








生物技术通报 ›› 2024, Vol. 40 ›› Issue (5): 310-320.doi: 10.13560/j.cnki.biotech.bull.1985.2023-1114
王鑫鑫1,2( ), 管玉祝1,2, 李晓苇1,2, 洪伟1,3, 吴道艳1,2, 康颖倩1,2, 刘永畅1,2, 陈峥宏1,2(
), 管玉祝1,2, 李晓苇1,2, 洪伟1,3, 吴道艳1,2, 康颖倩1,2, 刘永畅1,2, 陈峥宏1,2( ), 崔古贞1,2(
), 崔古贞1,2( )
)
收稿日期:2023-11-27
出版日期:2024-05-26
发布日期:2024-03-28
通讯作者:
崔古贞,男,博士,教授,研究方向:医学微生物学;E-mail: cuiguzhen@gmc.edu.cn;作者简介:王鑫鑫,女,硕士研究生,研究方向:医学微生物学;E-mail: wxx_999w@163.com
基金资助:
WANG Xin-xin1,2( ), GUAN Yu-zhu1,2, LI Xiao-wei1,2, HONG Wei1,3, WU Dao-yan1,2, KANG Ying-qian1,2, LIU Yong-chang1,2, CHEN Zheng-hong1,2(
), GUAN Yu-zhu1,2, LI Xiao-wei1,2, HONG Wei1,3, WU Dao-yan1,2, KANG Ying-qian1,2, LIU Yong-chang1,2, CHEN Zheng-hong1,2( ), CUI Gu-zhen1,2(
), CUI Gu-zhen1,2( )
)
Received:2023-11-27
Published:2024-05-26
Online:2024-03-28
摘要:
【目的】过氧化氢酶(KatA)是幽门螺杆菌编码的重要毒力因子,在抵抗宿主免疫杀伤和疫苗研发等方面具有重要功能。为了鉴定幽门螺杆菌 KatA 的酶学特性,并分析其在幽门螺杆菌氧化耐受中的功能。【方法】首先,从幽门螺杆菌临床耐药菌株 Hp_G272 基因组中分离 katA 基因,并在大肠杆菌中克隆表达、分离纯化KatAG272 蛋白;然后,利用 CAT 检测试剂盒体外分析 KatAG272 的过氧化氢酶活性;其次,利用同源重组技术构建 katA 基因工程菌株,包括敲除菌株和回补菌株;最后,通过比较野生菌株与工程菌株生长表型差异、分解过氧化氢能力差异及耐受过氧化氢能力差异,阐述 katA 基因的功能及其在幽门螺杆菌耐受氧化损伤中的作用。【结果】在大肠杆菌中成功克隆表达并分离纯化 KatAG272,体外酶学分析表明,KatAG272 是一类嗜酸性过氧化氢酶,基因敲除分析表明,敲除该基因幽门螺杆菌丧失分解和耐受过氧化氢的能力,而回补该基因幽门螺杆菌回复分解和耐受过氧化氢的能力。【结论】katA 基因是幽门螺杆菌分解和耐受过氧化氢的唯一功能基因,在幽门螺杆菌氧化耐受中具有重要功能。
王鑫鑫, 管玉祝, 李晓苇, 洪伟, 吴道艳, 康颖倩, 刘永畅, 陈峥宏, 崔古贞. 幽门螺杆菌katA基因功能及其在耐受氧化损伤中的作用分析[J]. 生物技术通报, 2024, 40(5): 310-320.
WANG Xin-xin, GUAN Yu-zhu, LI Xiao-wei, HONG Wei, WU Dao-yan, KANG Ying-qian, LIU Yong-chang, CHEN Zheng-hong, CUI Gu-zhen. Function of katA in Helicobacter pylori and Its Role in the Tolerance to Oxidative Damage[J]. Biotechnology Bulletin, 2024, 40(5): 310-320.
| 菌株及质粒Strain and plasmid | 相关特征Relevant characteristics | 来源Source |
|---|---|---|
| Strains E.coli | ||
| DH5α | Clone strain, F-lacZ ΔM15Δ(lacZYA-aragF)relA1 | This lab |
| DH5α::pSD | Derived fom DH5α, carrying plasmid pSD, kanR | This lab |
| DH5α:: pSD-KO-katA | Derived fom DH5α, carrying plasmid pSD-KO-katA, kanR | This work |
| DH5α:: pCD | Derived fom DH5α, carrying plasmid pCD, CmR | This work |
| DH5α:: pCD-EX-katA | Derived fom DH5α, carrying plasmid pCD-EX-katA, CmR | This work |
| DH5α:: pET-28a-katA | Derived fom DH5α, carrying plasmid pET-28a-katA, kanR | This work |
| BL21(DE3) | Expression strain, F-lon-11Δ(ompT-nfrA)885Δ(galM-ybhJ)884 | This lab |
| BL21(DE3)::pET-28a | Derived fom BL21(DE3), carrying plasmid pET-28a, kanR | This work |
| BL21(DE3)::pET-28a-katA | Derived fom BL21(DE3), carrying plasmid pET-28a-katA, kanR | This work |
| H.pylori | ||
| HpG272 | Clinical strains | This lab |
| Hp∆katA | H.pylori strain G272, knockout gene katA | This work |
| Hp∆katA::katA | H.pylori strain HpΔkatA::katA, anaplerosis gene katA | This work |
| Plasmids | ||
| pSD | From pill570, suicide plasmid, kanR | This lab |
| PCD | From pSD, suicide plasmid, CmR | This lab |
| pSD-KO-katA | Suicide plasmid, knockout gene katA, kanR | This work |
| pCD-EX-katA | HpΔkatA::katA replenishing plasmid, CmR | This work |
| pET-28a | N-T7, N-His, C-His, E. coli protein expression vector | This lab |
| pET-28a-katA | Carrying katA gene, E. coli protein expression vector | This work |
表1 菌株及质粒
Table 1 Strains and plasmids used in this study
| 菌株及质粒Strain and plasmid | 相关特征Relevant characteristics | 来源Source |
|---|---|---|
| Strains E.coli | ||
| DH5α | Clone strain, F-lacZ ΔM15Δ(lacZYA-aragF)relA1 | This lab |
| DH5α::pSD | Derived fom DH5α, carrying plasmid pSD, kanR | This lab |
| DH5α:: pSD-KO-katA | Derived fom DH5α, carrying plasmid pSD-KO-katA, kanR | This work |
| DH5α:: pCD | Derived fom DH5α, carrying plasmid pCD, CmR | This work |
| DH5α:: pCD-EX-katA | Derived fom DH5α, carrying plasmid pCD-EX-katA, CmR | This work |
| DH5α:: pET-28a-katA | Derived fom DH5α, carrying plasmid pET-28a-katA, kanR | This work |
| BL21(DE3) | Expression strain, F-lon-11Δ(ompT-nfrA)885Δ(galM-ybhJ)884 | This lab |
| BL21(DE3)::pET-28a | Derived fom BL21(DE3), carrying plasmid pET-28a, kanR | This work |
| BL21(DE3)::pET-28a-katA | Derived fom BL21(DE3), carrying plasmid pET-28a-katA, kanR | This work |
| H.pylori | ||
| HpG272 | Clinical strains | This lab |
| Hp∆katA | H.pylori strain G272, knockout gene katA | This work |
| Hp∆katA::katA | H.pylori strain HpΔkatA::katA, anaplerosis gene katA | This work |
| Plasmids | ||
| pSD | From pill570, suicide plasmid, kanR | This lab |
| PCD | From pSD, suicide plasmid, CmR | This lab |
| pSD-KO-katA | Suicide plasmid, knockout gene katA, kanR | This work |
| pCD-EX-katA | HpΔkatA::katA replenishing plasmid, CmR | This work |
| pET-28a | N-T7, N-His, C-His, E. coli protein expression vector | This lab |
| pET-28a-katA | Carrying katA gene, E. coli protein expression vector | This work |
| 引物Primer | 序列Sequence(5'-3') | 注释Notes |
|---|---|---|
| Kan-1 | GAATTCGAGCTCGGTACCCG | kanR 基因扩增和检测引物 |
| Kan-2 | CCCGGGTCATTATTCCCTCC | |
| Kat-1 | CTGCAGGGTATAAAAACTCACCGCCCCA | 敲除质粒及敲除菌株检测引物 |
| Kat-2 | ATCGATACCTAGTTTCAAGCCTTGCA | |
| KatA-1 | ATCGACTGCAGGGTATAAAAACTCACCGCCCCA | katA 基因敲除载体上游同源臂扩增 |
| KatA-2 | ATCGAGAATTCCGTAATTGACACTAAGCCGATTAC | |
| KatA-3 | ATCGAGGATCCCTTATTTTTTAGGAACGCTTTGTC | katA 基因敲除载体下游同源臂扩增 |
| KatA-4 | ATCGAATCGATACCTAGTTTCAAGCCTTGCA | |
| Chl-1 | TTAAAAAAATTACGCCCCGCCCTG | CmR 基因扩增和检测引物 |
| Chl-2 | AAATGGAGAAAAAAATCACTGGATATACCACCG | |
| KatA -5 | CCCCGGGGACCTGCAGGCCGCTTGCACGGGT | katA 基因回补载体上游同源臂 |
| KatA -6 | TTTTTTAAGGCAGTCTGCAGGCGACCAAAACCTCTCATGGAG | |
| KatA -7 | TTGACAGCTTATCATCGATGTGAGTCAGAATGTCTTTCACAACCC | katA 基因回补载体下游同源臂 |
| KatA -8 | CTCCTGAAAATCTCGGATCCGGGCGGTTTCACTGAGAAAACTT | |
| KatA -9 | CAGTGAAACCGCCCGGATCTTTATAAATTCTAAAGGGG | katA 基因回补载体Hp尿素酶启动子PureA 扩增 |
| KatA -10 | TTAGCTCCTGAAAATCTCGGATCCTTATTCTCCTATTCTTAAAGTG | |
| KatA -11 | AACACTTTAAGAATAGGAGAATAAGATGGTTAATAAAGATGTGAAACAAACTACTGC | 回补载体 katA 基因扩增 |
| KatA -12 | TTAGCTCCTGAAAATCTCGTTACTTTTTCTTTTTTGTGTGGTGCATGTCT | |
| Kat-3 | TTGACAGCTTATCATCGATGTGAGTCAGAATGTCTTTCACAACCC | 回补质粒及回补菌株检测引物 |
| Kat-4 | CCCCGGGGACCTGCAGGCCGCTTGCACGGGT | |
| KatA-R | GTCGACGGAGCTCGAATTCGTTACTTTTTCTTTTTTGTGTGGT | katA 基因扩增 |
| KatA-F | AGCAAATGGGTCGCGGATCATGGTTAATAAAGATGTGAAAC | |
| KatA-C1 | CAAAAAACCCCTCAAGACCCGTTTAG | KatAG272 蛋白表达质粒检测引物 |
| KatA-C2 | ACCGGCATACTCTGCGACATC |
表2 本研究所用引物
Table 2 Primers used in this study
| 引物Primer | 序列Sequence(5'-3') | 注释Notes |
|---|---|---|
| Kan-1 | GAATTCGAGCTCGGTACCCG | kanR 基因扩增和检测引物 |
| Kan-2 | CCCGGGTCATTATTCCCTCC | |
| Kat-1 | CTGCAGGGTATAAAAACTCACCGCCCCA | 敲除质粒及敲除菌株检测引物 |
| Kat-2 | ATCGATACCTAGTTTCAAGCCTTGCA | |
| KatA-1 | ATCGACTGCAGGGTATAAAAACTCACCGCCCCA | katA 基因敲除载体上游同源臂扩增 |
| KatA-2 | ATCGAGAATTCCGTAATTGACACTAAGCCGATTAC | |
| KatA-3 | ATCGAGGATCCCTTATTTTTTAGGAACGCTTTGTC | katA 基因敲除载体下游同源臂扩增 |
| KatA-4 | ATCGAATCGATACCTAGTTTCAAGCCTTGCA | |
| Chl-1 | TTAAAAAAATTACGCCCCGCCCTG | CmR 基因扩增和检测引物 |
| Chl-2 | AAATGGAGAAAAAAATCACTGGATATACCACCG | |
| KatA -5 | CCCCGGGGACCTGCAGGCCGCTTGCACGGGT | katA 基因回补载体上游同源臂 |
| KatA -6 | TTTTTTAAGGCAGTCTGCAGGCGACCAAAACCTCTCATGGAG | |
| KatA -7 | TTGACAGCTTATCATCGATGTGAGTCAGAATGTCTTTCACAACCC | katA 基因回补载体下游同源臂 |
| KatA -8 | CTCCTGAAAATCTCGGATCCGGGCGGTTTCACTGAGAAAACTT | |
| KatA -9 | CAGTGAAACCGCCCGGATCTTTATAAATTCTAAAGGGG | katA 基因回补载体Hp尿素酶启动子PureA 扩增 |
| KatA -10 | TTAGCTCCTGAAAATCTCGGATCCTTATTCTCCTATTCTTAAAGTG | |
| KatA -11 | AACACTTTAAGAATAGGAGAATAAGATGGTTAATAAAGATGTGAAACAAACTACTGC | 回补载体 katA 基因扩增 |
| KatA -12 | TTAGCTCCTGAAAATCTCGTTACTTTTTCTTTTTTGTGTGGTGCATGTCT | |
| Kat-3 | TTGACAGCTTATCATCGATGTGAGTCAGAATGTCTTTCACAACCC | 回补质粒及回补菌株检测引物 |
| Kat-4 | CCCCGGGGACCTGCAGGCCGCTTGCACGGGT | |
| KatA-R | GTCGACGGAGCTCGAATTCGTTACTTTTTCTTTTTTGTGTGGT | katA 基因扩增 |
| KatA-F | AGCAAATGGGTCGCGGATCATGGTTAATAAAGATGTGAAAC | |
| KatA-C1 | CAAAAAACCCCTCAAGACCCGTTTAG | KatAG272 蛋白表达质粒检测引物 |
| KatA-C2 | ACCGGCATACTCTGCGACATC |
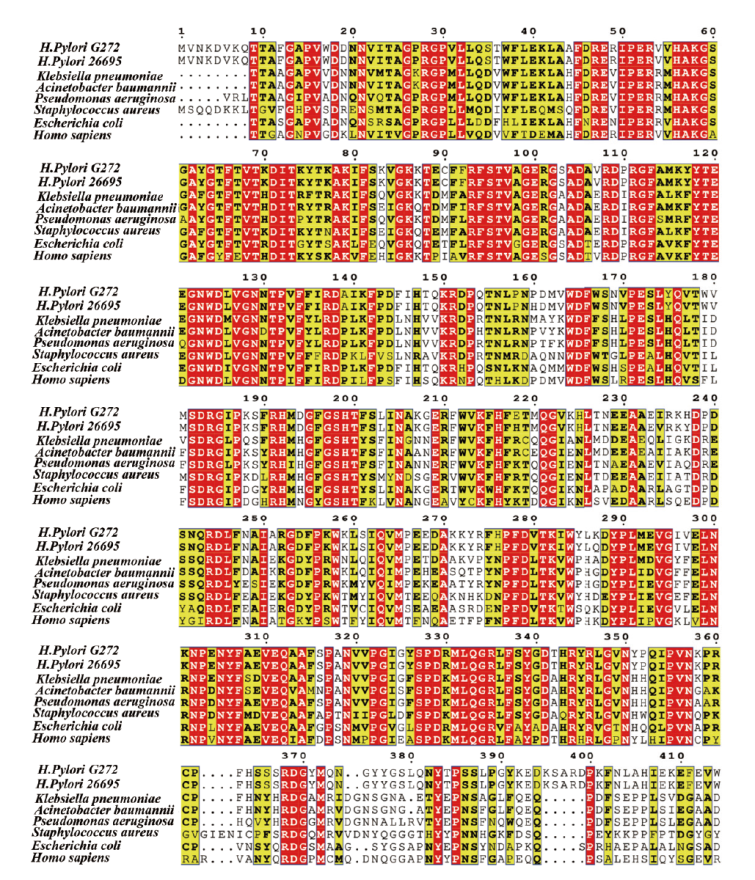
图1 katA 氨基酸序列保守性分析 红色:保守氨基酸;黄色:相对保守氨基酸;无色:不保守氨基酸
Fig. 1 Conservative analysis of amino acid sequence for katA Red: Conservative amino acids. Yellow: Relatively conserved amino acid. Colorless: Non conserved amino acids
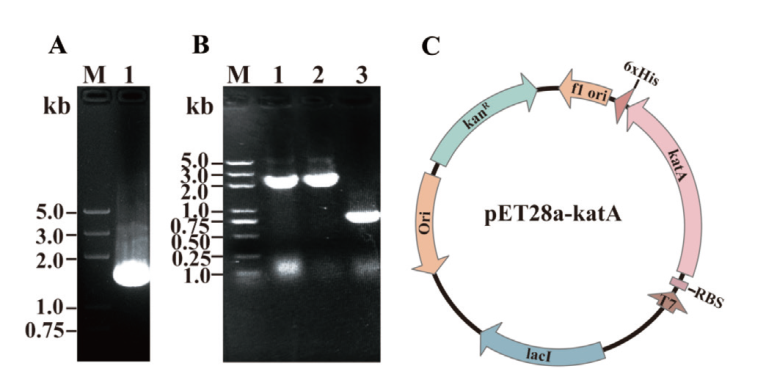
图2 katA 基因 PCR 扩增及表达载体构建 A:katA 基因 PCR 扩增;B:表达质粒 PCR 鉴定(泳道1-2:阳性质粒 PCR 鉴定;泳道3:pET28a 空白质粒对照,扩增引物为 KatA-C1/KatA-C2);C:pET28a-katA 表达质粒
Fig. 2 PCR amplification for katA and construction of expression vector A: PCR amplification for katA gene. B: Expression plasmid PCR identification (Lane 1-2: PCR identification for positive plasmid; lane 3: blank control, primers used in this identification are KatA-C1/KatA-C2). C: pET28a-katA expression plasmid
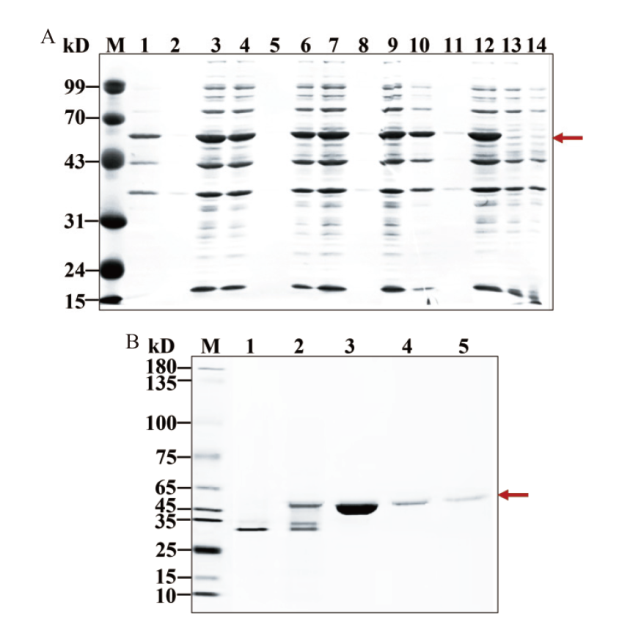
图3 KatAG272 蛋白表达及分离纯化 A:KatAG272 蛋白在大肠杆菌中的诱导表达(泳道 1、4、7、10:超声破碎前菌体沉淀,泳道 2、5、8、11:超声破碎前培养上清,泳道 3、6、9、12:超声破碎后上清,泳道 13-14:含 pET28a 载体的野生型超声破碎前菌体沉淀对照);B:KatAG272 蛋白分离纯化(泳道 1:流穿液,泳道 2:10 mmol/L咪唑漂洗液,泳道 3-5:200 mmol/L咪唑洗脱液)
Fig. 3 Protein expression, isolation and purification for KatAG272 A: The induced expression of KatAG272 protein in Escherichia coli(Lane 1, 4, 7, 10: Bacterial precipitation before ultrasound fragmentation; lane 2, 5, 8, 11: culture supernatant before ultrasound fragmentation; lane 3, 6, 9, 12: supernatant after ultrasound fragmentation; lane 13-14: control of bacterial precipitation before wild-type ultrasound fragmentation containing pET28a vector). B: KatAG272 protein isolation and purification(Lane 1: Flow through fluid; lane 2:10 mmol/L imidazole rinse; lane 3-5: 200 mmol/L imidazole elution)
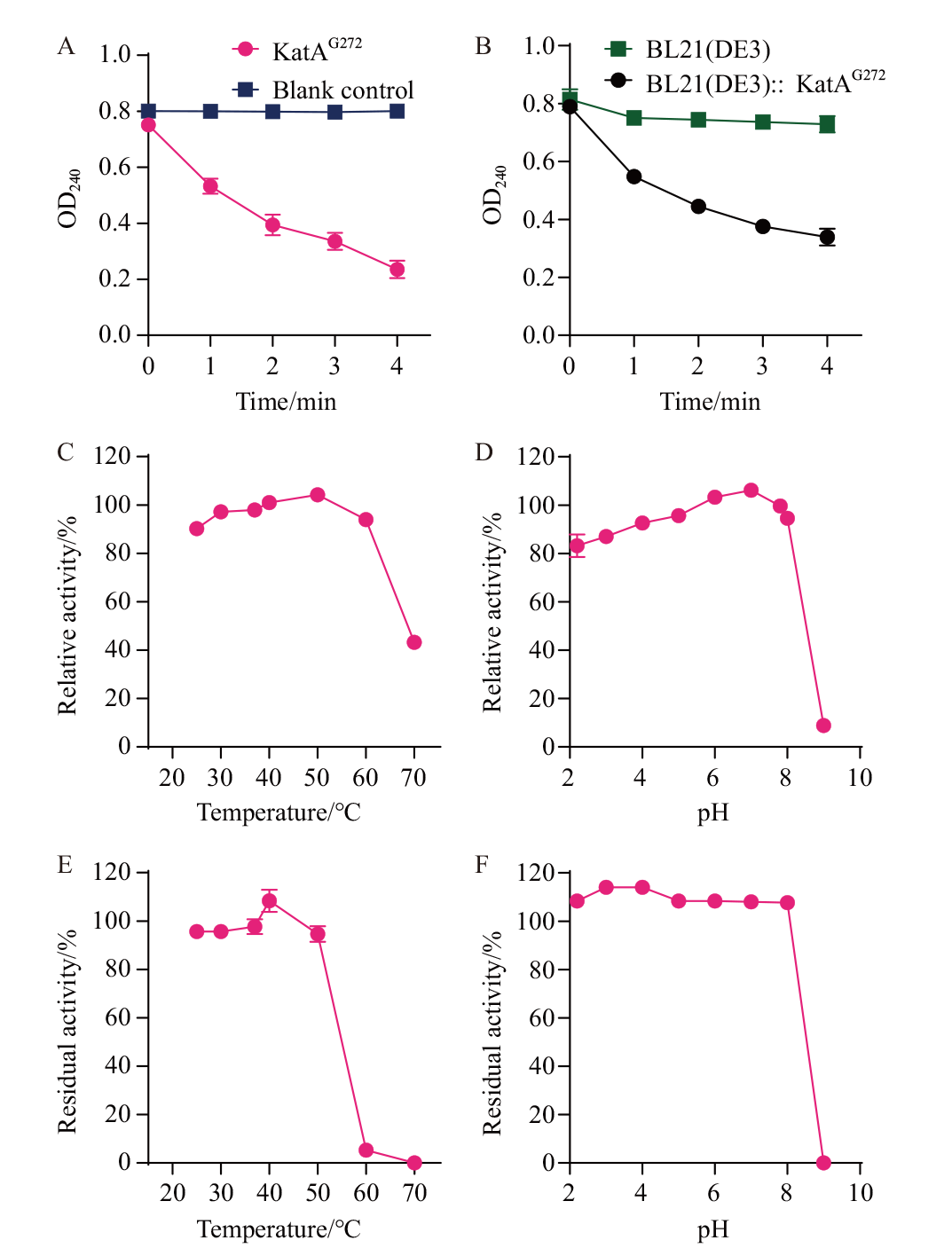
图4 过氧化氢酶活性分析及鉴定 A:KatAG272 纯酶的过氧化氢酶活性检测;B:重组大肠杆菌的 KatAG272 过氧化氢酶活性检测;C:温度对 KatAG272 活性的影响; D:pH 对KatAG272 活性的影响;E:KatAG272 蛋白温度耐受性;F:KatAG272 蛋白 pH 耐受性
Fig. 4 Analysis and identification of catalase activity A: Detection of catalase activity of KatAG272 pure enzyme. B: Detection of KatAG272 catalase activity in recombinant E. coli; C: The effect of temperature on the activity of KatAG272. D: The effect of pH on the activity of KatAG272. E: KatAG272 protein temperature tolerance. F: KatAG272 protein pH tolerance
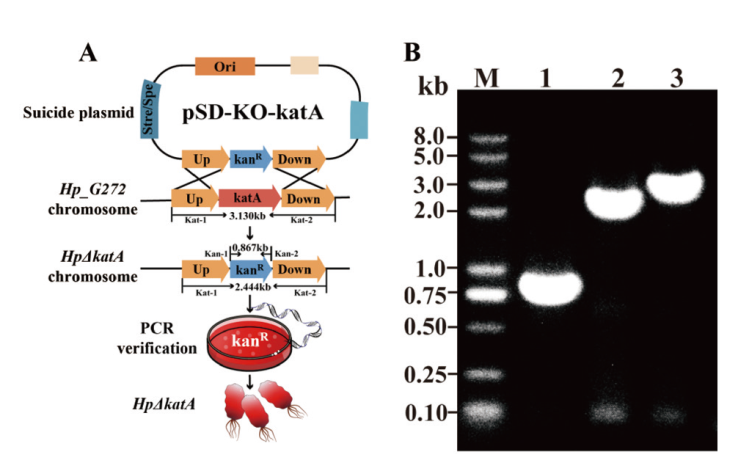
图5 敲除菌株(HpΔkatA)构建及鉴定 A:敲除菌株(HpΔkatA)构建示意图;B:敲除菌株(HpΔkatA)鉴定(泳道 1:引物 Kan-1 和 Kan-2,敲除菌株卡那霉素抗性验证,泳道 2:引物 Kat-1 和 Kat-2,敲除菌株卡那霉素抗性片段和上下游同源臂片段验证,泳道 3:引物 Kat-1 和 Kat-2,野生型对照)
Fig. 5 Construction and identification for katA-knockout strain HpΔkatA A: Schematics of HpΔkatA construction. B: Identification of HpΔkatA(Lane 1: primer Kan-1 and Kan-2, validation of kanamycin resistance in knockout strains; lane 2: primer Kat-1 and Kat-2, validation of kanamycin resistance fragments and upstream and downstream homologous arm fragments in knockout strains; lane 3: primer Kat-1 and Kat-2, wild-type control)
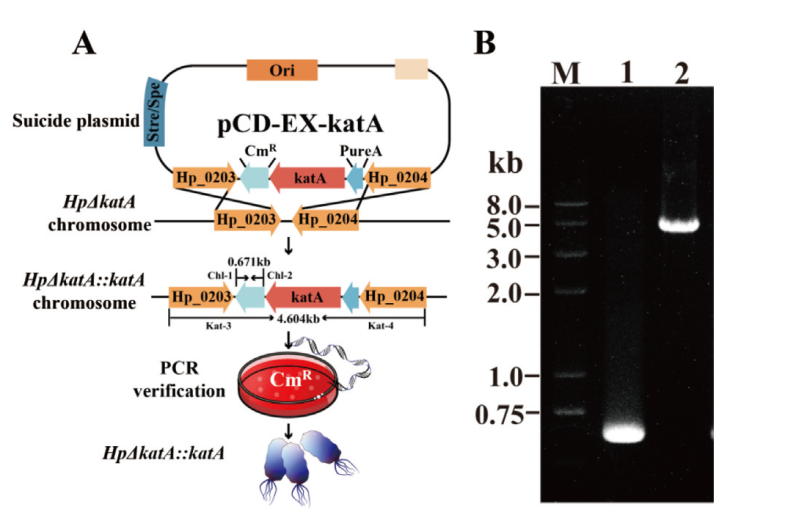
图6 回补菌株(HpΔkatA::katA)构建及鉴定 A:回补菌株(HpΔkatA::katA)构建示意图;B:回补菌株(HpΔkatA::katA)鉴定(泳道 1:引物 Chl-1 和 Chl-2,回补菌株氯霉素抗性验证,泳道 2:引物 Kat-3 和 Kat-4,回补菌株氯霉素抗性片段和上下游同源臂片段验证)
Fig. 6 Construction and identification for katA-complemented strain HpΔkatA::katA A: Schematics of HpΔkatA::katA construction; B: Identification HpΔkatA::katA(lane 1: primer Chl-1 and Chl-2, validation of chloramphenicol resistance of the complementary strain; lane 2: primer Kat-3 and Kat-4, validation of chloramphenicol resistance fragments and upstream and downstream homologous arm fragments of the complementary strain)
| 菌株Strain | 浓度Concentration/(mmol·L-1) | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 3 | 6 | 10 | 20 | 40 | 60 | 80 | 100 | ||
| Hp_G272 | + | + | + | + | + | + | - | - | - | |
| HpΔkatA | + | + | + | - | - | - | - | - | - | |
| HpΔkatA::katA | + | + | + | + | + | + | - | - | - | |
表3 Hp 对过氧化氢耐受性分析
Table 3 Analysis of tolerance of Hp to hydrogen peroxide
| 菌株Strain | 浓度Concentration/(mmol·L-1) | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 3 | 6 | 10 | 20 | 40 | 60 | 80 | 100 | ||
| Hp_G272 | + | + | + | + | + | + | - | - | - | |
| HpΔkatA | + | + | + | - | - | - | - | - | - | |
| HpΔkatA::katA | + | + | + | + | + | + | - | - | - | |
| [1] |
Malfertheiner P, Camargo MC, El-Omar E, et al. Helicobacter pylori infection[J]. Nat Rev Dis Primers, 2023, 9: 19.
doi: 10.1038/s41572-023-00431-8 pmid: 37081005 |
| [2] |
Robinson K, Atherton JC. The spectrum of Helicobacter-mediated diseases[J]. Annu Rev Pathol, 2021, 16: 123-144.
doi: 10.1146/annurev-pathol-032520-024949 pmid: 33197219 |
| [3] | Sung H, Ferlay J, Siegel RL, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries[J]. CA Cancer J Clin, 2021, 71(3): 209-249. |
| [4] | 中国疾病预防控制中心传染病预防控制所. 中国幽门螺杆菌感染防控白皮书[M]. 北京, 2023:1-47. |
| Institute for Infectious Disease Prevention and Control. China Center for Disease Control and Prevention White Paper on Prevention and Control of Helicobacter pylori Infection in China[M]. Beijing, 2023:1-47 | |
| [5] |
Tshibangu-Kabamba E, Yamaoka Y. Helicobacter pylori infection and antibiotic resistance—from biology to clinical implications[J]. Nat Rev Gastroenterol Hepatol, 2021, 18: 613-629.
doi: 10.1038/s41575-021-00449-x pmid: 34002081 |
| [6] |
Salama NR, Hartung ML, Müller A. Life in the human stomach: persistence strategies of the bacterial pathogen Helicobacter pylori[J]. Nat Rev Microbiol, 2013, 11(6): 385-399.
doi: 10.1038/nrmicro3016 pmid: 23652324 |
| [7] | Prashar A, Capurro MI, Jones NL. Under the radar: strategies used by Helicobacter pylori to evade host responses[J]. Annu Rev Physiol, 2022, 84: 485-506. |
| [8] | Sies H, Jones DP. Reactive oxygen species(ROS)as pleiotropic physiological signalling agents[J]. Nat Rev Mol Cell Biol, 2020, 21(7): 363-383. |
| [9] |
Alfonso-Prieto M, Biarnés X, Vidossich P, et al. The molecular mechanism of the catalase reaction[J]. J Am Chem Soc, 2009, 131(33): 11751-11761.
doi: 10.1021/ja9018572 pmid: 19653683 |
| [10] | Miyashita M, Joh T, Watanabe K, et al. Immune responses in mice to intranasal and intracutaneous administration of a DNA vaccine encoding Helicobacter pylori-catalase[J]. Vaccine, 2002, 20(17/18): 2336-2342. |
| [11] |
Radcliff FJ, Hazell SL, Kolesnikow T, et al. Catalase, a novel antigen for Helicobacter pylori vaccination[J]. Infect Immun, 1997, 65(11): 4668-4674.
doi: 10.1128/iai.65.11.4668-4674.1997 pmid: 9353048 |
| [12] | Harris AG, Wilson JE, Danon SJ, et al. Catalase(KatA)and KatA-associated protein(KapA)are essential to persistent colonization in the Helicobacter pylori SS1 mouse model[J]. Microbiology, 2003, 149(Pt 3): 665-672. |
| [13] | Harris AG, Hinds FE, Beckhouse AG, et al. Resistance to hydrogen peroxide in Helicobacter pylori: role of catalase(KatA)and Fur, and functional analysis of a novel gene product designated ‘KatA-associated protein', KapA(HP0874)[J]. Microbiology, 2002, 148(Pt 12): 3813-3825. |
| [14] | Cardenas VM, Dominguez DC, Puentes FA, et al. Evaluation of a novel stool native catalase antigen test for Helicobacter pylori infection in asymptomatic North American children[J]. J Pediatr Gastroenterol Nutr, 2008, 46(4): 399-402. |
| [15] | Okuda M, Osaki T, Kikuchi S, et al. Evaluation of a stool antigen test using a MAb for native catalase for diagnosis of Helicobacter pylori infection in children and adults[J]. J Med Microbiol, 2014, 63(Pt 12): 1621-1625. |
| [16] |
Shimoyama T, Sawaya M, Ishiguro A, et al. Applicability of a rapid stool antigen test, using monoclonal antibody to catalase, for the management of Helicobacter pylori infection[J]. J Gastroenterol, 2011, 46(4): 487-491.
doi: 10.1007/s00535-011-0371-4 pmid: 21264478 |
| [17] | Sato M, Shimoyama T, Takahashi R, et al. Characterization and usefulness of stool antigen tests using a monoclonal antibody to Helicobacter pylori catalase[J]. J Gastroenterol Hepatol, 2012, 27(Suppl 3): 23-28. |
| [18] | Zhang B, Li HL, Fan Q, et al. Serum Helicobacter pylori KatA and AhpC antibodies as novel biomarkers for gastric cancer[J]. World J Gastroenterol, 2016, 22(21): 5060-5067. |
| [19] | Robert X, Gouet P. Deciphering key features in protein structures with the new ENDscript server[J]. Nucleic Acids Res, 2014, 42(Web Server issue): W320-W324. |
| [20] |
Yoshida N, Granger DN, Evans DJ Jr, et al. Mechanisms involved in Helicobacter pylori-induced inflammation[J]. Gastroenterology, 1993, 105(5): 1431-1440.
pmid: 7901109 |
| [21] | Basu M, Czinn SJ, Blanchard TG. Absence of catalase reduces long-term survival of Helicobacter pylori in macrophage phagosomes[J]. Helicobacter, 2004, 9(3): 211-216. |
| [22] |
Ramarao N, Gray-Owen SD, Meyer TF. Helicobacter pylori induces but survives the extracellular release of oxygen radicals from professional phagocytes using its catalase activity[J]. Mol Microbiol, 2000, 38(1): 103-113.
pmid: 11029693 |
| [23] | Richter C, Mukherjee O, Ermert D, et al. Moonlighting of Helicobacter pylori catalase protects against complement-mediated killing by utilising the host molecule vitronectin[J]. Sci Rep, 2016, 6: 24391. |
| [24] | Usui Y, Taniyama Y, Endo M, et al. Helicobacter pylori, homologous-recombination genes, and gastric cancer[J]. N Engl J Med, 2023, 388(13): 1181-1190. |
| [25] | Shah SC, Tepler A, Chung CP, et al. Host genetic determinants associated with Helicobacter pylori eradication treatment failure: a systematic review and meta-analysis[J]. Gastroenterology, 2021, 161(5): 1443-1459. |
| [26] | Muñoz-Ramirez ZY, Pascoe B, Mendez-Tenorio A, et al. A 500-year tale of co-evolution, adaptation, and virulence: Helicobacter pylori in the Americas[J]. ISME J, 2021, 15: 78-92. |
| [27] |
Palamides P, Jolaiya T, Idowu A, et al. Helicobacter pylori patient isolates from South Africa and Nigeria differ in virulence factor pathogenicity profile and associated gastric disease outcome[J]. Sci Rep, 2020, 10(1): 11409.
doi: 10.1038/s41598-020-66128-0 pmid: 32651394 |
| [28] | Mi MH, Wu FC, Zhu J, et al. Heterogeneity of Helicobacter pylori strains isolated from patients with gastric disorders in Guiyang, China[J]. Infect Drug Resist, 2021, 14: 535-545. |
| [29] | Hosoda K, Wanibuchi K, Amgalanbaatar A, et al. A novel role of catalase in cholesterol uptake of Helicobacter pylori[J]. Steroids, 2023, 191: 109158. |
| [30] | Zhang Y, Li XY, Shan BE, et al. Perspectives from recent advances of Helicobacter pylori vaccines research[J]. Helicobacter, 2022, 27(6): e12926. |
| [1] | 高登科, 马白荣, 郭怡莹, 刘薇, 刘田, 靳亚平, 江舟, 陈华涛. 利用CRISPR/Cas9技术构建Quaking敲除的小鼠胚胎成纤维细胞株[J]. 生物技术通报, 2024, 40(2): 65-72. |
| [2] | 刘玉玲, 王梦瑶, 孙琦, 马利花, 朱新霞. 启动子RD29A对转雪莲SikCDPK1基因烟草抗逆性的影响[J]. 生物技术通报, 2023, 39(9): 168-175. |
| [3] | 赖瑞联, 冯新, 高敏霞, 路喻丹, 刘晓驰, 吴如健, 陈义挺. 猕猴桃过氧化氢酶基因家族全基因组鉴定与表达分析[J]. 生物技术通报, 2023, 39(4): 136-147. |
| [4] | 杨俊钊, 张新蕊, 赵国柱, 郑菲. 新型GH5家族多结构域纤维素酶的结构与功能研究[J]. 生物技术通报, 2023, 39(4): 71-80. |
| [5] | 李昕悦, 周明海, 樊亚超, 廖莎, 张风丽, 刘晨光, 孙悦, 张霖, 赵心清. 基于转运蛋白工程提升微生物菌株耐受性和生物制造效率的研究进展[J]. 生物技术通报, 2023, 39(11): 123-136. |
| [6] | 胡锦超, 沈文琦, 徐超业, 樊雅祺, 卢浩宇, 蒋雯杰, 李世龙, 晋洪晨, 骆健美, 王敏. 微生物酸胁迫耐受性能强化的研究进展[J]. 生物技术通报, 2023, 39(11): 137-149. |
| [7] | 晏雄鹰, 王振, 王霞, 杨世辉. 微生物硫代谢与抗逆性[J]. 生物技术通报, 2023, 39(11): 150-167. |
| [8] | 唐瑞琪, 赵心清, 朱笃, 汪涯. 大肠杆菌对木质纤维素水解液抑制物的胁迫耐受性[J]. 生物技术通报, 2023, 39(11): 205-216. |
| [9] | 王文韬, 冯颀, 刘晨光, 白凤武, 赵心清. 氧化还原敏感型基因元件增强酵母木质纤维素水解液抑制物胁迫耐受性[J]. 生物技术通报, 2023, 39(11): 360-372. |
| [10] | 刘晓黎, 童真艺, 赵亮, 尹丽, 刘晨光. 非抗生素类活性物质抗幽门螺杆菌研究进展[J]. 生物技术通报, 2022, 38(9): 96-105. |
| [11] | 史亚楠, 王德培, 王一川, 周昊, 薛鲜丽. 敲除msn2对米曲霉生长和发酵产曲酸的影响[J]. 生物技术通报, 2022, 38(8): 188-197. |
| [12] | 刘自然, 甄珍, 陈强, 李玥莹, 王泽, 逄洪波. 植物响应Cd胁迫研究进展[J]. 生物技术通报, 2022, 38(6): 13-26. |
| [13] | 薛鲜丽, 王静然, 毕杭杭, 王德培. 过表达Spt7对黑曲霉生长及抗逆性影响[J]. 生物技术通报, 2022, 38(5): 112-122. |
| [14] | 丁亚群, 丁宁, 谢深民, 黄梦娜, 张昱, 张勤, 姜力. Vps28基因敲除小鼠模型的构建及其对泌乳和免疫性状影响的研究[J]. 生物技术通报, 2022, 38(3): 164-172. |
| [15] | 燕炯, 冯晨毅, 高学坤, 许祥, 杨佳敏, 陈朝阳. 基于CRISPR/Cas9技术构建Plin1基因敲除小鼠模型及表型分析[J]. 生物技术通报, 2022, 38(3): 173-180. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||