








生物技术通报 ›› 2023, Vol. 39 ›› Issue (6): 88-101.doi: 10.13560/j.cnki.biotech.bull.1985.2022-1329
收稿日期:2022-10-27
出版日期:2023-06-26
发布日期:2023-07-07
通讯作者:
李陇平,男,博士,讲师,研究方向:动物营养与牛羊噬菌体;E-mail: llp_315@163.com作者简介:李托,女,硕士,讲师,研究方向:动物营养与健康养殖;E-mail: lituo0105@126.com
基金资助:
LI Tuo1,2( ), LI Long-ping1,2(
), LI Long-ping1,2( ), QU Lei1,2
), QU Lei1,2
Received:2022-10-27
Published:2023-06-26
Online:2023-07-07
摘要:
随着“超级耐药”细菌的不断出现和快速传播,噬菌体(细菌病毒)成为抗生素替代品研究的热点,是解决抗生素耐药难题、促进养殖业健康发展的新途径。噬菌体能够特异性裂解宿主菌是其发挥治疗功效的关键,然而噬菌体裂解细菌的特异性又取决于噬菌体受体结合蛋白与受体的识别与吸附。有尾噬菌体利用其受体结合蛋白(尾部纤维、尾钉和基板结构等)识别细菌表面受体(脂多糖、外膜蛋白、荚膜、鞭毛和菌毛等),最终将细菌裂解。本文综述了有尾噬菌体及其受体结合蛋白的类型和结构,以及噬菌体受体方面的研究进展,讨论了基于噬菌体与宿主菌互作研究基础上的噬菌体治疗制剂的选择策略,为后续深入研究噬菌体与其宿主菌互作机理、改造噬菌体和创制噬菌体生物杀菌制剂提供坚实的理论基础。
李托, 李陇平, 屈雷. 有尾噬菌体的结构及其受体研究进展[J]. 生物技术通报, 2023, 39(6): 88-101.
LI Tuo, LI Long-ping, QU Lei. Research Progress in the Structure of Tailed Bacteriophage and Its Receptors[J]. Biotechnology Bulletin, 2023, 39(6): 88-101.
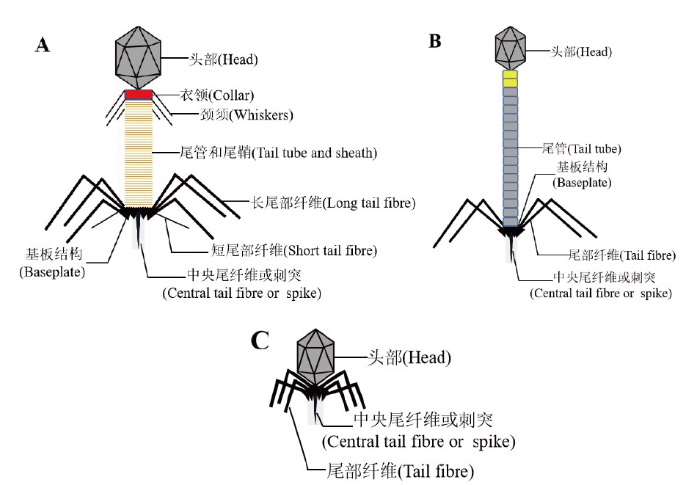
图1 有尾噬菌体的3种典型代表结构 A:肌尾科噬菌体;B:长尾科噬菌体;C:短尾科噬菌体
Fig. 1 Three typical representative structures of tailed bacteriophages A: Myophages; B:Siphophages; C: Podophages
| 噬菌体类型 Phage type | 特征 Features | 噬菌体 Phage species | 宿主细菌 Host bacterium | 噬菌体受体结合蛋白 Receptor-binding protein | 受体 Receptor | 参考文献 References |
|---|---|---|---|---|---|---|
| 肌尾科噬菌体 Myoviridae | 具有长的可收缩尾巴 Long contractile tail | T2、T4 | 大肠杆菌Escherichia coli | 尾部纤维 | OmpC、OmpF | [ |
| SPN3US | 沙门氏菌Salmonella enterica | 尾部纤维 | 鞭毛 | [ | ||
| MAM1 | 沙雷氏菌属、克鲁维菌属 Enterobacterial genera Serratia and Kluyvera | 尾钉 | 荚膜 | [ | ||
| Det7 | 沙门氏菌S. enterica | 尾钉 | 脂多糖 | [ | ||
| JG004 | 铜绿假单胞菌 Pseudomonas aeruginosa | - | 脂多糖 | [ | ||
| 长尾科噬菌体 Siphoviridae | 具有长的不可收缩尾巴 Long non-contractile tail | T5 | 大肠杆菌E. coli | 尾部纤维 | 脂多糖O抗原 | [ |
| SSU5 | 沙门氏菌S. enterica | 尾部纤维 | 脂多糖核心多糖 | [ | ||
| λ、TLS、H8、phi2013 | 大肠杆菌 E. coli | 尾部纤维 | LamB、TolC、FepA、FhuA | [ | ||
| iEPS5 | 沙门氏菌S. enterica | 尾部纤维 | 鞭毛 | [ | ||
| DMS3 | 铜绿假单胞菌P. aeruginosa | 尾部纤维 | 菌毛 | [ | ||
| Vi-II | 沙门氏菌S. enterica | 尾钉 | 荚膜 | [ | ||
| 9NA | 沙门氏菌S. enterica | 尾钉 | 脂多糖 | [ | ||
| phiChi13、phiCbK | Caulobacter crescentus | 衣壳 | 鞭毛 | [ | ||
| SPC35 | 沙门氏菌S. enterica | 尾部纤维 | BtuB | [ | ||
| SPN7C、9C、10H、12C、14、17T、18 | 沙门氏菌S. enterica | - | 鞭毛Flagella、BtuB、脂多糖O抗原 | [ | ||
| 短尾科噬菌体 Podoviridae | 具有短的可收缩尾巴 Short contractile tail | T3、T4、T7 | 大肠杆菌E. coli | 尾部纤维 | LPS, OmpC | [ |
| Bp7 | 大肠杆菌E. coli | - | LamB、OmpC、HepI | [ | ||
| Yep-phi | 鼠疫耶尔氏菌Yersinia pestis | 尾部纤维 | Ail、OmpF | [ | ||
| MPK7 | 铜绿假单胞菌P. aeruginosa | 尾部纤维 | IV型菌毛 | [ | ||
| phiK1-K5 | 大肠杆菌E. coli | 尾钉 | 荚膜 | [ | ||
| P22 | 沙门氏菌S. enterica | 尾钉 | 脂多糖O抗原 | [ | ||
| Sf6 | 志贺氏杆菌Shigella flexneri | 尾钉 | OmpA | [ |
表1 革兰氏阴性菌噬菌体受体结合蛋白与受体类型
Table 1 Gram-negative bacteria-specific bacteriophage receptor binding proteins and receptors
| 噬菌体类型 Phage type | 特征 Features | 噬菌体 Phage species | 宿主细菌 Host bacterium | 噬菌体受体结合蛋白 Receptor-binding protein | 受体 Receptor | 参考文献 References |
|---|---|---|---|---|---|---|
| 肌尾科噬菌体 Myoviridae | 具有长的可收缩尾巴 Long contractile tail | T2、T4 | 大肠杆菌Escherichia coli | 尾部纤维 | OmpC、OmpF | [ |
| SPN3US | 沙门氏菌Salmonella enterica | 尾部纤维 | 鞭毛 | [ | ||
| MAM1 | 沙雷氏菌属、克鲁维菌属 Enterobacterial genera Serratia and Kluyvera | 尾钉 | 荚膜 | [ | ||
| Det7 | 沙门氏菌S. enterica | 尾钉 | 脂多糖 | [ | ||
| JG004 | 铜绿假单胞菌 Pseudomonas aeruginosa | - | 脂多糖 | [ | ||
| 长尾科噬菌体 Siphoviridae | 具有长的不可收缩尾巴 Long non-contractile tail | T5 | 大肠杆菌E. coli | 尾部纤维 | 脂多糖O抗原 | [ |
| SSU5 | 沙门氏菌S. enterica | 尾部纤维 | 脂多糖核心多糖 | [ | ||
| λ、TLS、H8、phi2013 | 大肠杆菌 E. coli | 尾部纤维 | LamB、TolC、FepA、FhuA | [ | ||
| iEPS5 | 沙门氏菌S. enterica | 尾部纤维 | 鞭毛 | [ | ||
| DMS3 | 铜绿假单胞菌P. aeruginosa | 尾部纤维 | 菌毛 | [ | ||
| Vi-II | 沙门氏菌S. enterica | 尾钉 | 荚膜 | [ | ||
| 9NA | 沙门氏菌S. enterica | 尾钉 | 脂多糖 | [ | ||
| phiChi13、phiCbK | Caulobacter crescentus | 衣壳 | 鞭毛 | [ | ||
| SPC35 | 沙门氏菌S. enterica | 尾部纤维 | BtuB | [ | ||
| SPN7C、9C、10H、12C、14、17T、18 | 沙门氏菌S. enterica | - | 鞭毛Flagella、BtuB、脂多糖O抗原 | [ | ||
| 短尾科噬菌体 Podoviridae | 具有短的可收缩尾巴 Short contractile tail | T3、T4、T7 | 大肠杆菌E. coli | 尾部纤维 | LPS, OmpC | [ |
| Bp7 | 大肠杆菌E. coli | - | LamB、OmpC、HepI | [ | ||
| Yep-phi | 鼠疫耶尔氏菌Yersinia pestis | 尾部纤维 | Ail、OmpF | [ | ||
| MPK7 | 铜绿假单胞菌P. aeruginosa | 尾部纤维 | IV型菌毛 | [ | ||
| phiK1-K5 | 大肠杆菌E. coli | 尾钉 | 荚膜 | [ | ||
| P22 | 沙门氏菌S. enterica | 尾钉 | 脂多糖O抗原 | [ | ||
| Sf6 | 志贺氏杆菌Shigella flexneri | 尾钉 | OmpA | [ |
| 噬菌体类型 Phage type | 特征 Features | 噬菌体 Phage species | 宿主细菌 Host bacterium | 噬菌体受体结合蛋白 Receptor-binding protein | 受体 Receptor | 参考文献 References |
|---|---|---|---|---|---|---|
| 肌尾科噬菌体 Myoviridae | 具有长的可收缩尾巴 Long contractile tail | γ | 炭疽杆菌Bacillus anthracis | 尾部纤维 | GamR | [ |
| A511 | 李斯特菌Listeria monocytogenes | 尾部纤维 | 磷壁酸,肽聚糖 | [ | ||
| φ812、φK | 金黄色葡萄球菌 Staphylococcus aureus | - | 磷壁酸 | [ | ||
| 长尾科噬菌体 Siphoviridae | 具有长的不可收缩尾巴 Long non-contractile tail | IL-H | 德氏乳酸杆菌 Lactobacillus delbrueckii | 尾部纤维 | 磷壁酸 | [ |
| SPP1 | 枯草芽孢杆菌B. subtilis | 尾钉 | YueB | [ | ||
| φSLT | 金黄色葡萄球菌 Staphylococcus aureus | 中央尾尖 | 磷壁酸 | [ | ||
| 短尾科噬菌体 Podoviridae | 具有短的可收缩尾巴 Short contractile tail | phi29 | 枯草芽孢杆菌B. subtilis | 尾钉 | 磷壁酸 | [ |
| P2 | 乳酸乳球菌Lactococcus lactis | 尾部纤维 | 细胞壁 | [ |
表2 革兰氏阳性菌噬菌体受体结合蛋白与受体类型
Table 1 Gram-positive bacteria-specific bacteriophage receptor-binding proteins and receptor types
| 噬菌体类型 Phage type | 特征 Features | 噬菌体 Phage species | 宿主细菌 Host bacterium | 噬菌体受体结合蛋白 Receptor-binding protein | 受体 Receptor | 参考文献 References |
|---|---|---|---|---|---|---|
| 肌尾科噬菌体 Myoviridae | 具有长的可收缩尾巴 Long contractile tail | γ | 炭疽杆菌Bacillus anthracis | 尾部纤维 | GamR | [ |
| A511 | 李斯特菌Listeria monocytogenes | 尾部纤维 | 磷壁酸,肽聚糖 | [ | ||
| φ812、φK | 金黄色葡萄球菌 Staphylococcus aureus | - | 磷壁酸 | [ | ||
| 长尾科噬菌体 Siphoviridae | 具有长的不可收缩尾巴 Long non-contractile tail | IL-H | 德氏乳酸杆菌 Lactobacillus delbrueckii | 尾部纤维 | 磷壁酸 | [ |
| SPP1 | 枯草芽孢杆菌B. subtilis | 尾钉 | YueB | [ | ||
| φSLT | 金黄色葡萄球菌 Staphylococcus aureus | 中央尾尖 | 磷壁酸 | [ | ||
| 短尾科噬菌体 Podoviridae | 具有短的可收缩尾巴 Short contractile tail | phi29 | 枯草芽孢杆菌B. subtilis | 尾钉 | 磷壁酸 | [ |
| P2 | 乳酸乳球菌Lactococcus lactis | 尾部纤维 | 细胞壁 | [ |
| [14] |
Dobbins AT, George M Jr, Basham DA, et al. Complete genomic sequence of the virulent Salmonella bacteriophage SP6[J]. J Bacteriol, 2004, 186(7): 1933-1944.
doi: 10.1128/JB.186.7.1933-1944.2004 URL |
| [15] |
Walter M, Fiedler C, Grassl R, et al. Structure of the receptor-binding protein of bacteriophage det7: a podoviral tail spike in a myovirus[J]. J Virol, 2008, 82(5): 2265-2273.
doi: 10.1128/JVI.01641-07 pmid: 18077713 |
| [16] |
Olszak T, Shneider MM, Latka A, et al. The O-specific polysaccharide lyase from the phage LKA1 tailspike reduces Pseudomonas virulence[J]. Sci Rep, 2017, 7(1): 16302.
doi: 10.1038/s41598-017-16411-4 |
| [17] | Kutter EM, Skutt-Kakaria K, Blasdel B, et al. Characterization of a ViI-like phage specific to Escherichia coli O157: H7[J]. Virol J, 2011, 8: 430. |
| [18] |
Pickard D, Toribio AL, Petty NK, et al. A conserved acetyl esterase domain targets diverse bacteriophages to the Vi capsular receptor of Salmonella enterica serovar Typhi[J]. J Bacteriol, 2010, 192(21): 5746-5754.
doi: 10.1128/JB.00659-10 pmid: 20817773 |
| [19] |
Cerritelli ME, Wall JS, Simon MN, et al. Stoichiometry and domainal organization of the long tail-fiber of bacteriophage T4: a hinged viral adhesin[J]. J Mol Biol, 1996, 260(5): 767-780.
pmid: 8709154 |
| [20] |
Hashemolhosseini S, Stierhof YD, Hindennach I, et al. Characterization of the helper proteins for the assembly of tail fibers of coliphages T4 and lambda[J]. J Bacteriol, 1996, 178(21): 6258-6265.
pmid: 8892827 |
| [21] |
Bartual SG, Otero JM, Garcia-Doval C, et al. Structure of the bacteriophage T4 long tail fiber receptor-binding tip[J]. Proc Natl Acad Sci USA, 2010, 107(47): 20287-20292.
doi: 10.1073/pnas.1011218107 pmid: 21041684 |
| [22] |
Washizaki A, Yonesaki T, Otsuka Y. Characterization of the interactions between Escherichia coli receptors, LPS and OmpC, and bacteriophage T4 long tail fibers[J]. MicrobiologyOpen, 2016, 5(6): 1003-1015.
doi: 10.1002/mbo3.384 pmid: 27273222 |
| [23] |
van Raaij MJ, Schoehn G, Burda MR, et al. Crystal structure of a heat and protease-stable part of the bacteriophage T4 short tail fibre[J]. J Mol Biol, 2001, 314(5): 1137-1146.
pmid: 11743729 |
| [24] |
Mason WS. Product of T4 gene 12[J]. J Mol Biol, 1972, 66(3): 445-469.
pmid: 4624817 |
| [25] |
Burda MR, Miller S. Folding of coliphage T4 short tail fiber in vitro. Analysing the role of a bacteriophage-encoded chaperone[J]. Eur J Biochem, 1999, 265(2): 771-778.
pmid: 10504409 |
| [26] |
Leiman PG, Shneider MM, Mesyanzhinov VV, et al. Evolution of bacteriophage tails: structure of T4 gene product 10[J]. J Mol Biol, 2006, 358(3): 912-921.
pmid: 16554069 |
| [27] |
Garcia-Doval C, Luque D, Castón JR, et al. Crystallization of the C-terminal domain of the bacteriophage T5 L-shaped fibre[J]. Acta Crystallogr Sect F Struct Biol Cryst Commun, 2013, 69(Pt 12): 1363-1367.
doi: 10.1107/S1744309113028959 pmid: 24316831 |
| [28] |
Garcia-Doval C, van Raaij MJ. Structure of the receptor-binding carboxy-terminal domain of bacteriophage T7 tail fibers[J]. Proc Natl Acad Sci USA, 2012, 109(24): 9390-9395.
doi: 10.1073/pnas.1119719109 pmid: 22645347 |
| [29] |
Veesler D, Cambillau C. A common evolutionary origin for tailed-bacteriophage functional modules and bacterial machineries[J]. Microbiol Mol Biol Rev, 2011, 75(3): 423-433.
doi: 10.1128/MMBR.00014-11 URL |
| [30] |
Pires DP, Oliveira H, Melo LDR, et al. Bacteriophage-encoded depolymerases: their diversity and biotechnological applications[J]. Appl Microbiol Biotechnol, 2016, 100(5): 2141-2151.
doi: 10.1007/s00253-015-7247-0 pmid: 26767986 |
| [31] |
Boulanger P, Jacquot P, Plançon L, et al. Phage T5 straight tail fiber is a multifunctional protein acting as a tape measure and carrying fusogenic and muralytic activities[J]. J Biol Chem, 2008, 283(20): 13556-13564.
doi: 10.1074/jbc.M800052200 pmid: 18348984 |
| [32] |
Cumby N, Reimer K, Mengin-Lecreulx D, et al. The phage tail tape measure protein, an inner membrane protein and a periplasmic chaperone play connected roles in the genome injection process of E. coli phage HK97[J]. Mol Microbiol, 2015, 96(3): 437-447.
doi: 10.1111/mmi.2015.96.issue-3 URL |
| [33] |
Bhardwaj A, Olia AS, Walker-Kopp N, et al. Domain organization and polarity of tail needle GP26 in the portal vertex structure of bacteriophage P22[J]. J Mol Biol, 2007, 371(2): 374-387.
pmid: 17574574 |
| [1] |
Ackermann HW. 5500 Phages examined in the electron microscope[J]. Arch Virol, 2007, 152(2): 227-243.
doi: 10.1007/s00705-006-0849-1 pmid: 17051420 |
| [2] |
Turner D, Kropinski AM, Adriaenssens EM. A roadmap for genome-based phage taxonomy[J]. Viruses, 2021, 13(3): 506.
doi: 10.3390/v13030506 URL |
| [3] |
Leiman PG, Shneider MM. Contractile tail machines of bacteriophages[J]. Adv Exp Med Biol, 2012, 726: 93-114.
doi: 10.1007/978-1-4614-0980-9_5 pmid: 22297511 |
| [4] |
Nobrega FL, Vlot M, de Jonge PA, et al. Targeting mechanisms of tailed bacteriophages[J]. Nat Rev Microbiol, 2018, 16(12): 760-773.
doi: 10.1038/s41579-018-0070-8 pmid: 30104690 |
| [5] |
Cuervo A, Pulido-Cid M, Chagoyen M, et al. Structural characterization of the bacteriophage T7 tail machinery[J]. J Biol Chem, 2013, 288(36): 26290-26299.
doi: 10.1074/jbc.M113.491209 pmid: 23884409 |
| [6] |
González-García VA, Pulido-Cid M, Garcia-Doval C, et al. Conformational changes leading to T7 DNA delivery upon interaction with the bacterial receptor[J]. J Biol Chem, 2015, 290(16): 10038-10044.
doi: 10.1074/jbc.M114.614222 pmid: 25697363 |
| [7] | Hu B, Margolin W, Molineux IJ, et al. Structural remodeling of bacteriophage T4 and host membranes during infection initiation[J]. PNAS, 2015, 112(35): E4919-4928. |
| [8] |
Gordillo Altamirano FL, Barr JJ. Unlocking the next generation of phage therapy: the key is in the receptors[J]. Curr Opin Biotechnol, 2021, 68: 115-123.
doi: 10.1016/j.copbio.2020.10.002 URL |
| [9] |
Trojet SN, Caumont-Sarcos A, Perrody E, et al. The gp38 adhesins of the T4 superfamily: a complex modular determinant of the phage's host specificity[J]. Genome Biol Evol, 2011, 3: 674-686.
doi: 10.1093/gbe/evr059 pmid: 21746838 |
| [10] | Letarov AV, Kulikov EE. Adsorption of bacteriophages on bacterial cells[J]. Biochemistry(Mosc), 2017, 82(13): 1632-1658. |
| [11] |
Steinbacher S, Miller S, Baxa U, et al. Phage P22 tailspike protein: crystal structure of the head-binding domain at 2.3 A, fully refined structure of the endorhamnosidase at 1.56 A resolution, and the molecular basis of O-antigen recognition and cleavage[J]. J Mol Biol, 1997, 267(4): 865-880.
doi: 10.1006/jmbi.1997.0922 pmid: 9135118 |
| [12] |
Andres D, Hanke C, Baxa U, et al. Tailspike interactions with lipopolysaccharide effect DNA ejection from phage P22 particles in vitro[J]. J Biol Chem, 2010, 285(47): 36768-36775.
doi: 10.1074/jbc.M110.169003 URL |
| [13] |
Barbirz S, Müller JJ, Uetrecht C, et al. Crystal structure of Escherichia coli phage HK620 tailspike: podoviral tailspike endoglycosidase modules are evolutionarily related[J]. Mol Microbiol, 2008, 69(2): 303-316.
doi: 10.1111/mmi.2008.69.issue-2 URL |
| [34] |
Olia AS, Casjens S, Cingolani G. Structure of phage P22 cell envelope-penetrating needle[J]. Nat Struct Mol Biol, 2007, 14(12): 1221-1226.
doi: 10.1038/nsmb1317 |
| [35] |
Xiang Y, Morais MC, Cohen DN, et al. Crystal and cryoEM structural studies of a cell wall degrading enzyme in the bacteriophage phi29 tail[J]. Proc Natl Acad Sci USA, 2008, 105(28): 9552-9557.
doi: 10.1073/pnas.0803787105 pmid: 18606992 |
| [36] |
Xu JW, Gui M, Wang DH, et al. The bacteriophage ϕ29 tail possesses a pore-forming loop for cell membrane penetration[J]. Nature, 2016, 534(7608): 544-547.
doi: 10.1038/nature18017 |
| [37] |
Berkane E, Orlik F, Stegmeier JF, et al. Interaction of bacteriophage lambda with its cell surface receptor: an in vitro study of binding of the viral tail protein gpJ to LamB(Maltoporin)[J]. Biochemistry, 2006, 45(8): 2708-2720.
pmid: 16489764 |
| [38] |
Yoichi M, Abe M, Miyanaga K, et al. Alteration of tail fiber protein gp38 enables T2 phage to infect Escherichia coli O157: H7[J]. J Biotechnol, 2005, 115(1): 101-107.
doi: 10.1016/j.jbiotec.2004.08.003 URL |
| [39] |
Lee JH, Shin H, Kim H, et al. Complete genome sequence of Salmonella bacteriophage SPN3US[J]. J Virol, 2011, 85(24): 13470-13471.
doi: 10.1128/JVI.06344-11 URL |
| [40] |
Matilla MA, Salmond GPC. Bacteriophage ϕMAM1, a Viunalike-virus, is a broad-host-range, high-efficiency generalized transducer that infects environmental and clinical isolates of the enterobacterial genera Serratia and Kluyvera[J]. Appl Environ Microbiol, 2014, 80(20): 6446-6457.
doi: 10.1128/AEM.01546-14 URL |
| [41] |
Garbe J, Bunk B, Rohde M, et al. Sequencing and characterization of Pseudomonas aeruginosa phage JG004[J]. BMC Microbiol, 2011, 11: 102.
doi: 10.1186/1471-2180-11-102 |
| [42] |
Heller K, Braun V. Accelerated adsorption of bacteriophage T5 to Escherichia coli F, resulting from reversible tail fiber-lipopolysaccharide binding[J]. J Bacteriol, 1979, 139(1): 32-38.
doi: 10.1128/jb.139.1.32-38.1979 pmid: 378958 |
| [43] |
Kim M, Kim S, Park B, et al. Core lipopolysaccharide-specific phage SSU5 as an auxiliary component of a phage cocktail for Salmonella biocontrol[J]. Appl Environ Microbiol, 2014, 80(3): 1026-1034.
doi: 10.1128/AEM.03494-13 URL |
| [44] |
Rabsch W, Ma L, Wiley G, et al. FepA- and TonB-dependent bacteriophage H8: receptor binding and genomic sequence[J]. J Bacteriol, 2007, 189(15): 5658-5674.
pmid: 17526714 |
| [45] |
Wang J, Hofnung M, Charbit A. The C-terminal portion of the tail fiber protein of bacteriophage lambda is responsible for binding to LamB, its receptor at the surface of Escherichia coli K-12[J]. J Bacteriol, 2000, 182(2): 508-512.
doi: 10.1128/JB.182.2.508-512.2000 pmid: 10629200 |
| [46] |
German GJ, Misra R. The TolC protein of Escherichia coli serves as a cell-surface receptor for the newly characterized TLS bacteriophage[J]. J Mol Biol, 2001, 308(4): 579-585.
pmid: 11350161 |
| [47] |
Li DH, Zhang ZQ, Li YY, et al. Escherichia coli phage phi2013: genomic analysis and receptor identification[J]. Arch Virol, 2022, 167(12): 2689-2702.
doi: 10.1007/s00705-022-05617-1 |
| [48] |
Choi Y, Shin H, Lee JH, et al. Identification and characterization of a novel flagellum-dependent Salmonella-infecting bacteriophage, iEPS5[J]. Appl Environ Microbiol, 2013, 79(16): 4829-4837.
doi: 10.1128/AEM.00706-13 URL |
| [49] |
Budzik JM, Rosche WA, Rietsch A, et al. Isolation and characterization of a generalized transducing phage for Pseudomonas aeruginosa strains PAO1 and PA14[J]. J Bacteriol, 2004, 186(10): 3270-3273.
doi: 10.1128/JB.186.10.3270-3273.2004 URL |
| [50] |
Guerrero-Ferreira RC, Viollier PH, Ely B, et al. Alternative mechanism for bacteriophage adsorption to the motile bacterium Caulobacter crescentus[J]. Proc Natl Acad Sci USA, 2011, 108(24): 9963-9968.
doi: 10.1073/pnas.1012388108 pmid: 21613567 |
| [51] |
Kim M, Ryu S. Spontaneous and transient defence against bacteriophage by phase-variable glucosylation of O-antigen in Salmonella enterica serovar Typhimurium[J]. Mol Microbiol, 2012, 86(2): 411-425.
doi: 10.1111/j.1365-2958.2012.08202.x URL |
| [52] |
Shin H, Lee JH, Kim H, et al. Receptor diversity and host interaction of bacteriophages infecting Salmonella enterica serovar Typhimurium[J]. PLoS One, 2012, 7(8): e43392.
doi: 10.1371/journal.pone.0043392 URL |
| [53] |
Prehm P, Jann B, Jann K, et al. On a bacteriophage T3 and T4 receptor region within the cell wall lipopolysaccharide of Escherichia coli B[J]. J Mol Biol, 1976, 101(2): 277-281.
pmid: 772219 |
| [54] |
Suga A, Kawaguchi M, Yonesaki T, et al. Manipulating interactions between T4 phage long tail fibers and Escherichia coli receptors[J]. Appl Environ Microbiol, 2021, 87(13): e0042321.
doi: 10.1128/AEM.00423-21 URL |
| [55] | Chen PP, Sun HZ, Ren HY, et al. LamB, OmpC, and the core lipopolysaccharide of Escherichia coli K-12 function as receptors of bacteriophage Bp7[J]. J Virol, 2020, 94(12): e00325-e00320. |
| [56] |
Zhao XN, Cui YJ, Yan YF, et al. Outer membrane proteins ail and OmpF of Yersinia pestis are involved in the adsorption of T7-related bacteriophage Yep-phi[J]. J Virol, 2013, 87(22): 12260-12269.
doi: 10.1128/JVI.01948-13 URL |
| [57] | Bae HW, Cho YH. Complete genome sequence of Pseudomonas aeruginosa podophage MPK7, which requires type IV pili for infection[J]. Genome Announc, 2013, 1(5): e00744-e00713. |
| [58] |
Scholl D, Rogers S, Adhya S, et al. Bacteriophage K1-5 encodes two different tail fiber proteins, allowing it to infect and replicate on both K1 and K5 strains of Escherichia coli[J]. J Virol, 2001, 75(6): 2509-2515.
pmid: 11222673 |
| [59] |
Perez GL, Huynh B, Slater M, et al. Transport of phage P22 DNA across the cytoplasmic membrane[J]. J Bacteriol, 2009, 191(1): 135-140.
doi: 10.1128/JB.00778-08 pmid: 18978055 |
| [60] |
Parent KN, Erb ML, Cardone G, et al. OmpA and OmpC are critical host factors for bacteriophage Sf6 entry in Shigella[J]. Mol Microbiol, 2014, 92(1): 47-60.
doi: 10.1111/mmi.2014.92.issue-1 URL |
| [61] |
Davison S, Couture-Tosi E, Candela T, et al. Identification of the Bacillus anthracis(gamma)phage receptor[J]. J Bacteriol, 2005, 187(19): 6742-6749.
doi: 10.1128/JB.187.19.6742-6749.2005 pmid: 16166537 |
| [62] |
Habann M, Leiman PG, Vandersteegen K, et al. Listeria phage A511, a model for the contractile tail machineries of SPO1-related bacteriophages[J]. Mol Microbiol, 2014, 92(1): 84-99.
doi: 10.1111/mmi.2014.92.issue-1 URL |
| [63] |
Xia GQ, Corrigan RM, Winstel V, et al. Wall teichoic acid-dependent adsorption of staphylococcal siphovirus and myovirus[J]. J Bacteriol, 2011, 193(15): 4006-4009.
doi: 10.1128/JB.01412-10 pmid: 21642458 |
| [64] |
Munsch-Alatossava P, Alatossava T. The extracellular phage-host interactions involved in the bacteriophage LL-H infection of Lactobacillus delbrueckii ssp. lactis ATCC 15808[J]. Front Microbiol, 2013, 4: 408.
doi: 10.3389/fmicb.2013.00408 pmid: 24400001 |
| [65] |
Baptista C, Santos MA, São-José C. Phage SPP1 reversible adsorption to Bacillus subtilis cell wall teichoic acids accelerates virus recognition of membrane receptor YueB[J]. J Bacteriol, 2008, 190(14): 4989-4996.
doi: 10.1128/JB.00349-08 URL |
| [66] |
Kaneko J, Narita-Yamada S, Wakabayashi Y, et al. Identification of ORF636 in phage φSLT carrying Panton-Valentine leukocidin genes, acting as an adhesion protein for a poly(glycerophosphate)chain of lipoteichoic acid on the cell surface of Staphylococcus aureus[J]. J Bacteriol, 2009, 191(14): 4674-4680.
doi: 10.1128/JB.01793-08 pmid: 19429614 |
| [67] |
Bebeacua C, Tremblay D, Farenc C, et al. Structure, adsorption to host, and infection mechanism of virulent lactococcal phage p2[J]. J Virol, 2013, 87(22): 12302-12312.
doi: 10.1128/JVI.02033-13 pmid: 24027307 |
| [68] |
Kiljunen S, Datta N, Dentovskaya SV, et al. Identification of the lipopolysaccharide core of Yersinia pestis and Yersinia pseudotuberculosis as the receptor for bacteriophage φA1122[J]. J Bacteriol, 2011, 193(18): 4963-4972.
doi: 10.1128/JB.00339-11 pmid: 21764935 |
| [69] |
Pajunen M, Kiljunen S, Skurnik M. Bacteriophage phiYeO3-12, specific for Yersinia enterocolitica serotype O: 3, is related to coliphages T3 and T7[J]. J Bacteriol, 2000, 182(18): 5114-5120.
doi: 10.1128/JB.182.18.5114-5120.2000 pmid: 10960095 |
| [70] |
Bohm K, Porwollik S, Chu WP, et al. Genes affecting progression of bacteriophage P22 infection in Salmonella identified by transposon and single gene deletion screens[J]. Mol Microbiol, 2018, 108(3): 288-305.
doi: 10.1111/mmi.2018.108.issue-3 URL |
| [71] |
Le S, Yao XY, Lu SG, et al. Chromosomal DNA deletion confers phage resistance to Pseudomonas aeruginosa[J]. Sci Rep, 2014, 4: 4738.
doi: 10.1038/srep04738 |
| [72] |
Xu JL, Zhang JY, Lu X, et al. O antigen is the receptor of Vibrio cholerae serogroup O1 El Tor typing phage VP4[J]. J Bacteriol, 2013, 195(4): 798-806.
doi: 10.1128/JB.01770-12 URL |
| [73] |
Köhler T, Donner V, van Delden C. Lipopolysaccharide as shield and receptor for R-pyocin-mediated killing in Pseudomonas aeruginosa[J]. J Bacteriol, 2010, 192(7): 1921-1928.
doi: 10.1128/JB.01459-09 pmid: 20118263 |
| [74] |
Rakhuba DV, Kolomiets EI, Dey ES, et al. Bacteriophage receptors, mechanisms of phage adsorption and penetration into host cell[J]. Pol J Microbiol, 2010, 59(3): 145-155.
pmid: 21033576 |
| [75] |
Ricci V, Piddock LJV. Exploiting the role of TolC in pathogenicity: identification of a bacteriophage for eradication of Salmonella serovars from poultry[J]. Appl Environ Microbiol, 2010, 76(5): 1704-1706.
doi: 10.1128/AEM.02681-09 URL |
| [76] |
Breyton C, Flayhan A, Gabel F, et al. Assessing the conformational changes of Pb5, the receptor-binding protein of phage T5, upon binding to its Escherichia coli receptor FhuA[J]. J Biol Chem, 2013, 288(42): 30763-30772.
doi: 10.1074/jbc.M113.501536 pmid: 24014030 |
| [77] |
Porcek NB, Parent KN. Key residues of S. flexneri OmpA mediate infection by bacteriophage Sf6[J]. J Mol Biol, 2015, 427(10): 1964-1976.
doi: 10.1016/j.jmb.2015.03.012 URL |
| [78] |
Xu DL, Zhang JY, Liu J, et al. Outer membrane protein OmpW is the receptor for typing phage VP5 in the Vibrio cholerae O1 El Tor biotype[J]. J Virol, 2014, 88(12): 7109-7111.
doi: 10.1128/JVI.03186-13 URL |
| [79] |
Meyer JR, Dobias DT, Weitz JS, et al. Repeatability and contingency in the evolution of a key innovation in phage lambda[J]. Science, 2012, 335(6067): 428-432.
doi: 10.1126/science.1214449 pmid: 22282803 |
| [80] |
Müller JJ, Barbirz S, Heinle K, et al. An intersubunit active site between supercoiled parallel beta helices in the trimeric tailspike endorhamnosidase of Shigella flexneri Phage Sf6[J]. Structure, 2008, 16(5): 766-775.
doi: 10.1016/j.str.2008.01.019 URL |
| [81] |
Koebnik R, Locher KP, van Gelder P. Structure and function of bacterial outer membrane proteins: barrels in a nutshell[J]. Mol Microbiol, 2000, 37(2): 239-253.
doi: 10.1046/j.1365-2958.2000.01983.x pmid: 10931321 |
| [82] |
Li XH, Koç C, Kühner P, et al. An essential role for the baseplate protein Gp45 in phage adsorption to Staphylococcus aureus[J]. Sci Rep, 2016, 6: 26455.
doi: 10.1038/srep26455 |
| [83] |
São-José C, Lhuillier S, Lurz R, et al. The ectodomain of the viral receptor YueB forms a fiber that triggers ejection of bacteriophage SPP1 DNA[J]. J Biol Chem, 2006, 281(17): 11464-11470.
doi: 10.1074/jbc.M513625200 pmid: 16481324 |
| [84] |
Monteville MR, Ardestani B, Geller BL. Lactococcal bacteriophages require a host cell wall carbohydrate and a plasma membrane protein for adsorption and ejection of DNA[J]. Appl Environ Microbiol, 1994, 60(9): 3204-3211.
doi: 10.1128/aem.60.9.3204-3211.1994 URL |
| [85] |
Farenc C, Spinelli S, Vinogradov E, et al. Molecular insights on the recognition of a Lactococcus lactis cell wall pellicle by the phage 1358 receptor binding protein[J]. J Virol, 2014, 88(12): 7005-7015.
doi: 10.1128/JVI.00739-14 URL |
| [86] |
Tzipilevich E, Habusha M, Ben-Yehuda S. Acquisition of phage sensitivity by bacteria through exchange of phage receptors[J]. Cell, 2017, 168(1/2): 186-199.e12.
doi: 10.1016/j.cell.2016.12.003 URL |
| [87] |
Yen JY, Broadway KM, Scharf BE. Minimum requirements of flagellation and motility for infection of Agrobacterium sp. strain H13-3 by flagellotropic bacteriophage 7-7-1[J]. Appl Environ Microbiol, 2012, 78(20): 7216-7222.
doi: 10.1128/AEM.01082-12 URL |
| [88] |
Kim S, Rahman M, Seol SY, et al. Pseudomonas aeruginosa bacteriophage PA1Ø requires type IV pili for infection and shows broad bactericidal and biofilm removal activities[J]. Appl Environ Microbiol, 2012, 78(17): 6380-6385.
doi: 10.1128/AEM.00648-12 URL |
| [89] |
Raimondo LM, Lundh NP, Martinez RJ. Primary adsorption site of phage PBS1: the flagellum of Bacillus subtilis[J]. J Virol, 1968, 2(3): 256-264.
pmid: 4986906 |
| [90] | Evans TJ, Crow MA, Williamson NR, et al. Characterization of a broad-host-range flagellum-dependent phage that mediates high-efficiency generalized transduction in, and between, Serratia and Pantoea[J]. Microbiology(Reading), 2010, 156(Pt 1): 240-247. |
| [91] |
Pate JL, Petzold SJ, Umbreit TH. Two flagellotropic phages and one Pilus-specific phage active against Asticcacaulis biprosthecum[J]. Virology, 1979, 94(1): 24-37.
pmid: 18627889 |
| [92] |
Chibeu A, Ceyssens PJ, Hertveldt K, et al. The adsorption of Pseudomonas aeruginosa bacteriophage phiKMV is dependent on expression regulation of type IV pili genes[J]. FEMS Microbiol Lett, 2009, 296(2): 210-218.
doi: 10.1111/fml.2009.296.issue-2 URL |
| [93] |
Harvey H, Bondy-Denomy J, Marquis H, et al. Pseudomonas aeruginosa defends against phages through type IV Pilus glycosylation[J]. Nat Microbiol, 2018, 3(1): 47-52.
doi: 10.1038/s41564-017-0061-y |
| [94] |
Sørensen MC, van Alphen LB, Harboe A, et al. Bacteriophage F336 recognizes the capsular phosphoramidate modification of Campylobacter jejuni NCTC11168[J]. J Bacteriol, 2011, 193(23): 6742-6749.
doi: 10.1128/JB.05276-11 pmid: 21965558 |
| [95] |
Hsu CR, Lin TL, Pan YJ, et al. Isolation of a bacteriophage specific for a new capsular type of Klebsiella pneumoniae and characterization of its polysaccharide depolymerase[J]. PLoS One, 2013, 8(8): e70092.
doi: 10.1371/journal.pone.0070092 URL |
| [96] |
Scholl D, Adhya S, Merril C. Escherichia coli K1’s capsule is a barrier to bacteriophage T7[J]. Appl Environ Microbiol, 2005, 71(8): 4872-4874.
doi: 10.1128/AEM.71.8.4872-4874.2005 URL |
| [97] |
Oechslin F. Resistance development to bacteriophages occurring during bacteriophage therapy[J]. Viruses, 2018, 10(7): 351.
doi: 10.3390/v10070351 URL |
| [98] | Merabishvili M, Pirnay J-P, De Vos D. Guidelines to compose an ideal bacteriophage cocktail[J]. Bacteriophage Therapy, 2017, 1693: 99-110. |
| [99] |
Chan BK, Abedon ST, Loc-Carrillo C. Phage cocktails and the future of phage therapy[J]. Future Microbiol, 2013, 8(6): 769-783.
doi: 10.2217/fmb.13.47 pmid: 23701332 |
| [100] |
Yang YH, Shen W, Zhong Q, et al. Development of a bacteriophage cocktail to constrain the emergence of phage-resistant Pseudomonas aeruginosa[J]. Front Microbiol, 2020, 11: 327.
doi: 10.3389/fmicb.2020.00327 URL |
| [101] |
Tanji Y, Shimada T, Yoichi M, et al. Toward rational control of Escherichia coli O157: H7 by a phage cocktail[J]. Appl Microbiol Biotechnol, 2004, 64(2): 270-274.
pmid: 13680205 |
| [102] |
Gu JM, Liu XH, Li Y, et al. A method for generation phage cocktail with great therapeutic potential[J]. PLoS One, 2012, 7(3): e31698.
doi: 10.1371/journal.pone.0031698 URL |
| [103] |
Takeuchi I, Osada K, Azam AH, et al. The presence of two receptor-binding proteins contributes to the wide host range of staphylococcal twort-like phages[J]. Appl Environ Microbiol, 2016, 82(19): 5763-5774.
doi: 10.1128/AEM.01385-16 URL |
| [104] |
Bertozzi Silva J, Storms Z, Sauvageau D. Host receptors for bacteriophage adsorption[J]. FEMS Microbiol Lett, 2016, 363(4): fnw002.
doi: 10.1093/femsle/fnw002 URL |
| [105] |
Fernandes S, São-José C. Enzymes and mechanisms employed by tailed bacteriophages to breach the bacterial cell barriers[J]. Viruses, 2018, 10(8): 396.
doi: 10.3390/v10080396 URL |
| [106] |
Stockdale SR, Mahony J, Courtin P, et al. The lactococcal phages Tuc2009 and TP901-1 incorporate two alternate forms of their tail fiber into their virions for infection specialization[J]. J Biol Chem, 2013, 288(8): 5581-5590.
doi: 10.1074/jbc.M112.444901 pmid: 23300085 |
| [107] |
Stamereilers C, LeBlanc L, Yost D, et al. Comparative genomics of 9 novel Paenibacillus larvae bacteriophages[J]. Bacteriophage, 2016, 6(3): e1220349.
doi: 10.1080/21597081.2016.1220349 URL |
| [108] |
Ceyssens PJ, Miroshnikov K, Mattheus W, et al. Comparative analysis of the widespread and conserved PB1-like viruses infecting Pseudomonas aeruginosa[J]. Environ Microbiol, 2009, 11(11): 2874-2883.
doi: 10.1111/emi.2009.11.issue-11 URL |
| [109] |
Garbe J, Wesche A, Bunk B, et al. Characterization of JG024, a Pseudomonas aeruginosa PB1-like broad host range phage under simulated infection conditions[J]. BMC Microbiol, 2010, 10: 301.
doi: 10.1186/1471-2180-10-301 |
| [110] |
Li L, Shukla S, Meilleur F, et al. Neutron crystallographic studies of T4 lysozyme at cryogenic temperature[J]. Protein Sci, 2017, 26(10): 2098-2104.
doi: 10.1002/pro.3231 pmid: 28707382 |
| [111] |
Oliveira H, Pinto G, Oliveira A, et al. Characterization and genomic analyses of two newly isolated Morganella phages define distant members among Tevenvirinae and Autographivirinae subfamilies[J]. Sci Rep, 2017, 7: 46157.
doi: 10.1038/srep46157 pmid: 28387353 |
| [112] |
Koraimann G. Lytic transglycosylases in macromolecular transport systems of Gram-negative bacteria[J]. Cell Mol Life Sci, 2003, 60(11): 2371-2388.
doi: 10.1007/s00018-003-3056-1 pmid: 14625683 |
| [113] |
Majkowska-Skrobek G, Łątka A, Berisio R, et al. Capsule-targeting depolymerase, derived from Klebsiella KP36 phage, as a tool for the development of anti-virulent strategy[J]. Viruses, 2016, 8(12): 324.
doi: 10.3390/v8120324 URL |
| [114] |
Gutiérrez D, Briers Y, Rodríguez-Rubio L, et al. Role of the pre-neck appendage protein(Dpo7)from phage vB_SepiS-phiIPLA7 as an anti-biofilm agent in staphylococcal species[J]. Front Microbiol, 2015, 6: 1315.
doi: 10.3389/fmicb.2015.01315 pmid: 26635776 |
| [115] |
Knecht LE, Veljkovic M, Fieseler L. Diversity and function of phage encoded depolymerases[J]. Front Microbiol, 2020, 10: 2949.
doi: 10.3389/fmicb.2019.02949 URL |
| [116] |
Born Y, Fieseler L, Klumpp J, et al. The tail-associated depolymerase of Erwinia amylovora phage L1 mediates host cell adsorption and enzymatic capsule removal, which can enhance infection by other phage[J]. Environ Microbiol, 2014, 16(7): 2168-2180.
doi: 10.1111/emi.2014.16.issue-7 URL |
| [117] | Gontijo MTP, Jorge GP, Brocchi M. Current status of endolysin-based treatments against gram-negative bacteria[J]. Antibiotics(Basel), 2021, 10(10): 1143. |
| [118] |
Seed KD, Yen M, Shapiro BJ, et al. Evolutionary consequences of intra-patient phage predation on microbial populations[J]. eLife, 2014, 3: e03497.
doi: 10.7554/eLife.03497 URL |
| [119] |
Yen M, Cairns LS, Camilli A. A cocktail of three virulent bacteriophages prevents Vibrio cholerae infection in animal models[J]. Nat Commun, 2017, 8: 14187.
doi: 10.1038/ncomms14187 |
| [120] |
Chan BK, Sistrom M, Wertz JE, et al. Phage selection restores antibiotic sensitivity in MDR Pseudomonas aeruginosa[J]. Sci Rep, 2016, 6: 26717.
doi: 10.1038/srep26717 |
| [121] |
Filippov AA, Sergueev KV, He YX, et al. Bacteriophage-resistant mutants in Yersinia pestis: identification of phage receptors and attenuation for mice[J]. PLoS One, 2011, 6(9): e25486.
doi: 10.1371/journal.pone.0025486 URL |
| [122] |
Capparelli R, Nocerino N, Lanzetta R, et al. Bacteriophage-resistant Staphylococcus aureus mutant confers broad immunity against staphylococcal infection in mice[J]. PLoS One, 2010, 5(7): e11720.
doi: 10.1371/journal.pone.0011720 URL |
| [123] |
Gordillo Altamirano F, Forsyth JH, Patwa R, et al. Bacteriophage-resistant Acinetobacter baumannii are resensitized to antimicrobials[J]. Nat Microbiol, 2021, 6(2): 157-161.
doi: 10.1038/s41564-020-00830-7 pmid: 33432151 |
| [124] |
Lin H, Paff ML, Molineux IJ, et al. Therapeutic application of phage capsule depolymerases against K1, K5, and K30 capsulated E. coli in mice[J]. Front Microbiol, 2017, 8: 2257.
doi: 10.3389/fmicb.2017.02257 URL |
| [125] |
Majkowska-Skrobek G, Latka A, Berisio R, et al. Phage-borne depolymerases decrease Klebsiella pneumoniae resistance to innate defense mechanisms[J]. Front Microbiol, 2018, 9: 2517.
doi: 10.3389/fmicb.2018.02517 pmid: 30405575 |
| [126] |
Kim M, Ryu S. Characterization of a T5-like coliphage, SPC35, and differential development of resistance to SPC35 in Salmonella enterica serovar typhimurium and Escherichia coli[J]. Appl Environ Microbiol, 2011, 77(6): 2042-2050.
doi: 10.1128/AEM.02504-10 URL |
| [127] |
Sampson BA, Gotschlich EC. Elimination of the vitamin B12 uptake or synthesis pathway does not diminish the virulence of Escherichia coli K1 or Salmonella typhimurium in three model systems[J]. Infect Immun, 1992, 60(9): 3518-3522.
doi: 10.1128/iai.60.9.3518-3522.1992 pmid: 1500158 |
| [128] | Schooley RT, Biswas B, Gill JJ, et al. Development and use of personalized bacteriophage-based therapeutic cocktails to treat a patient with a disseminated resistant Acinetobacter baumannii infection[J]. Antimicrob Agents Chemother, 2017, 61(10): e00954-e00917. |
| [129] |
Philipson CW, Voegtly LJ, Lueder MR, et al. Characterizing phage genomes for therapeutic applications[J]. Viruses, 2018, 10(4): 188.
doi: 10.3390/v10040188 URL |
| [130] |
Leite DMC, Brochet X, Resch G, et al. Computational prediction of inter-species relationships through omics data analysis and machine learning[J]. BMC Bioinform, 2018, 19(S14): 420.
doi: 10.1186/s12859-018-2388-7 |
| [1] | 张曼, 张叶卓, 何其邹洪, 鄂一岚, 李晔. 植物细胞壁结构及成像技术研究进展[J]. 生物技术通报, 2023, 39(7): 113-122. |
| [2] | 胡海琳, 徐黎, 李晓旭, 王晨璨, 梅曼, 丁文静, 赵媛媛. 小肽激素调控植物生长发育及逆境生理研究进展[J]. 生物技术通报, 2023, 39(7): 13-25. |
| [3] | 崔学强, 黄昌艳, 邓杰玲, 李先民, 李秀玲, 张自斌. 基于SLAF-seq技术的石斛兰SNP标记开发及亲缘关系分析[J]. 生物技术通报, 2023, 39(6): 141-148. |
| [4] | 杨俊钊, 张新蕊, 赵国柱, 郑菲. 新型GH5家族多结构域纤维素酶的结构与功能研究[J]. 生物技术通报, 2023, 39(4): 71-80. |
| [5] | 周晞雯, 成柯, 朱鸿亮. 植物体内RNA二级结构探测方法的研究进展[J]. 生物技术通报, 2023, 39(2): 51-62. |
| [6] | 陈广霞, 李秀杰, 蒋锡龙, 单雷, 张志昌, 李勃. 植物小分子信号肽参与非生物逆境胁迫应答的研究进展[J]. 生物技术通报, 2023, 39(11): 61-73. |
| [7] | 陈泉冰, 曹伟洁, 李春, 吕波. GH79家族糖苷水解酶分子进化关系和蛋白结构研究[J]. 生物技术通报, 2023, 39(1): 104-114. |
| [8] | 罗皓天, 王龙, 王禹茜, 王月, 李佳祯, 杨梦珂, 张杰, 邓欣, 王红艳. 青狗尾草RNAi途径相关基因的全基因组鉴定和表达分析[J]. 生物技术通报, 2023, 39(1): 175-186. |
| [9] | 石成龙, 汪锡武, 李安琪, 钱森和, 王洲, 赵世光, 刘艳, 薛正莲. ε-聚赖氨酸对阪崎克罗诺杆菌细胞结构与生物被膜形成的影响[J]. 生物技术通报, 2022, 38(9): 147-157. |
| [10] | 李颖, 龙长梅, 蒋标, 韩丽珍. 两株PGPR菌株的花生定殖及对根际细菌群落结构的影响[J]. 生物技术通报, 2022, 38(9): 237-247. |
| [11] | 刘理慧, 储锦华, 隋雨欣, 陈杨, 程古月. 沙门氏菌中主要毒力因子的研究进展[J]. 生物技术通报, 2022, 38(9): 72-83. |
| [12] | 王子夜, 王志刚, 阎爱华. 不同树龄桑根际土壤原生生物群落组成多样性[J]. 生物技术通报, 2022, 38(8): 206-215. |
| [13] | 陈英, 王艺磊, 邹鹏飞. 大黄鱼TRAF6的克隆及表达分析[J]. 生物技术通报, 2022, 38(8): 233-243. |
| [14] | 王子寅, 刘秉儒, 李子豪, 赵晓玉. 荒漠草原柠条灌丛堆不同发育阶段土壤细菌群落结构特征[J]. 生物技术通报, 2022, 38(7): 205-214. |
| [15] | 王晨晨, 张凡丽, 陈珮琪, 翁思瑶, 王慧芳, 崔小娟. 哺乳动物DNA甲基转移酶DNMT1和DNMT3结构与功能的研究进展[J]. 生物技术通报, 2022, 38(7): 31-39. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||