








生物技术通报 ›› 2024, Vol. 40 ›› Issue (1): 262-269.doi: 10.13560/j.cnki.biotech.bull.1985.2023-0484
谢宏1( ), 周丽莹2, 李舒文1, 王梦迪1, 艾晔1, 晁跃辉1(
), 周丽莹2, 李舒文1, 王梦迪1, 艾晔1, 晁跃辉1( )
)
收稿日期:2023-05-23
出版日期:2024-01-26
发布日期:2024-02-06
通讯作者:
晁跃辉,男,博士,副教授,研究方向:草地植物生物技术;E-mail: chaoyuehui@bjfu.edu.cn作者简介:谢宏,女,硕士研究生,研究方向:草地植物生物技术;E-mail: xiehong20211014@163.com
基金资助:
XIE Hong1( ), ZHOU Li-ying2, LI Shu-wen1, WANG Meng-di1, AI Ye1, CHAO Yue-hui1(
), ZHOU Li-ying2, LI Shu-wen1, WANG Meng-di1, AI Ye1, CHAO Yue-hui1( )
)
Received:2023-05-23
Published:2024-01-26
Online:2024-02-06
摘要:
【目的】CIM(cytokinin induce messager)属于Expansin蛋白家族,该蛋白是一类在植物生长和发育过程中起关键作用的蛋白质。为分析蒺藜苜蓿MtCIM基因结构、表达、亚细胞定位及在植物生长和发育过程中的功能。【方法】利用生物信息学工具对其进行深入分子量、理论等电点、不稳定系数及蛋白质结构预测和分析;同时,利用RT-qPCR方法,对不同激素处理(IAA、ABA、GA3)下的基因表达进行了检测;并通过融合蛋白瞬时表达及酵母表达方法鉴定其亚细胞定位及转录自激活情况;对MtCIM转基因植株进行低磷环境的适应分析。【结果】生物信息学分析显示,CIM蛋白的相对分子量为29.563 kD,理论等电点为5.54,不稳定系数为41.62,其蛋白质性质不稳定。二级及三级结构预测表明,该蛋白主要由无规则卷曲构成。外施IAA能够显著提高MtCIM的表达,外施ABA和GA3后,MtCIM表达呈现先升高后降低的趋势。亚细胞定位结果显示,其定位于细胞质中。对MtCIM进行转录自激活活性分析显示,蒺藜苜蓿MtCIM不存在自激活活性。对获得的20株转基因株系进行分析表明,转基因株系叶片细胞壁更厚;转基因植株对低磷环境具有更好的适应能力。【结论】对蒺藜苜蓿MtCIM的全面分析,揭示了其作为Expansin蛋白家族成员在植物生长和发育中的关键作用,并提高了植物在低磷环境中适应能力。
谢宏, 周丽莹, 李舒文, 王梦迪, 艾晔, 晁跃辉. 蒺藜苜蓿MtCIM基因结构和功能分析[J]. 生物技术通报, 2024, 40(1): 262-269.
XIE Hong, ZHOU Li-ying, LI Shu-wen, WANG Meng-di, AI Ye, CHAO Yue-hui. Structural and Functional Analysis of MtCIM Gene in Medicago truncatula[J]. Biotechnology Bulletin, 2024, 40(1): 262-269.
| 引物名称 Prime name | 引物序列 Prime sequence(5'-3') |
|---|---|
| T7 | TAATACGACTCACTATAGG |
| 3-BD | TAAGAGTCACTTTAAAATTTGTAT |
| pGBKT7-CIM-F | GCTATTTCTACTGATTTTTCATGGCTCTTACACTTGAACA |
| pGBKT7-CIM-R | CAAAATCATGTCAAGGTCTTTTAAAAATTCACAATTGACC |
| 3302Y-F | TGACGCACAATCCCACTATCCTT |
| 3302Y-R | CCGTCCAGCTCGACCAGGAT |
| 3302Y-MtCIM-F | GAACACGGGGGACTCTTGACCATGGCTCTTACACTTGAACA |
| 3302Y-MtCIM-R | ACGCGTACTAGTCAGATCTACAAAATTCACAATTGACCTAT |
| MtActin-F | TGATCTGGCTGGTCGTGACCTTA |
| MtActin-R | ATGCCTGCTGCTTCCATTCCTAT |
| MtCIM-RT-F | TGGTGACGGAAGCGAAGGTGGTG |
| MtCIM-RT-R | CTTAACAGGGTTGCCTGAACATA |
表1 引物序列
Table 1 Primer sequences
| 引物名称 Prime name | 引物序列 Prime sequence(5'-3') |
|---|---|
| T7 | TAATACGACTCACTATAGG |
| 3-BD | TAAGAGTCACTTTAAAATTTGTAT |
| pGBKT7-CIM-F | GCTATTTCTACTGATTTTTCATGGCTCTTACACTTGAACA |
| pGBKT7-CIM-R | CAAAATCATGTCAAGGTCTTTTAAAAATTCACAATTGACC |
| 3302Y-F | TGACGCACAATCCCACTATCCTT |
| 3302Y-R | CCGTCCAGCTCGACCAGGAT |
| 3302Y-MtCIM-F | GAACACGGGGGACTCTTGACCATGGCTCTTACACTTGAACA |
| 3302Y-MtCIM-R | ACGCGTACTAGTCAGATCTACAAAATTCACAATTGACCTAT |
| MtActin-F | TGATCTGGCTGGTCGTGACCTTA |
| MtActin-R | ATGCCTGCTGCTTCCATTCCTAT |
| MtCIM-RT-F | TGGTGACGGAAGCGAAGGTGGTG |
| MtCIM-RT-R | CTTAACAGGGTTGCCTGAACATA |
| 顺式元件Cis-element | 基序Motif(5'-3') | 位置Position | 功能Function |
|---|---|---|---|
| ABRE | CACGTG | 348 | 脱落酸响应元件ABA response element |
| ACGTC | 349 | ||
| ACGTC | 708 | ||
| P-box | CCTTTTG | 53 | 赤霉素响应元件Gibberellin response element |
| TC-rich repeats | GTTTTCTTAC | 103 | 防御和胁迫反应相关的元件Defense and stress response related elements |
| TGA-element | AACGAC | 510 | 生长素响应元件Auxin response element |
| TCT-motif | TCTTAC | 103 | 光响应元件Light response element |
| ARE | AAACCA | 567 | 厌氧诱导元件Anaerobic induction element |
| TATA-box | TATTTAAA | 21 | 转录起始点-30附近的核心启动子元件 Core promoter element near trabscription starting point -30 |
| TATAA | 148 | ||
| TATTA | 633 | ||
| TATA | 502 | ||
| TATATA | 87 | ||
| TATA | 275 | ||
| G-BOX | CACGTG | 348 | 参与光反应的顺式作用元件Cis-acting elements involved in photoreactions |
| CACGTG | 707 |
表2 MtCIM启动子的主要顺式作用元件
Table 2 The main cis-acting elements of MtCIM promoter
| 顺式元件Cis-element | 基序Motif(5'-3') | 位置Position | 功能Function |
|---|---|---|---|
| ABRE | CACGTG | 348 | 脱落酸响应元件ABA response element |
| ACGTC | 349 | ||
| ACGTC | 708 | ||
| P-box | CCTTTTG | 53 | 赤霉素响应元件Gibberellin response element |
| TC-rich repeats | GTTTTCTTAC | 103 | 防御和胁迫反应相关的元件Defense and stress response related elements |
| TGA-element | AACGAC | 510 | 生长素响应元件Auxin response element |
| TCT-motif | TCTTAC | 103 | 光响应元件Light response element |
| ARE | AAACCA | 567 | 厌氧诱导元件Anaerobic induction element |
| TATA-box | TATTTAAA | 21 | 转录起始点-30附近的核心启动子元件 Core promoter element near trabscription starting point -30 |
| TATAA | 148 | ||
| TATTA | 633 | ||
| TATA | 502 | ||
| TATATA | 87 | ||
| TATA | 275 | ||
| G-BOX | CACGTG | 348 | 参与光反应的顺式作用元件Cis-acting elements involved in photoreactions |
| CACGTG | 707 |
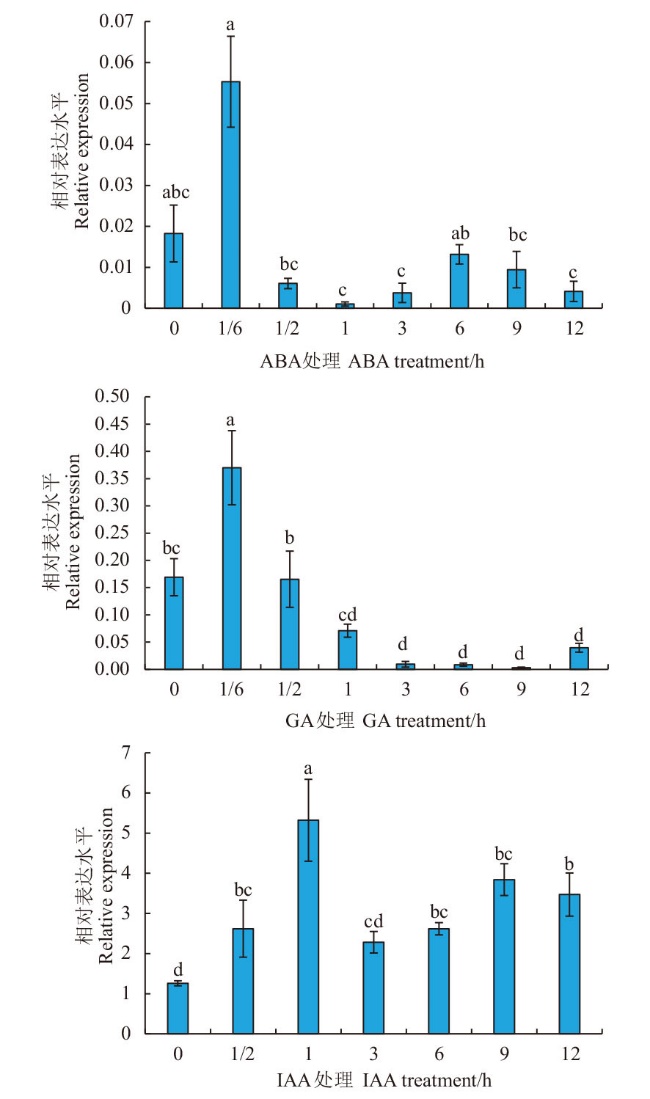
图4 不同激素处理下MtCIM在蒺藜苜蓿中的表达分析 不同小写字母表示差异显著(P<0.05)
Fig. 4 MtCIM expression analysis in Medicago truncatula under different hormone treatments Different lowercase letters indicate a significant difference at the 0.05 level
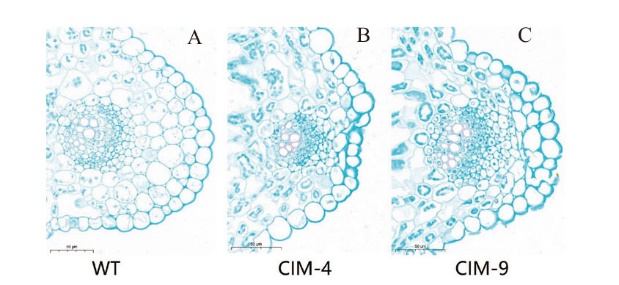
图6 番红固绿染色法对叶片结构的观察 A:野生型植株;B:转基因植株;C:转基因植株
Fig. 6 Observation of leaf structure by saffred solid green staining method A: Wild-type plants; B: transgenic plants; C: transgenic plants

图7 全磷和低磷处理下转基因和野生型植株差异 CIM:转基因;WT:野生型;A、C:全磷条件;B、D:低磷条件
Fig. 7 Difference between transgenic and wild type plants under total phosphorus and low phosphorus treatment CIM: Transgenic plants. WT: Wild types. A, C: Total phosphorus. B, D: Low phosphorus
| [1] |
McQueen-Mason S, Cosgrove DJ. Disruption of hydrogen bonding between plant cell wall polymers by proteins that induce wall extension[J]. PNAS, 1994. 91(14): 6574-6578.
doi: 10.1073/pnas.91.14.6574 pmid: 11607483 |
| [2] | 雷忠萍. 棉花扩展蛋白基因的鉴定及特征研究[D]. 杨凌: 西北农林科技大学, 2018. |
| Lei ZP. Identification and characterization of cotton expansin gene[D]. Yangling: Northwest A&F University, 2018. | |
| [3] |
Lee Y, Choi D, Kende H. Expansins: ever-expanding numbers and functions[J]. Curr Opin Plant Biol, 2001, 4(6): 527-532.
doi: 10.1016/s1369-5266(00)00211-9 pmid: 11641069 |
| [4] |
Yan A, Wu MJ, Yan LM. AtEXP2 is involved in seed germination and abiotic stress response in Arabidopsis[J]. PLoS One, 2014, 9(1): e85208.
doi: 10.1371/journal.pone.0085208 URL |
| [5] |
Cho HT, Cosgrove DJ. Altered expression of expansin modulates leaf growth and pedicel abscission in Arabidopsis thaliana[J]. PNAS, 2000, 97(17): 9783-9788.
doi: 10.1073/pnas.160276997 pmid: 10931949 |
| [6] |
Lee HW, Kim MJ, Kim NY, et al. LBD18 acts as a transcriptional activator that directly binds to the EXPANSIN14 promoter in promoting lateral root emergence of Arabidopsis[J]. Plant J, 2013. 73(2): 212-224.
doi: 10.1111/tpj.2013.73.issue-2 URL |
| [7] |
Lee HW, Kim J. EXPANSINA17 up-regulated by LBD18/ASL20 promotes lateral root formation during the auxin response[J]. Plant Cell Physiol, 2013, 54(10): 1600-1611.
doi: 10.1093/pcp/pct105 pmid: 23872272 |
| [8] |
Ma N, Wang Y, Qiu SC, et al. Overexpression of OsEXPA8, a root-specific gene, improves rice growth and root system architecture by facilitating cell extension[J]. PloS one, 2013, 8(10): e75997.
doi: 10.1371/journal.pone.0075997 URL |
| [9] |
Chen YH, Han YY, Zhang M, et al. Overexpression of the wheat expansin gene TaEXPA2 improved seed production and drought tolerance in transgenic tobacco plants[J]. PLos One, 2016, 11(4): e0153494.
doi: 10.1371/journal.pone.0153494 URL |
| [10] |
Kong YB, Wang B, Du H, et al. GmEXLB1, a soybean expansin-like B gene, alters root architecture to improve phosphorus acquisition in Arabidopsis[J]. Front Plant Sci, 2019, 10: 808.
doi: 10.3389/fpls.2019.00808 URL |
| [11] |
Yang ZJ, Gao Z, Zhou HW, et al. GmPTF1 modifies root architecture responses to phosphate starvation primarily through regulating GmEXPB2 expression in soybean[J]. Plant J, 2021, 107(2): 525-543.
doi: 10.1111/tpj.v107.2 URL |
| [12] |
Sampedro J, Cosgrove DJ. The expansin superfamily[J]. Genome Biol, 2005, 6(12): 242.
doi: 10.1186/gb-2005-6-12-242 pmid: 16356276 |
| [13] | 杨君娇. 扩展蛋白基因TaEXPA2在小麦耐旱性中的功能分析[D]. 泰安: 山东农业大学, 2020. |
| Yang JJ. Functional analysis of expandable protein gene TaEXPA2 in drought tolerance of wheat[D]. Tai'an: Shandong Agricultural University, 2020. | |
| [14] | Chen LJ, Zou WS, Fei CY, et al. α-expansin EXPA4 positively regulates abiotic stress tolerance but negatively regulates pathogen resistance in Nicotiana tabacum[J]. Plant Cell Physiol, 2018, 59(11): 2317-2330. |
| [15] |
Zhang BY, Chang L, Sun WN, et al. Overexpression of an expansin-like gene, GhEXLB2 enhanced drought tolerance in cotton[J]. Plant Physiol Biochem, 2021, 162: 468-475.
doi: 10.1016/j.plaphy.2021.03.018 URL |
| [16] |
Winey M, Meehl JB, O'Toole ET, et al. Conventional transmission electron microscopy[J]. Mol Biol Cell, 2014, 25(3): 319-323.
doi: 10.1091/mbc.E12-12-0863 pmid: 24482357 |
| [17] |
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method[J]. Methods, 2001, 25(4): 402-408.
doi: 10.1006/meth.2001.1262 pmid: 11846609 |
| [18] | 艾晔, 钱旭, 韩烈保, 等. 蒺藜苜蓿CIM基因的克隆及转化[J/OL]. 分子植物育种, 2021. https://kns.cnki.net/kcms/detail/46.1068.S.20210630.0948.004.html. |
| Ai Y, Qian X, Han LB, et al. Cloning and transformation of CIM gene from Medicago truncatula[J/OL]. Mol Plant Breed, 2021. https://kns.cnki.net/kcms/detail/46.1068.S.20210630.0948.004.html. | |
| [19] | 郑珊珊, 石卓兴, 代琦, 等. 蛋白质亚细胞定位预测研究进展[J]. 科技视界, 2014(15): 183-184. |
| Zheng SS, Shi ZX, Dai Q, et al. Research progress of subcellular localization prediction in protein[J]. Sci Technol Vis, 2014(15): 183-184. | |
| [20] | 李杰, 张福城, 王文泉, 等. 高等植物启动子的研究进展[J]. 生物技术通讯, 2006, 17(4): 658-661. |
| Li J, Zhang FC, Wang WQ, et al. Advance in the study of higher plant promoter[J]. Lett Biotechnol, 2006, 17(4): 658-661. | |
| [21] | 张春晓, 王文棋, 蒋湘宁, 等. 植物基因启动子研究进展[J]. 遗传学报, 2004, 31(12): 1455-1464. |
| Zhang CX, Wang WQ, Jiang XN, et al. Review on plant gene promoters[J]. Acta Genet Sin, 2004, 31(12): 1455-1464. | |
| [22] |
冯寒骞, 李超. 生长素信号转导研究进展[J]. 生物技术通报, 2018, 34(7): 24-30.
doi: 10.13560/j.cnki.biotech.bull.1985.2018-0488 |
| Feng HL, Li C. Research advances of auxin signal transduction[J]. 2018, 34(7): 24-30. | |
| [23] | 韩阳阳, 王天晓, 王玮. 扩展蛋白与激素调控的生长发育[J]. 生命的化学, 2015. 35(6): 778-784. |
| Han YY, Wang TX, Wang W. Research progress on the relationship between expansins and phytohormone induced plant growth and development[J]. Chemistry life, 2015, 35(6): 778-784. | |
| [24] | 孟永娇, 李季, 徐建, 等, 黄瓜扩张素蛋白基因CsEXPb1的克隆及表达分析[J]. 园艺学报, 2015, 42(4): 679-688. |
| Meng YJ, Li J, Xu J, et al. Cloning and expression analysis of the expansin gene CsEXPb1 in Cucumber[J]. Acta Hortic Sin, 2015, 42(4): 679-688. | |
| [25] |
Wu Y, Meeley RB, Cosgrove DJ. Analysis and expression of theα-expansin and β-expansin gene families in maize[J]. Plant Physiol, 2001, 126(1): 222-232.
pmid: 11351085 |
| [26] | 丁安明, 陈志华, 杨懿德, 等. NtabEXPA12基因过表达对烟草叶片发育及抗逆性的影响[J]. 中国烟草科学, 2021, 42(4): 58-66. |
| Ding AM, Chen ZH, Yang YD, et al. Overexpression of Ntab EXPA12 affects leaf development and abiotic stress tolerance in tobacco[J]. Chin tob sci, 2021, 42(4): 58-66. | |
| [27] |
Tan J, Wang ML, Shi ZY, et al. OsEXPA10 mediates the balance between growth and resistance to biotic stress in rice[J]. Plant Cell Rep, 2018, 37(7): 993-1002.
doi: 10.1007/s00299-018-2284-7 pmid: 29619515 |
| [28] |
Cheung AY, Cosgrove DJ, Hara-Nishimura I, et al. A rich and bountiful harvest: Key discoveries in plant cell biology[J]. Plant Cell, 2022, 34(1): 53-71.
doi: 10.1093/plcell/koab234 URL |
| [29] | 邢国芳, 郭平毅. 植物响应低磷胁迫的功能基因组研究进展[J]. 生物技术通报, 2013(7): 1-6. |
| Xing GF, Guo PY. Functional genomics approaches for the study of low phosphate responses in higher plants[J]. Biotechnol Bull, 2013(7): 1-6. |
| [1] | 朱毅, 柳唐镜, 宫国义, 张洁, 王晋芳, 张海英. 西瓜ClPP2C3克隆及表达分析[J]. 生物技术通报, 2024, 40(1): 243-249. |
| [2] | 唐伟林, 康琴, 汪霞, 谌明洋, 孙欣江, 王棵, 侯凯, 吴卫, 徐东北. 薄荷茉莉酸受体McCOI1a基因的克隆与表达模式分析[J]. 生物技术通报, 2024, 40(1): 270-280. |
| [3] | 吕秋谕, 孙培媛, 冉彬, 王佳蕊, 陈庆富, 李洪有. 苦荞转录因子基因FtbHLH3的克隆、亚细胞定位及表达分析[J]. 生物技术通报, 2023, 39(8): 194-203. |
| [4] | 王佳蕊, 孙培媛, 柯瑾, 冉彬, 李洪有. 苦荞糖基转移酶基因FtUGT143的克隆及表达分析[J]. 生物技术通报, 2023, 39(8): 204-212. |
| [5] | 李博, 刘合霞, 陈宇玲, 周兴文, 朱宇林. 金花茶CnbHLH79转录因子的克隆、亚细胞定位及表达分析[J]. 生物技术通报, 2023, 39(8): 241-250. |
| [6] | 孙明慧, 吴琼, 刘丹丹, 焦小雨, 王文杰. 茶树CsTMFs的克隆与表达分析[J]. 生物技术通报, 2023, 39(7): 151-159. |
| [7] | 赵雪婷, 高利燕, 王俊刚, 沈庆庆, 张树珍, 李富生. 甘蔗AP2/ERF转录因子基因ShERF3的克隆、表达及其编码蛋白的定位[J]. 生物技术通报, 2023, 39(6): 208-216. |
| [8] | 姜晴春, 杜洁, 王嘉诚, 余知和, 王允, 柳忠玉. 虎杖转录因子PcMYB2的表达特性和功能分析[J]. 生物技术通报, 2023, 39(5): 217-223. |
| [9] | 姚姿婷, 曹雪颖, 肖雪, 李瑞芳, 韦小妹, 邹承武, 朱桂宁. 火龙果溃疡病菌实时荧光定量PCR内参基因的筛选[J]. 生物技术通报, 2023, 39(5): 92-102. |
| [10] | 王艺清, 王涛, 韦朝领, 戴浩民, 曹士先, 孙威江, 曾雯. 茶树SMAS基因家族的鉴定及互作分析[J]. 生物技术通报, 2023, 39(4): 246-258. |
| [11] | 刘思佳, 王浩楠, 付宇辰, 闫文欣, 胡增辉, 冷平生. ‘西伯利亚’百合LiCMK基因克隆及功能分析[J]. 生物技术通报, 2023, 39(3): 196-205. |
| [12] | 王涛, 漆思雨, 韦朝领, 王艺清, 戴浩民, 周喆, 曹士先, 曾雯, 孙威江. CsPPR和CsCPN60-like在茶树白化叶片中的表达分析及互作蛋白验证[J]. 生物技术通报, 2023, 39(3): 218-231. |
| [13] | 庞强强, 孙晓东, 周曼, 蔡兴来, 张文, 王亚强. 菜心BrHsfA3基因克隆及其对高温胁迫的响应[J]. 生物技术通报, 2023, 39(2): 107-115. |
| [14] | 苗淑楠, 高宇, 李昕儒, 蔡桂萍, 张飞, 薛金爱, 季春丽, 李润植. 大豆GmPDAT1参与油脂合成和非生物胁迫应答的功能分析[J]. 生物技术通报, 2023, 39(2): 96-106. |
| [15] | 滕梦鑫, 徐亚, 何静, 汪奇, 乔飞, 李敬阳, 李新国. 香蕉MaMC6的克隆及原核表达分析[J]. 生物技术通报, 2023, 39(12): 179-186. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||