








生物技术通报 ›› 2023, Vol. 39 ›› Issue (11): 360-372.doi: 10.13560/j.cnki.biotech.bull.1985.2023-0858
收稿日期:2023-09-04
出版日期:2023-11-26
发布日期:2023-12-20
通讯作者:
刘晨光,男,博士,副教授,研究方向:微生物代谢工程与生物质炼制;E-mail: cg.liu@sjtu.edu.cn作者简介:王文韬,男,研究方向:微生物电化学和电发酵;E-mail: tenebrae@sjtu.edu.cn
基金资助:
WANG Wen-tao( ), FENG Qi, LIU Chen-guang(
), FENG Qi, LIU Chen-guang( ), BAI Feng-wu, ZHAO Xin-qing
), BAI Feng-wu, ZHAO Xin-qing
Received:2023-09-04
Published:2023-11-26
Online:2023-12-20
摘要:
纤维素乙醇作为一种清洁可再生的绿色能源,具有良好的应用前景。然而酿酒酵母利用木质纤维素原料生产乙醇的发酵过程易受多种抑制物胁迫的影响,因此提高其胁迫耐受性具有重要意义。本研究在细胞内设计了一种氧化还原敏感型基因元件,通过生物传感器Yap1感应胞内氧化还原状态,以调控抗胁迫基因智能表达。首先,分析了Yap1调控的天然内源启动子PTRR1、PTRX2和PMET16对木质纤维素水解液中典型抑制物的响应强度。其次,根据不同胁迫种类组合相应启动子与抗胁迫的效益基因,构建氧化还原敏感型基因元件提高了酿酒酵母的胁迫耐受性。最后,将表现较好的基因元件GP-CTT和GP-ADH串联整合到一起构建了双基因元件系统,在5-HMF和H2O2双重胁迫下细胞的死亡率与野生型相比下降了69.6%。相较于单基因元件GP-CTT,双基因元件整合菌株的比生长速率、葡萄糖消耗速率和乙醇生产速率分别提高了64.2%、60.1%和58.9%,重组菌株过氧化氢酶的酶活力提高了40.2%。本研究通过理性设计氧化还原敏感型基因元件的遗传回路,强化胞内关键抗氧化酶和醛降解途径,系统地提高了酿酒酵母的胁迫耐受性,为动态地提高酵母鲁棒性提供了新的见解。
王文韬, 冯颀, 刘晨光, 白凤武, 赵心清. 氧化还原敏感型基因元件增强酵母木质纤维素水解液抑制物胁迫耐受性[J]. 生物技术通报, 2023, 39(11): 360-372.
WANG Wen-tao, FENG Qi, LIU Chen-guang, BAI Feng-wu, ZHAO Xin-qing. Redox-sensitive Genetic Parts Improve the Tolerance of Yeast to Lignocellulosic Hydrolysate Inhibitors[J]. Biotechnology Bulletin, 2023, 39(11): 360-372.
| 名称 Name | 描述 Description | 来源 Source |
|---|---|---|
| E. coli DH5α | 质粒构建 | 实验室保存 |
| S. cerevisiaeS288C | 宿主细胞 | 购买于ATCC |
| SC-PGK1 | S288C, HO-yEGFP | 实验室保存 |
| SC-TRR1 | S288C, HO-TRR1 | 本工作构建 |
| SC-TRX2 | S288C, HO-TRX2 | 本工作构建 |
| SC-MET16 | S288C, HO-MET16 | 本工作构建 |
| SC-SOD | S288C, HO-SOD | 本工作构建 |
| SC-CTT | S288C, HO-CTT | 本工作构建 |
| SC-IDP | S288C, HO-IDP | 本工作构建 |
| SC-GLR | S288C, HO-GLR | 本工作构建 |
| SC-ADH | S288C, HO-ADH | 本工作构建 |
| SC-GRE | S288C, HO-GRE | 本工作构建 |
| SC-AC | S288C, HO-AC | 本工作构建 |
表1 使用的菌株
Table 1 Strains used
| 名称 Name | 描述 Description | 来源 Source |
|---|---|---|
| E. coli DH5α | 质粒构建 | 实验室保存 |
| S. cerevisiaeS288C | 宿主细胞 | 购买于ATCC |
| SC-PGK1 | S288C, HO-yEGFP | 实验室保存 |
| SC-TRR1 | S288C, HO-TRR1 | 本工作构建 |
| SC-TRX2 | S288C, HO-TRX2 | 本工作构建 |
| SC-MET16 | S288C, HO-MET16 | 本工作构建 |
| SC-SOD | S288C, HO-SOD | 本工作构建 |
| SC-CTT | S288C, HO-CTT | 本工作构建 |
| SC-IDP | S288C, HO-IDP | 本工作构建 |
| SC-GLR | S288C, HO-GLR | 本工作构建 |
| SC-ADH | S288C, HO-ADH | 本工作构建 |
| SC-GRE | S288C, HO-GRE | 本工作构建 |
| SC-AC | S288C, HO-AC | 本工作构建 |
| 名称 Name | 描述 Description | 来源 Source |
|---|---|---|
| GP-SOD | PTRR1-SOD1-TADH1 | 本工作构建 |
| GP-CTT | PTRR1-CTT1-TADH1 | 本工作构建 |
| GP-IDP | PTRR1-IDP1-TADH1 | 本工作构建 |
| GP-GLR | PTRR1-GLR1-TADH1 | 本工作构建 |
| GP-ADH | PMET16-ADH6-TADH1 | 本工作构建 |
| GP-GRE | PMET16-GRE2-TADH1 | 本工作构建 |
| GP-AC | PTRR1-SOD1-TADH1- PMET16-ADH6-TADH1 | 本工作构建 |
表2 使用的基因元件
Table 2 Genetic parts used
| 名称 Name | 描述 Description | 来源 Source |
|---|---|---|
| GP-SOD | PTRR1-SOD1-TADH1 | 本工作构建 |
| GP-CTT | PTRR1-CTT1-TADH1 | 本工作构建 |
| GP-IDP | PTRR1-IDP1-TADH1 | 本工作构建 |
| GP-GLR | PTRR1-GLR1-TADH1 | 本工作构建 |
| GP-ADH | PMET16-ADH6-TADH1 | 本工作构建 |
| GP-GRE | PMET16-GRE2-TADH1 | 本工作构建 |
| GP-AC | PTRR1-SOD1-TADH1- PMET16-ADH6-TADH1 | 本工作构建 |
| 名称 Name | 成分 Ingredient | 培养微生物 Cultured microorganism | 备注 Remark |
|---|---|---|---|
| LB | 5 g/L酵母粉、10 g/L胰蛋白胨和 10 g/L NaCl | 大肠杆菌 E. coli | 固体培养基添加20 g/L琼脂粉 |
| YPD生长 | 10 g/L酵母粉、20 g/L蛋白胨和 20 g/L葡萄糖 | 酿酒酵母 S. cerevisiae | |
| YPD发酵 | 3 g/L酵母粉、4 g/L蛋白胨和 100 g/L葡萄糖 | 酿酒酵母 S. cerevisiae |
表3 使用的培养基及组成成分
Table 3 Names and composition of the media used
| 名称 Name | 成分 Ingredient | 培养微生物 Cultured microorganism | 备注 Remark |
|---|---|---|---|
| LB | 5 g/L酵母粉、10 g/L胰蛋白胨和 10 g/L NaCl | 大肠杆菌 E. coli | 固体培养基添加20 g/L琼脂粉 |
| YPD生长 | 10 g/L酵母粉、20 g/L蛋白胨和 20 g/L葡萄糖 | 酿酒酵母 S. cerevisiae | |
| YPD发酵 | 3 g/L酵母粉、4 g/L蛋白胨和 100 g/L葡萄糖 | 酿酒酵母 S. cerevisiae |
| 启动子 Promoter | 长度 Length/bp | GC含量 GC content/% | Yap1结合位点 Yap1 binding site |
|---|---|---|---|
| PTRR1 | 545 | 39 | 2 |
| PTRX2 | 275 | 38 | 3 |
| PMET16 | 418 | 34 | 3 |
表4 启动子序列分析
Table 4 Sequence analysis of the promoters
| 启动子 Promoter | 长度 Length/bp | GC含量 GC content/% | Yap1结合位点 Yap1 binding site |
|---|---|---|---|
| PTRR1 | 545 | 39 | 2 |
| PTRX2 | 275 | 38 | 3 |
| PMET16 | 418 | 34 | 3 |
| 启动子 Promoter | EL | 抑制物:H2O2Inhibitor: H2O2 | 抑制物:糠醛 Inhibitor: Furfural | 抑制物:5-HMF Inhibitor: 5-HMF | |||
|---|---|---|---|---|---|---|---|
| 最大RS值 Max RS value | 对应浓度 Corresponding concentration | 最大RS值 Max RS value | 对应浓度 Corresponding concentration | 最大RS值 Max RS value | 对应浓度 Corresponding concentration | ||
| PTRR1 | 15.77 | 2.84 | 1 mmol/L | 1.64 | 10 mmol/L | 1.21 | 30 mmol/L |
| PTRX2 | 16.76 | 2.80 | 1 mmol/L | 2.40 | 10 mmol/L | 1.11 | 30 mmol/L |
| PMET16 | 1.26 | 1.27 | 1 mmol/L | 4.19 | 20 mmol/L | 1.54 | 30 mmol/L |
表5 启动子的响应表现
Table 5 Response performances of the promoters
| 启动子 Promoter | EL | 抑制物:H2O2Inhibitor: H2O2 | 抑制物:糠醛 Inhibitor: Furfural | 抑制物:5-HMF Inhibitor: 5-HMF | |||
|---|---|---|---|---|---|---|---|
| 最大RS值 Max RS value | 对应浓度 Corresponding concentration | 最大RS值 Max RS value | 对应浓度 Corresponding concentration | 最大RS值 Max RS value | 对应浓度 Corresponding concentration | ||
| PTRR1 | 15.77 | 2.84 | 1 mmol/L | 1.64 | 10 mmol/L | 1.21 | 30 mmol/L |
| PTRX2 | 16.76 | 2.80 | 1 mmol/L | 2.40 | 10 mmol/L | 1.11 | 30 mmol/L |
| PMET16 | 1.26 | 1.27 | 1 mmol/L | 4.19 | 20 mmol/L | 1.54 | 30 mmol/L |
| 菌株 Strain | 基因元件转录水平变化倍数 Fold-change in transcription level of gene element | 代谢速率变化(相比野生型) Metabolic rate change(Compared to wild type) | |||
|---|---|---|---|---|---|
| 与无胁迫比较 Compared with no stress | 胁迫下相比野生型 Compared to wild type under stress | 葡萄糖消耗速率 Glucose consumption rate/% | 乙醇生产速率 Ethanol production rate/% | 比生长速率 Specific growth rate/% | |
| SC-ADH | 3.84 | 1.12 | 27.23 | 39.99 | 20.46 |
| SC-GRE | 12.28 | 2.10 | 5.85 | 26.04 | -0.16 |
表9 5-HMF胁迫下重组菌株的代谢速率变化
Table 9 Metabolic rate changes in recombinant strains under 5-HMF stress
| 菌株 Strain | 基因元件转录水平变化倍数 Fold-change in transcription level of gene element | 代谢速率变化(相比野生型) Metabolic rate change(Compared to wild type) | |||
|---|---|---|---|---|---|
| 与无胁迫比较 Compared with no stress | 胁迫下相比野生型 Compared to wild type under stress | 葡萄糖消耗速率 Glucose consumption rate/% | 乙醇生产速率 Ethanol production rate/% | 比生长速率 Specific growth rate/% | |
| SC-ADH | 3.84 | 1.12 | 27.23 | 39.99 | 20.46 |
| SC-GRE | 12.28 | 2.10 | 5.85 | 26.04 | -0.16 |
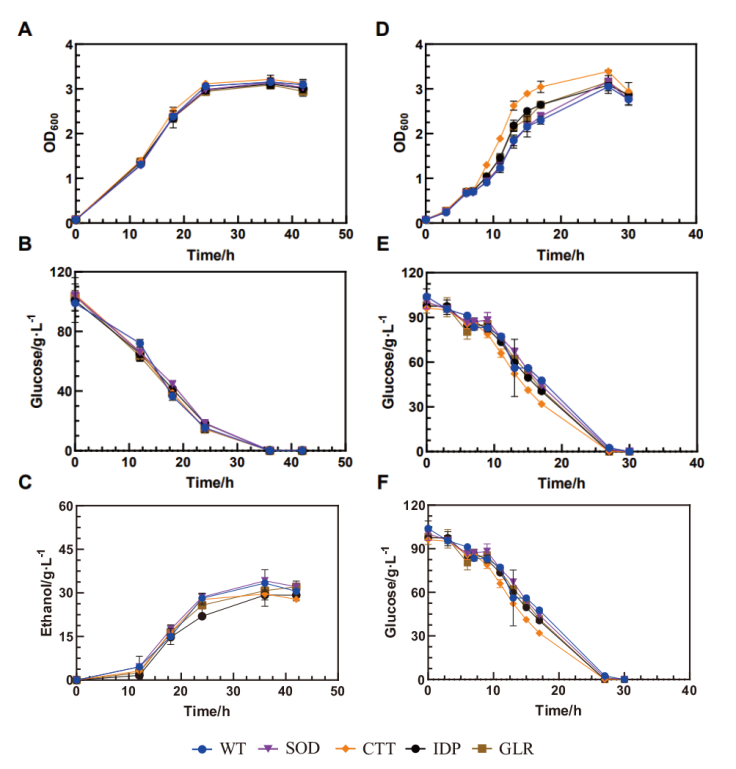
图1 无胁迫与氧化胁迫下重组菌株的生长情况 A:无胁迫下的生长曲线;B:无胁迫下的葡萄糖消耗曲线;C:无胁迫下的乙醇生产曲线;D:氧化胁迫下的生长曲线;E:氧化胁迫下的葡萄糖消耗曲线;F:氧化胁迫下的乙醇生产曲线
Fig. 1 Growths of the recombinant strains under stress-free and oxidative conditions A: Growth under no stress. B: Glucose consumption under no stress. C: Ethanol production under no stress. D: Growth under oxidative condition. E: Glucose consumption under oxidative condition. F: Ethanol production under oxidative condition
| 菌株 Strain | 基因元件转录水平变化倍数 Fold-change in transcription level of gene element | 代谢速率变化(相比野生型) Metabolic rate change(Compared to wild type) | |||||
|---|---|---|---|---|---|---|---|
| 与无胁迫比较 Compared with no stress | 胁迫下相比野生型 Compared to wild type under stress | 葡萄糖消耗速率 Glucose consumption rate | 乙醇生产速率 Ethanol production rate | 比生长速率 Specific growth rate | |||
| SC-CTT | 2.42 | 130 | 30.23% | 32.79% | 32.51% | ||
| SC-SOD | 4.34 | 5.33 | 6.47% | 4.37% | 3.79% | ||
| SC-IDP | 1.34 | 1.13 | 13.68% | 12.02% | 15.12% | ||
| SC-GLR | 1.57 | 6.56 | 11.67% | 16.34% | 15.03% | ||
表6 基因元件转录水平变化以及重组菌株的代谢速率变化
Table 6 Transcriptional level changes of genetic parts and metabolic rate changes in recombinant strains
| 菌株 Strain | 基因元件转录水平变化倍数 Fold-change in transcription level of gene element | 代谢速率变化(相比野生型) Metabolic rate change(Compared to wild type) | |||||
|---|---|---|---|---|---|---|---|
| 与无胁迫比较 Compared with no stress | 胁迫下相比野生型 Compared to wild type under stress | 葡萄糖消耗速率 Glucose consumption rate | 乙醇生产速率 Ethanol production rate | 比生长速率 Specific growth rate | |||
| SC-CTT | 2.42 | 130 | 30.23% | 32.79% | 32.51% | ||
| SC-SOD | 4.34 | 5.33 | 6.47% | 4.37% | 3.79% | ||
| SC-IDP | 1.34 | 1.13 | 13.68% | 12.02% | 15.12% | ||
| SC-GLR | 1.57 | 6.56 | 11.67% | 16.34% | 15.03% | ||
| 菌株 Strain | ROS荧光强度 ROS fluorescence intensity |
|---|---|
| WT | 2017±373 |
| SC-CTT | 1221±48 |
| SC-SOD | 1401±185 |
| SC-IDP | 1428±66 |
| SC-GLR | 1465±104 |
表7 菌株的ROS水平
Table 7 ROS levels in strains
| 菌株 Strain | ROS荧光强度 ROS fluorescence intensity |
|---|---|
| WT | 2017±373 |
| SC-CTT | 1221±48 |
| SC-SOD | 1401±185 |
| SC-IDP | 1428±66 |
| SC-GLR | 1465±104 |
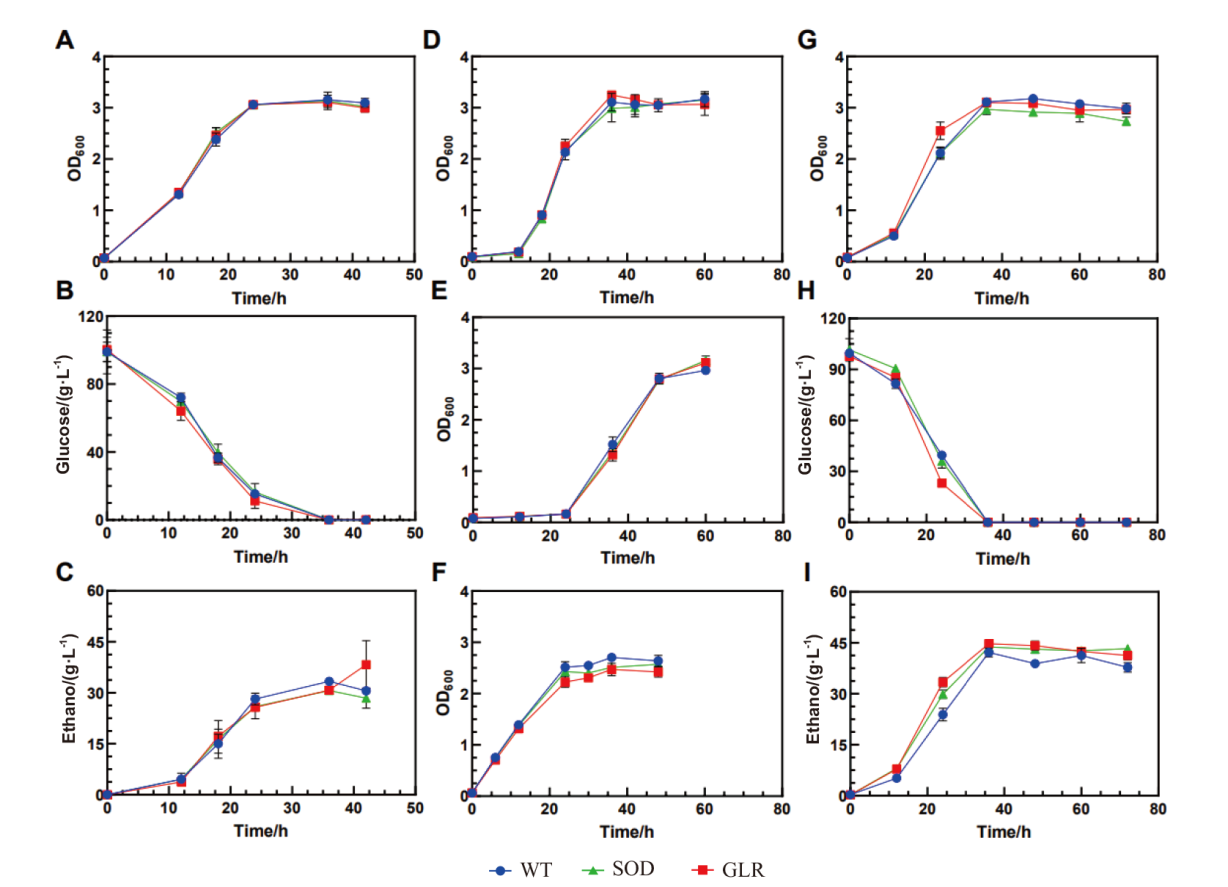
图2 无胁迫、糠醛胁迫以及5-HMF胁迫条件下重组菌株的生长情况 A:无胁迫下生长曲线;B:无胁迫下葡萄糖消耗曲线;C:无胁迫下乙醇生产曲线;D:3 g/L 糠醛胁迫的生长曲线;E:4 g/L 糠醛胁迫的生长曲线;F:4 g/L 糠醛的冲击实验;G:5-HMF胁迫下生长曲线;H:5-HMF胁迫下葡萄糖消耗曲线;I:5-HMF胁迫下乙醇生产曲线
Fig. 2 Growths of the recombinant strains under stress-free, furfural stress and 5-HMF stress conditions A: Growth under no stress. B: Glucose consumption under no stress. C: Ethanol production under no stress. D: Growth curve under 3 g/L furfural stress. E: Growth curve under 4 g/L furfural stress. F: Shock experiment with 4 g/L furfural. G: Growth under 5-HMF stress. H: Glucose consumption under 5-HMF stress. I: Ethanol production under 5-HMF stress
| 菌株 Strain | 糠醛胁迫下ROS荧光强度 ROS fluorescence intensity under furfural stress | 5-HMF胁迫下ROS荧光强度 ROS fluorescence intensity under 5-HMF stress |
|---|---|---|
| WT | 1118±219 | 1281±395 |
| SC-ADH | 1200±182 | 913±23 |
| SC-GRE | 1618±149 | 1135±203 |
表8 糠醛与5-HMF胁迫下的ROS水平
Table 8 ROS levels under furfural stress and 5-HMF stress
| 菌株 Strain | 糠醛胁迫下ROS荧光强度 ROS fluorescence intensity under furfural stress | 5-HMF胁迫下ROS荧光强度 ROS fluorescence intensity under 5-HMF stress |
|---|---|---|
| WT | 1118±219 | 1281±395 |
| SC-ADH | 1200±182 | 913±23 |
| SC-GRE | 1618±149 | 1135±203 |
| 菌株 Strain | 添加抑制物时期 Inhibitor addition period | 葡萄糖消耗速率 Glucose consumption rate/% | 乙醇生产速率 Ethanol production rate/% | 比生长速率 Specific growth rate/% |
|---|---|---|---|---|
| SC-ADH | 无胁迫 | 1.4 | 4.6 | 3.3 |
| 对数期 | 4.63 | 1.16 | 4.62 | |
| 稳定期 | 3.08 | 5.90 | 0.25 | |
| SC-CTT | 无胁迫 | 3.4 | -0.1 | 1.1 |
| 对数期 | 17.4 | 5.68 | 17.8 | |
| 稳定期 | 1.07 | 3.10 | 1.68 | |
| SC-AC | 无胁迫 | 6.7 | -0.8 | 0.7 |
| 对数期 | 31.0 | 41.7 | 24.3 | |
| 稳定期 | 3.07 | 4.86 | 1.26 |
表10 双基因元件应对复杂胁迫的性能提升(与野生型比较)
Table 10 Performance improvements of dual genetic parts in response to complex stress(Compared to the wild-type)
| 菌株 Strain | 添加抑制物时期 Inhibitor addition period | 葡萄糖消耗速率 Glucose consumption rate/% | 乙醇生产速率 Ethanol production rate/% | 比生长速率 Specific growth rate/% |
|---|---|---|---|---|
| SC-ADH | 无胁迫 | 1.4 | 4.6 | 3.3 |
| 对数期 | 4.63 | 1.16 | 4.62 | |
| 稳定期 | 3.08 | 5.90 | 0.25 | |
| SC-CTT | 无胁迫 | 3.4 | -0.1 | 1.1 |
| 对数期 | 17.4 | 5.68 | 17.8 | |
| 稳定期 | 1.07 | 3.10 | 1.68 | |
| SC-AC | 无胁迫 | 6.7 | -0.8 | 0.7 |
| 对数期 | 31.0 | 41.7 | 24.3 | |
| 稳定期 | 3.07 | 4.86 | 1.26 |
| 菌株 Strain | 总生物量 Total biomass | 葡萄糖消耗速率 Glucose consumption rate | 乙醇生产速率 Ethanol production rate | 比生长速率 Specific growth rate |
|---|---|---|---|---|
| SC-AC | 11.5% | 60.1% | 58.9% | 64.2% |
表11 双基因元件应对复杂胁迫的性能提升(延滞期添加抑制物,与SC-CTT比较)
Table 11 Performance improvements of dual genetic parts in response to complex stress(Adding inhibitors during the lag phase, compared to SC-CTT strain)
| 菌株 Strain | 总生物量 Total biomass | 葡萄糖消耗速率 Glucose consumption rate | 乙醇生产速率 Ethanol production rate | 比生长速率 Specific growth rate |
|---|---|---|---|---|
| SC-AC | 11.5% | 60.1% | 58.9% | 64.2% |
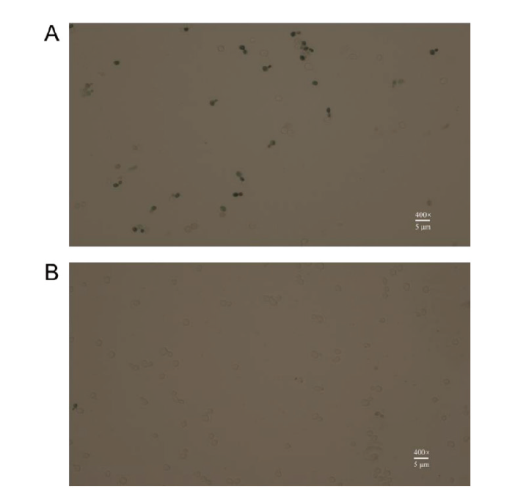

图4 美蓝染色后的光学显微形态 A:野生型;B:SC-AC(照片在400 ×光学显微镜下拍摄)
Fig. 4 Optical microscopic images of cells stained with methylene blue A: Wild-type. B: SC-AC(The photos were taken under 400 × optical microscope)
| 菌株 Strain | ROS level | NAD+/ NADH | NADP+/ NADPH | ATP content/ (nmol·mg-1Protein) | Total GSH content/(mmol·mg-1Protein) | SOD enzyme activity/(U·mg-1Protein) | CAT enzyme activity/(U·mg-1Protein) |
|---|---|---|---|---|---|---|---|
| SC-CTT | 324±5 | 0.32±0.04 | 0.26±0.02 | 11.54±0.33 | 16.98±3.29 | 295±15 | 26.10±1.22 |
| SC-AC | 312±2 | 0.30±0.14 | 0.28±0.01 | 8.34±0.52 | 23.37±4.45 | 242±98 | 36.60±0.09 |
表12 延滞期胁迫组重组菌株的生理状态表征
Table 12 Characterization of the physiological state of the recombinant strains in the lag phase stress group
| 菌株 Strain | ROS level | NAD+/ NADH | NADP+/ NADPH | ATP content/ (nmol·mg-1Protein) | Total GSH content/(mmol·mg-1Protein) | SOD enzyme activity/(U·mg-1Protein) | CAT enzyme activity/(U·mg-1Protein) |
|---|---|---|---|---|---|---|---|
| SC-CTT | 324±5 | 0.32±0.04 | 0.26±0.02 | 11.54±0.33 | 16.98±3.29 | 295±15 | 26.10±1.22 |
| SC-AC | 312±2 | 0.30±0.14 | 0.28±0.01 | 8.34±0.52 | 23.37±4.45 | 242±98 | 36.60±0.09 |
| [1] |
Rittmann BE. Opportunities for renewable bioenergy using microorganisms[J]. Biotechnology and Bioengineering, 2008, 100(2):203-212.
doi: 10.1002/bit.21875 pmid: 18431744 |
| [2] |
Zhang J, Wang YH, Du XJ, et al. Selective removal of lignin to enhance the process of preparing fermentable sugars and platform chemicals from lignocellulosic biomass[J]. Bioresource Technology, 2020, 303:122846.
doi: 10.1016/j.biortech.2020.122846 URL |
| [3] |
Zhang ZR, Song JL, Han BX. Catalytic transformation of lignocellulose into chemicals and fuel products in ionic liquids[J]. Chemical Reviews, 2017, 117(10):6834-6880.
doi: 10.1021/acs.chemrev.6b00457 pmid: 28535680 |
| [4] |
Chen CY, Zhao XQ, Yen HW, et al. Microalgae-based carbohydrates for biofuel production[J]. Biochemical Engineering Journal, 2013, 78:1-10.
doi: 10.1016/j.bej.2013.03.006 URL |
| [5] |
Zhao W, Zhao F, Zhang S, et al. Ethanol production by simultaneous saccharification and cofermentation of pretreated corn stalk[J]. Journal of Basic Microbiology, 2019, 59(7):744-753.
doi: 10.1002/jobm.201900117 pmid: 31087563 |
| [6] | Robak K, Balcerek M. Current state-of-the-art in ethanol production from lignocellulosic feedstocks[J]. Microbiology Research, 2020, 240:126534. |
| [7] | Deshavath NN, Mohan M, Veeranki VD, et al. Dilute acid pretreatment of sorghum biomass to maximize the hemicellulose hydrolysis with minimized levels of fermentative inhibitors for bioethanol production[J]. Biotechnology, 2017, 7(2):139. |
| [8] |
Allen SA, Clark W, Mccaffery JM, et al. Furfural induces reactive oxygen species accumulation and cellular damage in Saccharomyces cerevisiae[J]. Biotechnology for Biofuels, 2010, 3(1):2.
doi: 10.1186/1754-6834-3-2 |
| [9] |
Du X, Takagi H. N-Acetyltransferase Mpr1 confers ethanol tolerance on Saccharomyces cerevisiae by reducing reactive oxygen species[J]. Applied Microbiology and Biotechnology, 2007, 75(6):1343-1351.
doi: 10.1007/s00253-007-0940-x URL |
| [10] |
Herrero E, Ros J, Bellí G, et al. Redox control and oxidative stress in yeast cells[J]. Biochimica et Biophysica Acta, 2008, 1780(11):1217-1235.
doi: 10.1016/j.bbagen.2007.12.004 pmid: 18178164 |
| [11] | 马超颖, 戚薇, 杜连祥. 酿酒酵母氧化应激反应的研究进展[J]. 天津轻工业学院学报, 2002, 4: 14-17+31. |
| Ma CY, Qi W, Du LX. Research progress on oxidative stress response of Saccharomyces cerevisiae[J]. Journal of Tianjin University of Science and Technology, 2002, 4: 14-17+31. | |
| [12] |
Jayakody LN, Jin YS. In-depth understanding of molecular mechanisms of aldehyde toxicity to engineer robust Saccharomyces cerevisiae[J]. Applied Microbiology and Biotechnology, 2021, 105(7):2675-2692.
doi: 10.1007/s00253-021-11213-1 pmid: 33743026 |
| [13] | 杨雪雪, 蒋伶活. 酵母对木质纤维素酸解物中抑制物的应答及菌株开发[J]. 生命科学, 2012, 24(6):531-534. |
| Yang XX, Jiang LH. The response of yeast to inhibitors in lignocellulosic acidolysis and strain development[J]. Life Science, 2012, 24(6):531-534. | |
| [14] |
Vemuri GN, Altman E, Sangurdekar DP, et al. Overflow metabolism in Escherichia coli during steady-state growth: Transcriptional regulation and effect of the redox ratio[J]. Applied and Environmental Microbiology, 72(5):3653-3661.
doi: 10.1128/AEM.72.5.3653-3661.2006 URL |
| [15] |
Mason JT, Kim SK, Knaff DB, et al. Thermodynamic basis for redox regulation of the Yap1 signal transduction pathway[J]. Biochemistry, 2006, 45(45):13409-13417.
pmid: 17087494 |
| [16] |
Liu CG, Lin YH, Bai FW. Global gene expression analysis of Saccharomyces cerevisiae grown under redox potential controlled very-high-gravity conditions[J]. Biotechnology Journal, 2014, 8(11):1332-1340.
doi: 10.1002/biot.v8.11 URL |
| [17] |
Liu CG, Xue C, Lin YH, et al. Redox potential control and applications in microaerobic and anaerobic fermentations[J]. Biotechnology Advances, 2013, 31(2):257-265.
doi: 10.1016/j.biotechadv.2012.11.005 URL |
| [18] |
De La Torre-Ruiz MA, Pujol N, Sundaran V. Coping with oxidative stress. The yeast model[J]. Current Drug Targets, 2015, 16(1):2-12.
pmid: 25330032 |
| [19] | Hao X, Du B, Liu L, et al. Effect of ORP regulation on yeast fermentation with inhibitors of lignocellulose hydrolysate[J]. Ciesc Journal, 2015, 66(3):1066-1071. |
| [20] |
Li K, Xia J, Mehmood MA, et al. Extracellular redox potential regulation improves yeast tolerance to furfural[J]. Chemical Engineering Science, 2019, 196:54-63.
doi: 10.1016/j.ces.2018.11.059 URL |
| [21] |
Gietz RD, Schiestl RH. High-efficiency yeast transformation using the LiAc/SS carrier DNA/PEG method[J]. Nature Protocols, 2007, 2(1):31-34.
doi: 10.1038/nprot.2007.13 pmid: 17401334 |
| [22] |
Teixeira MC, Monteiro PT, Sa-Correia I. Predicting gene and genomic regulation in Saccharomyces cerevisiae, using the YEASTRACT Database: A step-by-step guided analysis[J]. Methods in Molecular Biology, 2016, 1361:391-404.
doi: 10.1007/978-1-4939-3079-1_22 pmid: 26483034 |
| [23] | 刘广新, 刘玲彦, 王林, 等. 啤酒酵母活性/活力检测方法比较及其与发酵性能的相关性研究[J]. 中外酒业, 2020, 9:40-54. |
| Liu GX, Liu LY, Wang L, et al. Comparison of assay methods for activity/vitality of brewer's yeast and its correlation with fermentation performance[J]. Chinese and foreign wine industry, 2020, 9:40-54. | |
| [24] |
Lee J, Godon C, Lagniel G, et al. Yap1 and Skn7 Control two specialized oxidative stress response regulons in yeast[J]. Journal of Biological Chemistry, 1999, 274(23):16040-16046.
doi: 10.1074/jbc.274.23.16040 pmid: 10347154 |
| [25] |
Kuge S, Jones N. YAP1 dependent activation of TRX2 is essential for the response of Saccharomyces cerevisiae to oxidative stress by hydroperoxides[J]. Embo Journal, 1994, 13(3):655-664
doi: 10.1002/j.1460-2075.1994.tb06304.x pmid: 8313910 |
| [26] |
Ouyang X, Tran QT, Goodwin S, et al. Yap1 activation by H2O2 or thiol-reactive chemicals elicits distinct adaptive gene responses[J]. Free Radical Biology and Medicine, 2011, 50(1):1-13.
doi: 10.1016/j.freeradbiomed.2010.10.697 pmid: 20971184 |
| [27] |
Thomas D, Barbey R, Surdin-Kerjan Y. Gene-enzyme relationship in the sulfate assimilation pathway of Saccharomyces cerevisiae. Study of the 3'-phosphoadenylylsulfate reductase structural gene[J]. Journal of Biological Chemistry, 1990, 265(26):15518-15524.
pmid: 2203779 |
| [28] |
Kim D, Hahn JS. Roles of the Yap1 transcription factor and antioxidants in Saccharomyces cerevisiae's tolerance to furfural and 5-hydroxymethylfurfural, which function as thiol-reactive electrophiles generating oxidative stress[J]. Applied and Environmental Microbiology, 2013, 79(16):5069-5077.
doi: 10.1128/AEM.00643-13 URL |
| [29] |
Benjaphokee S, Hasegawa D, Yokota D, et al. Highly efficient bioethanol production by a Saccharomyces cerevisiae strain with multiple stress tolerance to high temperature, acid and ethanol[J]. Nature Biotechnology, 2012, 29(3):379-386.
doi: 10.1038/nbt0511-379 |
| [30] |
Qiu Z, Jiang R. Improving Saccharomyces cerevisiae ethanol production and tolerance via RNA polymerase II subunit Rpb7[J]. Biotechnology for Biofuels, 2017, 10:125.
doi: 10.1186/s13068-017-0806-0 URL |
| [31] |
Gechev TS, Van Breusegem F, Stone JM, et al. Reactive oxygen species as signals that modulate plant stress responses and programmed cell death[J]. Bioessays, 2006, 28(11):1091-1101.
doi: 10.1002/bies.20493 pmid: 17041898 |
| [32] |
Azad GK, Singh V, Tomar RS. Assessment of the biological pathways targeted by isocyanate using N-succinimidyl N-methylcarbamate in budding yeast Saccharomyces cerevisiae[J]. PLoS One, 2014, 9(3):e92993.
doi: 10.1371/journal.pone.0092993 URL |
| [33] | Nishimoto T, Furuta M, Kataoka M, et al. Important role of catalase in the cellular response of the budding yeast Saccharomyces cerevisiae exposed to ionizing radiation[J]. Currunt Microbiology, 2015, 70(3):404-407. |
| [34] |
Qin J, Zhou YJ, Krivoruchko A, et al. Modular pathway rewiring of Saccharomyces cerevisiae enables high-level production of L-ornithine[J]. Nature Communications, 2015, 6:8224.
doi: 10.1038/ncomms9224 |
| [35] |
Ask M, Mapelli V, Höck H, et al. Engineering glutathione biosynthesis of Saccharomyces cerevisiae increases robustness to inhibitors in pretreated lignocellulosic materials[J]. Microbial Cell Factories, 2013, 12(1):87-87.
doi: 10.1186/1475-2859-12-87 |
| [36] |
Outten CE, Culotta VC. Alternative start sites in the Saccharomyces cerevisiae GLR1 gene are responsible for mitochondrial and cytosolic isoforms of glutathione reductase[J]. Journal of Biological Chemistry, 2004, 279(9):7785-7791.
doi: 10.1074/jbc.M312421200 pmid: 14672937 |
| [37] |
Daehwan K. Physico-chemical conversion of lignocellulose inhibitor effects and detoxification strategies: A mini review[J]. Molecules, 2018, 23(2):309-.
doi: 10.3390/molecules23020309 URL |
| [38] | Larroy C, Parés X, Biosca JA. Characterization of a Saccharomyces cerevisiae NADP(H)-dependent alcohol dehydrogenase(ADHVII), a member of the cinnamyl alcohol dehydrogenase family[J]. FEBS Journal, 2002, 269(22):5738-5745. |
| [39] |
Petersson A, Almeida JR, Modig T, et al. A 5-hydroxymethyl furfural reducing enzyme encoded by the Saccharomyces cerevisiae ADH6 gene conveys HMF tolerance[J]. Yeast, 2006, 23(6):455-464.
doi: 10.1002/yea.1370 pmid: 16652391 |
| [40] |
Moon J, Liu ZL. Engineered NADH-dependent GRE2 from Saccharomyces cerevisiae by directed enzyme evolution enhances HMF reduction using additional cofactor NADPH[J]. Enzyme and Microbial Technology, 2012, 50(2):115-120.
doi: 10.1016/j.enzmictec.2011.10.007 URL |
| [1] | 徐发迪, 徐康, 孙东明, 李萌蕾, 赵建志, 鲍晓明. 基于杨木(Populus sp.)的二代燃料乙醇技术研究进展[J]. 生物技术通报, 2023, 39(9): 27-39. |
| [2] | 成婷, 苑帅, 张晓元, 林良才, 李欣, 张翠英. 酿酒酵母异丁醇合成途径调控的研究进展[J]. 生物技术通报, 2023, 39(7): 80-90. |
| [3] | 李昕悦, 周明海, 樊亚超, 廖莎, 张风丽, 刘晨光, 孙悦, 张霖, 赵心清. 基于转运蛋白工程提升微生物菌株耐受性和生物制造效率的研究进展[J]. 生物技术通报, 2023, 39(11): 123-136. |
| [4] | 晏雄鹰, 王振, 王霞, 杨世辉. 微生物硫代谢与抗逆性[J]. 生物技术通报, 2023, 39(11): 150-167. |
| [5] | 祝瑛萱, 李克景, 何敏, 郑道琼. 酵母模型揭示胁迫因子驱动基因组变异的研究进展[J]. 生物技术通报, 2023, 39(11): 191-204. |
| [6] | 唐瑞琪, 赵心清, 朱笃, 汪涯. 大肠杆菌对木质纤维素水解液抑制物的胁迫耐受性[J]. 生物技术通报, 2023, 39(11): 205-216. |
| [7] | 孙言秋, 谢采芸, 汤岳琴. 耐高温酿酒酵母的构建与高温耐受机制解析[J]. 生物技术通报, 2023, 39(11): 226-237. |
| [8] | 翟旭航, 李霞, 元英进. 木质纤维素预处理及高值化技术研究进展[J]. 生物技术通报, 2021, 37(3): 162-174. |
| [9] | 胡芳, 董旭, 史长伟, 吴学栋. 超声波强化木质纤维素酶解的研究进展[J]. 生物技术通报, 2021, 37(10): 234-244. |
| [10] | 崔新刚, 孙雅欣, 崔晓静, 邓雁文, 孙恩浩, 王俊芳, 崔红晶. 酿酒酵母TAP42基因在细胞壁应激反应中的作用[J]. 生物技术通报, 2021, 37(10): 57-62. |
| [11] | 顾翰琦, 邵玲智, 刘冉, 刘晓光, 李玲, 刘倩, 李洁, 张雅丽. 酿酒酵母酚类抑制物耐受性脂质组学研究[J]. 生物技术通报, 2021, 37(1): 15-23. |
| [12] | 吴宇, 王金华, 赵筱. GLN1基因过表达对提高酿酒酵母糠醛耐受性的研究[J]. 生物技术通报, 2020, 36(8): 69-78. |
| [13] | 刘登, 刘均洪. 嗜热性木质纤维素酶在纤维素乙醇生产中的研究进展[J]. 生物技术通报, 2020, 36(8): 185-193. |
| [14] | 顾翰琦, 刘冉, 邵玲智, 徐岩岩, 王东岩, 张冬梅, 李洁. 酿酒酵母对酚类抑制物耐受性研究[J]. 生物技术通报, 2020, 36(6): 136-142. |
| [15] | 陈永灿, 张建志, 司同. 酿酒酵母中基于CRISPR/dCás9的基因转录调控工具的开发与应用[J]. 生物技术通报, 2020, 36(4): 1-12. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||