobm.v51.6 URL |
| [59] |
Yang T, Rao Z, Zhang X, et al. Improved production of 2, 3-butanediol in Bacillus amyloliquefaciens by over-expression of glyceraldehyde-3-phosphate dehydrogenase and 2, 3-butanediol dehydrogenase[J]. PLoS One, 2013, 8(10):e76149.
doi: 10.1371/journal.pone.0076149 URL |
| [60] |
Feng J, Gu YY, Quan YF, et al. Improved poly-γ-glutamic acid production in Bacillus amyloliquefaciens by modular pathway engineering[J]. Metab Eng, 2015, 32:106-115.
doi: S1096-7176(15)00121-4 pmid: 26410449 |
| [61] | 李彦岩, 张彩, 范熠, 等. 一株解淀粉芽孢杆菌产糖条件的优化[J]. 食品科学, 2013, 34(7):185-189. |
| Li YY, Zhang C, Fan Y, et al. Optimization of fermentation conditions for exopolysaccharide production by Bacillus amyloliquefaciens LPL061[J]. Food Sci, 2013, 34(7):185-189. | |
| [62] |
Sun J, Qian S, Lu J, et al. Knockout of rapC improves the bacillomycin D yield based on de novo genome sequencing of Bacillus amyloliquefaciens fmb[J]. J Agric Food Chem, 2018, 66(17):4422-4430.
doi: 10.1021/acs.jafc.8b00418 URL |
| [1] | 薛宁, 王瑾, 李世新, 刘叶, 程海娇, 张玥, 毛雨丰, 王猛. 多基因同步调控结合高通量筛选构建高产L-苯丙氨酸的谷氨酸棒杆菌工程菌株[J]. 生物技术通报, 2023, 39(9): 268-280. |
| [2] | 程亚楠, 张文聪, 周圆, 孙雪, 李玉, 李庆刚. 乳酸乳球菌生产2'-岩藻糖基乳糖的途径构建及发酵培养基优化[J]. 生物技术通报, 2023, 39(9): 84-96. |
| [3] | 赵思佳, 王晓璐, 孙纪录, 田健, 张杰. 代谢工程改造毕赤酵母生产赤藓糖醇[J]. 生物技术通报, 2023, 39(8): 137-147. |
| [4] | 李雨真, 梅天秀, 李治文, 王淇, 李俊, 邹岳, 赵心清. 红酵母基因组和代谢工程改造研究进展[J]. 生物技术通报, 2023, 39(7): 67-79. |
| [5] | 郁慧丽, 李爱涛. 细胞色素P450酶在香精香料绿色生物合成中的应用[J]. 生物技术通报, 2023, 39(4): 24-37. |
| [6] | 蒋晶晶, 周昭旭, 杜蕙, 吕昭龙, 王春明, 郭建国, 张新瑞, 李继平. 甘肃部分地区苹果褐腐病病原分离鉴定及拮抗细菌筛选[J]. 生物技术通报, 2023, 39(10): 209-218. |
| [7] | 马艳琴, 邱益彬, 李莎, 徐虹. 透明质酸的生物合成及其代谢工程的研究进展[J]. 生物技术通报, 2022, 38(2): 252-262. |
| [8] | 蔡国磊, 陆小凯, 娄水珠, 杨海英, 杜刚. 芽孢杆菌LM基于全基因组的分类鉴定及抑菌原理的研究[J]. 生物技术通报, 2021, 37(8): 176-185. |
| [9] | 叶健文, 陈江楠, 张旭, 吴赴清, 陈国强. 动态调控:一种高效的细胞工厂工程化代谢改造策略[J]. 生物技术通报, 2020, 36(6): 1-12. |
| [10] | 陈永灿, 张建志, 司同. 酿酒酵母中基于CRISPR/dCás9的基因转录调控工具的开发与应用[J]. 生物技术通报, 2020, 36(4): 1-12. |
| [11] | 郭鹤宝, 王星, 何山文, 张晓霞. 表型特征结合基因组分析鉴定不同菌落形态Bacillus velezensis ACCC 19742[J]. 生物技术通报, 2020, 36(2): 142-148. |
| [12] | 王世伟, 王卿惠. 解淀粉芽孢杆菌相关功能机制研究进展[J]. 生物技术通报, 2020, 36(1): 150-159. |
| [13] | 刘端木, 吴怿, 刘沄, 梁志宏. 一株产毒曲霉拮抗细菌的筛选、鉴定及抑菌活性研究[J]. 生物技术通报, 2019, 35(8): 42-50. |
| [14] | 刘超, 刘洪伟, 汪步青, 赵雯雅, 王雅娜, 张丽萍. 解淀粉芽孢杆菌BA-26抗菌物质分离及对灰葡萄孢抑菌作用研究[J]. 生物技术通报, 2019, 35(7): 83-89. |
| [15] | 李丽, 严月根, 吴华明. 解淀粉芽孢杆菌M1摇瓶发酵条件优化及脱氮性能评价[J]. 生物技术通报, 2019, 35(5): 102-108. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||