








生物技术通报 ›› 2023, Vol. 39 ›› Issue (4): 24-37.doi: 10.13560/j.cnki.biotech.bull.1985.2022-1248
收稿日期:2022-10-10
出版日期:2023-04-26
发布日期:2023-05-16
通讯作者:
李爱涛,男,博士,教授,研究方向:生物催化与酶工程;E-mail: aitaoli@hubu.eu.cn作者简介:郁慧丽,女,博士,副教授,研究方向:P450酶工程与生物催化;E-mail: huiliyu@hubu.edu.cn
基金资助:Received:2022-10-10
Published:2023-04-26
Online:2023-05-16
摘要:
细胞色素P450酶是一类含亚铁血红素的蛋白超家族,来源广泛,几乎存在于所有的生命形式中。P450酶被誉为“黄金催化剂”,主要参与生物体内源物质合成、异源物质代谢及天然产物的生物合成,具有底物谱广、催化反应类型多样、催化立体专一性强等优点,使得它们在药物/毒物代谢、工程化生物合成等方面受到越来越多的关注。其中,P450酶可以实现常温、常压、中性条件下特定惰性C-H键选择性羟化,在高附加值精细化学品的生物合成中具有重要应用价值。本文介绍了P450选择性羟化的催化机制及几种常见的P450电子传递系统,并结合蛋白质工程与代谢工程,以柠檬烯含氧衍生物、诺卡酮、檀香醇、4-羟基异佛尔酮、大环内酯麝香等高附加值化合物的生物合成为例,综述了单酶催化的生物转化或全细胞从头合成的方式在天然香精香料生物合成中的探索与应用,探讨其面临的挑战并展望了其应用前景。
郁慧丽, 李爱涛. 细胞色素P450酶在香精香料绿色生物合成中的应用[J]. 生物技术通报, 2023, 39(4): 24-37.
YU Hui-li, LI Ai-tao. Application of Cytochrome P450 in the Biosynthesis of Flavors and Fragrances[J]. Biotechnology Bulletin, 2023, 39(4): 24-37.
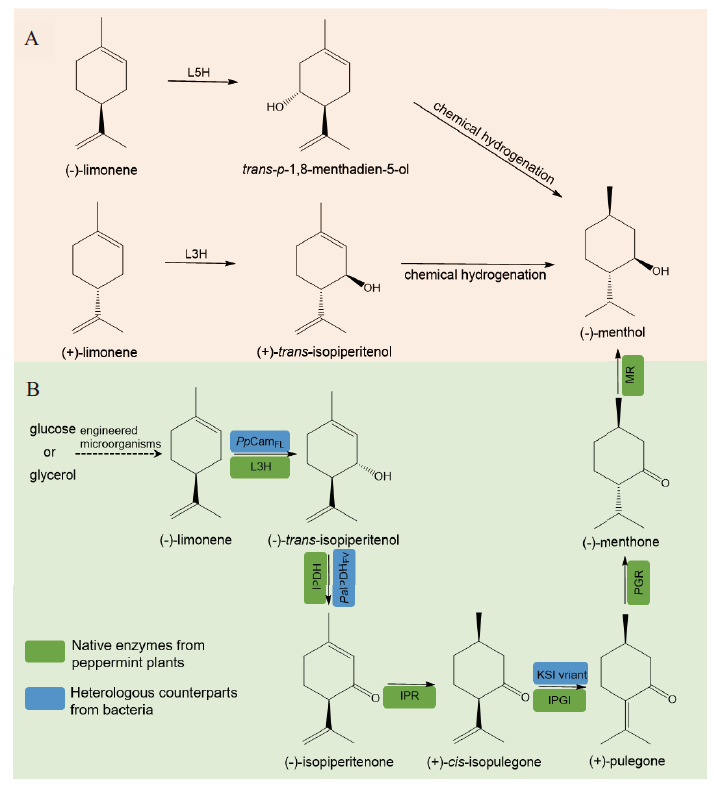
图5 P450参与的生物-化学两步转化(A)及生物法从头合成(-)-薄荷醇(B) L5H:柠檬烯-5-羟化酶;L3H:柠檬烯-3-羟化酶;IPDH:(-)-反式-异薄荷烯醇脱氢酶;IPR:(-)-异薄荷烯酮还原酶;IPGI:(+)-顺式-异胡薄荷酮异构酶;PGR:(+)-胡薄荷酮还原酶;MR:(-)-薄荷酮还原酶;PpCamFL:来源于恶臭假单胞菌的P450cam_Y96F/V247L 突变体;PaIPDHFV:来源于绿脓假单胞菌的IPDH_E95F/Y199V 突变体;KSI variant:来源于恶臭假单胞菌的Δ5-3-甾酮异构酶KSI_V88I/L99V/V101A/D103S 突变体
Fig. 5 P450 involved in the biotechnological-chemical processes(A)versus the de novo biosynthetic pathway for(-)-menthol production L5H: Limonene-5-hydroxylase; L3H: limonene-3-hydroxylase; IPDH:(-)-trans-isopiperitenol dehydrogenase; IPR:(-)-isopiperitenone reductase; IPGI:(+)-cis-isopulegone isomerase; PGR:(+)-pulegone reductase; MR:(-)-menthone reductase; PpCamFL: P450cam_Y96F/V247L variant from Pseudomonas putida; PaIPDHFV: IPDH_E95F/Y199V variant from Pseudomonas aeruginosa; KSI variant: Δ5-3-ketosteroid isomerase KSI_V88I/L99V/V101A/D103S variant from P. putida

图6 P450参与的生物法从头合成S-(-)-紫苏醇 AACT:乙酰CoA酰基转移酶;HMGS:HMG-CoA合酶;HMGR:HMG-CoA还原酶;MVK:甲羟戊酸激酶;PMVK:磷酸甲羟戊酸激酶;MVD:焦磷酸甲羟戊酸脱羧酶;IDI:IPP异构酶;GPPS:GPP合酶;LS:柠檬烯合酶
Fig. 6 P450 involved in the de novo biosynthesis of S-(-)-perillyl alcohol AACT: Acetyl-CoA C-acetyltransferase; HMGS: hydroxymethylglutaryl-CoA synthase; HMGR: hydroxymethylglutaryl-CoA reductase; MVK: mevalonate kinase; PMVK: phosphomevalonate kinase; MVD: diphosphomevalonate decarboxylase; IDI: isopentenyl-diphosphate delta-isomerase; GPPS: GPP synthase; LS: limonene synthase

图7 P450参与的单酶催化法(A)及全细胞从头合成(+)-诺卡酮(B) FPPS:FPP合酶;CnVS:来自阿拉斯加黄柏的瓦伦烯合酶;ADH:醇脱氢酶
Fig. 7 P450 involved in the biotransformation of(+)-valencene(A)and de novo biosynthesis of(+)-nootkatone(B) FPPS: FPP synthase; CnVS: valencene synthase from Chamaecyparis nootkatensis; ADH: alcohol dehydrogenase
| [1] |
Nelson DR, Koymans L, Kamataki T, et al. P450 superfamily: update on new sequences, gene mapping, accession numbers and nomenclature[J]. Pharmacogenetics, 1996, 6(1): 1-42.
doi: 10.1097/00008571-199602000-00002 pmid: 8845856 |
| [2] |
Omura T, Sato R. The carbon monoxide-binding pigment of liver microsomes[J]. J Biol Chem, 1964, 239(7): 2370-2378.
doi: 10.1016/S0021-9258(20)82244-3 URL |
| [3] |
Nelson DR. Cytochrome P450 diversity in the tree of life[J]. Biochim Biophys Acta Proteins Proteom, 2018, 1866(1): 141-154.
doi: 10.1016/j.bbapap.2017.05.003 URL |
| [4] |
Miles CS, Ost TWB, Noble MA, et al. Protein engineering of cytochromes P-450[J]. Biochim Biophys Acta Protein Struct Mol Enzymol, 2000, 1543(2): 383-407.
doi: 10.1016/S0167-4838(00)00236-3 URL |
| [5] | Werck-Reichhart D, Feyereisen R. Cytochromes P450: a success story[J]. Genome Biol, 2000, 1(6): REVIEWS3003. |
| [6] |
Urlacher VB, Eiben S. Cytochrome P450 monooxygenases: perspectives for synthetic application[J]. Trends Biotechnol, 2006, 24(7): 324-330.
doi: 10.1016/j.tibtech.2006.05.002 pmid: 16759725 |
| [7] |
Graham SE, Peterson JA. How similar are P450s and what can their differences teach us?[J]. Arch Biochem Biophys, 1999, 369(1): 24-29.
pmid: 10462437 |
| [8] |
Guengerich FP. Common and uncommon cytochrome P450 reactions related to metabolism and chemical toxicity[J]. Chem Res Toxicol, 2001, 14(6): 611-650.
doi: 10.1021/tx0002583 pmid: 11409933 |
| [9] |
Rittle J, Green MT. Cytochrome P450 compound I: capture, characterization, and C-H bond activation kinetics[J]. Science, 2010, 330(6006): 933-937.
doi: 10.1126/science.1193478 pmid: 21071661 |
| [10] |
Meunier B, de Visser SP, Shaik S. Mechanism of oxidation reactions catalyzed by cytochrome p450 enzymes[J]. Chem Rev, 2004, 104(9): 3947-3980.
doi: 10.1021/cr020443g pmid: 15352783 |
| [11] |
Li Z, Jiang YY, Guengerich FP, et al. Engineering cytochrome P450 enzyme systems for biomedical and biotechnological applications[J]. J Biol Chem, 2020, 295(3): 833-849.
doi: 10.1074/jbc.REV119.008758 pmid: 31811088 |
| [12] |
Whitehouse CJC, Bell SG, Wong LL. P450(BM3)(CYP102A1): connecting the dots[J]. Chem Soc Rev, 2012, 41(3): 1218-1260.
doi: 10.1039/C1CS15192D URL |
| [13] |
Degtyarenko KN, Kulikova TA. Evolution of bioinorganic motifs in P450-containing systems[J]. Biochem Soc Trans, 2001, 29(Pt 2): 139-147.
doi: 10.1042/bst0290139 URL |
| [14] |
Hannemann F, Bichet A, Ewen KM, et al. Cytochrome P450 systems—biological variations of electron transport chains[J]. Biochim Biophys Acta, 2007, 1770(3): 330-344.
pmid: 16978787 |
| [15] |
Sakaki T. Practical application of cytochrome P450[J]. Biol Pharm Bull, 2012, 35(6): 844-849.
pmid: 22687473 |
| [16] | Xu LH, Du YL. Rational and semi-rational engineering of cytochrome P450s for biotechnological applications[J]. Synth Syst Biotechnol, 2018, 3(4): 283-290. |
| [17] |
Roberts GA, Çelik A, Hunter DJB, et al. A self-sufficient cytochrome P450 with a primary structural organization that includes a flavin domain and a[2Fe-2S]redox center[J]. J Biol Chem, 2003, 278(49): 48914-48920.
doi: 10.1074/jbc.M309630200 pmid: 14514666 |
| [18] |
Daiber A, Shoun H, Ullrich V. Nitric oxide reductase(P450nor)from Fusarium oxysporum[J]. J Inorg Biochem, 2005, 99(1): 185-193.
doi: 10.1016/j.jinorgbio.2004.09.018 URL |
| [19] |
Choi KY, Jung E, Jung DH, et al. Engineering of daidzein 3'-hydroxylase P450 enzyme into catalytically self-sufficient cytochrome P450[J]. Microb Cell Fact, 2012, 11: 81.
doi: 10.1186/1475-2859-11-81 |
| [20] |
Li SY, Podust LM, Sherman DH. Engineering and analysis of a self-sufficient biosynthetic cytochrome P450 PikC fused to the RhFRED reductase domain[J]. J Am Chem Soc, 2007, 129(43): 12940-12941.
doi: 10.1021/ja075842d pmid: 17915876 |
| [21] |
Li SY, Du L, Bernhardt R. Redox partners: function modulators of bacterial P450 enzymes[J]. Trends Microbiol, 2020, 28(6): 445-454.
doi: S0966-842X(20)30048-2 pmid: 32396826 |
| [22] |
Coon MJ. Cytochrome P450: nature's most versatile biological catalyst[J]. Annu Rev Pharmacol Toxicol, 2005, 45: 1-25.
pmid: 15832443 |
| [23] |
Guengerich FP, Munro AW. Unusual cytochrome p450 enzymes and reactions[J]. J Biol Chem, 2013, 288(24): 17065-17073.
doi: 10.1074/jbc.R113.462275 pmid: 23632016 |
| [24] |
Lamb DC, Waterman MR. Unusual properties of the cytochrome P450 superfamily[J]. Philos Trans R Soc Lond B Biol Sci, 2013, 368(1612): 20120434.
doi: 10.1098/rstb.2012.0434 URL |
| [25] |
Zhang XW, Li SY. Expansion of chemical space for natural products by uncommon P450 reactions[J]. Nat Prod Rep, 2017, 34(9): 1061-1089.
doi: 10.1039/c7np00028f pmid: 28770915 |
| [26] |
Podust LM, Sherman DH. Diversity of P450 enzymes in the biosynthesis of natural products[J]. Nat Prod Rep, 2012, 29(10): 1251-1266.
doi: 10.1039/c2np20020a pmid: 22820933 |
| [27] |
Rudolf JD, Chang CY, Ma M, et al. Cytochromes P450 for natural product biosynthesis in Streptomyces: sequence, structure, and function[J]. Nat Prod Rep, 2017, 34(9): 1141-1172.
doi: 10.1039/C7NP00034K URL |
| [28] |
Bernhardt R, Urlacher VB. Cytochromes P450 as promising catalysts for biotechnological application: chances and limitations[J]. Appl Microbiol Biotechnol, 2014, 98(14): 6185-6203.
doi: 10.1007/s00253-014-5767-7 pmid: 24848420 |
| [29] |
Yasuda K, Sugimoto H, Hayashi K, et al. Protein engineering of CYP105s for their industrial uses[J]. Biochim Biophys Acta Proteins Proteom, 2018, 1866(1): 23-31.
doi: 10.1016/j.bbapap.2017.05.014 URL |
| [30] |
Paddon CJ, Westfall PJ, Pitera DJ, et al. High-level semi-synthetic production of the potent antimalarial artemisinin[J]. Nature, 2013, 496(7446): 528-532.
doi: 10.1038/nature12051 |
| [31] |
McLean KJ, Hans M, Meijrink B, et al. Single-step fermentative production of the cholesterol-lowering drug pravastatin via reprogramming of Penicillium chrysogenum[J]. Proc Natl Acad Sci USA, 2015, 112(9): 2847-2852.
doi: 10.1073/pnas.1419028112 URL |
| [32] |
Urlacher V B, Girhard M. Cytochrome P450 monooxygenases: an update on perspectives for synthetic application[J]. Trends Biotechnol, 2012, 30(1): 26-36.
doi: 10.1016/j.tibtech.2011.06.012 pmid: 21782265 |
| [33] |
Girvan HM, Munro AW. Applications of microbial cytochrome P450 enzymes in biotechnology and synthetic biology[J]. Curr Opin Chem Biol, 2016, 31: 136-145.
doi: 10.1016/j.cbpa.2016.02.018 pmid: 27015292 |
| [34] |
Wei YF, Ang EL, Zhao HM. Recent developments in the application of P450 based biocatalysts[J]. Curr Opin Chem Biol, 2018, 43: 1-7.
doi: S1367-5931(17)30120-5 pmid: 29100098 |
| [35] |
Kolwek J, Behrens C, Linke D, et al. Cell-free one-pot conversion of(+)-valencene to(+)-nootkatone by a unique dye-decolorizing peroxidase combined with a laccase from Funalia trogii[J]. J Ind Microbiol Biotechnol, 2018, 45(2): 89-101.
doi: 10.1007/s10295-017-1998-9 URL |
| [36] | Surburg H, Panten J. Natural raw materials in the flavor and fragrance industry[M]// Common Fragrance and Flavor Materials. Weinheim, Germany: Wiley-VCH Verlag GmbH & Co. KGaA, 2016: 193-264. |
| [37] |
Rottava I, Cortina PF, Martello E, et al. Optimization of α-terpineol production by the biotransformation of R-(+)-limonene and(-)-β-pinene[J]. Appl Biochem Biotechnol, 2011, 164(4): 514-523.
doi: 10.1007/s12010-010-9153-3 pmid: 21234702 |
| [38] |
Croteau RB, Davis EM, Ringer KL, et al. (-)-Menthol biosynthesis and molecular genetics[J]. Naturwissenschaften, 2005, 92(12): 562-577.
doi: 10.1007/s00114-005-0055-0 URL |
| [39] |
Kamatou GPP, Vermaak I, Viljoen AM, et al. Menthol: a simple monoterpene with remarkable biological properties[J]. Phytochemistry, 2013, 96: 15-25.
doi: 10.1016/j.phytochem.2013.08.005 pmid: 24054028 |
| [40] | Schempp FM, Strobel I, Etschmann MMW, et al. Identification of fungal limonene-3-hydroxylase for biotechnological menthol production[J]. Appl Environ Microbiol, 2021, 87(10): e02873-e02820. |
| [41] |
Lupien S, Karp F, Wildung M, et al. Regiospecific cytochrome P450 limonene hydroxylases from mint(Mentha)species: cDNA isolation, characterization, and functional expression of(-)-4S-limonene-3-hydroxylase and(-)-4S-limonene-6-hydroxylase[J]. Arch Biochem Biophys, 1999, 368(1): 181-192.
doi: 10.1006/abbi.1999.1298 pmid: 10415126 |
| [42] |
Schalk M, Croteau R. A single amino acid substitution(F363I)converts the regiochemistry of the spearmint(-)-limonene hydroxylase from a C6- to a C3-hydroxylase[J]. Proc Natl Acad Sci USA, 2000, 97(22): 11948-11953.
doi: 10.1073/pnas.97.22.11948 pmid: 11050228 |
| [43] | Bell SG, Sowden RJ, Wong LL. Engineering the haem monooxygenase cytochrome P450cam for monoterpene oxidation[J]. Chem Commun, 2001(7): 635-636. |
| [44] | Shou C, Zheng YC, Zhan JR, et al. Removing the obstacle to(-)-menthol biosynthesis by building a microbial cell factory of(+)- cis-isopulegone from(-)-limonene[J]. ChemSusChem, 2022, 15(9): e202101741. |
| [45] |
Currin A, Dunstan MS, Johannissen LO, et al. Engineering the “missing link” in biosynthetic(-)-menthol production: bacterial isopulegone isomerase[J]. ACS Catal, 2018, 8(3): 2012-2020.
doi: 10.1021/acscatal.7b04115 URL |
| [46] |
Willrodt C, Hoschek A, Bühler B, et al. Coupling limonene formation and oxyfunctionalization by mixed-culture resting cell fermentation[J]. Biotechnol Bioeng, 2015, 112(9): 1738-1750.
doi: 10.1002/bit.25592 pmid: 25786991 |
| [47] |
Zhang X, Liu X, Meng Y, et al. Combinatorial engineering of Saccharomyces cerevisiae for improving limonene production[J]. Biochem Eng J, 2021, 176: 108155.
doi: 10.1016/j.bej.2021.108155 URL |
| [48] |
Dusséaux S, Wajn WT, Liu YX, et al. Transforming yeast peroxisomes into microfactories for the efficient production of high-value isoprenoids[J]. Proc Natl Acad Sci USA, 2020, 117(50): 31789-31799.
doi: 10.1073/pnas.2013968117 pmid: 33268495 |
| [49] |
Cao X, Lv YB, Chen J, et al. Metabolic engineering of oleaginous yeast Yarrowia lipolytica for limonene overproduction[J]. Biotechnol Biofuels, 2016, 9: 214.
doi: 10.1186/s13068-016-0626-7 URL |
| [50] |
Pina LTS, Serafini MR, Oliveira MA, et al. Carvone and its pharmacological activities: a systematic review[J]. Phytochemistry, 2022, 196: 113080.
doi: 10.1016/j.phytochem.2021.113080 URL |
| [51] |
Morrish JLE, Brennan ET, Dry HC, et al. Enhanced bioproduction of carvone in a two-liquid-phase partitioning bioreactor with a highly hydrophobic biocatalyst[J]. Biotechnol Bioeng, 2008, 101(4): 768-775.
doi: 10.1002/bit.21941 pmid: 18478563 |
| [52] |
Davis EM, Ringer KL, McConkey ME, et al. Monoterpene metabolism. Cloning, expression, and characterization of menthone reductases from peppermint[J]. Plant Physiol, 2005, 137(3): 873-881.
doi: 10.1104/pp.104.053306 pmid: 15728344 |
| [53] |
Yoshida E, Kojima M, Suzuki M, et al. Increased carvone production in Escherichia coli by balancing limonene conversion enzyme expression via targeted quantification concatamer proteome analysis[J]. Sci Rep, 2021, 11(1): 22126.
doi: 10.1038/s41598-021-01469-y pmid: 34764337 |
| [54] |
Zhang LL, Fan G, Li X, et al. Identification of functional genes associated with the biotransformation of limonene to trans-dihydrocarvone in Klebsiella sp. O852[J]. J Sci Food Agric, 2022, 102(8): 3297-3307.
doi: 10.1002/jsfa.v102.8 URL |
| [55] |
Sun C, Dong XJ, Zhang RB, et al. Effectiveness of recombinant Escherichia coli on the production of(R)-(+)-perillyl alcohol[J]. BMC Biotechnol, 2021, 21: 3.
doi: 10.1186/s12896-020-00662-7 |
| [56] |
Mau CJD, Karp F, Ito M, et al. A candidate cDNA clone for(-)-limonene-7-hydroxylase from Perilla frutescens[J]. Phytochemistry, 2010, 71(4): 373-379.
doi: 10.1016/j.phytochem.2009.12.002 URL |
| [57] |
Fujiwara Y, Ito M. Molecular cloning and characterization of a Perilla frutescens cytochrome P450 enzyme that catalyzes the later steps of perillaldehyde biosynthesis[J]. Phytochemistry, 2017, 134: 26-37.
doi: S0031-9422(16)30259-X pmid: 27890582 |
| [58] |
van Beilen JB, Holtackers R, Lüscher D, et al. Biocatalytic production of perillyl alcohol from limonene by using a novel Mycobacterium sp. cytochrome P450 alkane hydroxylase expressed in Pseudomonas putida[J]. Appl Environ Microbiol, 2005, 71(4): 1737-1744.
doi: 10.1128/AEM.71.4.1737-1744.2005 URL |
| [59] |
Cornelissen S, Liu SS, Deshmukh AT, et al. Cell physiology rather than enzyme kinetics can determine the efficiency of cytochrome P450-catalyzed C-H-oxyfunctionalization[J]. J Ind Microbiol Biotechnol, 2011, 38(9): 1359-1370.
doi: 10.1007/s10295-010-0919-y URL |
| [60] |
Cornelissen S, Julsing MK, Volmer J, et al. Whole-cell-based CYP153A6-catalyzed(S)-limonene hydroxylation efficiency depends on host background and profits from monoterpene uptake via AlkL[J]. Biotechnol Bioeng, 2013, 110(5): 1282-1292.
doi: 10.1002/bit.24801 pmid: 23239244 |
| [61] |
Seifert A, Antonovici M, Hauer B, et al. An efficient route to selective bio-oxidation catalysts: an iterative approach comprising modeling, diversification, and screening, based on CYP102A1[J]. Chembiochem, 2011, 12(9): 1346-1351.
doi: 10.1002/cbic.201100067 pmid: 21591046 |
| [62] |
Alonso-Gutierrez J, Chan R, Batth TS, et al. Metabolic engineering of Escherichia coli for limonene and perillyl alcohol production[J]. Metab Eng, 2013, 19: 33-41.
doi: 10.1016/j.ymben.2013.05.004 pmid: 23727191 |
| [63] |
Bicas JL, Barros FF, Wagner R, et al. Optimization of R-(+)-α-terpineol production by the biotransformation of R-(+)-limonene[J]. J Ind Microbiol Biotechnol, 2008, 35(9): 1061-1070.
doi: 10.1007/s10295-008-0383-0 URL |
| [64] |
Tai YN, Xu M, Ren JN, et al. Optimisation of α-terpineol production by limonene biotransformation using Penicillium digitatum DSM 62840[J]. J Sci Food Agric, 2016, 96(3): 954-961.
doi: 10.1002/jsfa.7171 URL |
| [65] |
Molina G, Pessôa MG, Bicas JL, et al. Optimization of limonene biotransformation for the production of bulk amounts of α-terpineol[J]. Bioresour Technol, 2019, 294: 122180.
doi: 10.1016/j.biortech.2019.122180 URL |
| [66] |
Zhang LL, Huang W, Zhang YY, et al. Genomic and transcriptomic study for screening genes involved in the limonene biotransformation of Penicillium digitatum DSM 62840[J]. Front Microbiol, 2020, 11: 744.
doi: 10.3389/fmicb.2020.00744 URL |
| [67] |
Fraatz MA, Berger RG, Zorn H. Nootkatone—a biotechnological challenge[J]. Appl Microbiol Biotechnol, 2009, 83(1): 35-41.
doi: 10.1007/s00253-009-1968-x pmid: 19333595 |
| [68] |
Zhu BC, Henderson G, Chen F, et al. Nootkatone is a repellent for Formosan subterranean termite(Coptotermes formosanus)[J]. J Chem Ecol, 2001, 27(3): 523-531.
pmid: 11441443 |
| [69] |
Clarkson TC, Janich AJ, Sanchez-Vargas I, et al. Nootkatone is an effective repellent against Aedes aegypti and Aedes albopictus[J]. Insects, 2021, 12(5): 386.
doi: 10.3390/insects12050386 URL |
| [70] |
Waltz E. A biotech insect repellent, safe enough to eat[J]. Nat Biotechnol, 2020, 38(12): 1368-1369.
doi: 10.1038/s41587-020-00760-z pmid: 33273737 |
| [71] |
Seifert A, Vomund S, Grohmann K, et al. Rational design of a minimal and highly enriched CYP102A1 mutant library with improved regio-, stereo- and chemoselectivity[J]. Chembiochem, 2009, 10(5): 853-861.
doi: 10.1002/cbic.200800799 pmid: 19222039 |
| [72] |
Girhard M, Machida K, Itoh M, et al. Regioselective biooxidation of(+)-valencene by recombinant E. coli expressing CYP109B1 from Bacillus subtilis in a two-liquid-phase system[J]. Microb Cell Fact, 2009, 8: 36.
doi: 10.1186/1475-2859-8-36 pmid: 19591681 |
| [73] | Kokorin A, Urlacher VB. Artificial fusions between P450 BM3 and an alcohol dehydrogenase for efficient(+)-nootkatone production[J]. ChemBioChem, 2022, 23(12): e202200065. |
| [74] |
Gavira C, Höfer R, Lesot A, et al. Challenges and pitfalls of P450-dependent(+)-valencene bioconversion by Saccharomyces cerevisiae[J]. Metab Eng, 2013, 18: 25-35.
doi: 10.1016/j.ymben.2013.02.003 URL |
| [75] |
Cankar K, van Houwelingen A, Bosch D, et al. A chicory cytochrome P450 mono-oxygenase CYP71AV8 for the oxidation of(+)-valencene[J]. FEBS Lett, 2011, 585(1): 178-182.
doi: 10.1016/j.febslet.2010.11.040 URL |
| [76] |
Cankar K, van Houwelingen A, Goedbloed M, et al. Valencene oxidase CYP706M1 from Alaska cedar(Callitropsis nootkatensis)[J]. FEBS Lett, 2014, 588(6): 1001-1007.
doi: 10.1016/j.febslet.2014.01.061 URL |
| [77] |
Guo X, Sun J, Li D, et al. Heterologous biosynthesis of(+)-nootkatone in unconventional yeast Yarrowia lipolytica[J]. Biochem Eng J, 2018, 137: 125-131.
doi: 10.1016/j.bej.2018.05.023 URL |
| [78] |
Takahashi S, Yeo YS, Zhao YX, et al. Functional characterization of premnaspirodiene oxygenase, a cytochrome P450 catalyzing regio- and stereo-specific hydroxylations of diverse sesquiterpene substrates[J]. J Biol Chem, 2007, 282(43): 31744-31754.
doi: 10.1074/jbc.M703378200 pmid: 17715131 |
| [79] |
Wriessnegger T, Augustin P, Engleder M, et al. Production of the sesquiterpenoid(+)-nootkatone by metabolic engineering of Pichia pastoris[J]. Metab Eng, 2014, 24: 18-29.
doi: 10.1016/j.ymben.2014.04.001 pmid: 24747046 |
| [80] |
Cha YP, Li W, Wu T, et al. Probing the synergistic ratio of P450/CPR to improve(+)-nootkatone production in Saccharomyces cerevisiae[J]. J Agric Food Chem, 2022, 70(3): 815-825.
doi: 10.1021/acs.jafc.1c07035 URL |
| [81] |
Ye Z, Huang Y, Shi B, et al. Coupling cell growth and biochemical pathway induction in Saccharomyces cerevisiae for production of(+)-valencene and its chemical conversion to(+)-nootkatone[J]. Metab Eng, 2022, 72: 107-115.
doi: 10.1016/j.ymben.2022.03.005 URL |
| [82] |
Zha WL, An TY, Li T, et al. Reconstruction of the biosynthetic pathway of santalols under control of the GAL regulatory system in yeast[J]. ACS Synth Biol, 2020, 9(2): 449-456.
doi: 10.1021/acssynbio.9b00479 pmid: 31940436 |
| [83] |
Diaz-Chavez ML, Moniodis J, Madilao LL, et al. Biosynthesis of sandalwood oil: Santalum album CYP76F cytochromes P450 produce santalols and bergamotol[J]. PLoS One, 2013, 8(9): e75053.
doi: 10.1371/journal.pone.0075053 URL |
| [84] |
Celedon JM, Chiang A, Yuen MMS, et al. Heartwood-specific transcriptome and metabolite signatures of tropical sandalwood(Santalum album)reveal the final step of(Z)-santalol fragrance biosynthesis[J]. Plant J, 2016, 86(4): 289-299.
doi: 10.1111/tpj.2016.86.issue-4 URL |
| [85] |
Jiang LH, Dong C, Liu TF, et al. Improved functional expression of cytochrome P450s in Saccharomyces cerevisiae through screening a cDNA library from Arabidopsis thaliana[J]. Front Bioeng Biotechnol, 2021, 9: 764851.
doi: 10.3389/fbioe.2021.764851 URL |
| [86] |
Zha WL, Zhang F, Shao JQ, et al. Rationally engineering santalene synthase to readjust the component ratio of sandalwood oil[J]. Nat Commun, 2022, 13(1): 2508.
doi: 10.1038/s41467-022-30294-8 pmid: 35523896 |
| [87] |
Eggersdorfer M, Laudert D, Létinois U, et al. One hundred years of vitamins-a success story of the natural sciences[J]. Angew Chem Int Ed Engl, 2012, 51(52): 12960-12990.
doi: 10.1002/anie.201205886 URL |
| [88] |
Aranda C, Municoy M, Guallar V, et al. Selective synthesis of 4-hydroxyisophorone and 4-ketoisophorone by fungal peroxygenases[J]. Catal Sci Technol, 2019, 9(6): 1398-1405.
doi: 10.1039/C8CY02114G URL |
| [89] |
Zhong W, Mao L, Xu Q, et al. Allylic oxidation of α-isophorone to keto-isophorone with molecular oxygen catalyzed by copper chloride in acetylacetone[J]. Appl Catal A Gen, 2014, 486: 193-200.
doi: 10.1016/j.apcata.2014.08.005 URL |
| [90] |
Hennig M, Püntener K, Scalone M. Synthesis of(R)- and(S)-4-hydroxyisophorone by ruthenium-catalyzed asymmetric transfer hydrogenation of ketoisophorone[J]. Tetrahedron Asymmetry, 2000, 11(9): 1849-1858.
doi: 10.1016/S0957-4166(00)00141-5 URL |
| [91] |
Kaluzna I, Schmitges T, Straatman H, et al. Enabling selective and sustainable P450 oxygenation technology. production of 4-hydroxy-α-isophorone on kilogram scale[J]. Org Process Res Dev, 2016, 20(4): 814-819.
doi: 10.1021/acs.oprd.5b00282 URL |
| [92] |
Tavanti M, Parmeggiani F, Castellanos JRG, et al. One-pot biocatalytic double oxidation of α-isophorone for the synthesis of ketoisophorone[J]. ChemCatChem, 2017, 9(17): 3338-3348.
doi: 10.1002/cctc.201700620 URL |
| [93] |
Dezvarei S, Lee JHZ, Bell SG. Stereoselective hydroxylation of isophorone by variants of the cytochromes P450 CYP102A1 and CYP101A1[J]. Enzyme Microb Technol, 2018, 111: 29-37.
doi: 10.1016/j.enzmictec.2018.01.002 URL |
| [94] |
Bell SG, Chen XH, Sowden RJ, et al. Molecular recognition in(+)-α-pinene oxidation by cytochrome P450cam[J]. J Am Chem Soc, 2003, 125(3): 705-714.
doi: 10.1021/ja028460a URL |
| [95] |
Meng SQ, Guo J, Nie KL, et al. Chemoenzymatic synthesis of fragrance compounds from stearic acid[J]. Chembiochem, 2019, 20(17): 2232-2235.
doi: 10.1002/cbic.201900210 pmid: 30983113 |
| [96] |
Manning J, Tavanti M, Porter JL, et al. Regio- and enantio-selective chemo-enzymatic C-H-lactonization of decanoic acid to(S)-δ-decalactone[J]. Angew Chem Int Ed Engl, 2019, 58(17): 5668-5671.
doi: 10.1002/anie.v58.17 URL |
| [1] | 薛宁, 王瑾, 李世新, 刘叶, 程海娇, 张玥, 毛雨丰, 王猛. 多基因同步调控结合高通量筛选构建高产L-苯丙氨酸的谷氨酸棒杆菌工程菌株[J]. 生物技术通报, 2023, 39(9): 268-280. |
| [2] | 程亚楠, 张文聪, 周圆, 孙雪, 李玉, 李庆刚. 乳酸乳球菌生产2'-岩藻糖基乳糖的途径构建及发酵培养基优化[J]. 生物技术通报, 2023, 39(9): 84-96. |
| [3] | 赵思佳, 王晓璐, 孙纪录, 田健, 张杰. 代谢工程改造毕赤酵母生产赤藓糖醇[J]. 生物技术通报, 2023, 39(8): 137-147. |
| [4] | 叶云芳, 田清尹, 施婷婷, 王亮, 岳远征, 杨秀莲, 王良桂. 植物中β-紫罗兰酮生物合成及调控研究进展[J]. 生物技术通报, 2023, 39(8): 91-105. |
| [5] | 王玲, 卓燊, 付学森, 刘紫璇, 刘笑蓉, 王志辉, 周日宝, 刘湘丹. 莲生物碱生物合成途径及相关基因研究进展[J]. 生物技术通报, 2023, 39(7): 56-66. |
| [6] | 李雨真, 梅天秀, 李治文, 王淇, 李俊, 邹岳, 赵心清. 红酵母基因组和代谢工程改造研究进展[J]. 生物技术通报, 2023, 39(7): 67-79. |
| [7] | 姜晴春, 杜洁, 王嘉诚, 余知和, 王允, 柳忠玉. 虎杖转录因子PcMYB2的表达特性和功能分析[J]. 生物技术通报, 2023, 39(5): 217-223. |
| [8] | 周定定, 李辉虎, 汤兴涌, 余发新, 孔丹宇, 刘毅. 甘草酸和甘草苷生物合成与调控的研究进展[J]. 生物技术通报, 2023, 39(5): 44-53. |
| [9] | 张岩峰, 叶丽丹, 于洪巍. 氧化还原伴侣工程:P450低效问题的解决方案之一[J]. 生物技术通报, 2023, 39(4): 10-23. |
| [10] | 王慕镪, 陈琦, 马薇, 李春秀, 欧阳鹏飞, 许建和. 机器学习方法在酶定向进化中的应用进展[J]. 生物技术通报, 2023, 39(4): 38-48. |
| [11] | 魏倩, 刘小宁, 赵洁. 2-十三烷酮胁迫下棉铃虫FoxAl调控CYP6B6的表达[J]. 生物技术通报, 2022, 38(5): 84-92. |
| [12] | 李毅丹, 单晓辉. 赤霉素代谢调控与绿色革命[J]. 生物技术通报, 2022, 38(2): 195-204. |
| [13] | 邱益彬, 马艳琴, 沙媛媛, 朱逸凡, 苏二正, 雷鹏, 李莎, 徐虹. 解淀粉芽孢杆菌分子遗传操作及其应用研究进展[J]. 生物技术通报, 2022, 38(2): 205-217. |
| [14] | 马艳琴, 邱益彬, 李莎, 徐虹. 透明质酸的生物合成及其代谢工程的研究进展[J]. 生物技术通报, 2022, 38(2): 252-262. |
| [15] | 姚宇, 顾佳珺, 孙超, 申国安, 郭宝林. 植物类黄酮UDP-糖基转移酶研究进展[J]. 生物技术通报, 2022, 38(12): 47-57. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||
