








生物技术通报 ›› 2024, Vol. 40 ›› Issue (4): 167-178.doi: 10.13560/j.cnki.biotech.bull.1985.2023-0970
郭纯1,2( ), 宋桂梅1,2, 闫艳2, 邸鹏1,2(
), 宋桂梅1,2, 闫艳2, 邸鹏1,2( ), 王英平1,2
), 王英平1,2
收稿日期:2023-10-18
出版日期:2024-04-26
发布日期:2024-04-30
通讯作者:
邸鹏,男,副教授,硕士生导师,研究方向:分子生药学和药用植物育种;E-mail: di@jlau.edu.cn作者简介:郭纯,女,硕士研究生,研究方向:中药资源学;E-mail: 2069662120@qq.com
基金资助:
GUO Chun1,2( ), SONG Gui-mei1,2, YAN Yan2, DI Peng1,2(
), SONG Gui-mei1,2, YAN Yan2, DI Peng1,2( ), WANG Ying-ping1,2
), WANG Ying-ping1,2
Received:2023-10-18
Published:2024-04-26
Online:2024-04-30
摘要:
【目的】碱性亮氨酸拉链转录因子 (the basic leucine zipper,bZIP) 是植物中最大的转录因子家族之一 ,参与植物生长发育及激素调控等多个进程,对西洋参bZIP基因家族进行鉴定并对其结构及表达模式进行分析,为深入研究西洋参bZIP基因家族功能提供理论依据。【方法】利用生物信息学对西洋参bZIP基因进行理化性质、系统发育关系、基因结构、保守基序、顺式作用元件、染色体定位、进化选择压力分析、以及组织特异性表达分析,同时利用外源激素ABA处理西洋参后进行转录组数据分析及荧光定量验证。【结果】西洋参中共鉴定出121个bZIPs基因;根据系统发育树,这些基因被分为12个亚组;基因结构和保守基序分析表明,同一亚组的蛋白具有相似的保守基序和结构域,保守基序1在西洋参bZIP蛋白质中分布最广,保守性最强;顺式作用元件分析表明,bZIP上游启动子元件存在激素响应元件、防御和应激反应元件,其中ABA激素响应元件占比最大;染色体定位分析显示bZIP基因不均匀的分布在24条染色体上;进行进化选择压力分析,发现纯化选择在西洋参bZIP基因家族进化过程中起着重要作用;组织特异性表达分析,PqbZIPs基因在花和根中的表达量相对较高;转录组数据发现bZIP转录因子积极响应ABA激素处理,随后进行荧光定量实验验证,发现基因表达变化趋势基本一致。【结论】PqbZIP基因家族成员具有组织特异性表达模式且多数成员响应ABA激素处理。
郭纯, 宋桂梅, 闫艳, 邸鹏, 王英平. 西洋参bZIP基因家族全基因组鉴定和表达分析[J]. 生物技术通报, 2024, 40(4): 167-178.
GUO Chun, SONG Gui-mei, YAN Yan, DI Peng, WANG Ying-ping. Genome Wide Identification and Expression Analysis of the bZIP Gene Family in Panax quinquefolius[J]. Biotechnology Bulletin, 2024, 40(4): 167-178.
| 基因 Gene | 上游引物Forward primer(5'-3') | 下游引物Reverse primer(5'-3') |
|---|---|---|
| PqbZIP1778 | TGATACATCTTCTGTGTCACCAATTCC | TGCCTCCTCTCAACAACCTTCTC |
| PqbZIP3076 | GCCAGAGCCAGCAACAAGTTC | GATCTCTTCATCACTCTCCATACCTTC |
| PqbZIP53695 | GTGCCACAACTCAACCAGAAGG | TCCGACTCCATCCAAGGTGTTC |
| PqbZIP53753 | TGATGAACGATAACGGCATTGATAGG | TCCAGTAGAATATCCGCCAAGAGG |
| GAPDH1-F | GCTATCAAGGAGGAGTCGGAGGG | CTGGACAGATGCCATGTGAACAA |
表1 PqbZIP基因家族实时荧光定量引物
Table 1 Real-time fluorescence quantitative primers of PqbZIP gene family
| 基因 Gene | 上游引物Forward primer(5'-3') | 下游引物Reverse primer(5'-3') |
|---|---|---|
| PqbZIP1778 | TGATACATCTTCTGTGTCACCAATTCC | TGCCTCCTCTCAACAACCTTCTC |
| PqbZIP3076 | GCCAGAGCCAGCAACAAGTTC | GATCTCTTCATCACTCTCCATACCTTC |
| PqbZIP53695 | GTGCCACAACTCAACCAGAAGG | TCCGACTCCATCCAAGGTGTTC |
| PqbZIP53753 | TGATGAACGATAACGGCATTGATAGG | TCCAGTAGAATATCCGCCAAGAGG |
| GAPDH1-F | GCTATCAAGGAGGAGTCGGAGGG | CTGGACAGATGCCATGTGAACAA |

图2 PqbZIP家族中的基因结构示意图和保留的基序模式(A)及顺式作用元件分析(B)
Fig. 2 Schematic diagram of gene structure and retained motif patterns in the PqbZIP family (A) and Cis-acting element analysis (B)

图3 PqbZIP家族保守结构域的可视化 每个位置的字母堆的总高度表示该位置的序列守恒。字母堆中单个字母的高度代表该位置对应氨基酸的相对频率
Fig. 3 Visualization of conservative domains in the PqbZIP family The total height of the letter stack at each position indicates the conservation of the sequence at that position. The height of a single letter in the letter stack indicates the relative frequency of the corresponding amino acid at that position

图4 西洋参bZIP基因家族的染色体定位及共线性分析 绿线表示重复的西洋参bZIP基因对
Fig. 4 Chromosomal localization and collinearity analysis of the bZIP gene family in P. quinquefolius The green line indicates duplicate pairs of bZIP genes in P. quinquefolius
| Sequence | Method | Ka | Ks | Ka/Ks | Length/bp | Sequence | Method | Ka | Ks | Ka/Ks | Length/bp | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| bZIP2-bZIP43 | MA | 0.117946 | 0.165607 | 0.712204 | 960 | bZIP67-bZIP1 | MA | 0.097071 | 0.323816 | 0.299772 | 1 152 | |
| bZIP3-bZIP44 | MA | 0.0130427 | 0.0128818 | 1.01249 | 489 | bZIP70-bZIP19 | MA | 0.0142327 | 0.0409637 | 0.347448 | 1 332 | |
| bZIP10-bZIP38 | MA | 0.100549 | 0.140421 | 0.716056 | 846 | bZIP73-bZIP14 | MA | 0.0547414 | 0.0672358 | 0.81417 | 702 | |
| bZIP10-bZIP112 | MA | 0.11451 | 0.521399 | 0.219622 | 879 | bZIP76-bZIP36 | MA | 0.110057 | 0.324559 | 0.339097 | 564 | |
| bZIP11-bZIP37 | MA | 0.0942089 | 0.0981423 | 0.959922 | 957 | bZIP79-bZIP104 | MA | 0.0904191 | 0.339624 | 0.266233 | 462 | |
| bZIP13-bZIP76 | MA | 0.104919 | 0.30191 | 0.347518 | 564 | bZIP87-bZIP81 | MA | 0.317343 | 0.931453 | 0.340696 | 1 065 | |
| bZIP13-bZIP109 | MA | 0.101853 | 0.290082 | 0.351118 | 564 | bZIP92-bZIP51 | MA | 0.015582 | 0.0313318 | 0.497324 | 903 | |
| bZIP15-bZIP54 | MA | 0.105643 | 0.321516 | 0.328579 | 972 | bZIP98-bZIP52 | MA | 0.0483138 | 0.0459249 | 1.05202 | 1 023 | |
| bZIP15-bZIP55 | MA | 0.156553 | 0.426652 | 0.366934 | 1 197 | bZIP100-bZIP80 | MA | 0.0367017 | 0.113648 | 0.322942 | 1 086 | |
| bZIP16-bZIP80 | MA | 0.0682639 | 0.322732 | 0.211519 | 1 128 | bZIP101-bZIP83 | MA | 0.0497949 | 0.0973565 | 0.51147 | 1 368 | |
| bZIP16-bZIP100 | MA | 0.0627594 | 0.322994 | 0.194305 | 1 083 | bZIP105-bZIP60 | MA | 0.0758902 | 0.29116 | 0.260647 | 1 218 | |
| bZIP36-bZIP109 | MA | 0.106955 | 0.312518 | 0.342235 | 564 | bZIP108-bZIP53 | MA | 0.0113524 | 0.0549189 | 0.206712 | 1 002 | |
| bZIP45-bZIP20 | MA | 0.0126196 | 0.0519384 | 0.242973 | 435 | bZIP114-bZIP26 | MA | 0.0106575 | 0.0442045 | 0.241095 | 846 | |
| bZIP46-bZIP24 | MA | 0.0182878 | 0.0429762 | 0.425534 | 1 533 | bZIP115-bZIP81 | MA | 0.242357 | 0.752788 | 0.321946 | 1 068 | |
| bZIP47-bZIP23 | MA | 0.018861 | 0.0298436 | 0.631996 | 993 | bZIP115-bZIP82 | MA | 0.16807 | 0.619685 | 0.271218 | 1 308 | |
| bZIP48-bZIP21 | MA | 0.0890243 | 0.194441 | 0.457848 | 663 | bZIP118-bZIP85 | MA | 0.114442 | 0.317447 | 0.360506 | 894 | |
| bZIP50-bZIP95 | MA | 0.119225 | 0.411079 | 0.29003 | 843 | bZIP118-bZIP103 | MA | 0.0895535 | 0.261122 | 0.342957 | 885 | |
| bZIP61-bZIP91 | MA | 0.168025 | 0.327658 | 0.512807 | 771 | bZIP119-bZIP32 | MA | 0.0231439 | 0.0836963 | 0.276523 | 996 |
表2 PqbZIPs大规模复制事件的复制模式
Table 2 Replication mode of PqbZIPs for large-scale replication events
| Sequence | Method | Ka | Ks | Ka/Ks | Length/bp | Sequence | Method | Ka | Ks | Ka/Ks | Length/bp | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| bZIP2-bZIP43 | MA | 0.117946 | 0.165607 | 0.712204 | 960 | bZIP67-bZIP1 | MA | 0.097071 | 0.323816 | 0.299772 | 1 152 | |
| bZIP3-bZIP44 | MA | 0.0130427 | 0.0128818 | 1.01249 | 489 | bZIP70-bZIP19 | MA | 0.0142327 | 0.0409637 | 0.347448 | 1 332 | |
| bZIP10-bZIP38 | MA | 0.100549 | 0.140421 | 0.716056 | 846 | bZIP73-bZIP14 | MA | 0.0547414 | 0.0672358 | 0.81417 | 702 | |
| bZIP10-bZIP112 | MA | 0.11451 | 0.521399 | 0.219622 | 879 | bZIP76-bZIP36 | MA | 0.110057 | 0.324559 | 0.339097 | 564 | |
| bZIP11-bZIP37 | MA | 0.0942089 | 0.0981423 | 0.959922 | 957 | bZIP79-bZIP104 | MA | 0.0904191 | 0.339624 | 0.266233 | 462 | |
| bZIP13-bZIP76 | MA | 0.104919 | 0.30191 | 0.347518 | 564 | bZIP87-bZIP81 | MA | 0.317343 | 0.931453 | 0.340696 | 1 065 | |
| bZIP13-bZIP109 | MA | 0.101853 | 0.290082 | 0.351118 | 564 | bZIP92-bZIP51 | MA | 0.015582 | 0.0313318 | 0.497324 | 903 | |
| bZIP15-bZIP54 | MA | 0.105643 | 0.321516 | 0.328579 | 972 | bZIP98-bZIP52 | MA | 0.0483138 | 0.0459249 | 1.05202 | 1 023 | |
| bZIP15-bZIP55 | MA | 0.156553 | 0.426652 | 0.366934 | 1 197 | bZIP100-bZIP80 | MA | 0.0367017 | 0.113648 | 0.322942 | 1 086 | |
| bZIP16-bZIP80 | MA | 0.0682639 | 0.322732 | 0.211519 | 1 128 | bZIP101-bZIP83 | MA | 0.0497949 | 0.0973565 | 0.51147 | 1 368 | |
| bZIP16-bZIP100 | MA | 0.0627594 | 0.322994 | 0.194305 | 1 083 | bZIP105-bZIP60 | MA | 0.0758902 | 0.29116 | 0.260647 | 1 218 | |
| bZIP36-bZIP109 | MA | 0.106955 | 0.312518 | 0.342235 | 564 | bZIP108-bZIP53 | MA | 0.0113524 | 0.0549189 | 0.206712 | 1 002 | |
| bZIP45-bZIP20 | MA | 0.0126196 | 0.0519384 | 0.242973 | 435 | bZIP114-bZIP26 | MA | 0.0106575 | 0.0442045 | 0.241095 | 846 | |
| bZIP46-bZIP24 | MA | 0.0182878 | 0.0429762 | 0.425534 | 1 533 | bZIP115-bZIP81 | MA | 0.242357 | 0.752788 | 0.321946 | 1 068 | |
| bZIP47-bZIP23 | MA | 0.018861 | 0.0298436 | 0.631996 | 993 | bZIP115-bZIP82 | MA | 0.16807 | 0.619685 | 0.271218 | 1 308 | |
| bZIP48-bZIP21 | MA | 0.0890243 | 0.194441 | 0.457848 | 663 | bZIP118-bZIP85 | MA | 0.114442 | 0.317447 | 0.360506 | 894 | |
| bZIP50-bZIP95 | MA | 0.119225 | 0.411079 | 0.29003 | 843 | bZIP118-bZIP103 | MA | 0.0895535 | 0.261122 | 0.342957 | 885 | |
| bZIP61-bZIP91 | MA | 0.168025 | 0.327658 | 0.512807 | 771 | bZIP119-bZIP32 | MA | 0.0231439 | 0.0836963 | 0.276523 | 996 |

图5 西洋参与人参、西洋参与三七之间的bZIP同源性分析 背景上的灰色线条表示西洋参和其他植物基因组中的共线块;绿线突出相同基因的西洋参bZIP基因对
Fig. 5 Homology analysis of bZIPs between P. quinquefoli-us and Panax ginseng as well as P. quinquefolius and Panax notoginseng The gray lines on the background indicate collinear blocks in the genomes of P. quinquefolius and other plants; the green line highlights the bZIP gene pair of homologous P. quinquefolius
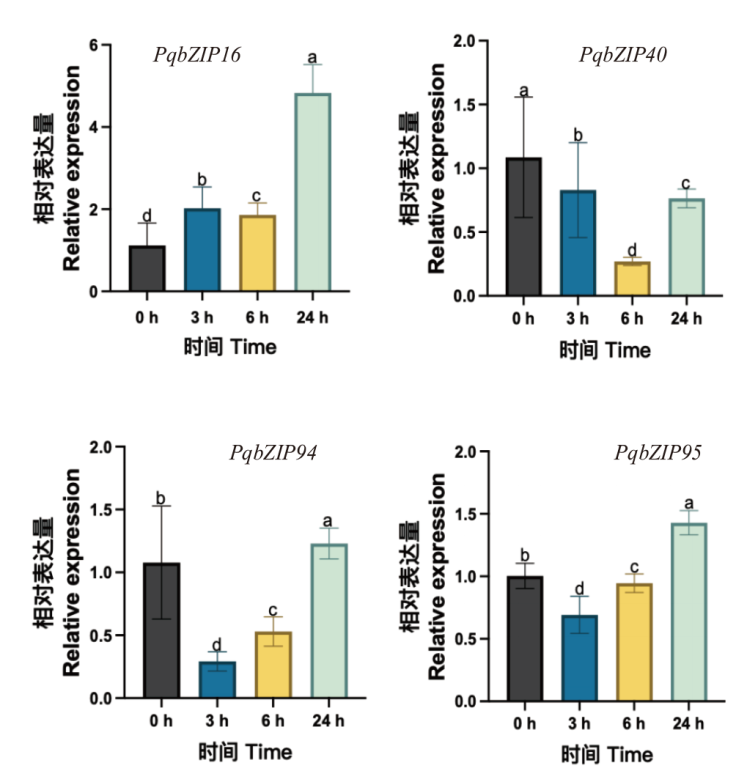
图9 在ABA处理下4个PqbZIPs基因的相对表达量 不同小写字母表示P < 0.05时差异显著
Fig. 9 Relative expressions of four PqbZIPs genes under ABA treatment Different lowercase letters indicate significant differences when P<0.05
| [1] |
Jakoby M, Weisshaar B, Dröge-Laser W, et al. bZIP transcription factors in Arabidopsis[J]. Trends Plant Sci, 2002, 7(3): 106-111.
doi: 10.1016/s1360-1385(01)02223-3 pmid: 11906833 |
| [2] | Pérez-Rodríguez P, Riaño-Pachón DM, Corrêa LGG, et al. PlnTFDB: updated content and new features of the plant transcription factor database[J]. Nucleic Acids Res, 2010, 38(Database issue): D822-D827. |
| [3] | Nijhawan A, Jain M, Tyagi AK, et al. Genomic survey and gene expression analysis of the basic leucine zipper transcription factor family in rice[J]. Plant Physiol, 2008, 146(2): 323-324. |
| [4] | Zhang Y, Gao WL, Li HT, et al. Genome-wide analysis of the bZIP gene family in Chinese jujube(Ziziphus jujuba Mill.)[J]. BMC Genom, 2020, 21(1): 483. |
| [5] |
Fukazawa J, Sakai T, Ishida S, et al. Repression of shoot growth, a bZIP transcriptional activator, regulates cell elongation by controlling the level of gibberellins[J]. Plant Cell, 2000, 12(6): 901-915.
doi: 10.1105/tpc.12.6.901 pmid: 10852936 |
| [6] |
Yin Y. RF2a, a bZIP transcriptional activator of the phloem-specific rice tungro bacilliform virus promoter, functions in vascular development[J]. EMBO J, 1997, 16(17): 5247-5259.
pmid: 9311985 |
| [7] |
Baena-González E, Rolland F, Thevelein JM, et al. A central integrator of transcription networks in plant stress and energy signalling[J]. Nature, 2007, 448(7156): 938-942.
doi: 10.1038/nature06069 |
| [8] |
Lara P, Oñate-Sánchez L, Abraham Z, et al. Synergistic activation of seed storage protein gene expression in Arabidopsis by ABI3 and two bZIPs related to OPAQUE2[J]. J Biol Chem, 2003, 278(23): 21003-21011.
doi: 10.1074/jbc.M210538200 URL |
| [9] |
Chinnusamy V, Ohta M, Kanrar S, et al. ICE1: a regulator of cold-induced transcriptome and freezing tolerance in Arabidopsis[J]. Genes Dev, 2003, 17(8): 1043-1054.
doi: 10.1101/gad.1077503 URL |
| [10] |
Kiribuchi K, Sugimori M, Takeda M, et al. RERJ1, a jasmonic acid-responsive gene from rice, encodes a basic helix-loop-helix protein[J]. Biochem Biophys Res Commun, 2004, 325(3): 857-863.
doi: 10.1016/j.bbrc.2004.10.126 URL |
| [11] |
Liao Y, Zou HF, Wei W, et al. Soybean GmbZIP44, GmbZIP62 and GmbZIP78 genes function as negative regulator of ABA signaling and confer salt and freezing tolerance in transgenic Arabidopsis[J]. Planta, 2008, 228(2): 225-240.
doi: 10.1007/s00425-008-0731-3 URL |
| [12] | Yang SQ, Xu K, Chen SJ, et al. A stress-responsive bZIP transcription factor OsbZIP62 improves drought and oxidative tolerance in rice[J]. BMC Plant Biol, 2019, 19(1): 260. |
| [13] |
Ma HZ, Liu C, Li ZX, et al. ZmbZIP4 contributes to stress resistance in maize by regulating ABA synthesis and root development[J]. Plant Physiol, 2018, 178(2): 753-770.
doi: 10.1104/pp.18.00436 pmid: 30126870 |
| [14] |
Tu MX, Wang XH, Feng TY, et al. Expression of a grape(Vitis vinifera)bZIP transcription factor, VlbZIP36, in Arabidopsis thaliana confers tolerance of drought stress during seed germination and seedling establishment[J]. Plant Sci, 2016, 252: 311-323.
doi: 10.1016/j.plantsci.2016.08.011 URL |
| [15] |
An JP, Yao JF, Wang XN, et al. MdHY5 positively regulates cold tolerance via CBF-dependent and CBF-independent pathways in apple[J]. J Plant Physiol, 2017, 218: 275-281.
doi: 10.1016/j.jplph.2017.09.001 URL |
| [16] |
Zhang Y, Xu ZC, Ji AJ, et al. Genomic survey of bZIP transcription factor genes related to tanshinone biosynthesis in Salvia miltiorrhiza[J]. Acta Pharm Sin B, 2018, 8(2): 295-305.
doi: 10.1016/j.apsb.2017.09.002 pmid: 29719790 |
| [17] |
Liu JX, Srivastava R, Howell SH. Stress-induced expression of an activated form of AtbZIP17 provides protection from salt stress in Arabidopsis[J]. Plant Cell Environ, 2008, 31(12): 1735-1743.
doi: 10.1111/pce.2008.31.issue-12 URL |
| [18] |
Li DY, Fu FY, Zhang HJ, et al. Genome-wide systematic characterization of the bZIP transcriptional factor family in tomato(Solanum lycopersicum L.)[J]. BMC Genomics, 2015, 16: 771.
doi: 10.1186/s12864-015-1990-6 URL |
| [19] |
Wei KF, Chen J, Wang YM, et al. Genome-wide analysis of bZIP-encoding genes in maize[J]. DNA Res, 2012, 19(6): 463-476.
doi: 10.1093/dnares/dss026 pmid: 23103471 |
| [20] | Wang XL, Chen XL, Yang TB, et al. Genome-wide identification of bZIP family genes involved in drought and heat stresses in strawberry(Fragaria vesca)[J]. Int J Genomics, 2017, 2017: 3981031. |
| [21] |
Wang JZ, Zhou JX, Zhang BL, et al. Genome-wide expansion and expression divergence of the basic leucine zipper transcription factors in higher plants with an emphasis on Sorghum[J]. J Integr Plant Biol, 2011, 53(3): 212-231.
doi: 10.1111/j.1744-7909.2010.01017.x |
| [22] |
Wang YP, Choi HK, Brinckmann JA, et al. Chemical analysis of Panax quinquefolius(North American ginseng): a review[J]. J Chromatogr A, 2015, 1426: 1-15.
doi: 10.1016/j.chroma.2015.11.012 URL |
| [23] | Wang ZH, Wang XF, Lu TY, et al. Reshuffling of the ancestral core-eudicot genome shaped chromatin topology and epigenetic modification in Panax[J]. Nat Commun, 2022, 13(1): 1902. |
| [24] | Jin JP, Tian F, Yang DC, et al. PlantTFDB 4.0: toward a central hub for transcription factors and regulatory interactions in plants[J]. Nucleic Acids Res, 2017, 45(D1): D1040-D1045. |
| [25] |
Wilkins MR, Gasteiger E, Bairoch A, et al. Protein identification and analysis tools in the ExPASy server[J]. Methods Mol Biol, 1999, 112: 531-552.
pmid: 10027275 |
| [26] |
Kumar S, Stecher G, Li M, et al. MEGA X: molecular evolutionary genetics analysis across computing platforms[J]. Mol Biol Evol, 2018, 35(6): 1547-1549.
doi: 10.1093/molbev/msy096 pmid: 29722887 |
| [27] |
Nguyen LT, Schmidt HA, von Haeseler A, et al. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies[J]. Mol Biol Evol, 2015, 32(1): 268-274.
doi: 10.1093/molbev/msu300 URL |
| [28] | Subramanian B, Gao SH, Lercher MJ, et al. Evolview v3: a webserver for visualization, annotation, and management of phylogenetic trees[J]. Nucleic Acids Res, 2019, 47(W1): W270-W275. |
| [29] | Bailey TL, Williams N, Misleh C, et al. MEME: discovering and analyzing DNA and protein sequence motifs[J]. Nucleic Acids Res, 2006, 34(Web Server issue): W369-W373. |
| [30] |
Chen CJ, Chen H, Zhang Y, et al. TBtools: an integrative toolkit developed for interactive analyses of big biological data[J]. Mol Plant, 2020, 13(8): 1194-1202.
doi: S1674-2052(20)30187-8 pmid: 32585190 |
| [31] |
Lescot M, Déhais P, Thijs G, et al. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences[J]. Nucleic Acids Res, 2002, 30(1): 325-327.
doi: 10.1093/nar/30.1.325 URL |
| [32] |
Krzywinski M, Schein J, Birol İ, et al. Circos: an information aesthetic for comparative genomics[J]. Genome Res, 2009, 19(9): 1639-1645.
doi: 10.1101/gr.092759.109 pmid: 19541911 |
| [33] |
Tang HB, Bowers JE, Wang XY, et al. Synteny and collinearity in plant genomes[J]. Science, 2008, 320(5875): 486-488.
doi: 10.1126/science.1153917 pmid: 18436778 |
| [34] |
Zhang Z. KaKs_Calculator 3.0: calculating selective pressure on coding and non-coding sequences[J]. Genom Proteom Bioinform, 2022, 20(3): 536-540.
doi: 10.1016/j.gpb.2021.12.002 URL |
| [35] |
Zhang WW, Xu F, Cheng SY, et al. Characterization and functional analysis of a MYB gene(GbMYBFL)related to flavonoid accumulation in Ginkgo biloba[J]. Genes Genom, 2018, 40(1): 49-61.
doi: 10.1007/s13258-017-0609-5 |
| [36] | Liu X, Chu ZQ. Genome-wide evolutionary characterization and analysis of bZIP transcription factors and their expression profiles in response to multiple abiotic stresses in Brachypodium distachyon[J]. BMC Genom, 2015, 16(1): 227. |
| [37] | Ji CL, Mao X, Hao JY, et al. Analysis of bZIP transcription factor family and their expressions under salt stress in Chlamydomonas reinhardtii[J]. Int J Mol Sci, 2018, 19(9): 2800. |
| [38] |
Gai WX, Ma X, Qiao YM, et al. Characterization of the bZIP transcription factor family in pepper(Capsicum annuum L.): CabZIP25 positively modulates the salt tolerance[J]. Front Plant Sci, 2020, 11: 139.
doi: 10.3389/fpls.2020.00139 URL |
| [39] |
Han YX, Hou ZN, He QL, et al. Genome-wide characterization and expression analysis of bZIP gene family under abiotic stress in Glycyrrhiza uralensis[J]. Front Genet, 2021, 12: 754237.
doi: 10.3389/fgene.2021.754237 URL |
| [1] | 李慧, 文钰芳, 王悦, 纪超, 石国优, 罗英, 周勇, 李志敏, 吴晓玉, 杨有新, 刘建萍. 盐胁迫下辣椒CaPIF4的表达特性与功能分析[J]. 生物技术通报, 2024, 40(4): 148-158. |
| [2] | 肖雅茹, 贾婷婷, 罗丹, 武喆, 李丽霞. 黄瓜CsERF025L转录因子的克隆及表达分析[J]. 生物技术通报, 2024, 40(4): 159-166. |
| [3] | 钟匀, 林春, 刘正杰, 董陈文华, 毛自朝, 李兴玉. 芦笋皂苷合成相关糖基转移酶基因克隆及原核表达分析[J]. 生物技术通报, 2024, 40(4): 255-263. |
| [4] | 杨淇, 魏子迪, 宋娟, 童堃, 杨柳, 王佳涵, 刘海燕, 栾维江, 马轩. 水稻组蛋白H1三突变体的创建和转录组学分析[J]. 生物技术通报, 2024, 40(4): 85-96. |
| [5] | 谢倩, 江来, 贺进, 刘玲玲, 丁明月, 陈清西. 不同鲜食品质橄榄果实转录组测序及酚类代谢途径相关调控基因挖掘[J]. 生物技术通报, 2024, 40(3): 215-228. |
| [6] | 陈艳梅. 蛋白质翻译后修饰之间的互作关系及其协同调控机理[J]. 生物技术通报, 2024, 40(2): 1-8. |
| [7] | 杨艳, 胡洋, 刘霓如, 殷璐, 杨锐, 王鹏飞, 穆霄鹏, 张帅, 程春振, 张建成. ‘红满堂’苹果MbbZIP43基因的克隆与功能研究[J]. 生物技术通报, 2024, 40(2): 146-159. |
| [8] | 路喻丹, 刘晓驰, 冯新, 陈桂信, 陈义挺. 猕猴桃BBX基因家族成员鉴定与转录特征分析[J]. 生物技术通报, 2024, 40(2): 172-182. |
| [9] | 邹修为, 岳佳妮, 李志宇, 戴良英, 李魏. 水稻热激转录因子HsfA2b调控非生物胁迫抗性的功能分析[J]. 生物技术通报, 2024, 40(2): 90-98. |
| [10] | 周会汶, 吴兰花, 韩德鹏, 郑伟, 余跑兰, 吴杨, 肖小军. 甘蓝型油菜种子硫苷含量全基因组关联分析[J]. 生物技术通报, 2024, 40(1): 222-230. |
| [11] | 吴圳, 张明英, 闫锋, 李依民, 高静, 颜永刚, 张岗. 掌叶大黄(Rheum palmatum L.)WRKY基因家族鉴定与分析[J]. 生物技术通报, 2024, 40(1): 250-261. |
| [12] | 王斌, 袁晓, 蒋园园, 王玉昆, 肖艳辉, 何金明. bHLH96的克隆及其在薄荷萜烯生物合成调控中的功能[J]. 生物技术通报, 2024, 40(1): 281-293. |
| [13] | 陈治民, 李翠, 韦继天, 李昕然, 刘峄, 郭强. 绿原酸生物合成调控及其应用研究进展[J]. 生物技术通报, 2024, 40(1): 57-71. |
| [14] | 林红妍, 郭晓蕊, 刘迪, 李慧, 陆海. 转录组分析转录因子AtbHLH68调控细胞壁发育的分子机制[J]. 生物技术通报, 2023, 39(9): 105-116. |
| [15] | 黄小龙, 孙贵连, 马丹丹, 闫慧清. 水稻幼苗酵母单杂文库构建及LAZY1上游调控因子筛选[J]. 生物技术通报, 2023, 39(9): 126-135. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||