








生物技术通报 ›› 2026, Vol. 42 ›› Issue (2): 293-305.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0417
李勒松1,2,3( ), 张金金1,2,3, 贺佳蝶1,2,3, 梁艳丽1,2, 杨生超1,3, 栗孟飞4, 赵艳1,2,3(
), 张金金1,2,3, 贺佳蝶1,2,3, 梁艳丽1,2, 杨生超1,3, 栗孟飞4, 赵艳1,2,3( )
)
收稿日期:2025-04-20
出版日期:2026-02-26
发布日期:2026-03-17
通讯作者:
赵艳,女,博士,教授,研究方向 :药用植物学;E-mail: zhaoyankm@126.com作者简介:李勒松,女,硕士研究生,研究方向 :药用植物学;E-mail: lls15126334094@163.com
基金资助:
LI Le-song1,2,3( ), ZHANG Jin-jin1,2,3, HE Jia-die1,2,3, LIANG Yan-li1,2, YANG Sheng-chao1,3, LI Meng-fei4, ZHAO Yan1,2,3(
), ZHANG Jin-jin1,2,3, HE Jia-die1,2,3, LIANG Yan-li1,2, YANG Sheng-chao1,3, LI Meng-fei4, ZHAO Yan1,2,3( )
)
Received:2025-04-20
Published:2026-02-26
Online:2026-03-17
摘要:
目的 探究4-香豆酸∶辅酶A连接酶(4-coumarate:coenzyme A ligase, 4CL)在当归根部木质素生物合成中的作用,为通过遗传改良或栽培手段定向调控当归药材品质奠定基础。 方法 通过生物信息学方法鉴定当归4CL基因家族,开展蛋白理化性质、二级结构预测、基因结构、启动子顺式作用元件以及系统进化分析,通过大肠杆菌异源表达和体外酶活验证基因功能,利用实时荧光定量PCR分析基因表达特征。 结果 在当归基因组中共鉴定到12个As4CLs,编码415‒572个氨基酸,相对分子质量在45‒62 kD,等电点范围为5.47‒9.06,二级结构以无规则卷曲和α-螺旋为主。系统进化分析将As4CL6与As4CL7归在Group I,其余As4CLs均属于Group Ⅲ。氨基酸序列比对结果显示As4CLs含有2个功能保守结构域Box I和Box Ⅱ。启动子分析结果显示,As4CLs基因含有丰富的光响应元件、激素响应元件及非生物胁迫响应元件;体外酶活实验证明As4CL6与As4CL7均能够催化4-香豆酸、咖啡酸和阿魏酸生成相应的辅酶A硫酯,且两个酶均对阿魏酸的亲和力最强;RT-qPCR结果显示As4CL6和As4CL7在当归抽薹根部的表达量高于未抽薹根部。 结论 成功克隆了As4CL6与As4CL7基因,通过体外酶活实验证明2个重组蛋白对3种底物均具有催化活性。
李勒松, 张金金, 贺佳蝶, 梁艳丽, 杨生超, 栗孟飞, 赵艳. 当归4-香豆酸∶辅酶A连接酶基因家族的鉴定与功能表征[J]. 生物技术通报, 2026, 42(2): 293-305.
LI Le-song, ZHANG Jin-jin, HE Jia-die, LIANG Yan-li, YANG Sheng-chao, LI Meng-fei, ZHAO Yan. Identification and Functional Characterization of the 4-Coumarate∶coenzyme A Ligase Gene Family in Angelica sinensis (Oliv.) Diels[J]. Biotechnology Bulletin, 2026, 42(2): 293-305.
引物名称 Primer name | 引物序列 Primer sequence (5′-3′) | 备注 Note |
|---|---|---|
| As4CL6-F | cagcaaatgggtcgcggatccATGGGGGATTGTGTAGCACC | 基因克隆 |
| As4CL6-R | cagtggtggtggtggtggtgTTATTTGGGAAGATCACCGGATGC | Gene cloning |
| As4CL7-F | cagcaaatgggtcgcggatccATGGATACCAAAACAAACAAGAAACAAG | |
| As4CL7-R | cagtggtggtggtggtggtgTTAGTTTTGAGGAAGATCGCCAGC | |
| q-As4CL6-F | TTCGTCAGGTACCACAGGAC | 实时荧光定量 |
| q-As4CL6-R | CCCCTGCTCTAAGTCCACAA | RT-qPCR |
| q-As4CL7-F | ATAAAATCGGGTGCGTGTGG | |
| q-As4CL7-R | TGCAGCCAACCTTCTTTGTC | |
| EEF1G-F | GTCCCAGCAGCCAAAAAGTC | 内参基因 |
| EEF1G-R | TCTGCCTTGGGCAATTCCTT | Reference genes |
表1 引物序列
Table 1 Primer sequences
引物名称 Primer name | 引物序列 Primer sequence (5′-3′) | 备注 Note |
|---|---|---|
| As4CL6-F | cagcaaatgggtcgcggatccATGGGGGATTGTGTAGCACC | 基因克隆 |
| As4CL6-R | cagtggtggtggtggtggtgTTATTTGGGAAGATCACCGGATGC | Gene cloning |
| As4CL7-F | cagcaaatgggtcgcggatccATGGATACCAAAACAAACAAGAAACAAG | |
| As4CL7-R | cagtggtggtggtggtggtgTTAGTTTTGAGGAAGATCGCCAGC | |
| q-As4CL6-F | TTCGTCAGGTACCACAGGAC | 实时荧光定量 |
| q-As4CL6-R | CCCCTGCTCTAAGTCCACAA | RT-qPCR |
| q-As4CL7-F | ATAAAATCGGGTGCGTGTGG | |
| q-As4CL7-R | TGCAGCCAACCTTCTTTGTC | |
| EEF1G-F | GTCCCAGCAGCCAAAAAGTC | 内参基因 |
| EEF1G-R | TCTGCCTTGGGCAATTCCTT | Reference genes |
基因名 Gene name | 基因ID Gene ID | 氨基酸数量 Number of amino acids (aa) | 分子量 Molecular weight (Da) | 等电点 Isoelectric point | 不稳定系数 Instability index | 亲水系数 Hydrophilicity coefficient | 亚细胞定位Subcellular localization |
|---|---|---|---|---|---|---|---|
| As4CL1 | AS01G01317 | 541 | 59 348.05 | 8.90 | 33.77 | 0.068 | 过氧化物酶体 |
| As4CL2 | AS02G00709 | 502 | 54 877.60 | 6.84 | 35.33 | 0.109 | 过氧化物酶体 |
| As4CL3 | AS04G00278 | 572 | 62 011.86 | 6.22 | 54.87 | -0.061 | 过氧化物酶体 |
| As4CL4 | AS04G02110 | 470 | 50 783.66 | 7.60 | 40.56 | 0.077 | 过氧化物酶体 |
| As4CL5 | AS05G03160 | 415 | 45 242.42 | 6.16 | 33.56 | 0.134 | 过氧化物酶体 |
| As4CL6 | AS06G02531 | 548 | 60 207.84 | 5.47 | 34.15 | 0.057 | 过氧化物酶体 |
| As4CL7 | AS07G02833 | 549 | 60 205.70 | 5.87 | 33.28 | -0.013 | 过氧化物酶体 |
| As4CL8 | AS07G03471 | 475 | 51 047.82 | 9.06 | 45.65 | -0.004 | 过氧化物酶体 |
| As4CL9 | AS08G00981 | 568 | 62 771.36 | 6.71 | 37.89 | 0.065 | 过氧化物酶体 |
| As4CL10 | AS09G00363 | 543 | 59 783.05 | 8.76 | 39.09 | -0.042 | 过氧化物酶体 |
| As4CL11 | AS09G01563 | 544 | 59 805.70 | 7.08 | 37.46 | -0.100 | 过氧化物酶体 |
| As4CL12 | ASUnG00581 | 550 | 60 229.63 | 5.70 | 35.50 | 0.024 | 过氧化物酶体 |
表2 As4CLs蛋白理化性质
Table 2 Physicochemical properties of As4CLs protein
基因名 Gene name | 基因ID Gene ID | 氨基酸数量 Number of amino acids (aa) | 分子量 Molecular weight (Da) | 等电点 Isoelectric point | 不稳定系数 Instability index | 亲水系数 Hydrophilicity coefficient | 亚细胞定位Subcellular localization |
|---|---|---|---|---|---|---|---|
| As4CL1 | AS01G01317 | 541 | 59 348.05 | 8.90 | 33.77 | 0.068 | 过氧化物酶体 |
| As4CL2 | AS02G00709 | 502 | 54 877.60 | 6.84 | 35.33 | 0.109 | 过氧化物酶体 |
| As4CL3 | AS04G00278 | 572 | 62 011.86 | 6.22 | 54.87 | -0.061 | 过氧化物酶体 |
| As4CL4 | AS04G02110 | 470 | 50 783.66 | 7.60 | 40.56 | 0.077 | 过氧化物酶体 |
| As4CL5 | AS05G03160 | 415 | 45 242.42 | 6.16 | 33.56 | 0.134 | 过氧化物酶体 |
| As4CL6 | AS06G02531 | 548 | 60 207.84 | 5.47 | 34.15 | 0.057 | 过氧化物酶体 |
| As4CL7 | AS07G02833 | 549 | 60 205.70 | 5.87 | 33.28 | -0.013 | 过氧化物酶体 |
| As4CL8 | AS07G03471 | 475 | 51 047.82 | 9.06 | 45.65 | -0.004 | 过氧化物酶体 |
| As4CL9 | AS08G00981 | 568 | 62 771.36 | 6.71 | 37.89 | 0.065 | 过氧化物酶体 |
| As4CL10 | AS09G00363 | 543 | 59 783.05 | 8.76 | 39.09 | -0.042 | 过氧化物酶体 |
| As4CL11 | AS09G01563 | 544 | 59 805.70 | 7.08 | 37.46 | -0.100 | 过氧化物酶体 |
| As4CL12 | ASUnG00581 | 550 | 60 229.63 | 5.70 | 35.50 | 0.024 | 过氧化物酶体 |
基因名 Gene name | α-螺旋 Alpha helix (%) | β-折叠 Beta turn (%) | 延伸链 Extended strand (%) | 无规则卷曲 Random coil (%) |
|---|---|---|---|---|
| As4CL1 | 30.87 | 7.76 | 19.59 | 41.77 |
| As4CL2 | 33.27 | 7.97 | 20.32 | 38.45 |
| As4CL3 | 29.72 | 6.82 | 17.66 | 45.80 |
| As4CL4 | 32.77 | 7.66 | 19.79 | 39.79 |
| As4CL5 | 30.60 | 7.95 | 20.00 | 41.45 |
| As4CL6 | 29.96 | 6.99 | 19.49 | 43.57 |
| As4CL7 | 30.28 | 6.97 | 19.45 | 43.30 |
| As4CL8 | 30.95 | 8.21 | 20.21 | 40.63 |
| As4CL9 | 28.87 | 6.51 | 17.96 | 46.65 |
| As4CL10 | 31.68 | 7.00 | 19.15 | 42.17 |
| As4CL11 | 30.33 | 6.99 | 19.67 | 43.01 |
| As4CL12 | 30.55 | 6.91 | 19.45 | 43.09 |
表3 As4CLs蛋白质二级结构分析
Table 3 Secondary structure analysis of As4CLs protein
基因名 Gene name | α-螺旋 Alpha helix (%) | β-折叠 Beta turn (%) | 延伸链 Extended strand (%) | 无规则卷曲 Random coil (%) |
|---|---|---|---|---|
| As4CL1 | 30.87 | 7.76 | 19.59 | 41.77 |
| As4CL2 | 33.27 | 7.97 | 20.32 | 38.45 |
| As4CL3 | 29.72 | 6.82 | 17.66 | 45.80 |
| As4CL4 | 32.77 | 7.66 | 19.79 | 39.79 |
| As4CL5 | 30.60 | 7.95 | 20.00 | 41.45 |
| As4CL6 | 29.96 | 6.99 | 19.49 | 43.57 |
| As4CL7 | 30.28 | 6.97 | 19.45 | 43.30 |
| As4CL8 | 30.95 | 8.21 | 20.21 | 40.63 |
| As4CL9 | 28.87 | 6.51 | 17.96 | 46.65 |
| As4CL10 | 31.68 | 7.00 | 19.15 | 42.17 |
| As4CL11 | 30.33 | 6.99 | 19.67 | 43.01 |
| As4CL12 | 30.55 | 6.91 | 19.45 | 43.09 |
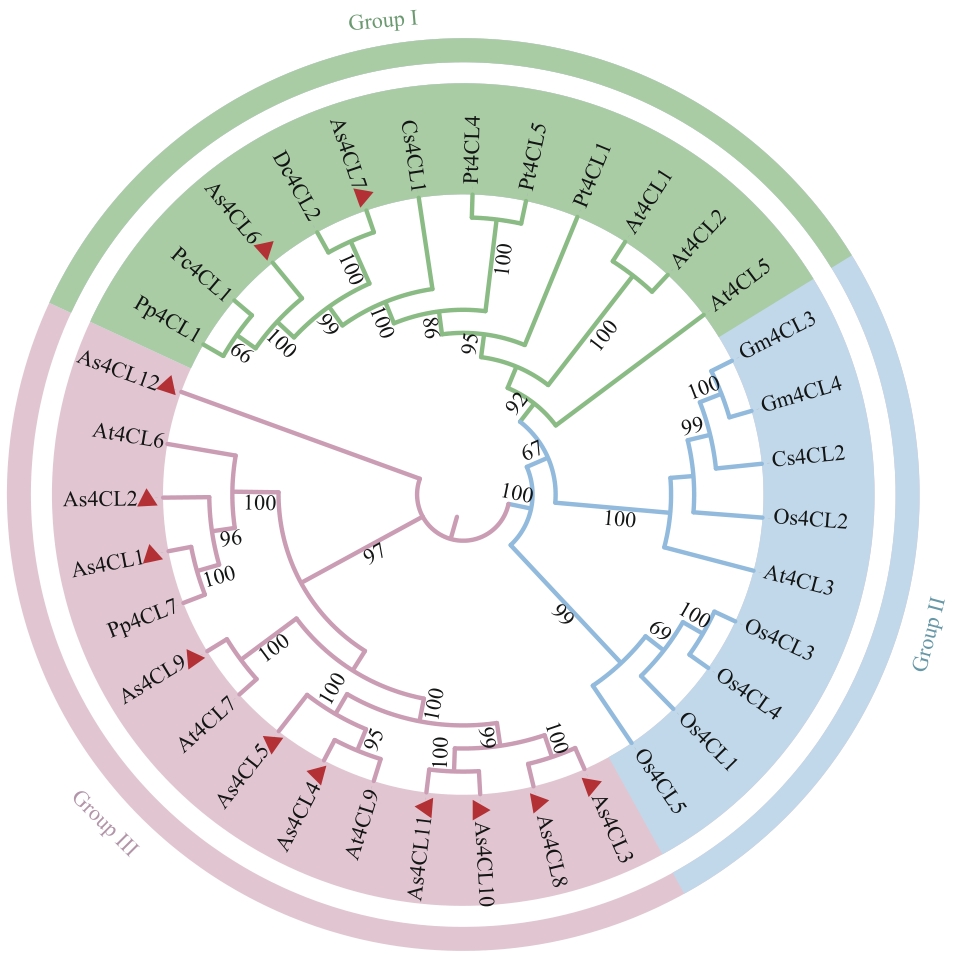
图3 当归As4CLs基因家族的系统发育分析当归As4CLs家族成员用三角形标注。At:拟南芥;Os:水稻;Gm:大豆;Pt:毛白杨;Cs:茶树;Pp:白花前胡;Pc:欧芹;Dc:胡萝卜。蛋白名称及ID号见附表1
Fig. 3 Phylogenetic analysis of the As4CLs gene family in A. sinensisMembers of the As4CLs family in A. sinensis are labeled with triangles. At: Arabidopsis thaliana; Os: Oryza sativa; Gm: Glycine max; Pt: Populus tomentosa; Cs: Camellia sinensis; Pp: Peucedanum praeruptorum; Pc: Petroselinum crispum; Dc: Daucus carota. Protein names and ID numbers are listed in Supplementary table 1
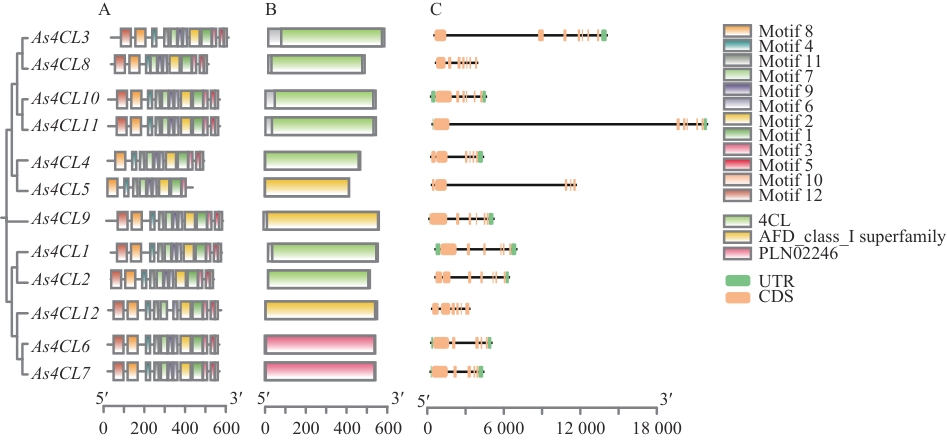
图4 当归As4CLs基因结构和保守结构域A:基因结构;B:保守结构域;C:内含子与外显子结构
Fig. 4 The structure and conserved domains of the As4CLs gene in A. sinensisA: Gene structure. B: Conserved domain. C: Intron and exon structures
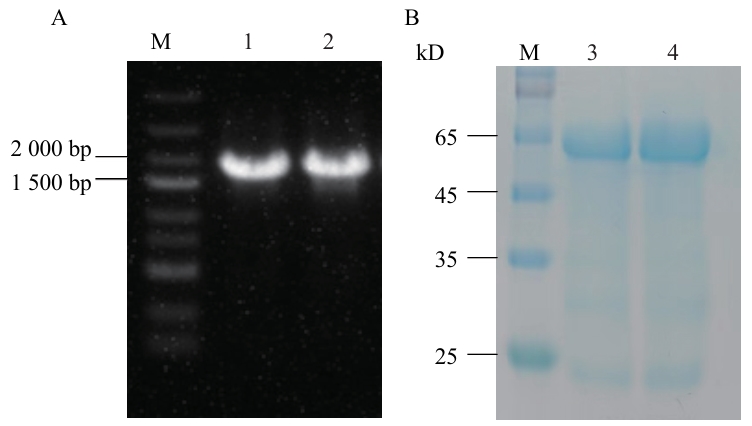
图7 电泳检测分析A:琼脂糖凝胶电泳检测扩增产物;B:As4CLs蛋白的SDS-PAGE分析;M:Marker;1:As4CL6;2:As4CL7;3:pET-28a-As4CL6重组蛋白;4:pET-28a-As4CL7重组蛋白
Fig. 7 Electrophoretic detection analysisA: Detection of amplified products by agarose gel electrophoresis; B: SDS-PAGE analysis of As4CLs protein. M: Marker; 1: As4CL6; 2: As4CL7; 3: purified pET-28a-As4CL6 protein; 4: purified pET-28a-As4CL7 protein
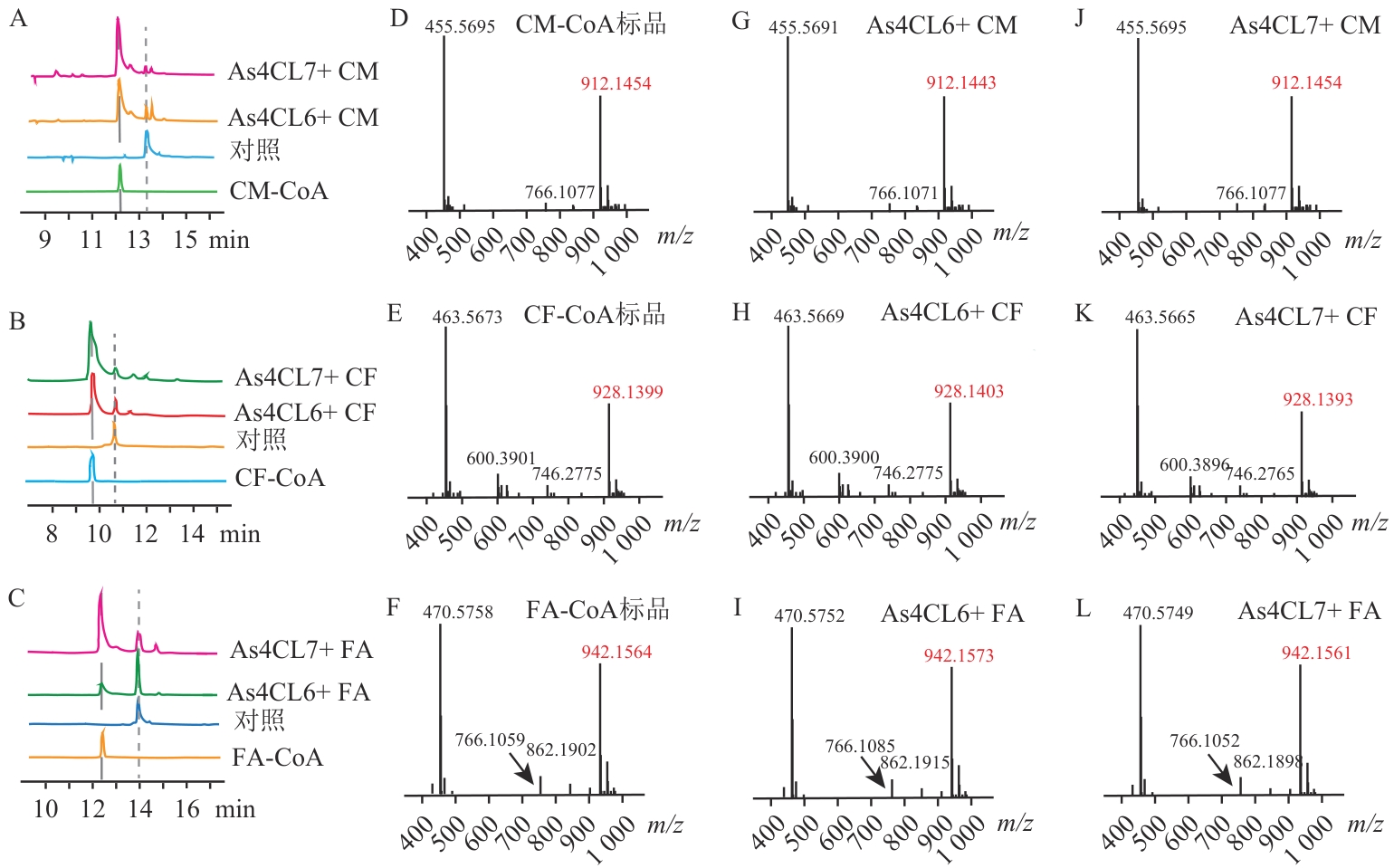
图8 As4CL6与As4CL7的功能表征A‒C:As4CL6、As4CL7体外酶促反应产物的HPLC分析;D‒F:产物标准品的LC-MS分析;G‒L:As4CL6、As4CL7体外酶促反应产物的LC-MS分析。CM、CF、FA分别代表4-香豆酸、咖啡酸和阿魏酸;CM-CoA、CF-CoA、FA-CoA分别代表4-香豆酰辅酶A、咖啡酰辅酶A和阿魏酰辅酶A
Fig. 8 Functional profiling of As4CL6 and As4CL7A‒C: HPLC analysis of in vitro enzymatic reaction products of As4CL6 and As4CL7; D-F: LC-MS analysis of product standards; G‒L: LC-MS analysis of in vitro enzymatic reaction products of As4CL6 and As4CL7. CM, CF, and FA refers to 4-coumaric acid, caffeic acid, and ferulic acid; CM-CoA, CF-CoA, and FA-CoA represent 4-coumaroyl CoA, caffeoyl CoA, and feruloyl CoA, respectively
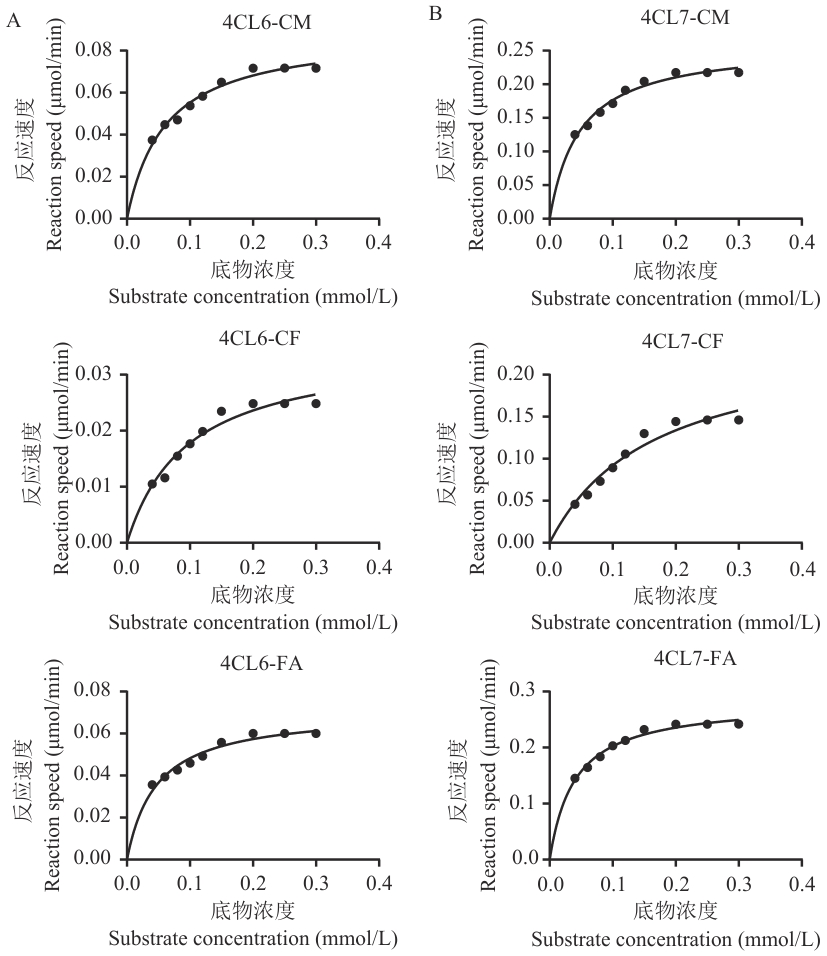
图9 As4CL6与As4CL7的酶动力学分析A:4CL6对不同底物的非线性回归曲线;B:4CL7对不同底物的非线性回归曲线。CM、CF、FA分别代表4-香豆酸、咖啡酸和阿魏酸
Fig. 9 Enzyme kinetic analysis of As4CL6 and As4CL7A: Nonlinear regression curve of 4CL6 on different substrates. B: Nonlinear regression curve of 4CL7 on different substrates. CM, CF, and FA stand for 4-coumaric acid, caffeic acid, and ferulic acid
酶 Enzyme | 底物 Substrate | Km (μmol/L) | Vmax (μmol·L-1·min-1) | Vmax/Km (10-3/min) |
|---|---|---|---|---|
| As4CL6 | 4-香豆酸 | 60.73 | 0.088 83 | 1.462 7 |
| 咖啡酸 | 96.28 | 0.034 96 | 0.363 1 | |
| 阿魏酸 | 46.77 | 0.070 77 | 1.513 1 | |
| As4CL7 | 4-香豆酸 | 47.55 | 0.260 20 | 5.472 1 |
| 咖啡酸 | 166.10 | 0.244 70 | 1.473 2 | |
| 阿魏酸 | 39.61 | 0.282 20 | 7.124 5 |
表4 As4CL6与As4CL7的动力学参数
Table 4 Kinetic parameters of As4CL6 and As4CL7
酶 Enzyme | 底物 Substrate | Km (μmol/L) | Vmax (μmol·L-1·min-1) | Vmax/Km (10-3/min) |
|---|---|---|---|---|
| As4CL6 | 4-香豆酸 | 60.73 | 0.088 83 | 1.462 7 |
| 咖啡酸 | 96.28 | 0.034 96 | 0.363 1 | |
| 阿魏酸 | 46.77 | 0.070 77 | 1.513 1 | |
| As4CL7 | 4-香豆酸 | 47.55 | 0.260 20 | 5.472 1 |
| 咖啡酸 | 166.10 | 0.244 70 | 1.473 2 | |
| 阿魏酸 | 39.61 | 0.282 20 | 7.124 5 |
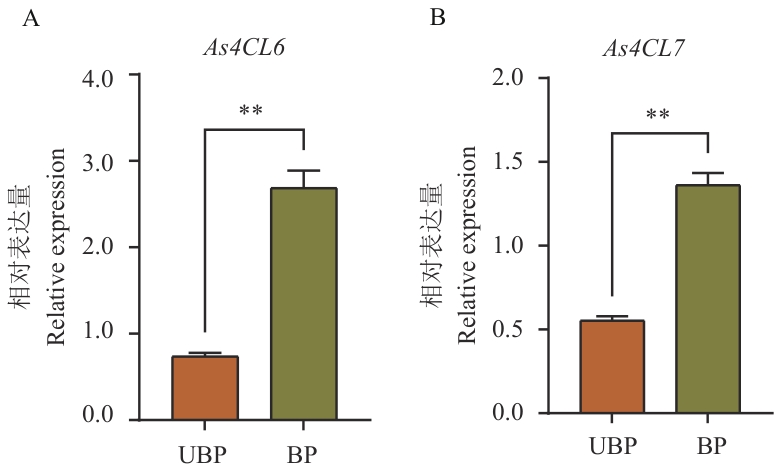
图10 As4CL6和As4CL7表达量分析UBP:未抽薹植株根部;BP:抽薹植株根部;** P<0.01
Fig. 10 Analysis of As4CL6 and As4CL7 expressionsUBP: Roots of unbolting plants; BP: roots of bolting plants;** P<0.01
| [1] | 严辉, 段金廒, 尚尔鑫, 等. 当归不同部位入药功效取向差异的化学物质基础与药性关联性研究 [J]. 中草药, 2014, 45(21): 3208-3212. |
| Yan H, Duan JA, Shang EX, et al. Study on chemical materials and drug nature association of efficacy orientation of different parts from Angelicae Sinensis Radix [J]. Chin Tradit Herb Drugs, 2014, 45(21): 3208-3212. | |
| [2] | 董涵. 低温调控当归生长及主效活性物质生物合成的机制研究 [D]. 兰州: 甘肃中医药大学, 2022. |
| Dong H. Mechanism of cool temperature regulating growth and main metabolites biosynthesis in Angelica sinensis [D]. Lanzhou: Gansu University of Chinese Medicine, 2022. | |
| [3] | 吕成龙, 李会会, 史永洁, 等. 中药当归现代研究进展及其质量标志物的预测分析 [J]. 中国中药杂志, 2022, 47(19): 5140-5157. |
| Lyu CL, Li HH, Shi YJ, et al. Research progress of Angelicae sinensis and predictive analysis on its quality markers [J]. China J Chin Mater Med, 2022, 47(19): 5140-5157. | |
| [4] | 赵静, 夏晓培. 当归的化学成分及药理作用研究现状 [J]. 临床合理用药杂志, 2020, 13(6): 172-174. |
| Zhao J, Xia XP. Research status of chemical constituents and pharmacological effects of Angelica sinensis [J]. Chin J Clin Ration Drug Use, 2020, 13(6): 172-174. | |
| [5] | 国家药典委员会. 中华人民共和国药典-一部: 2020年版 [M]. 北京: 中国医药科技出版社, 2020. |
| National Pharmacopoeia Committee. People’s republic of China (PRC) pharmacopoeia-part I: 2020 edition [M]. Beijing: China Medical Science Press, 2020. | |
| [6] | 梁盈, 袭晓娟, 刘巧丽, 等. 阿魏酸及其衍生物的生理活性及应用研究进展 [J]. 食品与生物技术学报, 2018, 37(5): 449-454. |
| Liang Y, Xi XJ, Liu QL, et al. Research progress on the physiological activity and application of ferulic acid and its derivatives [J]. J Food Sci Biotechnol, 2018, 37(5): 449-454. | |
| [7] | Kitashiba H, Yokoi S. Genes for bolting and flowering [M]. Compendium of Plant Genomes, The Radish Genome, 2017: 151-163. |
| [8] | Li ML, Cui XW, Jin L, et al. Bolting reduces ferulic acid and flavonoid biosynthesis and induces root lignification in Angelica sinensis [J]. Plant Physiol Biochem, 2022, 170: 171-179. |
| [9] | Liu L, Liu C, Hou XL, et al. FTIP1 is an essential regulator required for florigen transport [J]. PLoS Biol, 2012, 10(4): e1001313. |
| [10] | 黎洁. 光周期阶段当归抽薹开花的调控机制研究 [D]. 兰州: 甘肃农业大学, 2021. |
| Li J. Study on regulation mechanism of bolting and flowering of Angelica sinensis at photoperiodic stage [D]. Lanzhou: Gansu Agricultural University, 2021. | |
| [11] | Sulis DB, Jiang X, Yang CM, et al. Multiplex CRISPR editing of wood for sustainable fiber production [J]. Science, 2023, 381(6654): 216-221. |
| [12] | Chanoca A, de Vries L, Boerjan W. Lignin engineering in forest trees [J]. Front Plant Sci, 2019, 10: 912. |
| [13] | Liu QQ, Luo L, Zheng LQ. Lignins: Biosynthesis and biological functions in plants [J]. Int J Mol Sci, 2018, 19(2): 335. |
| [14] | 吴萍, 王晓宇, 郭俊霞, 等. 早期抽薹对白芷生长发育和品质的影响 [J]. 中国农业科技导报, 2023, 25(12): 58-66. |
| Wu P, Wang XY, Guo JX, et al. Effects of early bolting on growth and quality of Angelica dahurica [J]. J Agric Sci Technol, 2023, 25(12): 58-66. | |
| [15] | 刘双利, 许永华, 王晓慧, 等. 防风抽薹开花的研究进展 [J]. 人参研究, 2016, 28(6): 52-56. |
| Liu SL, Xu YH, Wang XH, et al. Research progress on the early bolting and flowering of Divaricate saposhnikovia Root [J]. Ginseng Res, 2016, 28(6): 52-56. | |
| [16] | Lavhale SG, Kalunke RM, Giri AP. Structural, functional and evolutionary diversity of 4-coumarate-CoA ligase in plants [J]. Planta, 2018, 248(5): 1063-1078. |
| [17] | Sun SC, Xiong XP, Zhang XL, et al. Characterization of the Gh4CL gene family reveals a role of Gh4CL7 in drought tolerance [J]. BMC Plant Biol, 2020, 20(1): 125. |
| [18] | Liu H, Guo ZH, Gu FW, et al. 4-Coumarate-CoA ligase-like gene OsAAE3 negatively mediates the rice blast resistance, floret development and lignin biosynthesis [J]. Front Plant Sci, 2017, 7: 2041. |
| [19] | Trantas E, Panopoulos N, Ververidis F. Metabolic engineering of the complete pathway leading to heterologous biosynthesis of various flavonoids and stilbenoids in Saccharomyces cerevisiae [J]. Metab Eng, 2009, 11(6): 355-366. |
| [20] | Wang YC, Yi H, Wang M, et al. Structural and kinetic analysis of the unnatural fusion protein 4-coumaroyl-CoA ligase: stilbene synthase [J]. J Am Chem Soc, 2011, 133(51): 20684-20687. |
| [21] | Lin YH, Sun XX, Yuan QP, et al. Combinatorial biosynthesis of plant-specific coumarins in bacteria [J]. Metab Eng, 2013, 18: 69-77. |
| [22] | Li Y, Kim JI, Pysh L, et al. Four isoforms of Arabidopsis 4-coumarate: CoA ligase have overlapping yet distinct roles in phenylpropanoid metabolism [J]. Plant Physiol, 2015, 169(4): 2409-2421. |
| [23] | Sun HY, Li Y, Feng SQ, et al. Analysis of five rice 4-coumarate: coenzyme A ligase enzyme activity and stress response for potential roles in lignin and flavonoid biosynthesis in rice [J]. Biochem Biophys Res Commun, 2013, 430(3): 1151-1156. |
| [24] | Reinprecht Y, Yadegari Z, Perry GE, et al. In silico comparison of genomic regions containing genes coding for enzymes and transcription factors for the phenylpropanoid pathway in Phaseolus vulgaris L. and Glycine max L. Merr [J]. Front Plant Sci, 2013, 4: 317. |
| [25] | Rao GD, Pan X, Xu F, et al. Divergent and overlapping function of five 4-coumarate/coenzyme a ligases from Populus tomentosa [J]. Plant Mol Biol Report, 2015, 33(4): 841-854. |
| [26] | Li MZ, Guo LL, Wang YR, et al. Molecular and biochemical characterization of two 4-coumarate: CoA ligase genes in tea plant (Camellia sinensis) [J]. Plant Mol Biol, 2022, 109(4/5): 579-593. |
| [27] | 侯冲, 晁楠, 戴明洁, 等. 桑树4CL基因家族的筛选和Mm4CL2的功能研究 [J]. 蚕业科学, 2022, 48(1): 18-24. |
| Hou C, Chao N, Dai MJ, et al. Screening of 4CL family genes in mulberry and functional study of Mm4CL2 [J]. Acta Sericologica Sin, 2022, 48(1): 18-24. | |
| [28] | Gong JM, Sun SC, Zhu QH, et al. Gh4CL20/20A involved in flavonoid biosynthesis is essential for male fertility in cotton (Gossypium hirsutum L.) [J]. Plant Physiol Biochem, 2024, 208: 108484. |
| [29] | Zhou X, Cao J, Liu XM, et al. Cloning and functional analysis of Gb4CL1 and Gb4CL2 from Ginkgo biloba [J]. Plant Genome, 2024, 17(2): e20440. |
| [30] | Douglas C, Hoffmann H, Schulz W, et al. Structure and elicitor or u.v.-light-stimulated expression of two 4-coumarate: CoA ligase genes in parsley [J]. EMBO J, 1987, 6(5): 1189-1195. |
| [31] | Liu TT, Yao RL, Zhao YC, et al. Cloning, functional characterization and site-directed mutagenesis of 4-coumarate: coenzyme a ligase (4CL) involved in coumarin biosynthesis in Peucedanum praeruptorum Dunn [J]. Front Plant Sci, 2017, 8: 4. |
| [32] | 张苛苛, 谭宇萍, 徐欢欢, 等. 菘蓝4-香豆酸: 辅酶A连接酶全基因家族的鉴定及表达分析 [J]. 中国中药杂志, 2024, 49(2): 361-369. |
| Zhang KK, Tan YP, Xu HH, et al. Identification and expression analysis of whole gene family of Isatis indigotica 4-coumarate: CoA ligase [J]. China J Chin Mater Med, 2024, 49(2): 361-369. | |
| [33] | 朋冬琴, 罗蜜蜜, 郭欣慰, 等. 当归实时荧光定量PCR内参基因筛选 [J]. 中草药, 2024, 55(1): 269-278. |
| Peng DQ, Luo MM, Guo XW, et al. Selection of reference genes for quantitative real-time PCR analysis in Angelica sinensis [J]. Chin Tradit Herb Drugs, 2024, 55(1): 269-278. | |
| [34] | 何磊, 严希, 袁圆, 等. 辣椒4CL基因家族成员的鉴定与生物信息学分析 [J]. 分子植物育种, 2022, 20(8): 2478-2484. |
| He L, Yan X, Yuan Y, et al. Identification and bioinformatics analysis of the 4CL gene family members in pepper (Capsicum annuum) [J]. Mol Plant Breed, 2022, 20(8): 2478-2484. | |
| [35] | 郑飞雄, 陈俊宇, 江林琪, 等. 掌叶覆盆子4CL基因家族鉴定及表达分析 [J]. 农业生物技术学报, 2024, 32(2): 311-321. |
| Zheng FX, Chen JY, Jiang LQ, et al. Identification and expression analysis of 4CL gene family in Rubus chingii [J]. J Agric Biotechnol, 2024, 32(2): 311-321. | |
| [36] | Wang YY, Guo LH, Zhao YJ, et al. Systematic analysis and expression profiles of the 4-coumarate: CoA ligase (4CL) gene family in pomegranate (Punica granatum L.) [J]. Int J Mol Sci, 2022, 23(7): 3509. |
| [37] | Kienow L, Schneider K, Bartsch M, et al. Jasmonates meet fatty acids: functional analysis of a new acyl-coenzyme A synthetase family from Arabidopsis thaliana [J]. J Exp Bot, 2008, 59(2): 403-419. |
| [38] | De Azevedo Souza C, Barbazuk B, Ralph SG, et al. Genome-wide analysis of a land plant-specific acyl: coenzymeA synthetase (ACS) gene family in Arabidopsis, poplar, rice and Physcomitrella [J]. New Phytol, 2008, 179(4): 987-1003. |
| [39] | Ehlting J, Büttner D, Wang Q, et al. Three 4-coumarate: coenzyme A ligases in Arabidopsis thaliana represent two evolutionarily divergent classes in angiosperms [J]. Plant J, 1999, 19(1): 9-20. |
| [40] | Hamberger B, Hahlbrock K. The 4-coumarate: CoA ligase gene family in Arabidopsis thaliana comprises one rare, sinapate-activating and three commonly occurring isoenzymes [J]. Proc Natl Acad Sci USA, 2004, 101(7): 2209-2214. |
| [41] | Lavhale SG, Joshi RS, Kumar Y, et al. Functional insights into two Ocimum kilimandscharicum 4-coumarate-CoA ligases involved in phenylpropanoid biosynthesis [J]. Int J Biol Macromol, 2021, 181: 202-210. |
| [42] | Zhang GH, Yu ZM, Yao B, et al. SsMYB113, a Schima superba MYB transcription factor, regulates the accumulation of flavonoids and functions in drought stress tolerance by modulating ROS generation [J]. Plant Soil, 2022, 478(1): 427-444. |
| [43] | Yuan CX, Li LS, Zhou PH, et al. Decoding the root lignification mechanism of Angelica sinensis through genome-wide DNA methylation analysis [J]. J Exp Bot, 2025, 76(9): 2573-2589. |
| [44] | Lindermayr C, Möllers B, Fliegmann J, et al. Divergent members of a soybean (Glycine max L.) 4-coumarate: coenzyme A ligase gene family [J]. Eur J Biochem, 2002, 269(4): 1304-1315. |
| [45] | Zhao Y, Liu GZ, Yang F, et al. Multilayered regulation of secondary metabolism in medicinal plants [J]. Mol Hortic, 2023, 3(1): 11. |
| [46] | Gao QQ, Zhang JJ, Cao JT, et al. MetaDb: a database for metabolites and their regulation in plants with an emphasis on medicinal plants [J]. Mol Hortic, 2024, 4(1): 17. |
| [1] | 刘保财, 胡学博, 张武君, 赵云青, 黄颖桢, 陈菁瑛. 石仙桃内参基因筛选与应用[J]. 生物技术通报, 2026, 42(2): 158-168. |
| [2] | 倪莹, 李雷, 汪进萱, 马波, 孟昕, 冷平生, 吴静, 胡增辉. 紫丁香So4CL的克隆及功能分析[J]. 生物技术通报, 2026, 42(1): 139-149. |
| [3] | 向春繁, 李勒松, 王娟, 梁艳丽, 杨生超, 栗孟飞, 赵艳. 当归肉桂醇脱氢酶AsCAD功能鉴定及表达分析[J]. 生物技术通报, 2025, 41(2): 295-308. |
| [4] | 杨晨欣, 李梦秀, 姜唐, 张文娥, 潘学军. 美人蕉R2R3-MYB基因家族的鉴定及与花青素相关成员的表达分析[J]. 生物技术通报, 2025, 41(12): 201-213. |
| [5] | 蒋小刚, 余凯迪, 郭晓亮. 重齿毛当归核心种质构建及其有效成分关联分析[J]. 生物技术通报, 2025, 41(11): 236-246. |
| [6] | 杜品廷, 吴国江, 王振国, 李岩, 周伟, 周亚星. 高粱CPP基因家族鉴定及表达分析[J]. 生物技术通报, 2025, 41(1): 132-142. |
| [7] | 孔小平, 陈利文, 刘思思, 严湘萍. 胡萝卜抽薹相关性状全基因组关联分析[J]. 生物技术通报, 2024, 40(5): 120-130. |
| [8] | 钟匀, 林春, 刘正杰, 董陈文华, 毛自朝, 李兴玉. 芦笋皂苷合成相关糖基转移酶基因克隆及原核表达分析[J]. 生物技术通报, 2024, 40(4): 255-263. |
| [9] | 杨淇, 魏子迪, 宋娟, 童堃, 杨柳, 王佳涵, 刘海燕, 栾维江, 马轩. 水稻组蛋白H1三突变体的创建和转录组学分析[J]. 生物技术通报, 2024, 40(4): 85-96. |
| [10] | 田姗姗, 黄诗宇, 杨天为, 高曼熔, 张尚文, 何龙飞, 张向军, 李婷, 石前. 高温干旱复合胁迫下铁皮石斛MYB基因家族鉴定及表达分析[J]. 生物技术通报, 2024, 40(12): 145-159. |
| [11] | 谢田朋, 张佳宁, 董永骏, 张建, 景明. 早期抽薹对当归根际土壤微环境的影响[J]. 生物技术通报, 2023, 39(7): 206-218. |
| [12] | 滕梦鑫, 徐亚, 何静, 汪奇, 乔飞, 李敬阳, 李新国. 香蕉MaMC6的克隆及原核表达分析[J]. 生物技术通报, 2023, 39(12): 179-186. |
| [13] | 谢田朋, 柳娜, 刘越敏, 曲馨, 薄双琴, 景明. 化肥减量配施中药源植物生长调节剂对当归质量和根际土壤细菌群落的影响[J]. 生物技术通报, 2022, 38(3): 79-91. |
| [14] | 王雁,汤纳平,邱云良,南雅萍,马璟. 血浆/血清MicroRNAs定量分析的校正策略[J]. 生物技术通报, 2013, 29(10): 52-57. |
| [15] | 于光;段金廒;宋秉生;何子清;. cDNA-AFLP比较当归早薹基因转录差异反应体系的建立[J]. , 2010, 0(09): 134-137. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||