








生物技术通报 ›› 2022, Vol. 38 ›› Issue (9): 116-126.doi: 10.13560/j.cnki.biotech.bull.1985.2021-1544
• 细菌耐药性专题(专题主编: 刘雅红 教授 孙坚 教授) • 上一篇 下一篇
胡雪莹1,2( ), 张越2, 郭雅杰2, 仇天雷2, 高敏2, 孙兴滨1(
), 张越2, 郭雅杰2, 仇天雷2, 高敏2, 孙兴滨1( ), 王旭明2(
), 王旭明2( )
)
收稿日期:2021-12-14
出版日期:2022-09-26
发布日期:2022-10-11
作者简介:胡雪莹,女,硕士研究生,研究方向:环境抗生素抗性;E-mail: 基金资助:
HU Xue-ying1,2( ), ZHANG Yue2, GUO Ya-jie2, QIU Tian-lei2, GAO Min2, SUN Xing-bin1(
), ZHANG Yue2, GUO Ya-jie2, QIU Tian-lei2, GAO Min2, SUN Xing-bin1( ), WANG Xu-ming2(
), WANG Xu-ming2( )
)
Received:2021-12-14
Published:2022-09-26
Online:2022-10-11
摘要:
为了揭示有机肥和化肥长期施用对农田土壤噬菌体携带的抗生素抗性基因(antibiotic resistance genes,ARGs)多样性和丰度的影响,并与土壤细菌携带的ARGs进行对比,本文将土壤抗生素抗性分为细菌和噬菌体两个部分,利用微滴数字PCR(droplet digital PCR,ddPCR)技术定量分析了土壤噬菌体和细菌DNA中25种ARGs亚型和I类整合子(intl1)的丰度。结果表明,土壤噬菌体中ARGs的检出率和总丰度以及intl1丰度均低于土壤细菌,其中噬菌体中检测到20种ARGs亚型,在不施肥、单施化肥和单施有机肥土壤的噬菌体中,目标ARGs的检出率分别为68%、72%和76%。土壤噬菌体中ARGs的总丰度在有机肥施用土壤中显著高于不施肥和化肥施用土壤(P<0.05),其中多耐药类、大环内酯-林肯酰胺-链阳性菌素B(MLSB)类和β-内酰胺类抗性基因丰度占显著优势。除了β-内酰胺类抗性基因blaTEM,噬菌体中其他ARGs亚型的丰度均显著低于细菌(P<0.05)。噬菌体与细菌携带ARGs在不同施肥处理中均存在显著正相关(P<0.05)。冗余分析结果显示,施肥可能通过改变土壤pH、重金属和营养因子水平来影响细菌和噬菌体中ARGs的赋存特征。本研究结果表明,噬菌体是除细菌之外的农田土壤另一个重要ARGs储存库,施用有机肥能同时显著提高土壤细菌和噬菌体中ARGs的多样性和丰度。
胡雪莹, 张越, 郭雅杰, 仇天雷, 高敏, 孙兴滨, 王旭明. 不同施肥处理农田土壤中噬菌体与细菌携带抗生素抗性基因的比较[J]. 生物技术通报, 2022, 38(9): 116-126.
HU Xue-ying, ZHANG Yue, GUO Ya-jie, QIU Tian-lei, GAO Min, SUN Xing-bin, WANG Xu-ming. Comparison in Antibiotic Resistance Genes Carried by Bacteriophages and Bacteria in Farmland Soil Amended with Different Fertilizers[J]. Biotechnology Bulletin, 2022, 38(9): 116-126.
| 处理Treatment | 施肥量Fertilizer amount/(kg· m-2) |
|---|---|
| 不施肥(CK) Without fertilizer | 0 |
| 单施化肥(IF) Inorganic fertilizer amendment | Urea 0.04、(NH4)2HPO4 0.05和K2SO4 0.03 |
| 单施有机肥(OF) Organic fertilizer amendment | Compost from chicken manure 2.38 |
表1 不同施肥处理的施肥量
Table 1 Fertilizer amount of different treatments
| 处理Treatment | 施肥量Fertilizer amount/(kg· m-2) |
|---|---|
| 不施肥(CK) Without fertilizer | 0 |
| 单施化肥(IF) Inorganic fertilizer amendment | Urea 0.04、(NH4)2HPO4 0.05和K2SO4 0.03 |
| 单施有机肥(OF) Organic fertilizer amendment | Compost from chicken manure 2.38 |
| 抗生素抗性基因ARGs | 引物序列Primer sequences | 退火温度Annealing temperature Tm/oC | 参考文献Reference | |
|---|---|---|---|---|
| 氨基糖苷类 Aminoglycosides | strA | CCGGTGGCATTTGAGAAAAA | 60 | [ |
| GTGGCTCAACCTGCGAAAAG | ||||
| strB | GCTCGGTCGTGAGAACAATCT | 60 | [ | |
| CAATTTCGGTCGCCTGGTAGT | ||||
| aadA-01 | GTTGTGCACGACGACATCATT | 60 | [ | |
| GGCTCGAAGATACCTGCAAGAA | ||||
| β-内酰胺类 β-lactams | blaOXA-20 | TGATGATTGTCGAAGCCAAA | 60 | [ |
| GCCTGTAGGCCACTCTACCC | ||||
| blaTEM | AGCATCTTACGGATGGCATGA | 60 | [ | |
| TCCTCCGATCGTTGTCAGAAGT | ||||
| blaCTX-M | CAGATTCGGTTCGCTTTCAC | 55 | [ | |
| GCAAATACTTTATCGTGCTGATG | ||||
| 大环内酯-林肯酰胺-链阳性菌素B MLSB | ermA | TTGAGAAGGGATTTGCGAAAAG | 60 | [ |
| ATATCCATCTCCACCATTAATAGTAAACC | ||||
| ermB | TAAAGGGCATTTAACGACGAAACT | 60 | [ | |
| TTTATACCTCTGTTTGTTAGGGAATTGAA | ||||
| mphA-01 | CTGACGCGCTCCGTGTT | 60 | [ | |
| GGTGGTGCATGGCGATCT | ||||
| oleC | CCCGGAGTCGATGTTCGA | 60 | [ | |
| GCCGAAGACGTACACGAACAG | ||||
| 磺胺类 Sulfonamides | sul1 | CAGCGCTATGCGCTCAAG | 60 | [ |
| ATCCCGCTGCGCTGAGT | ||||
| sul2 | TCATCTGCCAAACTCGTCGTTA | 60 | [ | |
| GTCAAAGAACGCCGCAATGT | ||||
| 四环素类Tetracyclines | tetA | CTCACCAGCCTGACCTCGAT | 60 | [ |
| CACGTTGTTATAGAAGCCGCATAG | ||||
| tetW | ATGAACATTCCCACCGTTATCTTT | 60 | [ | |
| ATATCGGCGGAGAGCTTATCC | ||||
| tetM | GGAGCGATTACAGAATTAGGAAGC | 60 | [ | |
| TCCATATGTCCTGGCGTGTC | ||||
| tetX | AAATTTGTTACCGACACGGAAGTT | 60 | [ | |
| CATAGCTGAAAAAATCCAGGACAGTT | ||||
| 多耐药类 Multi-drug resistance | acrA-05 | CGTGCGCGAACGAACA | 60 | [ |
| ACTTTGCGCGCCATCTTC | ||||
| emrD | CTCAGCAGTATGGTGGTAAGCATT | 60 | [ | |
| ACCAGGCGCCGAAGAAC | ||||
| mepA | ATCGGTCGCTCTTCGTTCAC | 60 | [ | |
| ATAAATAGGATCGAGCTGCTGGAT | ||||
| mexF | CCGCGAGAAGGCCAAGA | 60 | [ | |
| TTGAGTTCGGCGGTGATGA | ||||
| 喹诺酮类 Quinolones | qnrA | AGGATTTCTCACGCCAGGATT | 60 | [ |
| CCGCTTTCAATGAAACTGCAA | ||||
| qnrS | CGACGTGCTAACTTGCGTGA | 60 | [ | |
| GGCATTGTTGGAAACTTGCA | ||||
| 万古霉素类 Vancomycins | vanHB | GAGGTTTCCGAGGCGACAA | 60 | [ |
| CTCTCGGCGGCAGTCGTAT | ||||
| vanA | GGGCTGTGAGGTCGGTTG | 60 | [ | |
| TTCAGTACAATGCGGCCGTTA | ||||
| 多黏菌素类 Polymyxins | mcr-1 | CACATCGACGGCGTATTCTG | 60 | [ |
| CAACGAGCATACCGACATCG | ||||
| 可移动遗传元件 MGEs | intl1 | CCGTAGAACAAGCAGGCATCA | 55 | [ |
| GCGTTGAAATCATCGTCGTAGAG | ||||
表2 ARGs及I类整合子intl1 PCR引物信息
Table 2 PCR primer information of ARGs and class I integrase gene(intl1)
| 抗生素抗性基因ARGs | 引物序列Primer sequences | 退火温度Annealing temperature Tm/oC | 参考文献Reference | |
|---|---|---|---|---|
| 氨基糖苷类 Aminoglycosides | strA | CCGGTGGCATTTGAGAAAAA | 60 | [ |
| GTGGCTCAACCTGCGAAAAG | ||||
| strB | GCTCGGTCGTGAGAACAATCT | 60 | [ | |
| CAATTTCGGTCGCCTGGTAGT | ||||
| aadA-01 | GTTGTGCACGACGACATCATT | 60 | [ | |
| GGCTCGAAGATACCTGCAAGAA | ||||
| β-内酰胺类 β-lactams | blaOXA-20 | TGATGATTGTCGAAGCCAAA | 60 | [ |
| GCCTGTAGGCCACTCTACCC | ||||
| blaTEM | AGCATCTTACGGATGGCATGA | 60 | [ | |
| TCCTCCGATCGTTGTCAGAAGT | ||||
| blaCTX-M | CAGATTCGGTTCGCTTTCAC | 55 | [ | |
| GCAAATACTTTATCGTGCTGATG | ||||
| 大环内酯-林肯酰胺-链阳性菌素B MLSB | ermA | TTGAGAAGGGATTTGCGAAAAG | 60 | [ |
| ATATCCATCTCCACCATTAATAGTAAACC | ||||
| ermB | TAAAGGGCATTTAACGACGAAACT | 60 | [ | |
| TTTATACCTCTGTTTGTTAGGGAATTGAA | ||||
| mphA-01 | CTGACGCGCTCCGTGTT | 60 | [ | |
| GGTGGTGCATGGCGATCT | ||||
| oleC | CCCGGAGTCGATGTTCGA | 60 | [ | |
| GCCGAAGACGTACACGAACAG | ||||
| 磺胺类 Sulfonamides | sul1 | CAGCGCTATGCGCTCAAG | 60 | [ |
| ATCCCGCTGCGCTGAGT | ||||
| sul2 | TCATCTGCCAAACTCGTCGTTA | 60 | [ | |
| GTCAAAGAACGCCGCAATGT | ||||
| 四环素类Tetracyclines | tetA | CTCACCAGCCTGACCTCGAT | 60 | [ |
| CACGTTGTTATAGAAGCCGCATAG | ||||
| tetW | ATGAACATTCCCACCGTTATCTTT | 60 | [ | |
| ATATCGGCGGAGAGCTTATCC | ||||
| tetM | GGAGCGATTACAGAATTAGGAAGC | 60 | [ | |
| TCCATATGTCCTGGCGTGTC | ||||
| tetX | AAATTTGTTACCGACACGGAAGTT | 60 | [ | |
| CATAGCTGAAAAAATCCAGGACAGTT | ||||
| 多耐药类 Multi-drug resistance | acrA-05 | CGTGCGCGAACGAACA | 60 | [ |
| ACTTTGCGCGCCATCTTC | ||||
| emrD | CTCAGCAGTATGGTGGTAAGCATT | 60 | [ | |
| ACCAGGCGCCGAAGAAC | ||||
| mepA | ATCGGTCGCTCTTCGTTCAC | 60 | [ | |
| ATAAATAGGATCGAGCTGCTGGAT | ||||
| mexF | CCGCGAGAAGGCCAAGA | 60 | [ | |
| TTGAGTTCGGCGGTGATGA | ||||
| 喹诺酮类 Quinolones | qnrA | AGGATTTCTCACGCCAGGATT | 60 | [ |
| CCGCTTTCAATGAAACTGCAA | ||||
| qnrS | CGACGTGCTAACTTGCGTGA | 60 | [ | |
| GGCATTGTTGGAAACTTGCA | ||||
| 万古霉素类 Vancomycins | vanHB | GAGGTTTCCGAGGCGACAA | 60 | [ |
| CTCTCGGCGGCAGTCGTAT | ||||
| vanA | GGGCTGTGAGGTCGGTTG | 60 | [ | |
| TTCAGTACAATGCGGCCGTTA | ||||
| 多黏菌素类 Polymyxins | mcr-1 | CACATCGACGGCGTATTCTG | 60 | [ |
| CAACGAGCATACCGACATCG | ||||
| 可移动遗传元件 MGEs | intl1 | CCGTAGAACAAGCAGGCATCA | 55 | [ |
| GCGTTGAAATCATCGTCGTAGAG | ||||
| 抗生素抗性基因亚型 ARG subtypes | 细菌Bacteria | 噬菌体acteriophages | ||||||
|---|---|---|---|---|---|---|---|---|
| CK | IF | OF | CK | IF | OF | |||
| strA | + | + | + | - | + | + | ||
| strB | + | + | + | + | + | + | ||
| aadA-01 | + | + | + | + | + | + | ||
| blaOXA-20 | - | - | - | - | - | - | ||
| blaTEM | + | + | + | + | + | + | ||
| blaCTX-M | - | + | + | - | - | + | ||
| ermA | + | + | + | - | - | - | ||
| ermB | + | + | + | - | - | - | ||
| mphA-01 | + | + | + | + | + | + | ||
| oleC | + | + | + | + | + | + | ||
| sul1 | + | + | + | - | - | - | ||
| sul2 | + | + | + | + | + | + | ||
| tetA | + | + | + | + | + | + | ||
| tetW | + | + | + | + | + | + | ||
| tetM | + | + | + | + | + | + | ||
| tetX | + | + | + | + | + | + | ||
| acrA-05 | + | + | + | + | + | + | ||
| emrD | + | + | + | + | + | + | ||
| mepA | + | + | + | + | + | + | ||
| mexF | + | + | + | + | + | + | ||
| qnrA | - | - | - | - | - | - | ||
| qnrS | + | + | + | - | + | - | ||
| vanHB | + | + | + | + | + | + | ||
| vanA | + | + | + | + | + | + | ||
| mcr-1 | + | + | + | + | - | + | ||
| 检出率Detection rate/% | 88 | 92 | 92 | 68 | 72 | 76 | ||
| intl1 | + | + | + | + | + | + | ||
表3 不同施肥处理农田土壤中噬菌体和细菌携带ARGs及intl1的检出情况
Table 3 ARGs and intl1 carried by bacteriophages and bacteria in farmland soil under different ferti-lization treatments
| 抗生素抗性基因亚型 ARG subtypes | 细菌Bacteria | 噬菌体acteriophages | ||||||
|---|---|---|---|---|---|---|---|---|
| CK | IF | OF | CK | IF | OF | |||
| strA | + | + | + | - | + | + | ||
| strB | + | + | + | + | + | + | ||
| aadA-01 | + | + | + | + | + | + | ||
| blaOXA-20 | - | - | - | - | - | - | ||
| blaTEM | + | + | + | + | + | + | ||
| blaCTX-M | - | + | + | - | - | + | ||
| ermA | + | + | + | - | - | - | ||
| ermB | + | + | + | - | - | - | ||
| mphA-01 | + | + | + | + | + | + | ||
| oleC | + | + | + | + | + | + | ||
| sul1 | + | + | + | - | - | - | ||
| sul2 | + | + | + | + | + | + | ||
| tetA | + | + | + | + | + | + | ||
| tetW | + | + | + | + | + | + | ||
| tetM | + | + | + | + | + | + | ||
| tetX | + | + | + | + | + | + | ||
| acrA-05 | + | + | + | + | + | + | ||
| emrD | + | + | + | + | + | + | ||
| mepA | + | + | + | + | + | + | ||
| mexF | + | + | + | + | + | + | ||
| qnrA | - | - | - | - | - | - | ||
| qnrS | + | + | + | - | + | - | ||
| vanHB | + | + | + | + | + | + | ||
| vanA | + | + | + | + | + | + | ||
| mcr-1 | + | + | + | + | - | + | ||
| 检出率Detection rate/% | 88 | 92 | 92 | 68 | 72 | 76 | ||
| intl1 | + | + | + | + | + | + | ||
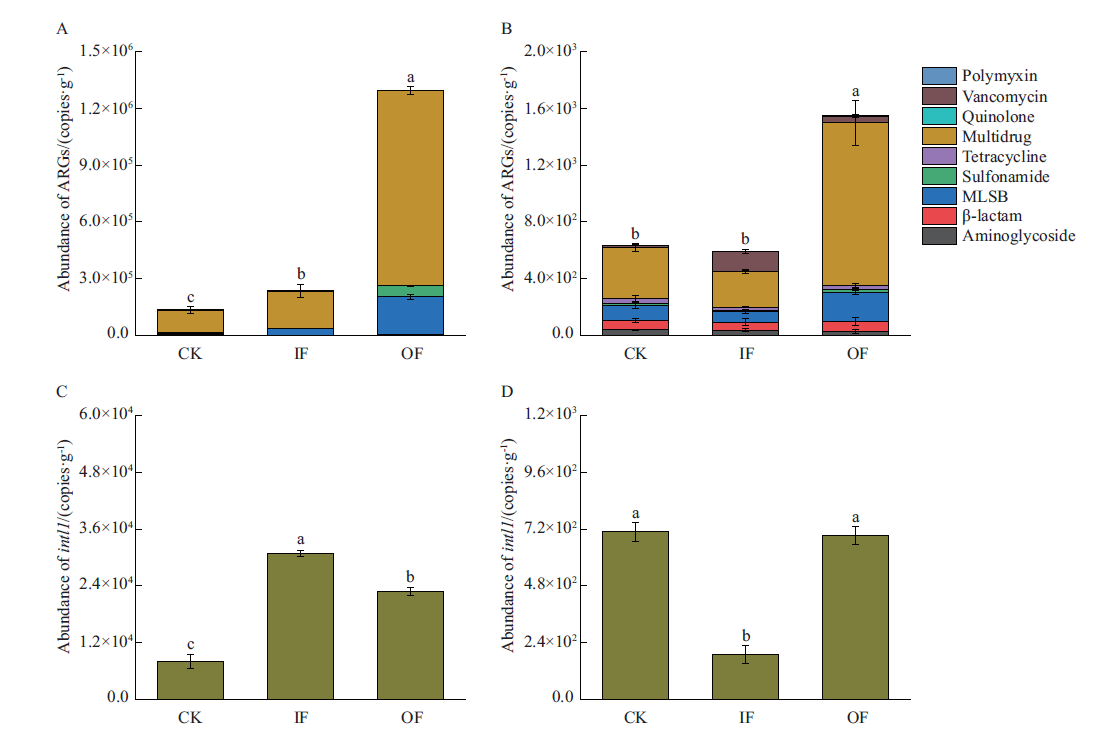
图1 不同施肥处理农田土壤细菌(A、C)和噬菌体(B、D)中ARGs与intl1丰度 不同小写字母表示差异显著(P<0.05),下同
Fig. 1 Abundances of ARGs and intl1 in bacteria(A,C)and bacteriophages(B,D)in farmland soil under different fertilization treatments Different lowercase letters indicate the statistically significant difference(P<0.05). The same below
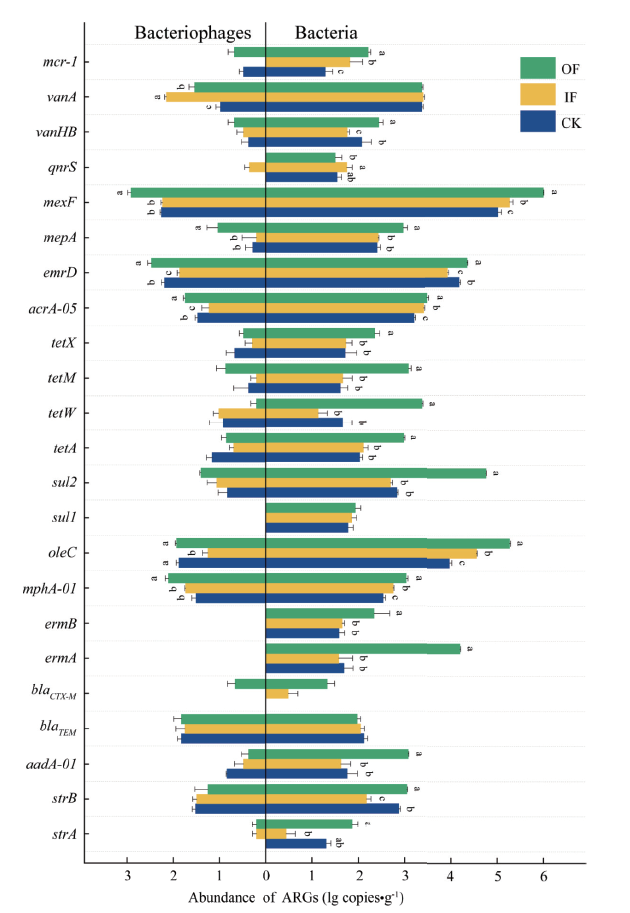
图2 土壤细菌和噬菌体中不同ARGs亚型的丰度
Fig. 2 Abundances of ARG subtypes in the bacteria and bacteriophages in farmland soil under different fertilization treatments
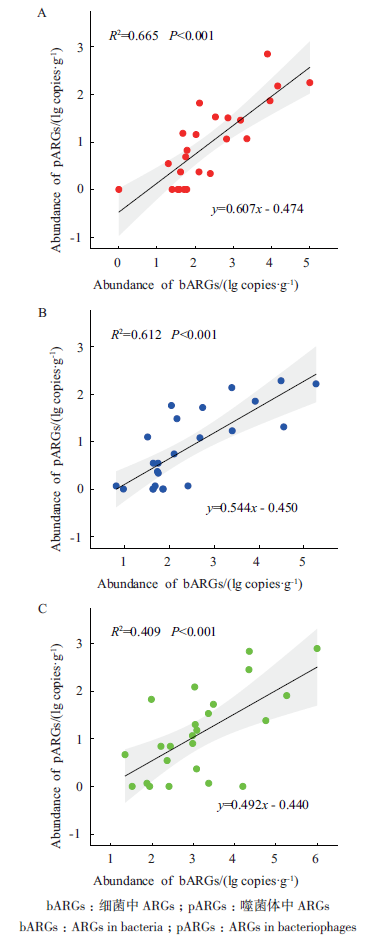
图3 不同施肥处理土壤噬菌体与细菌中ARGs丰度的相关性(A:CK;B:IF;C:OF)
Fig. 3 Correlation of the abundances of ARGs between bacteria and bacteriophages fractions in farmland soil under different fertilization treatments(A:CK;B:IF;C:OF)
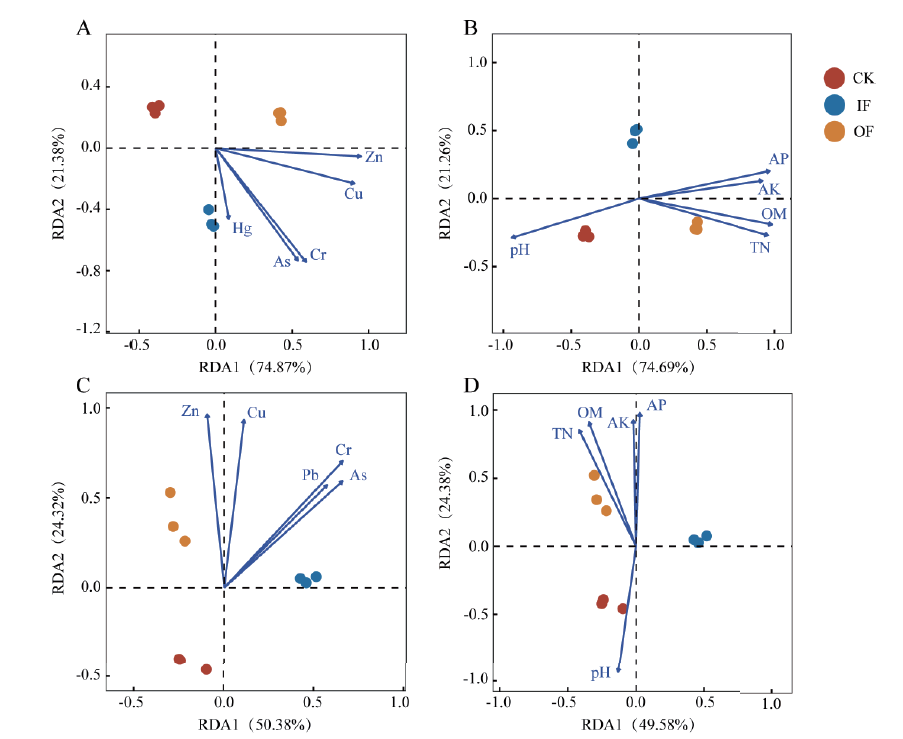
图4 细菌(A、B)与噬菌体(C、D)中ARGs与环境因子的冗余分析
Fig.4 Redundancy analysis depicting the relationship between ARGs in bacteria(A,B)and bacteriophages (C,D)and environmental factors
| [1] |
Pruden A, Pei RT, Storteboom H, et al. Antibiotic resistance genes as emerging contaminants:studies in northern Colorado[J]. Environ Sci Technol, 2006, 40(23):7445-7450.
pmid: 17181002 |
| [2] | 朱冬, 陈青林, 丁晶, 等. 土壤生态系统中抗生素抗性基因与星球健康:进展与展望[J]. 中国科学:生命科学, 2019, 49(12):1652-1663. |
|
Zhu D, Chen QL, Ding J, et al. Antibiotic resistance genes in the soil ecosystem and planetary health:Progress and prospect[J]. Sci Sin Vitae, 2019, 49(12):1652-1663.
doi: 10.1360/SSV-2019-0267 URL |
|
| [3] |
D'Costa VM, King CE, Kalan L, et al. Antibiotic resistance is ancient[J]. Nature, 2011, 477(7365):457-461.
doi: 10.1038/nature10388 URL |
| [4] |
Zhu YG, Johnson TA, Su JQ, et al. Diverse and abundant antibiotic resistance genes in Chinese swine farms[J]. PNAS, 2013, 110(9):3435-3440.
doi: 10.1073/pnas.1222743110 URL |
| [5] |
Udikovic-Kolic N, Wichmann F, Broderick NA, et al. Bloom of resident antibiotic-resistant bacteria in soil following manure fertilization[J]. Proc Natl Acad Sci USA, 2014, 111(42):15202-15207.
doi: 10.1073/pnas.1409836111 URL |
| [6] |
Xie WY, Shen Q, Zhao FJ. Antibiotics and antibiotic resistance from animal manures to soil:a review[J]. Eur J Soil Sci, 2018, 69(1):181-195.
doi: 10.1111/ejss.12494 URL |
| [7] |
Balcázar JL. How do bacteriophages promote antibiotic resistance in the environment?[J]. Clin Microbiol Infect, 2018, 24(5):447-449.
doi: 10.1016/j.cmi.2017.10.010 URL |
| [8] |
Muniesa M, Colomer-Lluch M, Jofre J. Potential impact of environmental bacteriophages in spreading antibiotic resistance genes[J]. Future Microbiol, 2013, 8(6):739-751.
doi: 10.2217/fmb.13.32 pmid: 23701331 |
| [9] |
Dion MB, Oechslin F, Moineau S. Phage diversity, genomics and phylogeny[J]. Nat Rev Microbiol, 2020, 18(3):125-138.
doi: 10.1038/s41579-019-0311-5 pmid: 32015529 |
| [10] |
Yang YX, Shi WJ, Lu SY, et al. Prevalence of antibiotic resistance genes in bacteriophage DNA fraction from Funan River water in Sichuan, China[J]. Sci Total Environ, 2018, 626:835-841.
doi: 10.1016/j.scitotenv.2018.01.148 URL |
| [11] |
Larrañaga O, Brown-Jaque M, Quirós P, et al. Phage particles harboring antibiotic resistance genes in fresh-cut vegetables and agricultural soil[J]. Environ Int, 2018, 115:133-141.
doi: S0160-4120(18)30048-5 pmid: 29567433 |
| [12] |
Sun MM, Ye M, Jiao WT, et al. Changes in tetracycline partitioning and bacteria/phage-comediated ARGs in microplastic-contaminated greenhouse soil facilitated by sophorolipid[J]. J Hazard Mater, 2018, 345:131-139.
doi: S0304-3894(17)30858-0 pmid: 29175125 |
| [13] |
Wang MZ, Liu P, Zhou Q, et al. Estimating the contribution of bacteriophage to the dissemination of antibiotic resistance genes in pig feces[J]. Environ Pollut, 2018, 238:291-298.
doi: S0269-7491(17)34918-7 pmid: 29573711 |
| [14] |
Calero-Cáceres W, Ye M, Balcázar JL. Bacteriophages as environmental reservoirs of antibiotic resistance[J]. Trends Microbiol, 2019, 27(7):570-577.
doi: S0966-842X(19)30059-9 pmid: 30905524 |
| [15] |
Oliveira J, Mahony J, Hanemaaijer L, et al. Detecting Lactococcus lactis prophages by mitomycin C-mediated induction coupled to flow cytometry analysis[J]. Front Microbiol, 2017, 8:1343.
doi: 10.3389/fmicb.2017.01343 pmid: 28769907 |
| [16] | Lekunberri I, Subirats J, Borrego CM, et al. Exploring the contribution of bacteriophages to antibiotic resistance[J]. Environ Pollut, 2017, 220(Pt B):981-984. |
| [17] |
Ouyang WY, Huang FY, Zhao Y, et al. Increased levels of antibiotic resistance in urban stream of Jiulongjiang River, China[J]. Appl Microbiol Biotechnol, 2015, 99(13):5697-5707.
doi: 10.1007/s00253-015-6416-5 URL |
| [18] |
Bert F, Branger C, Lambert-Zechovsky N. Identification of PSE and OXA beta-lactamase genes in Pseudomonas aeruginosa using PCR-restriction fragment length polymorphism[J]. J Antimicrob Chemother, 2002, 50(1):11-18.
doi: 10.1093/jac/dkf069 URL |
| [19] |
Chen ZY, Zhang W, Yang LX, et al. Antibiotic resistance genes and bacterial communities in cornfield and pasture soils receiving swine and dairy manures[J]. Environ Pollut, 2019, 248:947-957.
doi: S0269-7491(18)34775-4 pmid: 30861417 |
| [20] |
Colomer-Lluch M, Jofre J, Muniesa M. Quinolone resistance genes(qnrA and qnrS)in bacteriophage particles from wastewater samples and the effect of inducing agents on packaged antibiotic resistance genes[J]. J Antimicrob Chemother, 2014, 69(5):1265-1274.
doi: 10.1093/jac/dkt528 pmid: 24458509 |
| [21] |
Kenzaka T, Tani K, Sakotani A, et al. High-frequency phage-mediated gene transfer among Escherichia coli cells, determined at the single-cell level[J]. Appl Environ Microbiol, 2007, 73(10):3291-3299.
doi: 10.1128/AEM.02890-06 URL |
| [22] |
Subirats J, Sànchez-Melsió A, Borrego CM, et al. Metagenomic analysis reveals that bacteriophages are reservoirs of antibiotic resistance genes[J]. Int J Antimicrob Agents, 2016, 48(2):163-167.
doi: 10.1016/j.ijantimicag.2016.04.028 URL |
| [23] | 黄福义, 周曙仡聃, 王佳妮, 等. 不同作物农田土壤抗生素抗性基因多样性[J]. 环境科学, 2021, 42(6):2975-2980. |
| Huang FY, Zhou S, Wang JN, et al. Profiling of antibiotic resistance genes in different croplands[J]. Environ Sci, 2021, 42(6):2975-2980. | |
| [24] |
Forsberg KJ, Patel S, Gibson MK, et al. Bacterial phylogeny structures soil resistomes across habitats[J]. Nature, 2014, 509(7502):612-616.
doi: 10.1038/nature13377 URL |
| [25] |
Sui QW, Zhang JY, Chen MX, et al. Fate of microbial pollutants and evolution of antibiotic resistance in three types of soil amended with swine slurry[J]. Environ Pollut, 2019, 245:353-362.
doi: S0269-7491(18)32705-2 pmid: 30448505 |
| [26] |
Debroas D, Siguret C. Viruses as key reservoirs of antibiotic resistance genes in the environment[J]. ISME J, 2019, 13(11):2856-2867.
doi: 10.1038/s41396-019-0478-9 pmid: 31358910 |
| [27] |
Lood R, Ertürk G, Mattiasson B. Revisiting antibiotic resistance spreading in wastewater treatment plants - bacteriophages as a much neglected potential transmission vehicle[J]. Front Microbiol, 2017, 8:2298.
doi: 10.3389/fmicb.2017.02298 pmid: 29209304 |
| [28] |
Anand T, Bera BC, Vaid RK, et al. Abundance of antibiotic resistance genes in environmental bacteriophages[J]. J Gen Virol, 2016, 97(12):3458-3466.
doi: 10.1099/jgv.0.000639 pmid: 27902329 |
| [29] |
Ross J, Topp E. Abundance of antibiotic resistance genes in bacteriophage following soil fertilization with dairy manure or municipal biosolids, and evidence for potential transduction[J]. Appl Environ Microbiol, 2015, 81(22):7905-7913.
doi: 10.1128/AEM.02363-15 URL |
| [30] |
Yang YX, Xie XJ, Tang MJ, et al. Exploring the profile of antimicrobial resistance genes harboring by bacteriophage in chicken feces[J]. Sci Total Environ, 2020, 700:134446.
doi: 10.1016/j.scitotenv.2019.134446 URL |
| [31] |
Brown-Jaque M, Calero-Cáceres W, Espinal P, et al. Antibiotic resistance genes in phage particles isolated from human faeces and induced from clinical bacterial isolates[J]. Int J Antimicrob Agents, 2018, 51(3):434-442.
doi: 10.1016/j.ijantimicag.2017.11.014 URL |
| [32] |
Calero-Cáceres W, Muniesa M. Persistence of naturally occurring antibiotic resistance genes in the bacteria and bacteriophage fractions of wastewater[J]. Water Res, 2016, 95:11-18.
doi: 10.1016/j.watres.2016.03.006 pmid: 26978717 |
| [33] | Yang YW, Xing SC, Chen YX, et al. Profiles of bacteria/phage-comediated ARGs in pig farm wastewater treatment plants in China:association with mobile genetic elements, bacterial communities and environmental factors[J]. J Hazard Mater, 2021, 404(Pt B):124149. |
| [34] |
Dickinson AW, Power A, Hansen MG, et al. Heavy metal pollution and co-selection for antibiotic resistance:a microbial palaeontology approach[J]. Environ Int, 2019, 132:105117.
doi: 10.1016/j.envint.2019.105117 URL |
| [35] | Zhao ZL, Wang J, Han Y, et al. Nutrients, heavy metals and microbial communities co-driven distribution of antibiotic resistance genes in adjacent environment of mariculture[J]. Environ Pollut, 2017, 220(Pt B):909-918. |
| [1] | 周振超, 郑吉, 帅馨怡, 林泽俊, 陈红. 高通量分析人类粪便、皮肤和水环境中共享抗生素抗性基因的分布[J]. 生物技术通报, 2023, 39(7): 288-297. |
| [2] | 李托, 李陇平, 屈雷. 有尾噬菌体的结构及其受体研究进展[J]. 生物技术通报, 2023, 39(6): 88-101. |
| [3] | 文畅, 刘晨, 卢诗韵, 许忠兵, 艾超凡, 廖汉鹏, 周顺桂. 一株新的多重耐药福氏志贺菌噬菌体生物学特性及基因组分析[J]. 生物技术通报, 2022, 38(9): 127-135. |
| [4] | 徐重新, 张霄, 刘媛, 仲建锋, 谢雅晶, 卢莉娜, 高美静, 刘贤金. 靶向模拟Bt Cry1C蛋白抗虫功能的人源化基因工程抗体筛选及鉴定[J]. 生物技术通报, 2022, 38(5): 191-200. |
| [5] | 王加利, 和似琦, 康子茜, 王建勋. 噬菌体抗体展示技术及其在抗新冠病毒抗体发现中的应用[J]. 生物技术通报, 2022, 38(5): 248-256. |
| [6] | 张俊锋, 李孟珂, 吴志浩, 崔晓龙, 肖炜, 张仕颖. 噬菌体DCEAV-31和DCEIV-9对溶藻菌溶藻特性的影响[J]. 生物技术通报, 2022, 38(11): 250-257. |
| [7] | 黄景晓, 尚俊康, 陈慧敏, 沈嘉旻, 黎圆圆, 喻玉立, 倪进东, 林伯坤. 一株烈性沙门氏菌噬菌体的生物学特性及基因组分析[J]. 生物技术通报, 2021, 37(6): 136-146. |
| [8] | 王孝芳, 侯玉刚, 杨可铭, 王佳宁, 韦中, 徐阳春, 沈其荣. 一株青枯菌专性噬菌体的分离及应用效果研究[J]. 生物技术通报, 2020, 36(9): 194-201. |
| [9] | 戚家明, 杨娜, 孙杉杉, 明艳超, 郭亮, 张东旭, 徐志文. 一株具有噬菌体抗性的芽孢杆菌BS-2的鉴定及葡萄糖流加工艺优化[J]. 生物技术通报, 2019, 35(3): 210-216. |
| [10] | 耿慧君, 邹伟, 崔惠敬, 李晓宇, 王丽丽, 徐永平. 基于转录组学的金黄色葡萄球菌噬菌体安全性评估[J]. 生物技术通报, 2019, 35(12): 64-75. |
| [11] | 田琳, 张珣. 雌酮胁迫农田土壤微生物群落结构变化研究[J]. 生物技术通报, 2017, 33(6): 230-236. |
| [12] | 刘秀侠, 徐海燕, 辛国芹, 穆熙军, 孙学森, 谷巍. 一株枯草芽孢杆菌噬菌体的生物学特性分析及抗性菌株的诱变筛选[J]. 生物技术通报, 2017, 33(2): 143-148. |
| [13] | 李想, 孙岩, 王新珍, 刘俊杰, 王光华. 一种新的噬菌体基因多样性分子标记基因phoH研究进展[J]. 生物技术通报, 2017, 33(10): 40-45. |
| [14] | 李明源, 王继莲, 任羽, 季秀玲, 古丽巴哈尔·萨吾提. 三株假单胞菌低温噬菌体生物学特性比较研究[J]. 生物技术通报, 2016, 32(9): 123-130. |
| [15] | 吴蒙, 陆海荣, 黄青山. 金黄色葡萄球菌噬菌体裂解酶Ply187的CHAP结构域的表达及抗菌活性分析[J]. 生物技术通报, 2016, 32(9): 232-238. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||